INTRODUCTION
Antibacterial resistance in clinically important Gram-negative pathogens is transmitted easily among individuals in various community settings via water, sanitation, hygiene, and food pathways [Reference Fornasini1–Reference Amaya6]. In both urban and rural settings, there are many means by which resistant pathogens are disseminated, including human migration, overcrowding, chemical pollution, waste water, and untreated groundwater [Reference McKeon, Calabrese and Bissonnette3, Reference Hawkey and Jones5–Reference Eisenberg9]. An important mode of transmission of resistant Gram-negative bacteria includes animal vectors, as observational data have clearly shown a similarity in the resistance genes and enterotoxins of Escherichia coli isolated from companion animals and horses, and extended-spectrum β-lactamase (ESBL)-producing isolates described in humans [Reference Dierikx10]. Faecal carriage of antibiotic-resistant pathogens in humans and animals predisposes to further contamination of food and water supplies to complete a transmission cycle [Reference Woerther4, Reference Carattoli11–Reference Moreno16].
Latin America fulfils all of the above-mentioned criteria required to drive the spread of antibacterial drug resistance. As in other world regions, high antibacterial use and misuse (e.g. inappropriate drug selection, suboptimal dosing, poor patient adherence) may also drive bacterial resistance in Latin America [Reference Amábile-Cuevas7, Reference Rossi17]. Consequently, for many pathogens, rates of antimicrobial resistance in Latin America appear to be high relative to other regions of the world [Reference Reinert8, Reference Baquero18].
The unique set of medical, societal, and ecological circumstances in Latin America has underpinned a dynamic epidemiology of Gram-negative infections in the outpatient setting over the last 15 years. In particular, community-associated infections caused by multidrug-resistant Enterobacteriaceae and other Gram-negative bacteria are an important public health concern [Reference Andrade19–Reference Zurita25]. Foremost of the pathogens causing concern are strains of E. coli-producing ESBLs, which are an important cause of urinary tract infection (UTI) and intra-abdominal infection (IAI) sometimes associated with bacteraemia [Reference Andrade19–Reference Casellas21, Reference Gales, Sader and Jones23, Reference Pitout and Laupland26, Reference Rodríguez-Baño27]. Because ESBL-producing bacteria now cause many infections in the community setting, the medical community is increasingly reliant on multilevel microbial surveillance to inform treatment decisions, identify major problems, and guide adequate control measures [Reference Andrade19–Reference Alvarez, Espino and Contreras22, Reference Pitout and Laupland26, Reference O'Brien and Stelling28].
This narrative review stems from the 10th Meeting of the Latin America Working Group on Bacterial Resistance in São Paulo, Brazil, 20–21 May 2012, at which the Working Group members reflected on the increasing recognition of the clinical importance of multidrug-resistant Gram-negative infections in the community setting in Latin America. Using primary data from published studies and policy documents, the aim of this review is to report on the epidemiology of Gram-negative bacteria isolated in Latin America over the last 10 years, describe the importance of detection in outpatients, and consider the implications for prescribing decisions.
METHODS
In order to review the published clinical data relating to UTIs and IAIs due to Gram-negative pathogens in the community setting of Latin America, a systematic search of the biomedical literature was conducted. The title/abstract fields of Pubmed were searched, limited by the dates 1 January 2005 to 12 November 2012, for articles using the following terms and Boolean logic: (‘Latin America’ or ‘South America’ or ‘Central America’ or Mexico or Guatemala or Honduras or Nicaragua or ‘Costa Rica’ or Cuba or ‘Dominican Republic’ or Panama or Colombia or Venezuela or Guyana or Suriname or ‘French Guiana’ or Brazil or Ecuador or Peru or Bolivia or Paraguay or Uruguay or Chile or Argentina) and (‘Gram-negative infection’ or ‘Gram-negative pathogen’ or ‘Gram-negative bacilli’ or ‘Escherichia coli’ or ‘Klebsiella pneumoniae’ or ‘Proteus mirabilis’ orCitrobacter orSerratia or ‘urinary tract infection’ or ‘intra-abdominal infection’). In addition, we searched the Scientific Electronic Library Online (SciELO), which publishes health science data specifically from Latin America. Searches on SciELO were limited to policy statements. A small number of additional references were identified from the reference lists of published articles.
The titles and abstracts of all references obtained in the search were screened by the authors. We included all original research articles that reported information on the: (1) susceptibilities of Gram-negative pathogens causing UTIs and IAIs in Latin American primary care; (2) proportion of Gram-negative clinical isolates harbouring ESBL genes or expressing ESBLs; and (3) presence of circulating ESBLs and first reports of novel ESBLs. Only studies reporting data for >40 isolates were selected for assessing drug resistance.
EPIDEMIOLOGY
Overview
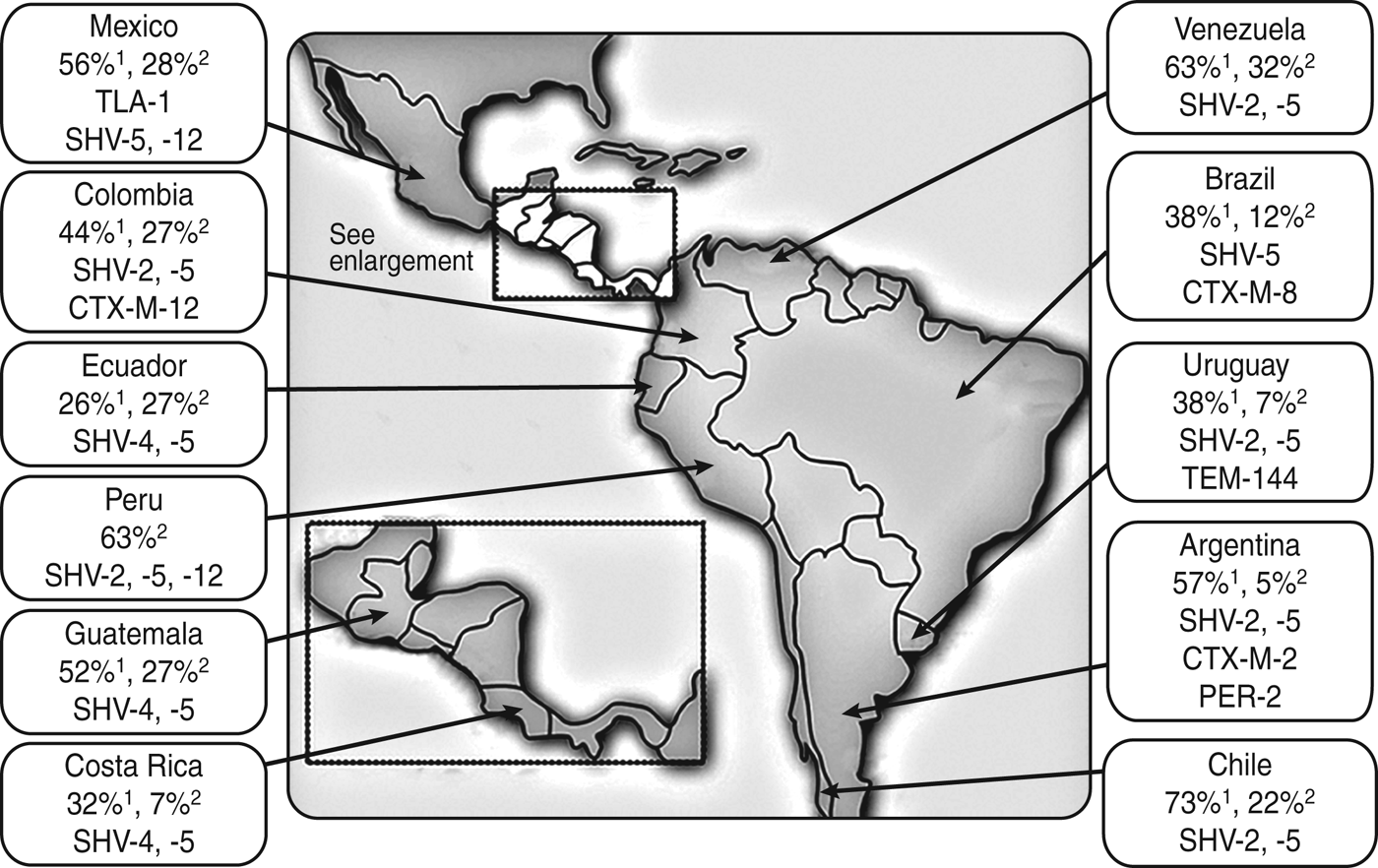
ESBLs are found predominantly in Klebsiella spp. and E. coli, but have also been described in other genera of Enterobacteriaceae including species of Citrobacter, Serratia, Proteus, Salmonella, and Enterobacter [Reference Bradford29, Reference Minarini30]. Until the turn of the century, Klebsiella pneumoniae harbouring Temoneira (TEM)-type and sulfhydryl variable (SHV)-type ESBLs were prevalent in the nosocomial setting only, and cefotaxime-resistant (CTX-M) β-lactamase-producing organisms were rarely isolated (Fig. 1) [Reference Minarini30–Reference Mattar and Martinez32]. However, in the first years of this century, Latin America became the first continent where CTX-M variants began to displace TEM and SHV variants as the most common type of ESBL, mainly in E. coli [Reference Cantón and Coque31]. CTX-M-type ESBLs share only 40% homology with TEM or SHV enzymes and are considered unrelated [Reference Minarini30].

Fig. 1. Prevalence (and type) of extended spectrum β-lactamases harboured by K. pneumoniae 1 and E. coli 2 in Latin American clinical isolates during the late 1990s [Reference Mattar and Martinez32].
The CTX-M-type ESBLs have now reached endemic proportions in South America aided by the location of bla CTX-M genes on plasmids and transposons, which engenders dissemination of CTX-M-producing strains in different Enterobacteriaceae [Reference Minarini30, Reference Cantón and Coque31, Reference Cantón, González-Alba and Galán33]. The spread of mobile genetic elements, mainly conjugative plasmids belonging to classic incompatibility groups, and the dispersion of specific clones have been responsible for the increase in ESBL-producing isolates and for the spread of many specific ESBLs, including CTX-M. ESBLs are often associated with co-resistance to fluoroquinolones, aminoglycosides, and trimethoprim/sulfamethoxazole, which may also have contributed to the current epidemiological ESBL scenario [Reference Cantón, González-Alba and Galán33].
Also over the same time period, E. coli replaced Klebsiella spp. as the predominant species of ESBL-producing Enterobacteriaceae in the community, largely because of the ease of CTX-M mobilization [Reference Cantón and Coque31]. Perhaps the high prevalence of community-acquired infection by E. coli-harbouring ESBLs was to be expected, given that these strains have been isolated from numerous sources such as domestic animals, food products, well water, sewage, and stool samples from healthy individuals [Reference Amaya6, Reference Ben-Ami34]. Importantly, there is evidence that community-associated E. coli ESBLs have infiltrated hospital settings [Reference Cantón and Coque31, Reference Ben-Ami34].
The most common community-acquired infection by Gram-negative bacteria is UTI, but clinicians are confronted with many different infection types including pneumonia and IAI [Reference Andrade19, Reference Bours20, Reference Casellas35]. E. coli, K. pneumoniae, and Proteus mirabilis are the most common organisms causing UTIs in ambulatory patients in Latin America [Reference Andrade19–Reference Casellas21, Reference Casellas35]. The aetiology of community-associated IAIs depends on the distribution of microflora at the anatomical site, although infections by enterobacteria (primarily E. coli and K. pneumoniae) tend to predominate [Reference Hawser36].
UTI
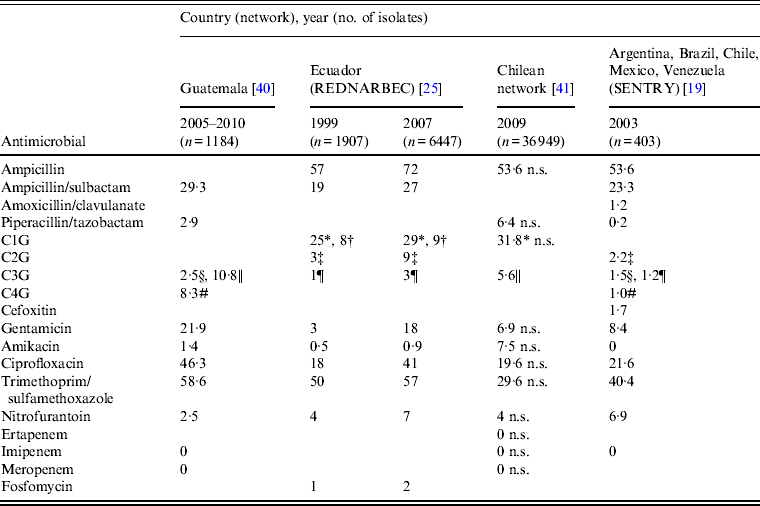
The current rate of clinical failure associated with community-source uropathogens is unacceptably high, coincident with high levels of resistance to commonly used antimicrobials (Table 1, Fig. 2) [Reference Andrade19, Reference Zurita25, Reference Rocha, Tuon and Johnson37–Reference Silva41]. Overall, data from four network studies [Reference Andrade19, Reference Zurita25, Reference Gordillo and Mejía40, Reference Silva41] as well as data from the Pan-American Health Organization (PAHO) surveillance system for 2009 and 2010 [38, 39], indicated that continent-wide resistance of E. coli to trimethoprim/sulfamethoxazole was high, resistance to quinolones was variable, and resistance to second-generation cephalosporins and gentamicin was routinely greater than 20%. In contrast, rates of E. coli resistance to nitrofurantoin and fosfomycin were generally low.

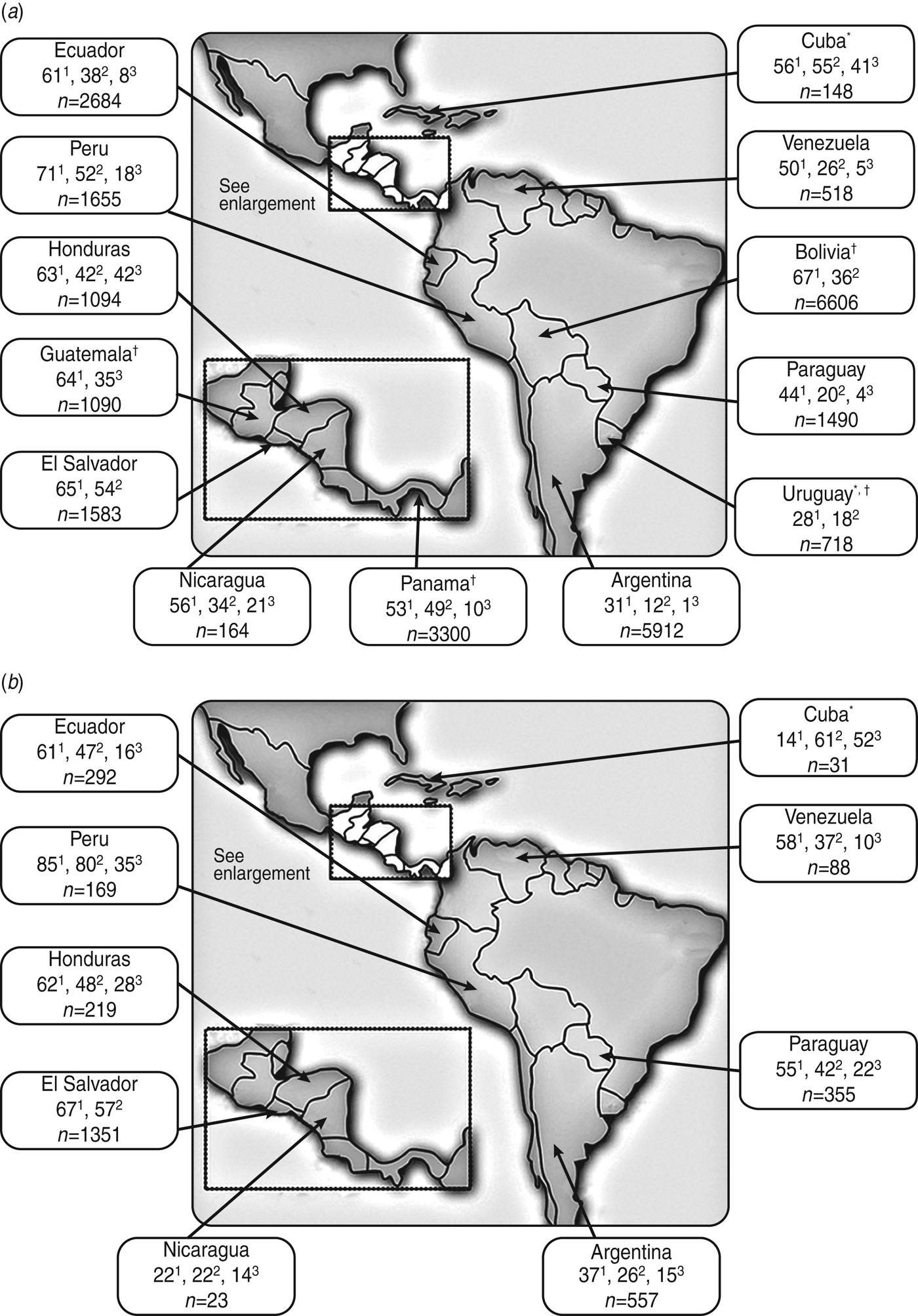
Fig. 2. Percentage (no. of isolates) of urinary tract E. coli isolates collected from (a) women and (b) men in Latin America (2010 PAHO report) that were resistant to trimethoprim/sulfamethoxazole1, ciprofloxacin2, and the second-generation cephalosporin cefuroxime3 [39]. * Data from the 2009 PAHO report [38] is given for this country because 2010 data were not reported; † all adults (women and men).
Table 1. Percentage of drug-resistant community-acquired urinary tract E. coli isolates collected during five surveillance network studies

REDNARBEC, Red Nacional de Resistencia Bacteriana de Ecuador; C1G, First-generation cephalosporins; C2G, second-generation cephalosporins; C3G, third-generation cephalosporins; C4G, fourth-generation cephalosporins; n.s., non-susceptible.
* Cefalotin; † cefazolin; ‡ cefuroxime; § ceftazidime; || cefotaxime; ¶ ceftriaxone; # cefepime.
Rates of ESBL production by community-source isolates have been reported only infrequently in the literature. In the SENTRY 2003 study, which collected data from Argentina, Brazil, Chile, Mexico and Venezuela, rates of ESBL production were 1·7%, 16·3%, and 5·1% in urinary isolates of E. coli, Klebsiella spp., and P. mirabilis, respectively [Reference Andrade19]. The rate of ESBL production in E. coli isolates was 16% in outpatients in Guatemala during the period 2005–2010 [Reference Gordillo and Mejía40].
Central America
The PAHO 2010 data for isolates of community origin reported high resistance rates of E. coli to trimethoprim/sulfamethoxazole and ciprofloxacin throughout Central America (Fig. 2), as well as low resistance rates of E. coli to nitrofurantoin (⩽17%). Resistance of E. coli to cefuroxime ranged from 10% (in Panama) to 42% (females in Honduras); resistance to gentamicin ranged from 4% (in Guatemala) to 28% (males in Honduras) [39]. In the 4 years since PAHO 2006 surveillance data were collected, PAHO 2010 surveillance data indicated that resistance had edged higher to most antimicrobial drug classes in those countries with available data (i.e. El Salvador, Honduras, Nicaragua).
Resistance rates of E. coli isolates collected in a surveillance study in Guatemala in 2005–2010 are shown in Table 1. In Mexican data from the SENTRY 2003 study, antimicrobial susceptibility rates of urinary E. coli isolates were as follows: ampicillin, 22%; nalidixic acid, 25%; levofloxacin, 28%; gatifloxacin, 31%; trimethoprim/sulfamethoxazole, 39%; amoxicillin/clavulanate, 56%; cefuroxime, 64%; and nitrofurantoin, 81% [Reference Andrade19]. Although the nitrofurantoin susceptibility data in SENTRY 2003 data for Mexico were encouraging, increased utilization of this antimicrobial agent as a recommended first-line agent appeared to induce resistance in a study of 304 patients with suspicion of UTI at the university hospital and primary health centres of León, Nicaragua [Reference Bours20]. In the 5 years after the introduction of the therapeutic guidelines, resistance of E. coli against nitrofurantoin increased from 0% to 7% [Reference Bours20]. In this study [Reference Bours20], high resistance rates of E. coli were also observed in 2008 with ampicillin (61%), cefalotin (46%), trimethoprim/sulfamethoxazole (39%), ciprofloxacin (32%), gentamicin (25%), and ceftriaxone (20%). Thirteen (30%) of 44 E. coli strains were suspected of producing ESBLs, with resistance rates in ESBL-producing E. coli significantly higher to ampicillin (85% vs. 52%), amoxicillin/clavulanate (46% vs. 6%), cefalotin (85% vs. 29%), ceftriaxone (69% vs. 0%), and ciprofloxacin (62% vs. 19%) compared to pathogens that did not produce ESBLs [Reference Bours20].
South America
The PAHO 2010 data for isolates of community origin revealed that resistance of E. coli to trimethoprim/sulfamethoxazole ranged from 31% to 85% across South American countries, and resistance to ciprofloxacin ranged from 12% to 80%, with the lowest and highest resistance rates for both antimicrobial agents occurring in Argentina and Peru (Fig. 2) [39]. E. coli resistance to nitrofurantoin was low (i.e. <20%) except in Argentinian males (31%). In contrast, resistance to gentamicin was erratic, ranging from 4% in Uruguay to 29% in Peruvian males [39].
Eight-year temporal data reported in the Red Nacional de Resistencia Bacteriana de Ecuador (REDNARBEC) study are shown in Table 1. Of the South American countries that participated in SENTRY 2003 (Argentina, Brazil, Chile, Venezuela; Table 1), susceptibility rates were generally comparable across countries [Reference Andrade19]. The greatest difference was for the fluoroquinolones, with susceptibility to levofloxacin ranging from 72% in Venezuela to 91% in Brazil [Reference Andrade19].
Results of a large Brazilian study conducted in Curitiba found that few suitable empirical treatment options for community-source UTIs were available for women aged >60 years and males of any age [Reference Rocha, Tuon and Johnson37]. Of 9798 consecutive, non-duplicate, community-source urine isolates from ambulatory patients aged >13 years during 2009, E. coli (66%) was by far the most prevalent Gram-negative pathogen followed by Klebsiella spp. (6%), P. mirabilis (4%), and Enterobacter spp. (3%) [Reference Rocha, Tuon and Johnson37]. Susceptibility of E. coli varied widely by drug class, being very low for ampicillin and trimethoprim/sulfamethoxazole (56% and 66%, respectively), suboptimal for fluoroquinolones (82%), and acceptable for gentamicin, ceftriaxone/cefotaxime and nitrofurantoin (93%, 97% and 96%, respectively) [Reference Rocha, Tuon and Johnson37]. Importantly, the susceptibility rates of E. coli urinary isolates were 3–4% higher for fluoroquinolones and gentamicin, and at least 30% higher for nitrofurantoin and extended-spectrum cephalosporins, compared to susceptibility rates for other pathogens causing community-source urinary infections [Reference Rocha, Tuon and Johnson37].
IAI
The Study for Monitoring Antimicrobial Resistance Trends (SMART) monitors the activity of key antimicrobial drug classes against Gram-negative bacteria isolated from IAIs to ensure that the current susceptibility patterns of these organisms are well understood and reported effectively [Reference Hawser36]. The number of participating sites in Latin America increased from six in 2002 to 13 in 2007, with 740 isolates sent for analysis on average each year [Reference Hawser36]. In 2008, there were 23 centres in ten Latin American countries (Argentina, Brazil, Chile, Colombia, Dominican Republic, Guatemala, Mexico, Panama, Peru, Venezuela) that participated in SMART, with each centre collecting up to 100 consecutive non-duplicate clinical isolates from patients with IAI [Reference Villegas42]. Of 1003 Gram-negative bacilli isolates collected from IAIs, E. coli (50%), K. pneumoniae (15%), and Enterobacter cloacae (7%) were the most commonly isolated pathogens (4% of isolates were P. mirabilis) [Reference Villegas42]. More than one quarter of E. coli (27%) and more than one third of K. pneumoniae (38%) isolates were ESBL-positive. The prevalence of ESBL-producing E. coli and Klebsiella spp. associated with community-acquired infections in particular was 29% [Reference Villegas42].
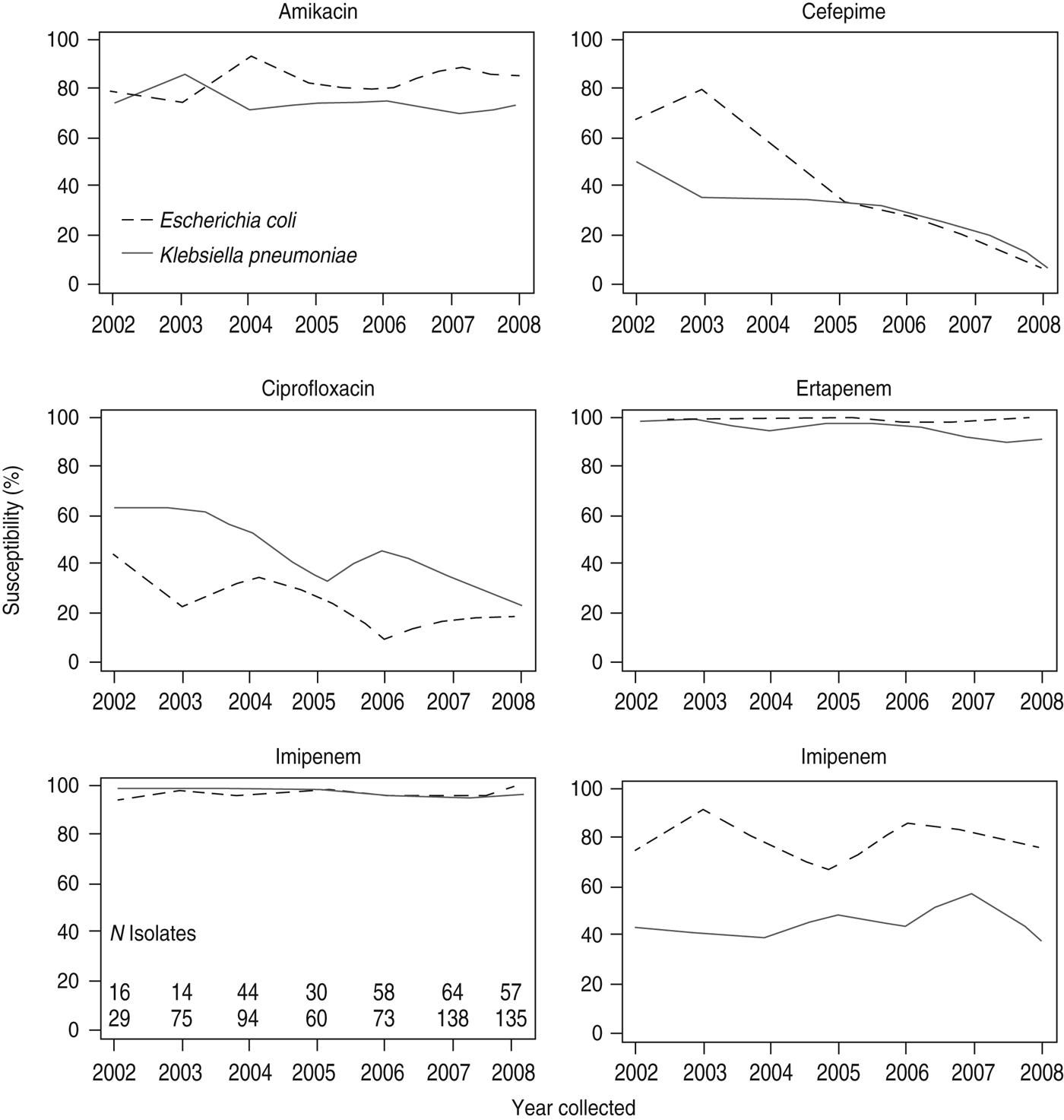
In the SMART study, susceptibilities of ESBL-producing strains to antibacterial agents commonly used in the community such as third-generation cephalosporins, fluoroquinolones, and ampicillin/sulbactam were low (Table 2) [Reference Villegas42]. Six-year temporal data from SMART indicate that the percentages of susceptible ESBL-positive E. coli and K. pneumoniae isolates to ciprofloxacin and cefepime are declining, whereas susceptibilities to amikacin are steady (Fig. 3) [Reference Villegas42].

Fig. 3. Antimicrobial susceptibilities of ESBL-producing E. coli and K. pneumoniae intra-abdominal isolates in Latin America (2002–2008). Susceptibilities are based on in vitro minimum inhibitory concentration data. (Reprinted from Villegas et al. [Reference Villegas42], copyright © 2011, with permission from Elsevier.)
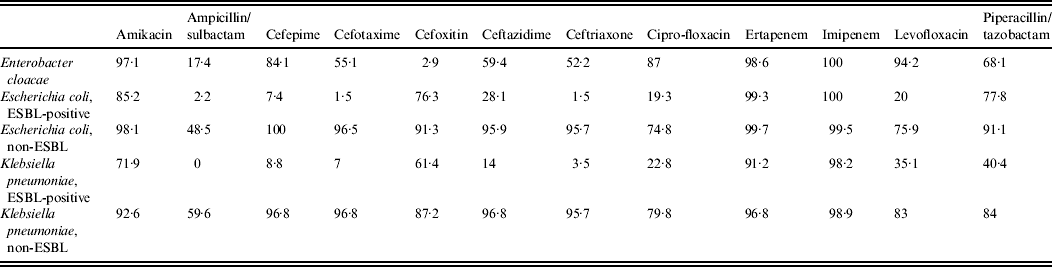
Table 2. Antimicrobial susceptibilities of the most commonly isolated pathogens (>50 isolates) recovered from intra-abdominal infections of Latin American patients participating in SMART, 2008 [Reference Villegas42]

ESBL, Extended-spectrum β-lactamase.
Reprinted from Villegas et al. [Reference Villegas42], copyright © 2011, with permission from Elsevier.
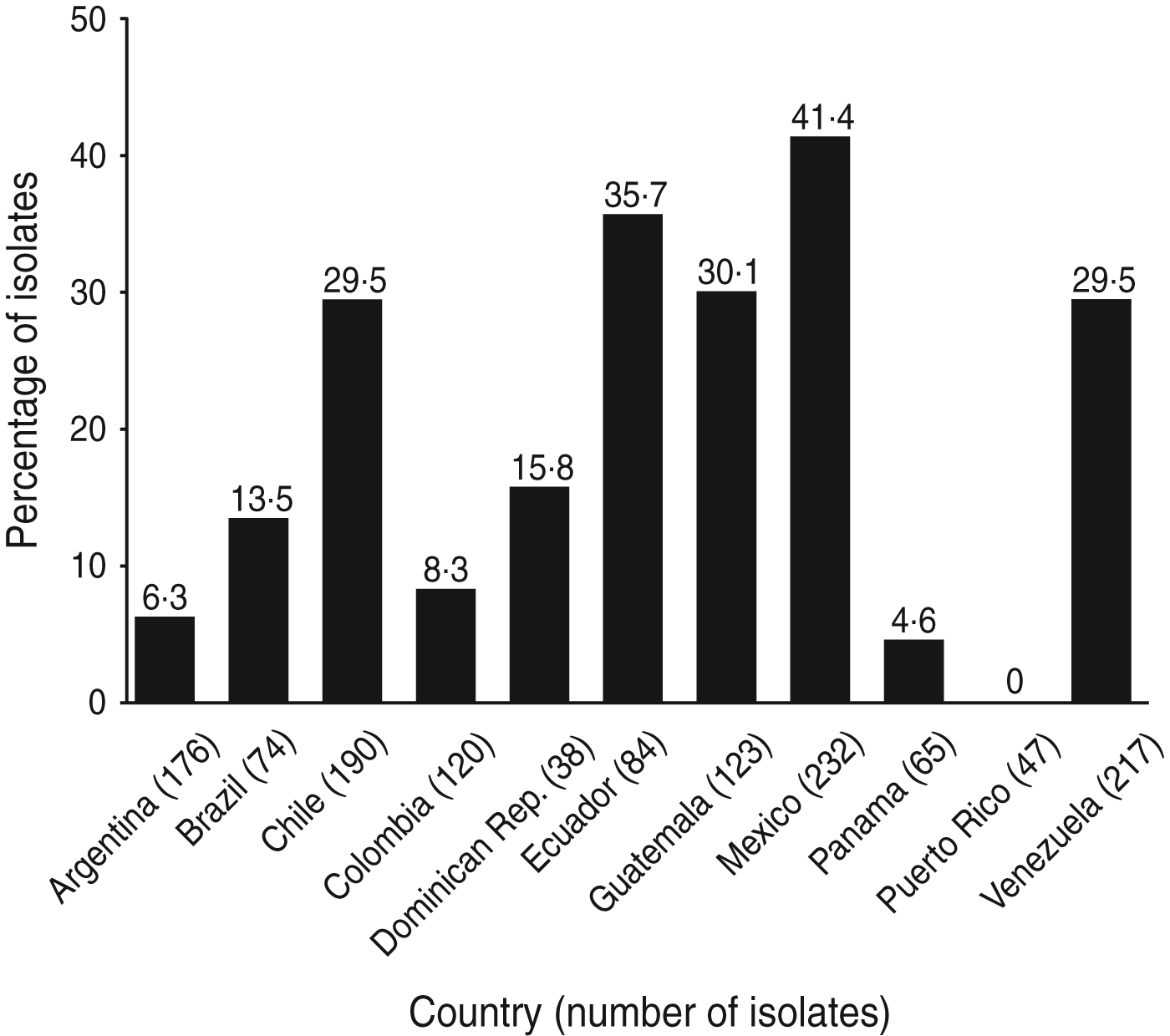
The phenotypes of E. coli isolated from 19 SMART investigator sites in 11 Latin American countries during 2008–2009 were determined [Reference Hawser43]. Of the 1366 isolates, 323 (24%) produced ESBLs, which is an increase from 12% in 2004 and 22% in 2005–2007. The proportion of isolates that were ESBL-producing varied widely in Latin American countries (Fig. 4) [Reference Hawser43]. ESBL production had a major deleterious effect on activity against ampicillin/sulbactam, cephalosporins, and fluoroquinolones, as well as on the activity of amikacin, perhaps indicating a co-resistance phenomenon (Table 3) [Reference Hawser43]. It has been postulated that fluoroquinolone resistance is a harbinger of broader antimicrobial resistance, including ESBL selection [Reference van der Starre44].

Fig. 4. Proportion of E. coli isolates from intra-abdominal infections in Latin America that were extended-spectrum β-lactamase positive (SMART, 2008–2009) [Reference Hawser43].
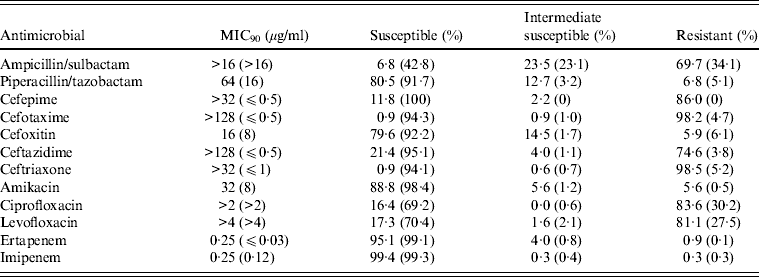
Table 3. Antimicrobial susceptibilities of 323 ESBL-positive intra-abdominal E. coli isolates tested in SMART 2008–2009 based on Clinical and Laboratory Standards Institute breakpoints [Reference Hawser43]

ESBL, Extended-spectrum β-lactamase.
Values in parentheses are the corresponding data pertaining to the 1043 ESBL-negative isolates tested in SMART 2008–2009.
RISK FACTORS
Early recognition of patients at a heightened risk for infection with multidrug-resistant bacteria is necessary to provide appropriate empirical treatment and institute measures that control or limit disease transmission. This observation is especially true in cases of IAI since ampicillin/sulbactam, third-generation cephalosporins, and fluoroquinolones have a limited role as first-line treatments for IAIs. Risk factors for infection with ESBL-producing Enterobacteriaceae have been identified in several studies of hospitalized patients [Reference Cohen-Nahum45–Reference Saely49]. Risk factors identified in multivariate analyses in these studies include previous antibiotic use (specifically, use of quinolones [Reference Cassier47], cephalosporins [Reference Cohen-Nahum45], oxyimino-cephalosporins [Reference Wu46], piperacillin/tazobactam [Reference Cohen-Nahum45], or β-lactams with an oxyimino group [Reference Paterson48]), recurrent infections [Reference Cassier47], haemodialysis [Reference Saely49], urinary catheterization [Reference Wu46], artificial nutrition [Reference Cassier47], and residence in a nursing home [Reference Saely49].
The relevance of risk factors in the hospital setting to the community setting is unclear. According to Pitout & Laupland [Reference Pitout and Laupland26], repeated episodes of UTI and underlying renal pathology, previous use of antibiotics including cephalosporins and quinolones, previous hospitalization, nursing home residence, older age, presence of diabetes mellitus, and underlying liver pathology are risk factors for community-onset infections caused by ESBL-producing bacteria [Reference Pitout and Laupland26]. A meta-analysis of epidemiological studies of infection caused by ESBL-producing Enterobacteriaceae in non-hospitalized patients from six centres in Europe, Asia, and North America revealed that recent use of any antibiotic, residence in a long-term care facility, recent hospitalization, age ⩾65 years, and male sex were statistically significant risk factors for infection by an ESBL-producing organism [Reference Ben-Ami34]. However, it can be argued that emergence of ESBL-producing bacteria, primarily CTX-M-type β-lactamases, confounds infection control strategies based on traditional risk factors, since one third of ESBL-producing isolates in the meta-analysis were obtained from patients with no recent healthcare contact [Reference Ben-Ami34]. Multivariate analysis of a nested case-control study of 787 consecutive patients presenting with febrile UTI in 2004–2009 in The Netherlands revealed recent hospitalization, presence of a urinary catheter, and fluoroquinolone use in the past 6 months to be independent risk factors for infection due to fluoroquinolone-resistant E. coli [Reference van der Starre44]. In summary, risk factors for infection by ESBL-producing bacteria in the community setting are similar to those in the hospital setting and are fairly consistent across studies.
SURVEILLANCE AND DETECTION
With the advent of widespread dispersion of ESBL-producing Enterobacteriaceae in communities of Latin America, empirical prescribing of antimicrobial agents according to general principles can no longer ensure effective treatment. Rather, local susceptibility patterns attained within each institution and maintenance of longitudinal surveillance programmes are needed to inform treatment decisions. Surveillance also aids patient diagnosis and facilitates infection control strategies. Despite an investment to survey bacterial resistance in Latin America through PAHO, SENTRY, SMART and other programmes, further efforts are required to more fully integrate these resources in a practical way so that real-time data regarding antimicrobial susceptibility and genotypic patterns are available to healthcare professionals [Reference O'Brien and Stelling28].
Aside from these overarching logistical issues, there are also major limitations in our capability to detect ESBL-producing Enterobacteriaceae. These strains confound traditional surveillance strategies because ESBL production cannot be inferred from the antimicrobial resistance profile alone. Hence, the emergence, maintenance, and dissemination of ESBLs must be characterized and closely monitored by implementation of integrated phenotypic and genotypic testing strategies. To give some scale to the magnitude of the task ahead, the number of CTX-M variants alone numbered 138 as of March 2013 [50].
Phenotypic methods
Most clinical microbiology laboratories in Latin America conduct susceptibility testing as it is easy, automated, inexpensive, and accessible; however, phenotypic methods cannot provide information on the type of ESBL produced. The value of non-molecular phenotypic methods is premised on the fact that most ESBLs hydrolyse third-generation cephalosporins and are inhibited by clavulanate [Reference Pitout and Laupland26]. Numerous methods have been developed to detect or confirm ESBL production by Enterobacteriaceae. Automated systems screen isolates and confirm ESBL production based on minimum inhibitory concentrations of cefotaxime and ceftazidime with and without clavulanic acid.
However, the most appropriate choice of extended-spectrum cephalosporins and type of confirmatory test that provides the greatest assay sensitivity have yet to be determined. For instance, in early testing, the E-test ESBL screen test with ceftazidime or cefepime as substrate, and use of Oxoid combination discs with cefotaxime and ceftazidime, were useful for demonstrating the presence of Enterobacteriaceae potentially producing ESBLs but were also associated with clinically significant false-susceptible and false-resistant results [Reference Mendes51, Reference Cormican, Marshall and Jones52]. Current guidance from the Clinical and Laboratory Standards Institute (CLSI) describes laboratory detection of ESBL produced by E. coli, P. mirabilis, and Klebsiella spp., but not for species with inducible AmpC β-lactamases (such as Enterobacter spp.) [53]. Specifically, for ESBL detection in Enterobacteriaceae, CLSI recommends initial screening with 8 μg/ml of cefpodoxime; 1 μg/ml each of cefotaxime, ceftazidime, ceftriaxone, or aztreonam; followed by confirmatory tests (including the E-test ESBL strips) with both cefotaxime and ceftazidime in combination with clavulanate at a concentration of 4 μg/ml [54]. As of 2007, high sensitivities of up to 94% and specificity of 98% for detecting ESBLs in E. coli, Klebsiella spp, and Proteus spp. were expected if these techniques were adhered to [Reference Wiegand55].
Recent emergence of plasmidic AmpC β-lactamases harboured by E. coli and K. pneumoniae may result in changes to CLSI recommendations. The phenotypic detection of ESBLs in bacteria other than E. coli, Klebsiella spp., and Proteus spp. will remain problematical because of the possible association of resistance mechanisms, presence of more than one β-lactamase [especially carbapenemases of the K. pneumoniae carbapenemase (KPC) type], and reduced permeability.
Genotypic methods
Genotypic methods use molecular biology techniques to detect the gene responsible for ESBL production, with the aim of distinguishing between resistance genes, genetic elements, and strains. The primary technology used is polymerase chain reaction (PCR) amplification of bla TEM, bla SHV, and blaCTX-M genes with oligonucleotide primers followed by sequencing to discriminate between non-ESBL parent enzymes and their ESBL variants [Reference Pitout and Laupland26]. ESBLs have also been characterized by PCR restriction fragment-length polymorphism and single-strand conformational polymorphism, and by restriction site insertion PCR, real-time PCR, and ligase chain reaction [Reference Pitout and Laupland26]. A limitation of PCR is that it detects ESBL genes but does not inform on ESBL production. In addition, few clinical microbiology laboratories in Latin America are equipped to perform recombinant DNA techniques, which are complicated, time-consuming, and expensive in part because of the presence of multiple copies of ESBLs in any given clinical isolate.
Whole-genome sequencing and multilocus sequence typing has tremendous utility, as it enables microbiologists to characterize the population biology of a species, and thus track the evolution and spread of clones. Currently, these techniques are confined to reference laboratories and have little influence on local and regional infection control programmes.
Clonality in Latin America
The CTX-M family includes a heterogeneous group of ESBLs divided into five groups based on primary structure (CTX-M-1, CTX-M-2, CTX-M-8, CTX-M-9, CTX-M-25) [Reference Pitout and Laupland26]. Although CTX-Ms are produced by a wide variety of Enterobacteriaceae strains, CTX-M globalization is associated with a few clones of E. coli and K. pneumoniae, which underscores the selective advantage of expressing these enzymes. Most clinical isolates are not clonally related, but clonal outbreaks have been described in several countries [Reference Pitout and Laupland26, Reference Manges56]. It has been estimated that at least 10–20% of all UTIs are caused by clonally related E. coli, which are often co-resistant to aminoglycosides and trimethoprim/sulfamethoxazole [Reference Cantón, González-Alba and Galán33, Reference Manges56]. Detecting successful strains or epidemic clones from large volumes of isolates with the same phenotype is challenging [Reference O'Brien and Stelling28].
E. coli belonging to phylogroup B2 [sequence type 131 (ST131)] and phylogroup D (ST405) are the most infamous community-associated, high-risk clones because of their rapid globalization coupled with high virulence and multidrug-resistant IncFII plasmids [Reference Rogers, Sidjabat and Paterson15, Reference Coque57]. E. coli ST131 has been implicated in severe community-acquired infections, including septicaemia [Reference Pitout and Laupland26, Reference Pitout58]. In Latin America, E. coli ST131 was initially detected in the Colombian and Brazilian hospital setting in 2008 [Reference Leal59, Reference Peirano60] and subsequently in the Colombian community setting in 2010 (along with clone ST405) and possibly in 2011 [Reference Ruiz61, Reference Martinez, Garzón and Mattar62]. The ease with which E. coli ST131 diffuses through communities via infected or colonized family members, wildlife, foodstuffs, and companions represents a major public health concern [Reference Rogers, Sidjabat and Paterson15].
The composition of a different high-risk clone of E. coli causing community-acquired UTIs in Rio de Janeiro, Brazil, between 2005 and 2006 was determined by a cross-sectional study of 344 women seeking care in one public walk-in clinic [Reference Dias63]. More than half (54%) of the women had a documented UTI, of which 63% were caused by E. coli. Of the 50% of E. coli isolates resistant to ampicillin and trimethoprim/sulfamethoxazole, most (81%) belonged to 19 enterobacterial repetitive intergenic consensus (ERIC2) clonal groups. All isolates in the largest clonal group (n = 15 isolates) belonged to multilocus sequence typing group ST69 and phylogenetic group D, and had 89% similarity to a clonal group A (CgA) reference strain from the USA [Reference Dias63]. These data indicate that uropathogenic E. coli CgA strains have mobilized from North America and Europe to Latin America, where they have the potential to cause multidrug-resistant outbreaks.
Brazil has a particularly high diversity of CTX-M enzymes harboured by clinical isolates of K. pneumoniae and E. coli. In a community and hospital setting in Rio de Janeiro (2000–2001), analysis of the epidemiological features of 41 E. coli isolates resistant to third-generation cephalosporins and/or non-β-lactam antibiotics revealed a high prevalence of CTX-M-2 production [Reference Queiroz64]. CTX-M-9 and CTX-M-59 (a variant of CTX-M-2) were also identified. Of note, the CTX-M-producing E. coli in this study belonged to different phylogroups/sequence types that were associated with IncA/C plasmids implicated in the facilitation of CTX-M globalization and evolution [Reference Queiroz64].
The first report of a Citrobacter freundii-producing CTX-M-14 isolated from a woman in Venezuela with a recurrent UTI highlights the problem of plasmid-mediated dissemination of these β-lactamases into other bacterial populations [Reference Millán65].
PROPOSALS FOR LATIN AMERICA
When reviewing the data, we recognize that better-designed studies are urgently required to more accurately quantify the epidemiology of Gram-negative community-associated infections in Latin America. Rather than use local criteria, these studies should report on a standardized definition for community-associated UTIs and IAIs and use approved methods for phenotypic and genotypic testing. Furthermore, local and national susceptibility data are required on a far greater number of isolates before treatment algorithms can be developed.
Even taking into consideration the gaps in our knowledge regarding the epidemiology of community-associated UTIs and IAIs by Gram-negative bacteria, the weight of evidence suggests that primary-care physicians in Latin America should consider the potential for involvement of multidrug-resistant bacteria when managing cases of UTI and IAI. Published guidelines (including recent guidelines of the Infectious Diseases Society of America for treatment of acute uncomplicated cystitis [Reference Gupta66]) can be consulted for general guidance, but local conditions, including availability of specific agents, will influence treatment. (Fosfomycin, for example, may be used in the correct formulation for treatment of UTI in women.) Severity of infection as well as risk factors for infection by multidrug-resistant Enterobacteriaceae can be used to inform treatment decisions. If available, knowledge of local antimicrobial susceptibility data should be used as a reference guide. However, we recommend that urine cultures be collected for all recurrent or relapsing UTIs, complicated UTIs, and UTI cases presenting to the emergency room. This assertion is supported by the high levels of resistance of urinary E. coli isolates to trimethoprim/sulfamethoxazole and quinolones across the continent, rendering prescribing decisions difficult without supportive microbiological data [Reference Andrade19, 39]. Urine cultures are inexpensive to perform and are readily available in most hospitals of Latin America. Furthermore, obtaining cultures is necessary to implement antimicrobial stewardship programmes, in which de-escalation is a very important method to decrease selective pressure on broad-spectrum antibiotics. Primary-care physicians, including gynaecologists, should also be educated that asymptomatic patients who receive a positive test for bacteria growth in urine should not be treated with antibiotics, as the positive test usually indicates colonization and not infection.
CONCLUSION
Overall, data describing the microbiology of community-associated UTIs and IAIs caused by Gram-negative bacteria in Latin America over the last 10 years indicate high rates of continent-wide resistance to trimethoprim/sulfamethoxazole, quinolones, second-generation cephalosporins, and gentamicin, and low levels of resistance to third- and fourth-generation cephalosporins, nitrofurantoin, and fosfomycin. We report E. coli resistance rates to quinolones routinely greater than 20% (and up to 80%), which is higher than the 2009 national average in the USA (19·5%) [67]. Widespread use of quinolones in humans and in animal husbandry in Latin America may account for this difference. The concern around the endemicity of quinolone-resistant E. coli in Latin America is the association with plasmid-mediated quinolone resistance, which accelerates the rate at which other Enterobacteriaceae develop fluoroquinolone resistance. The extremely high rate of E. coli resistance to trimethoprim/sulfamethoxazole probably reflects widespread use and misuse of this low-cost antimicrobial agent for treatment of community-associated UTI in the region. Findings from the publications assessed in this review tentatively indicate that ESBL rates in E. coli-causing IAIs are variable but increasing over time, although not enough data are available to confirm the seriousness of this problem [Reference Hawser43]. The rate of ESBLs harboured by urinary isolates of E. coli requires further study, given the ease of CTX-M mobilization, high potential for clonal outbreaks, and increase in reports of UTIs by ceftriaxone-resistant E. coli.
Given the high resistance rates in Enterobacteriaceae causing community-acquired UTIs and IAIs and lack of therapeutic options, we recommend that antimicrobial prescribing be guided by considering infection severity, established patient risk factors for multidrug-resistant infections, acquaintance with local antimicrobial susceptibility data, and culture collection.
APPENDIX: Latin America Working Group on Bacterial Resistance
Carlos Alvarez (Hospital Universitario San Ignacio and Pontificia Universidad Javeriana, Bogotá, Colombia), Luis Bavestrello (Clinica Reñaca, Viña Del Mar, Chile), Eitan Berezin (Santa Casa de São Paulo School of Medicine, Brazil), Eduardo Gotuzzo (Universidad Peruana Cayetano Heredia, Lima, Perú), Manuel Guzmán-Blanco (Hospital Privado Centro Médico de Caracas, Venezuela), Jaime A. Labarca (Pontificia Universidad Católica de Chile, Santiago, Chile), Carlos M. Luna (Hospital de Clínicas José de San Martin Hospital, Universidad de Buenos Aires, Argentina), Carlos Mejía (Hospital Roosevelt, Guatemala City, Guatemala), Simone Nouer (Hospital Universitario Clementino Fraga Filho, Rio de Janeiro, Brazil), Eduardo Rodríguez-Noriega (Hospital Civil de Guadalajara Fray Antonio Alcalde, Guadalajara, Mexico), Mauro José Costa Salles (Hospital Irmandade da Santa Casa de Misericórdia de São Paulo, Brazil), Carlos Seas (Universidad Peruana Cayetano Heredia, Lima, Perú), Fortino Solórzano Santos (Hospital de Pediatría Centro Médico Nacional Siglo XXI, Mexico City, Mexico), Maria Virginia Villegas [International Center for Medical Research and Training (CIDEIM), Cali, Colombia], Jeannete Zurita (Hospital Vozandes and Pontificia Universidad Católica del Ecuador, Quito, Ecuador).
ACKNOWLEDGEMENTS
This publication was funded by Pfizer Inc. Medical writing support was provided by Malcolm Darkes and Lisa Baker of UBC Scientific Solutions and was funded by Pfizer Inc.
DECLARATION OF INTEREST
Dr Mauro José Costa Salles is an advisory board member for Pfizer and Novartis; a consultant or speaker for Pfizer, Merck Sharp & Dohme, Novartis and United Medicals; and a recipient of research grants from Pfizer and Novartis. Dr Jeannete Zurita is an Advisory Board member and consultant for Pfizer, and received research grants from Wyeth (now part of Pfizer) and from Merck. Dr Carlos Mejía participated in advisory groups for HIV treatment in association with Abbott, GlaxoSmithKline, and Stendhal. He participated in the SMART study, which was financially supported by Merck, and is a member of an advisory group for Pfizer. Dr Maria Virginia Villegas has received consulting fees and research grants from Merck Sharp & Dohme, Pfizer SA, Janssen-Cilag SA, Novartis, and AstraZeneca Colombia SA.