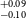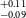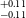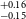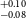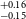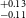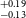2 results
VaTEST III: Validation of eight potential super-earths from TESS data
- Priyashkumar Mistry, Aniket Prasad, Mousam Maity, Kamlesh Pathak, Sarvesh Gharat, Georgios Lekkas, Surendra Bhattarai, Dhruv Kumar, Jack J. Lissauer, Joseph D. Twicken, Abderahmane Soubkiou, Francisco J. Pozuelos, Jon Jenkins, Keith Horne, Steven Giacalone, Khalid Barkaoui, Mathilde Timmermans, Cristilyn N. Watkins, Ramotholo Sefako, Karen A. Collins, David R. Ciardi, Catherine A. Clark, Boris S. Safonov, Avi Shporer, Joshua E. Schlieder, Zouhair Benkhaldoun, Chris Stockdale, Carl Ziegler, Emily A. Gilbert, Jehin Emmanuël, Felipe Murgas, Ian J. M. Crossfield, Martin Paegert, Michael B. Lund, Norio Narita, Richard P. Schwarz, Robert F. Goeke, Sergio B. Fajardo-Acosta, Steve B. Howell, Thiam-Guan Tan, Thomas Barclay, Yugo Kawai
-
- Journal:
- Publications of the Astronomical Society of Australia / Volume 41 / 2024
- Published online by Cambridge University Press:
- 11 April 2024, e030
-
- Article
- Export citation
-
NASA’s all-sky survey mission, the Transiting Exoplanet Survey Satellite (TESS), is specifically engineered to detect exoplanets that transit bright stars. Thus far, TESS has successfully identified approximately 400 transiting exoplanets, in addition to roughly 6 000 candidate exoplanets pending confirmation. In this study, we present the results of our ongoing project, the Validation of Transiting Exoplanets using Statistical Tools (VaTEST). Our dedicated effort is focused on the confirmation and characterisation of new exoplanets through the application of statistical validation tools. Through a combination of ground-based telescope data, high-resolution imaging, and the utilisation of the statistical validation tool known as TRICERATOPS, we have successfully discovered eight potential super-Earths. These planets bear the designations: TOI-238b (1.61
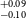 $^{+0.09} _{-0.10}$ R
$^{+0.09} _{-0.10}$ R $_\oplus$), TOI-771b (1.42
$_\oplus$), TOI-771b (1.42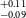 $^{+0.11} _{-0.09}$ R
$^{+0.11} _{-0.09}$ R $_\oplus$), TOI-871b (1.66
$_\oplus$), TOI-871b (1.66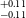 $^{+0.11} _{-0.11}$ R
$^{+0.11} _{-0.11}$ R $_\oplus$), TOI-1467b (1.83
$_\oplus$), TOI-1467b (1.83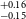 $^{+0.16} _{-0.15}$ R
$^{+0.16} _{-0.15}$ R $_\oplus$), TOI-1739b (1.69
$_\oplus$), TOI-1739b (1.69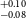 $^{+0.10} _{-0.08}$ R
$^{+0.10} _{-0.08}$ R $_\oplus$), TOI-2068b (1.82
$_\oplus$), TOI-2068b (1.82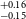 $^{+0.16} _{-0.15}$ R
$^{+0.16} _{-0.15}$ R $_\oplus$), TOI-4559b (1.42
$_\oplus$), TOI-4559b (1.42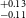 $^{+0.13} _{-0.11}$ R
$^{+0.13} _{-0.11}$ R $_\oplus$), and TOI-5799b (1.62
$_\oplus$), and TOI-5799b (1.62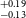 $^{+0.19} _{-0.13}$ R
$^{+0.19} _{-0.13}$ R $_\oplus$). Among all these planets, six of them fall within the region known as ‘keystone planets’, which makes them particularly interesting for study. Based on the location of TOI-771b and TOI-4559b below the radius valley we characterised them as likely super-Earths, though radial velocity mass measurements for these planets will provide more details about their characterisation. It is noteworthy that planets within the size range investigated herein are absent from our own solar system, making their study crucial for gaining insights into the evolutionary stages between Earth and Neptune.
$_\oplus$). Among all these planets, six of them fall within the region known as ‘keystone planets’, which makes them particularly interesting for study. Based on the location of TOI-771b and TOI-4559b below the radius valley we characterised them as likely super-Earths, though radial velocity mass measurements for these planets will provide more details about their characterisation. It is noteworthy that planets within the size range investigated herein are absent from our own solar system, making their study crucial for gaining insights into the evolutionary stages between Earth and Neptune.
New Single Molecule Approaches to Genomic Analysis: Optical Mapping
- Schwartz. D.C, Anantharaman T, Cai W, Clarke V, Delobette S, Dimalanta E, Edington J, Giacalone J, Hiort C, Hu X, Huff E, Jing J, Lai Z, Lee E, Mishra B, Murti J.R, Porter B., Qi R, Rabbah R, Ramanathan A, Reed J, Samad A, Shenker A, Skiadas Y, Tankhoveya K, Wang W, Lin J
-
- Journal:
- Microscopy and Microanalysis / Volume 3 / Issue S2 / August 1997
- Published online by Cambridge University Press:
- 02 July 2020, pp. 199-200
- Print publication:
- August 1997
-
- Article
- Export citation
-
Current molecular biological approaches were developed primarily for characterization of single genes, not entire genomes, and, as such, are not ideally suited to analysis of complex traits and population-based molecular genetics. Despite rapid progress in the human genome project effort, there is little doubt that radically new conceptual approaches are needed before routine whole genome-based analyses can be undertaken by both basic research and clinical laboratories.
Physical mapping of genomes, using restriction endonucleases, has played a major role in the identification and characterizing various loci, for example, by aiding clone contig formation and by characterizing genetic lesions. Restriction maps provide precise genomic distances, unlike ordered sequence-based landmarks such as Sequence Tagged Sites (STSs), that are essential for optimizing the efficiency of sequencing efforts, and for determining the spatial relationships of specific loci. When compared to tedious hybridization-based fingerprinting approaches, ordered restriction maps offer relatively unambiguous clone characterization that is useful in contig formation, establishment of minimal tiling paths for sequencing, and preliminary characterization of sequence lesions.