2 results
Characterization of carbapenem-resistant gram-negative bacteria collected in the Sentinel Surveillance Program, 2018–2019
- Lori Spicer, Davina Campbell, J. Kristie Johnson, Cynthia Longo, Thomas Balbuena, Thomas Ewing, Maria Karlsson, J. Kamile Rasheed, Christopher Elkins, Amy Gargis, Joseph Lutgring
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 2 / Issue S1 / July 2022
- Published online by Cambridge University Press:
- 16 May 2022, p. s52
-
- Article
-
- You have access Access
- Open access
- Export citation
-
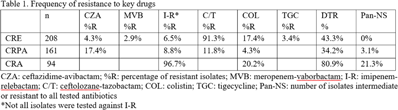
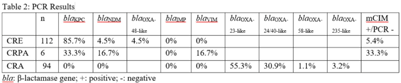
Background: Carbapenem resistance in gram-negative organisms is an important public health problem. The CDC conducted Sentinel surveillance in 2018–2019 to characterize these organisms from 9 facilities in 9 different states. Methods: Carbapenem-resistant Enterobacterales (CRE), Pseudomonas aeruginosa (CRPA), and Acinetobacter spp (CRA) obtained from clinical samples of patients in acute-care or long-term care facilities were submitted to the CDC. Identification was confirmed using matrix-assisted laser desorption ionization time-of-flight (MALDI-TOF), and antimicrobial susceptibility testing (AST) was performed via broth microdilution for 27 antibiotics. All confirmed CRE and CRPA were tested for carbapenemase production (CP) using the modified carbapenem inactivation method (mCIM). The isolates that were mCIM-positive were assessed by real-time PCR for presence of blaKPC, blaNDM, blaVIM, and blaIMP. CP-CRE were also assessed for blaOXA-48-like. All confirmed CRA were tested for the same genes as CRPA and blaOXA-23–like, blaOXA-24/40-like, blaOXA-58–like, and blaOXA-235–like genes. Difficult-to-treat resistance (DTR) was defined as resistance to all β-lactams (excluding newer β-lactam combination agents) and quinolones tested. Results: The CDC confirmed 208 CRE, 161 CRPA, and 94 CRA. Table 1 summarizes AST results for a selection of drugs. We identified 112 (53.8%) mCIM-positive CRE and 6 (3.7%) mCIM-positive CRPA. The PCR results are summarized in Table 2. One mCIM-positive and PCR-negative isolate was positive in a metallo-β-lactamase screen. Conclusions: Resistance among CRE and CRPA to newer β-lactam combination agents was detected. Options for treating CRA are limited. Of 112 CP-CRE, 85.7% harbored blaKPC; CP-CRPA were rare (3.7%); and most CRA harbored blaOXA-23-like (55.3%) or blaOXA-24/40-like (30.9%). Whole-genome sequencing is planned to better understand gene variants, sequence types, and additional resistance markers present among the isolates.
Funding: None
Disclosures: None
New Approaches to Colonization Screening in Response to Emerging Antimicrobial Resistance
- Karen Anderson, Maria Karlsson, Sandra Boyd, Natashia Reese, Uzma Ansari, Davina Campbell, Amelia Bhatnagar, Paige Gable, Stephanie Swint, Cynthia Longo, Sarah Gilbert, Lori Spicer, Jake Cochran, David Lonsway
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, p. s330
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: The capacity to monitor the emergence of carbapenemase-producing organisms (CPO) is critical in limiting transmission. CPO-colonized patients can be identified by screening rectal specimens for carbapenemase genes and the Cepheid GeneXpert Carba-R (XCR), the only FDA-approved test, is limited to 5 carbapenemase genes and cannot identify the bacterial species. Objective: We describe the development and validation of culture-based methods for the detection of CPO in rectal cultures (RCs) and nonrectal cultures (NRCs) of tracheal aspirate and axilla-groin swabs. Methods: Colonization screening was performed at 3 US healthcare facilities; specimens of RC swabs and NRC ESwabs were collected. Each specimen was inoculated to a MacConkey broth enrichment tube for overnight incubation then were subcultured to MacConkey agar with meropenem and ertapenem 10 µg disks (BEMA) and CHROMagar KPC (KCHR) or CHROMagar Acinetobacter (ACHR). All media were evaluated for the presence of carbapenem-resistant organisms; suspect colonies were screened by real-time PCR for the most common carbapenemase genes. MALDI-TOF was performed for species identification. BEMA, a previously validated method, was the comparator for 52 RCs; clinical culture (CC) served as the comparator method for 66 NRCs. Select CPO-positive and -negative specimens underwent reproducibility testing. Results: Among 56 patients undergoing colonization screening, 12 (21%) carried a CPO. Only 1 patient had CPO solely from RC. Also, 6 patients had both CPO-positive RC and NRC, and 5 patients only had a CPO-positive NRC. Of the latter, 4 had a CPO-positive tracheal specimen, and 1 had a positive culture from both tracheal and axilla-groin specimens. Sensitivity of BEMA (70%) for NRC was lower than for KCHR (96%) and ACHR (88 %) for all specimens. All methods showed a specificity of 100% and reproducibility of 92%. The detected CPO included OXA-23–positive Acinetobacter baumannii, NDM-positive Escherichia coli, KPC-positive Pseudomonas aeruginosa and 4 genera of KPC-positive Enterobacteriaceae. Conclusions:The addition of nonrectal specimens and use of selective media contributed to increased sensitivity and enhanced identification of CPO-colonized patients. Positive cultures were equally distributed among the 3 specimen types. The addition of the nonrectal specimens resulted in the identification of more colonized patients. The culture-based method was successful in detecting an array of different CPOs and target genes, including genes not detected by the Carba-R assay (eg, blaOXA-23-like). Enhanced isolation and characterization of CPOs will be key in aiding epidemiologic investigations and strengthening targeted guidance for containment strategies.
Funding: None
Disclosures: We discuss the drug combination aztreonam-avibactam and acknowledge that this drug combination is not currently FDA approved.





