7 results
Characterization of carbapenem-resistant gram-negative bacteria collected in the Sentinel Surveillance Program, 2018–2019
- Lori Spicer, Davina Campbell, J. Kristie Johnson, Cynthia Longo, Thomas Balbuena, Thomas Ewing, Maria Karlsson, J. Kamile Rasheed, Christopher Elkins, Amy Gargis, Joseph Lutgring
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 2 / Issue S1 / July 2022
- Published online by Cambridge University Press:
- 16 May 2022, p. s52
-
- Article
-
- You have access Access
- Open access
- Export citation
-
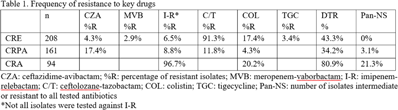
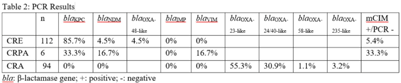
Background: Carbapenem resistance in gram-negative organisms is an important public health problem. The CDC conducted Sentinel surveillance in 2018–2019 to characterize these organisms from 9 facilities in 9 different states. Methods: Carbapenem-resistant Enterobacterales (CRE), Pseudomonas aeruginosa (CRPA), and Acinetobacter spp (CRA) obtained from clinical samples of patients in acute-care or long-term care facilities were submitted to the CDC. Identification was confirmed using matrix-assisted laser desorption ionization time-of-flight (MALDI-TOF), and antimicrobial susceptibility testing (AST) was performed via broth microdilution for 27 antibiotics. All confirmed CRE and CRPA were tested for carbapenemase production (CP) using the modified carbapenem inactivation method (mCIM). The isolates that were mCIM-positive were assessed by real-time PCR for presence of blaKPC, blaNDM, blaVIM, and blaIMP. CP-CRE were also assessed for blaOXA-48-like. All confirmed CRA were tested for the same genes as CRPA and blaOXA-23–like, blaOXA-24/40-like, blaOXA-58–like, and blaOXA-235–like genes. Difficult-to-treat resistance (DTR) was defined as resistance to all β-lactams (excluding newer β-lactam combination agents) and quinolones tested. Results: The CDC confirmed 208 CRE, 161 CRPA, and 94 CRA. Table 1 summarizes AST results for a selection of drugs. We identified 112 (53.8%) mCIM-positive CRE and 6 (3.7%) mCIM-positive CRPA. The PCR results are summarized in Table 2. One mCIM-positive and PCR-negative isolate was positive in a metallo-β-lactamase screen. Conclusions: Resistance among CRE and CRPA to newer β-lactam combination agents was detected. Options for treating CRA are limited. Of 112 CP-CRE, 85.7% harbored blaKPC; CP-CRPA were rare (3.7%); and most CRA harbored blaOXA-23-like (55.3%) or blaOXA-24/40-like (30.9%). Whole-genome sequencing is planned to better understand gene variants, sequence types, and additional resistance markers present among the isolates.
Funding: None
Disclosures: None
Epidemiology of extended-spectrum β-lactamase–producing Enterobacterales in five US sites participating in the Emerging Infections Program, 2017
- Nadezhda Duffy, Maria Karlsson, Hannah E. Reses, Davina Campbell, Jonathan Daniels, Richard A. Stanton, Sarah J. Janelle, Kyle Schutz, Wendy Bamberg, Paulina A. Rebolledo, Chris Bower, Rebekah Blakney, Jesse T. Jacob, Erin C. Phipps, Kristina G. Flores, Ghinwa Dumyati, Hannah Kopin, Rebecca Tsay, Marion A. Kainer, Daniel Muleta, Benji Byrd-Warner, Julian E. Grass, Joseph D. Lutgring, J. Kamile Rasheed, Christopher A. Elkins, Shelley S. Magill, Isaac See
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 43 / Issue 11 / November 2022
- Published online by Cambridge University Press:
- 14 February 2022, pp. 1586-1594
- Print publication:
- November 2022
-
- Article
- Export citation
-
Objective
The incidence of infections from extended-spectrum β-lactamase (ESBL)–producing Enterobacterales (ESBL-E) is increasing in the United States. We describe the epidemiology of ESBL-E at 5 Emerging Infections Program (EIP) sites.
MethodsDuring October–December 2017, we piloted active laboratory- and population-based (New York, New Mexico, Tennessee) or sentinel (Colorado, Georgia) ESBL-E surveillance. An incident case was the first isolation from normally sterile body sites or urine of Escherichia coli or Klebsiella pneumoniae/oxytoca resistant to ≥1 extended-spectrum cephalosporin and nonresistant to all carbapenems tested at a clinical laboratory from a surveillance area resident in a 30-day period. Demographic and clinical data were obtained from medical records. The Centers for Disease Control and Prevention (CDC) performed reference antimicrobial susceptibility testing and whole-genome sequencing on a convenience sample of case isolates.
ResultsWe identified 884 incident cases. The estimated annual incidence in sites conducting population-based surveillance was 199.7 per 100,000 population. Overall, 800 isolates (96%) were from urine, and 790 (89%) were E. coli. Also, 393 cases (47%) were community-associated. Among 136 isolates (15%) tested at the CDC, 122 (90%) met the surveillance definition phenotype; 114 (93%) of 122 were shown to be ESBL producers by clavulanate testing. In total, 111 (97%) of confirmed ESBL producers harbored a blaCTX-M gene. Among ESBL-producing E. coli isolates, 52 (54%) were ST131; 44% of these cases were community associated.
ConclusionsThe burden of ESBL-E was high across surveillance sites, with nearly half of cases acquired in the community. EIP has implemented ongoing ESBL-E surveillance to inform prevention efforts, particularly in the community and to watch for the emergence of new ESBL-E strains.
New Approaches to Colonization Screening in Response to Emerging Antimicrobial Resistance
- Karen Anderson, Maria Karlsson, Sandra Boyd, Natashia Reese, Uzma Ansari, Davina Campbell, Amelia Bhatnagar, Paige Gable, Stephanie Swint, Cynthia Longo, Sarah Gilbert, Lori Spicer, Jake Cochran, David Lonsway
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, p. s330
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: The capacity to monitor the emergence of carbapenemase-producing organisms (CPO) is critical in limiting transmission. CPO-colonized patients can be identified by screening rectal specimens for carbapenemase genes and the Cepheid GeneXpert Carba-R (XCR), the only FDA-approved test, is limited to 5 carbapenemase genes and cannot identify the bacterial species. Objective: We describe the development and validation of culture-based methods for the detection of CPO in rectal cultures (RCs) and nonrectal cultures (NRCs) of tracheal aspirate and axilla-groin swabs. Methods: Colonization screening was performed at 3 US healthcare facilities; specimens of RC swabs and NRC ESwabs were collected. Each specimen was inoculated to a MacConkey broth enrichment tube for overnight incubation then were subcultured to MacConkey agar with meropenem and ertapenem 10 µg disks (BEMA) and CHROMagar KPC (KCHR) or CHROMagar Acinetobacter (ACHR). All media were evaluated for the presence of carbapenem-resistant organisms; suspect colonies were screened by real-time PCR for the most common carbapenemase genes. MALDI-TOF was performed for species identification. BEMA, a previously validated method, was the comparator for 52 RCs; clinical culture (CC) served as the comparator method for 66 NRCs. Select CPO-positive and -negative specimens underwent reproducibility testing. Results: Among 56 patients undergoing colonization screening, 12 (21%) carried a CPO. Only 1 patient had CPO solely from RC. Also, 6 patients had both CPO-positive RC and NRC, and 5 patients only had a CPO-positive NRC. Of the latter, 4 had a CPO-positive tracheal specimen, and 1 had a positive culture from both tracheal and axilla-groin specimens. Sensitivity of BEMA (70%) for NRC was lower than for KCHR (96%) and ACHR (88 %) for all specimens. All methods showed a specificity of 100% and reproducibility of 92%. The detected CPO included OXA-23–positive Acinetobacter baumannii, NDM-positive Escherichia coli, KPC-positive Pseudomonas aeruginosa and 4 genera of KPC-positive Enterobacteriaceae. Conclusions:The addition of nonrectal specimens and use of selective media contributed to increased sensitivity and enhanced identification of CPO-colonized patients. Positive cultures were equally distributed among the 3 specimen types. The addition of the nonrectal specimens resulted in the identification of more colonized patients. The culture-based method was successful in detecting an array of different CPOs and target genes, including genes not detected by the Carba-R assay (eg, blaOXA-23-like). Enhanced isolation and characterization of CPOs will be key in aiding epidemiologic investigations and strengthening targeted guidance for containment strategies.
Funding: None
Disclosures: We discuss the drug combination aztreonam-avibactam and acknowledge that this drug combination is not currently FDA approved.
Molecular Typing of Invasive Staphylococcus aureus from the Emerging Infections Program (EIP) Using Whole-Genome Sequencing
- Davina Campbell, Gillian McAllister, Kelly Jackson, Isaac See, Alison Halpin, Joseph Lutgring, Erin Epson, Susan Petit, Susan Ray, William Schaffner, Ghinwa Dumyati, Thomas Ewing, Michelle Adamczyk, Amy Gargis
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s71-s72
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: The CDC has performed surveillance for invasive Staphylococcus aureus (iSA) infections through the Emerging Infections Program (EIP) since 2004. SCCmec and spa typing for clonal complex (CC) assignment and genomic markers have been used to characterize isolates. In 2019, whole-genome sequencing (WGS) of isolates began, allowing for high-resolution assessment of genomic diversity. Here, we evaluate the reliability of SCCmec typing, spa typing, and CC assignment using WGS data compared to traditional methods to ensure that backwards compatibility is maintained. Methods:S. aureus isolates were obtained from a convenience sample of iSA cases reported through the EIP surveillance system. Overall, 78 iSA isolates with diverse spa repeat patterns, CCs, SCCmec types, and antimicrobial susceptibility profiles were sequenced (MiSeq, Illumina). Real-time PCR and Sanger sequencing were used as the SCCmec and spa typing reference methods, respectively. spa-MLST mapping (Ridom SpaServer) served as the reference method for CC assignment. WGS assembly and multilocus sequence typing (MLST) were performed using the CDC QuAISAR-H pipeline. WGS-based MLST CCs were assigned using eBURST and SCCmec types using SCCmecFinder. spa types were assigned from WGS assemblies using BioNumerics. For isolate subtyping, previously published and validated canonical single-nucleotide polymorphisms (canSNPs) as well as the presence of the Panton-Valentine leukocidin (PVL) toxin and arginine catabolic mobile element (ACME) virulence factor were assessed for all genome assemblies. Results: All isolates were assigned WGS-based spa types, which were 100% concordant (78 of 78) with Sanger-based spa typing. SCCmecFinder assigned 91% of isolates (71 of 78) SCCmec types, which were 100% concordant with reference method results. Also, 7 isolates had multiple cassettes predicted or an incomplete SCCmec region assembly. Using WGS data, 96% (75 of 78) of isolates were assigned CCs; 3 isolates had unknown sequence types that were single-locus variants of established sequence types. Overall, 70 isolates had CCs assigned by the reference method; 100% (70 of 70) concordance was observed with WGS-based CCs. Analysis of canSNPs placed 42% (33 of 78) of isolates into CC8, with 17 (52%) of these isolates classified as USA300. PVL and ACME were not accurate markers for inferring the USA300 subtype as 24% (4 of 17) of isolates did not contain these markers. Conclusions:S. aureus CCs, SCCmec, and spa types can be reliably determined using WGS. Incorporation of canSNP analysis represents a more efficient method for CC8 assignment than the use of genomic markers alone. WGS allows for the replacement of multiple typing methods for increased laboratory efficiency, while maintaining backward compatibility with historical typing nomenclature.
Funding: None
Disclosures: None
Whole-Genome Sequencing Reveals Diversity of Carbapenem-Resistant Pseudomonas aeruginosa Collected Through the Emerging Infections Program
- Richard Stanton, Jonathan Daniels, Erin Breaker, Davina Campbell, Joseph Lutgring, Maria Karlsson, Kyle Schutz, Jesse Jacob, Lucy Wilson, Elisabeth Vaeth, Linda Li, Ruth Lynfield, Erin C. Phipps, Emily Hancock, Ghinwa Dumyati, Rebecca Tsay, P. Maureen Cassidy, Jacquelyn Mounsey, Julian Grass, Maroya Walters, Alison Halpin
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s513-s514
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: Carbapenem-resistant Pseudomonas aeruginosa (CRPA) is a frequent cause of healthcare-associated infections (HAIs). The CDC Emerging Infections Program (EIP) conducted population and laboratory-based surveillance of CRPA in selected areas in 8 states from August 1, 2016, through July 31, 2018. We aimed to describe the molecular epidemiology and mechanisms of resistance of CRPA isolates collected through this surveillance. Methods: We defined a case as the first isolate of P. aeruginosa resistant to imipenem, meropenem, or doripenem from the lower respiratory tract, urine, wounds, or normally sterile sites identified from a resident of the EIP catchment area in a 30-day period; EIP sites submitted a systematic random sample of isolates to CDC for further characterization. Of 1,021 CRPA clinical isolates submitted, 707 have been sequenced to date using an Illumina MiSeq. Sequenced genomes were classified using the 7-gene multilocus sequence typing (MLST) scheme, and a core genome MLST (cgMLST) scheme was used to determine phylogeny. Antimicrobial resistance genes were identified using publicly available databases, and chromosomal mechanisms of carbapenem resistance were determined using previously validated genetic markers. Results: There were 189 sequence types (STs) among the 707 sequenced genomes (Fig. 1). The most frequently occurring were high-risk clones ST235 (8.5%) and ST298 (4.7%), which were found across all EIP sites. Carbapenemase genes were identified in 5 (<1%) isolates. Overall, 95.6% of the isolates had chromosomal mutations associated with carbapenem resistance: 93.2% had porinD-associated mutations that decrease membrane permeability to the drugs; 24.8% had mutations associated with overexpression of the multidrug efflux pump MexAB-OprM; and 22.9% had mutations associated with overexpression of the endogenous β-lactamase ampC. More than 1 such chromosomal resistance mutation type was present in 37.8% of the isolates. Conclusions: The diversity of the sequence types demonstrates that HAIs caused by CRPA can arise from a variety of strains and that high-risk clones are broadly disseminated across the EIP sites but are a minority of CRPA strains overall. Carbapenem resistance in P. aeruginosa was predominantly driven by chromosomal mutations rather than acquired mechanisms (ie, carbapenemases). The diversity of the CRPA isolates and the lack of carbapenemase genes suggest that this ubiquitous pathogen can readily evolve chromosomal resistance mechanisms, but unlike carbapenemases, these cannot be easily spread through horizontal transfer.
Funding: None
Disclosures: None
Epidemiologic Characteristics of ESBL-Producing ST131 E. coli Identified Through the Emerging Infections Program, 2017
- Nadezhda Duffy, Davina Campbell, Hannah E. Reses, Elizabeth Basiliere, Chris Bower, Ghinwa Dumyati, Marion Kainer, Daniel Muleta, Benji Byrd-Warner, Richard Stanton, Jonathan Daniels, Alison Halpin, Joseph Lutgring, Isaac See
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s214-s215
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: Extended-spectrum β-lactamase–producing (ESBL) Escherichia coli infection incidence is increasing in the United States. This increase may be due to the rapid expansion of ST131, which is now the predominant ESBL strain globally, often multidrug resistant, and has been shown to establish longer-term human colonization than other E. coli strains. We assessed potential risk factors that distinguish ST131 from other ESBL E. coli. Methods: From October 1 through December 31, 2017, 5 CDC Emerging Infections Program (EIP) sites pilot tested active, laboratory-based surveillance in selected counties in Colorado, Georgia, New Mexico, New York, and Tennessee. An E. coli case was defined as the first isolation from a normally sterile body site or urine in a surveillance area resident in a 30-day period resistant to 1 extended-spectrum cephalosporin antibiotic and susceptible or intermediate to all carbapenem antibiotics tested. Epidemiologic data were collected from case patients’ medical records. A convenience sample of 117 E. coli isolates from case patients was collected. All isolates underwent whole-genome sequencing to determine sequence type and the presence of ESBL genes. We compared ST131 E. coli epidemiology to other ESBL E. coli. Results: Among 117 E. coli isolates, 97 (83%) were ESBL producers. Of the 97 ESBL E. coli, 52 (54%) were ST131 (range, for 4 EIP sites submitting >10 isolates: 25%–88%; P < .001). Other common STs were ST38 (12%) and ST10 (5%). ST131 infections were more likely to be healthcare-associated than non-ST131 (56% vs 36%; P = .05) (Table 1). Among specific prior healthcare exposures, only residence in long-term care facilities (LTCFs) in the year before culture was more common among ST131 case patients (29% vs 11%; P = .03). Notably, 85% of ESBL E. coli collected from LTCF residents were ST131. ST131 E. coli were more common among patients with underlying medical conditions (81% vs 60%; P = .02). No statistically significant difference by sex, race, age, culture source, location of culture collection, and frequency of antibiotic use in the prior 30 days was observed. Conclusions:The prevalence of ST131 E. coli varies regionally. The association between ST131 and LTCFs suggests that these may be particularly important settings for ST131 acquisition. Improving infection control measures that limit ESBL transmission in these settings and preventing dissemination in facilities receiving patients from LTCFs may be necessary to contain ST131 spread.
Funding: None
Disclosures: None
Antimicrobial Nonsusceptibility Among Invasive MRSA USA300 Strains by Healthcare Exposure, Three Sites, 2005–2016
- Kelly Jackson, Runa Gokhale, Davina Campbell, Amy Gargis, Susan Ray, Ruth Lynfield, William Schaffner, Joseph Lutgring, Isaac See
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s120-s121
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: Incidence of community-associated (CA) and healthcare-associated, community-onset (HACO) USA300 methicillin-resistant Staphylococcus aureus (MRSA) bloodstream infections has remained unchanged in recent years. Traditionally considered a CA strain, USA300 is increasingly associated with healthcare settings. We examined whether antimicrobial nonsusceptibility among USA300 strains could distinguish epidemiologic class (community vs hospital), and whether divergences in susceptibility were occurring over time. Methods: We used data on invasive MRSA infections from active, population, and laboratory-based surveillance during 2005–2016 from 11 counties in 3 states. Invasive cases were defined as MRSA isolation from a normally sterile site in a surveillance area resident. Cases were considered hospital-onset (HO) if the culture was obtained >3 days after hospitalization and HACO if ≥1 of the following risk factors was present: hospitalization, surgery, dialysis, or residence in a long-term care facility in the past year; or central vascular catheter ≤2 days before culture. Otherwise, cases were considered CA. Sites submitted a convenience sample of clinical MRSA isolates for molecular typing and antimicrobial susceptibility testing. Molecular typing was performed by pulsed-field gel electrophoresis until 2008, when typing was inferred using a validated algorithm based on molecular characteristics. Reference broth microdilution was performed for 8 antimicrobials and interpreted based on CLSI interpretive criteria. We compared USA300 nonsusceptibility for HO and CA isolates. For antimicrobials with >5% nonsusceptibility and for which HO isolates had greater nonsusceptibility than CA isolates, we compared nonsusceptibility for HACO and CA and analyzed annual trends in nonsusceptibility within each epidemiologic class (ie, CA, HACO, and HO) using linear regression. Results: Of 17,947 MRSA cases during 2005–2016, isolates were available for 6,685 (37%), and 2,120 were USA300 (34% CA, 52% HACO, 14% HO). HO isolates had more nonsusceptibility than CA isolates to gentamicin (2.2% vs 0.6%; P = .03), levofloxacin (47.8% vs 39.7%; P = .02), rifampin (3.7 vs 1.1%; P = .01), and trimethoprim-sulfamethoxazole (3.4% vs 0.6%; P = .04). HACO isolates also had more nonsusceptibility than CA isolates to levofloxacin (50.9% vs 39.7%; P < .01). Levofloxacin nonsusceptibility increased during 2005–2016 for HACO and CA isolates (P < .01), but not among HO isolates (P = .36) (Fig. 1). Conclusions: Overall, nonsusceptibility across drugs cannot distinguish USA300 isolates causing HO versus CA disease. Although HO isolates had higher levofloxacin nonsusceptibility than CA and HACO isolates early on, USA300 MRSA HACO isolates now have levofloxacin nonsusceptibility most similar to that of HO isolates. Further study could help to explore whether increases in fluoroquinolone nonsusceptibility among CA and HACO cases may be contributing to the persistence of USA300 strains.
Disclosures: None
Funding: None





