Introduction
Several technological advances have been critical in the development of the scanning transmission electron microscope (STEM) from its inception (von Ardenne, Reference von Ardenne1938) to its current status as one of the most important techniques for high-resolution imaging of materials. Specifically, improved vacuum systems, field emission sources, aberration correction, and the introduction of annular dark field (ADF) detectors (Crewe, Reference Crewe1966; Crewe et al., Reference Crewe, Wall and Welter1968) were all crucial developments. ADF detectors are typically formed of one or more PN diode segments or scintillator photomultiplier tube arrangements. These are placed in the far field of the objective lens and sample an angular range of the diffraction pattern of the area of the sample illuminated by the electron beam. Such devices are well suited for use in STEM due to their fast readout, and imaging with pixel dwell times measured in microseconds is normal. ADF imaging was initially understood as being based on Z-contrast (Reference CreweCrewe, Reference Crewe1970a, Reference Crewe1970b), though understanding of the contrast mechanism evolved over time (Donald & Craven, Reference Donald and Craven1979), in turn influencing the design of such detectors. In particular, later contributions demonstrated that the inner angle of ADF detectors had to be relatively high to exclude coherent diffraction from dominating the signal (Pennycook & Jesson, Reference Pennycook and Jesson1991; Hartel et al., Reference Hartel, Rose and Dinges1996). Other refinements of this arrangement have been introduced over the years, including the use of split detectors for differential phase contrast (Dekkers & de Lang, Reference Dekkers and de Lang1977; Chapman, Reference Chapman, Batson, Waddell and Ferrier1978; Chapman et al., Reference Chapman, McFadyen and McVitie1990; McGrouther et al., Reference McGrouther, Benitez Romero, McFadzean and McVitie2014), multiple annular detectors (Shibata et al., Reference Shibata, Kohno, Findlay, Sawada, Kondo and Ikuhara2010, Reference Shibata, Seki, Sánchez-Santolino, Findlay, Kohno, Matsumoto, Ishikawa and Ikuhara2017), and the use of bright field or annular bright field imaging (Hammel & Rose, Reference Hammel and Rose1995; LeBeau et al., Reference LeBeau, D'Alfonso, Findlay, Stemmer and Allen2009; Findlay et al., Reference Findlay, Shibata, Sawada, Okunishi, Kondo and Ikuhara2010; MacLaren et al., Reference MacLaren, Wang, Morris, Craven, Stamps, Schaffer, Ramasse, Miao, Kalantari, Sterianou and Reaney2015). However, all these detector configurations integrate over large angular ranges of the back focal plane, resulting in the loss of most of the information contained in the diffraction pattern. Furthermore, space constraints in the microscope's camera chamber can limit which detectors can be used simultaneously in an individual experiment, so that data acquisition may have to be repeated several times from the same area using different detectors to collect all the signals of interest. This can lead to difficulties in correlating the information contained in images acquired in successive experiments due to drift, and results in a higher overall dose to the sample, which is undesirable for beam-sensitive samples.
Recently, building upon technologies developed for particle physics (Turala, Reference Turala2005; Wermes, Reference Wermes2005; Turchetta et al., Reference Turchetta, Fant, Gasiorek, Esbrand, Griffiths, Metaxas, Royle, Speller, Venanzi, van der Stelt, Verheij, Li, Theodoridis, Georgiou, Cavouras, Hall, Noy, Jones, Leaver, Machin, Greenwood, Khaleeq, Schulerud, Østby, Triantis, Asimidis, Bolanakis, Manthos, Longo and Bergamaschi2007; Delpierre, Reference Delpierre2014), pixelated detectors developed for X-ray imaging have been adopted for electron imaging (Clough et al., Reference Clough, Moldovan and Kirkland2014; McMullan et al., Reference McMullan, Faruqi, Clare and Henderson2014; McGrouther et al., Reference McGrouther, Krajnak, MacLaren, Maneuski, O'Shea and Nellist2015; Tate et al., Reference Tate, Purohit, Chamberlain, Nguyen, Hovden, Chang, Deb, Turgut, Heron, Schlom, Ralph, Fuchs, Shanks, Philipp, Muller and Gruner2016; Mir et al., Reference Mir, Clough, MacInnes, Gough, Plackett, Shipsey, Sawada, MacLaren, Ballabriga, Maneuski, O'Shea, McGrouther and Kirkland2017; Tinti et al., Reference Tinti, Fröjdh, van Genderen, Gruene, Schmitt, de Winter, Weckhuysen and Abrahams2018). Compared with charge coupled device (CCD)-based detectors, these direct electron detectors (DEDs) typically offer much lower noise levels, improved detector quantum efficiency (DQE), and modulation transfer function (MTF), some degree of radiation hardness, and crucially, fast readout of the images. This allows the efficient recording of the entire diffraction pattern at each scan position using millisecond or sub-millisecond dwell times, enabling either improvements in or the use of different imaging modes, such as nanobeam STEM diffraction (Mahr et al., Reference Mahr, Müller-Caspary, Ritz, Simson, Grieb, Schowalter, Krause, Lackmann, Soltau, Wittstock and Rosenauer2019), position averaged convergent beam electron diffraction (LeBeau et al., Reference LeBeau, Findlay, Allen and Stemmer2010; Ophus et al., Reference Ophus, Ercius, Huijben and Ciston2017), atomically resolved electrostatic field mapping (Hachtel et al., Reference Hachtel, Idrobo and Chi2018; Fang et al., Reference Fang, Wen, Allen, Ophus, Han, Kirkland, Kaxiras and Warner2019), improved magnetic (Krajnak et al., Reference Krajnak, McGrouther, Maneuski, O'Shea and McVitie2016) induction characterization, determination of crystal periodicity along the beam direction (Nord et al., Reference Nord, Ross, McGrouther, Barthel, Moreau, Hallsteinsen, Tybell and MacLaren2019a), fluctuation electron microscopy (Banerjee et al., Reference Banerjee, Baker, Doye, Nord, Heath, Erotokritou, Bosworth, Barber, MacLaren and Hadfield2017), and ptychography (Pennycook et al., Reference Pennycook, Lupini, Yang, Murfitt, Jones and Nellist2015; Yang et al., Reference Yang, Rutte, Jones, Simson, Sagawa, Ryll, Huth, Pennycook, Green, Soltau, Kondo, Davis and Nellist2016). Additionally, work is in progress to also improve scanning precession electron diffraction (SPED) (Rauch et al., Reference Rauch, Portillo, Nicolopoulos, Bultreys, Rouvimov and Moeck2010) using such DEDs, especially because of their better noise performance that optically coupled CCDs (MacLaren et al., Reference MacLaren, Frutos-Myro, McGrouther, McFadzean, Weiss, Cosart, Portillo, Robins, Nicolopoulos, del Busto and Skogeby2020). Indeed, pixelated detectors are increasingly regarded as “universal” detectors (Tate et al., Reference Tate, Purohit, Chamberlain, Nguyen, Hovden, Chang, Deb, Turgut, Heron, Schlom, Ralph, Fuchs, Shanks, Philipp, Muller and Gruner2016; Yang et al., Reference Yang, Rutte, Jones, Simson, Sagawa, Ryll, Huth, Pennycook, Green, Soltau, Kondo, Davis and Nellist2016; Hachtel et al., Reference Hachtel, Idrobo and Chi2018; Fang et al., Reference Fang, Wen, Allen, Ophus, Han, Kirkland, Kaxiras and Warner2019; Ophus, Reference Ophus2019) capable of imaging under multiple modes.
Along with the many advantages that fast pixelated detectors bring, many practical limitations arise from their use, such as the ability to get real-time information from the data stream produced from a scan to enable navigation and identification of relevant sample features, and the storage and processing of very large datasets, often much larger than the available computer memory. In this paper (Part I), we present solutions for the hardware control, data acquisition, real-time processing and visualization, and storage of data from fast pixelated detectors. The majority of the software solutions presented in this work are made available under the free and open source GPLv3 licence, allowing transparency of the implemented algorithms, and the ability for anyone to use and to further improve upon them. The names of the software packages, modules, classes, and functions we present are given in typewriter font.
Most of the libraries reported here are implemented in Python. Python, being an open and free programming language, is rapidly becoming the standard language for many aspects of scientific computing (Oliphant, Reference Oliphant2007; Gouillart et al., Reference Gouillart, Nunez-Iglesias and van der Walt2016). In addition to its comparative ease of use, which lowers the barrier for people to contribute and minimizes developer time, Python has an extensive standard library and a large ecosystem of external libraries, including ones for optimized numerical (Oliphant, Reference Oliphant2006) and scientific (Jones et al., Reference Jones, Oliphant and Peterson2001) computing, image processing (van der Walt, Reference van der Walt, Schönberger, Nunez-Iglesias, Boulogne, Warner, Yager, Gouillart and Yu2014), data visualization (Hunter, Reference Hunter2007), and work flow documentation (Kluyver et al., Reference Kluyver, Ragan-Kelley, Pérez, Granger, Bussonnier, Frederic, Kelley, Hamrick, Grout, Corlay, Ivanov, Avila, Abdalla, Willing, Loizides and Schmidt2016). Furthermore, it is straightforward to link Python to low-level C-code, allowing the development of optimized routines or use of external libraries (Behnel et al., Reference Behnel, Bradshaw, Citro, Dalcin, Seljebotn and Smith2011).
Within the electron microscopy community, a number of Python packages have also been developed. One example of this is HyperSpy (de la Peña et al., Reference de la Peña, Ostasevicius, Fauske, Burdet, Prestat, Jokubauskas, Nord, Sarahan, MacArthur, Johnstone, Taillon, Caron, Migunov, Furnival, Eljarrat, Mazzucco, Aarholt, Walls, Slater, Winkler, Martineau, Donval, McLeod, Hoglund, Alxneit, Hjorth, Henninen, Zagonel and Garmannslund2018), which contains functionality for processing data from a wide range of TEM techniques: electron energy loss spectroscopy, energy-dispersive X-ray spectroscopy, electron holography, and more standard imaging. It also serves as a base for several other packages, such as pyXem for analyzing SPED data (Johnstone et al., Reference Johnstone, Crout, Høgås, Martineau, Smeets, Laulainen, Collins, Morzy, Prestat, Ånes, Doherty, Ostasevicius and Bergh2019), Atomap for processing atomic resolution STEM data (Nord et al., Reference Nord, Vullum, MacLaren, Tybell and Holmestad2017), and pixStem for working with data from fast pixelated STEM detectors (pixStem devs, 2015). Several other packages exist, like rigidRegistration for doing rigid image registration of atomic resolution image stacks (Savitzky et al., Reference Savitzky, Baggari, Clement, Waite, Goodge, Baek, Sheckelton, Pasco, Nair, Schreiber, Hoffman, Admasu, Kim, Cheong, Bhattacharya, Schlom, McQueen, Hovden and Kourkoutis2018), and wrappers for doing STEM simulations, like PyPrismatic (Ophus, Reference Ophus2017). Other packages for processing data from fast pixelated STEM detectors include py4DSTEM (Savitzky et al., Reference Savitzky, Zeltmann, Barnard, Brown, Henderson and Ginsburg2019), LiberTEM (Clausen et al., Reference Clausen, Weber, Caron, Nord, Müller-Caspary, Ophus, Dunin-Borkowski, Ruzaeva, Chandra, Shin and van Schyndel2019), pycroscopy (Somnath et al., Reference Somnath, Smith, Laanait, Vasudevan, Ievlev, Belianinov, Lupini, Shankar, Kalinin and Jesse2019), and fpd (fpd devs, 2015).
The post-acquisition visualization and processing of data from fast pixelated detectors using the fpd and pixStemFootnote 1 libraries for the structural characterization of materials will be reported in Part II of this work (Paterson et al., Reference Paterson, Webster, Ross, Paton, Macgregor, McGrouther, MacLaren and Nord2020b). A third and final part (yet to be submitted) will cover aspects related to differential phase contrast analysis. Throughout all parts, we provide examples using data from a Medipix3 detector (Ballabriga et al., Reference Ballabriga, Alozy, Blaj, Campbell, Fiederle, Frojdh, Heijne, Llopart, Pichotka, Procz, Tlustos and Wong2013). Although some sections of the codebase are specific to the use of this detector, such as aspects relating to data acquisition, many of the issues discussed and the techniques and tools presented in this part are applicable to a wide range of other detectors, while the data processing described in the forthcoming parts II and III of this paper series are applicable to data from any detector.
This paper is organized as follows. In the section "Medipix3 Detector", the Medipix3 detector is briefly introduced. Methodologies for acquiring data from it are discussed in the section "Medipix3 Data Acquisition". In the section "Live Data Processing", an architecture developed to process a live data stream from a fast pixelated detector is outlined. In the section "Data Storage", the issues around data storage are discussed and our implementation is presented. The source data and scripts to analyze the data and produce the results presented here are publicly available (Nord et al., Reference Nord, Webster, Paton, McVitie, McGrouther, MacLaren and Paterson2019b).
Medipix3 Detector
All pixelated data reported in this work is from a ![]() $256\times 256$ pixel Medipix3RX (henceforth referred to as Medipix3) detector (Ballabriga et al., Reference Ballabriga, Alozy, Blaj, Campbell, Fiederle, Frojdh, Heijne, Llopart, Pichotka, Procz, Tlustos and Wong2013) affixed to a Merlin 1R retractable Medipix3 mount from Quantum Detectors (Harwell, Oxfordshire, UK). The Medipix3 detector is a radiation hard hybrid counting DED, where active analog and digital signal processing circuitry in each 55 µm pixel is bump-bonded to a relatively thick sensor layer. Si sensor layers of 500 µm are needed for operation at primary electron energies of 300 keV. In our case, a 300 µm silicon sensor layer was used for all data except that in Figure 1, where a 500 µm layer was used instead.
$256\times 256$ pixel Medipix3RX (henceforth referred to as Medipix3) detector (Ballabriga et al., Reference Ballabriga, Alozy, Blaj, Campbell, Fiederle, Frojdh, Heijne, Llopart, Pichotka, Procz, Tlustos and Wong2013) affixed to a Merlin 1R retractable Medipix3 mount from Quantum Detectors (Harwell, Oxfordshire, UK). The Medipix3 detector is a radiation hard hybrid counting DED, where active analog and digital signal processing circuitry in each 55 µm pixel is bump-bonded to a relatively thick sensor layer. Si sensor layers of 500 µm are needed for operation at primary electron energies of 300 keV. In our case, a 300 µm silicon sensor layer was used for all data except that in Figure 1, where a 500 µm layer was used instead.
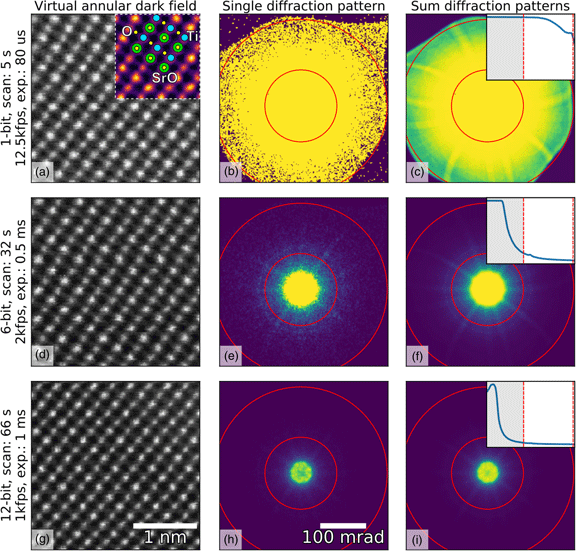
Fig. 1. Imaging of SrTiO3 along the [110] direction using different bit depths and probe dwell times (in rows) with the Medipix3 detector in a continuous read-write mode. The bit depth, total scan time, and the detector frame acquisition rate and exposure are shown in the annotations. HAADF images (left column) were calculated by summing all counts inside a virtual aperture defined over the collection angles 80–192 mrad (assuming a linear mapping of pixel count to diffracted angle, which may not be entirely true in an image-corrected microscope), shown by the red lines in the diffraction images (middle column), using the pixStem library (pixStem devs, 2015). The colored section of the 1-bit HAADF image in (a) is Fourier filtered with a schematic overlay of the atomic columns imaged: green: Sr, yellow: O, and blue: Ti. The third column shows the summed diffraction patterns, with the insets displaying their radial distributions from 0 to 192 mrad. The dip in intensity in the centre of the direct spot in the 12-bit mode data in (h) and (i) is due to the higher bit depth, now allowing the details of the primary beam and low-order diffraction discs to be seen.
In electron microscopy applications, an incident electron produces electron-hole pairs in the sensor layer in sufficient numbers (Scholze et al., Reference Scholze, Rabus and Ulm1998) for the signal due to a primary electron to be clearly distinguishable from noise in the detector. This makes the detector capable of noiseless operation by the setting of an appropriate threshold for counting, and the detector is thus able to detect individual electrons. As a consequence, the Medipix3 detector is of potential use in time-resolved electron microscopy experiments, where sub-100 ns time resolution has been recently demonstrated (Paterson et al., Reference Paterson, Lamb, Ballabriga, Maneuski, O'Shea and McGrouther2020a).
Each pixel can operate independently, with its active circuitry processing only the signal induced in that pixel, in a mode of operation known as single pixel mode (SPM). Alternatively, in so-called charge summing mode (CSM), neighboring pixels can pool their circuitry and collectively process the signals induced in each pixel (Ballabriga et al., Reference Ballabriga, Alozy, Blaj, Campbell, Fiederle, Frojdh, Heijne, Llopart, Pichotka, Procz, Tlustos and Wong2013). CSM attempts to account for charge spread between pixels due to electron-matter interactions in the thick sensor layer. At an acceleration voltage of up to 80 kV, the Medipix3 has a near-perfect DQE and MTF when imaging electrons (Mir et al., Reference Mir, Clough, MacInnes, Gough, Plackett, Shipsey, Sawada, MacLaren, Ballabriga, Maneuski, O'Shea, McGrouther and Kirkland2017). The use of alternative high-Z sensor-layer materials is expected to improve the performance at higher acceleration voltages (McMullan et al., Reference McMullan, Cattermole, Chen, Henderson, Llopart, Summerfield, Tlustos and Faruqi2007) and is currently being investigated.
Another notable feature of the Medipix3 detector is the ability to operate in a continuous read-write mode, where one of the two sets of counters in each pixel is used to readout the data while the other takes over counting. This gapless recording maximizes dose efficiency, which is important for beam-sensitive samples, and also enables faster acquisitions, which is important for minimizing artifacts due to microscope instabilities, particularly when imaging with atomic resolution.
The Medipix3 detector can be operated in 1-, 6-, 12-, and 24-bit depth modes, allowing the compromise between readout time, file size, and dynamic range to be varied. The clock on the Medipix3 was designed to be driven at frequencies up to 200 MHz but, with additional cooling, it can be overclocked to allow faster operation. With the 120 MHz clock rate of the Merlin readout system (Plackett et al., Reference Plackett, Horswell, Gimenez, Marchal, Omar and Tartoni2013) used here, the readout times are 70.8 µs, 412 µs, 822 µs, and 1.64 ms, for 1, 6, 12, and 24-bit modes, respectively. While the 24-bit mode is ideal for very high dynamic range diffraction studies (Mir et al., Reference Mir, Clough, MacInnes, Gough, Plackett, Shipsey, Sawada, MacLaren, Ballabriga, Maneuski, O'Shea, McGrouther and Kirkland2017), the higher readout rates of the lower bit depth modes are more generally useful across a wide range of imaging conditions (it would take ![]() $\gt$4 ms to exceed 12 bits at 1 MHz count rates per pixel, so 24 bits are only needed for long counting times or high arrival rates on some pixels).
$\gt$4 ms to exceed 12 bits at 1 MHz count rates per pixel, so 24 bits are only needed for long counting times or high arrival rates on some pixels).
To demonstrate the use of different bit depths, atomic resolution data from SrTiO3 imaged along the [110] direction were acquired on a Medipix3 detector at bit depths of 1, 6, and 12, giving maximum counts of 1, 63, and 4095, respectively. The data were acquired on a JEOL ARM 300CF using an acceleration voltage of 200 kV and a convergence angle of 22.4 mrad, with the Medipix3 operated in SPM with continuous read–write enabled. High-angle ADF (HAADF) images produced from these datasets are shown in the left-hand column of Figure 1, with the bit depth increasing from top (a) to bottom (g). The atomic resolution contrast in these images arises mostly from incoherent scattering of the electrons, similar to that in regular HAADF imaging with dedicated annular detectors. The middle- and right-hand columns show individual and summed diffraction patterns from each scan, respectively. The circular red lines in the diffraction patterns mark the edges of the virtual aperture used within which pixel counts were added up to give the intensity used for each pixel of the real-space images, while the insets in the third column show the radial distributions. The non-round “shadow” easily visible at the outer edges of the 1-bit diffraction pattern (Fig. 1a) is due to high-angle cutoff in the microscope due to the image corrector. Although the 1-bit diffraction patterns (Figs. 1b, 1c) do not seem to contain much information, the ADF data (Fig. 1a) show that high-quality atomic resolution imaging is possible, with the SrO, Ti, and O2 columns (Abramov et al., Reference Abramov, Tsirelson, Zavodnik, Ivanov and Brown1995) all resolved, as shown in the inset schematic.
The very high frame rates of 12,500 frames per second with 1-bit data in a continuous read–write mode that are achievable makes this acquisition configuration particularly suitable for navigation during setup or in especially beam-sensitive materials. One image of ![]() $256\times 256$ probe positions takes about 5 s to complete at this rate, but smaller scan sizes are often adequate for navigation. In the experiment, the beam current was maintained and this unavoidably resulted in the central portion of the diffraction pattern being saturated when recording at bit depths of 1 and 6. As shown by the radial distributions, we have selected the scattering angles where the detector is not saturated and contrast can be extracted. With shorter exposures or lower beam currents, regions closer to the central spot of the diffraction pattern will not be saturated and would produce useable image contrast. In 6-bit mode, more features of the diffraction pattern are visible than in the 1-bit mode, and the darkfield image (Fig. 1d) is better defined. This trend continues to the 12-bit mode (Fig. 1g) where the direct beam is no longer saturated, as shown in the inset in Figure 1i. However, more atomic columns are present in the image as a result of larger spatial drift during the longer acquisition (9 or 10 Sr columns per row in the 12-bit data compared with 8 or 9 columns per row for the 1-bit data). The principal benefit of higher bit depth imaging in this context is that a greater range of scattering angles may be used for virtual aperture imaging post-acquisition, and the signal-to-noise ratio (SNR) is generally higher, even if there is a cost in acquisition time and consequent drift.
$256\times 256$ probe positions takes about 5 s to complete at this rate, but smaller scan sizes are often adequate for navigation. In the experiment, the beam current was maintained and this unavoidably resulted in the central portion of the diffraction pattern being saturated when recording at bit depths of 1 and 6. As shown by the radial distributions, we have selected the scattering angles where the detector is not saturated and contrast can be extracted. With shorter exposures or lower beam currents, regions closer to the central spot of the diffraction pattern will not be saturated and would produce useable image contrast. In 6-bit mode, more features of the diffraction pattern are visible than in the 1-bit mode, and the darkfield image (Fig. 1d) is better defined. This trend continues to the 12-bit mode (Fig. 1g) where the direct beam is no longer saturated, as shown in the inset in Figure 1i. However, more atomic columns are present in the image as a result of larger spatial drift during the longer acquisition (9 or 10 Sr columns per row in the 12-bit data compared with 8 or 9 columns per row for the 1-bit data). The principal benefit of higher bit depth imaging in this context is that a greater range of scattering angles may be used for virtual aperture imaging post-acquisition, and the signal-to-noise ratio (SNR) is generally higher, even if there is a cost in acquisition time and consequent drift.
The selection of higher scattering angles by saturating the central spot in 1-bit and 6-bit modes is possible here because, unlike CCDs, the Medipix3 is not damaged by the very intense direct beam, and because the noise-free readout enables each single electron hit to be accurately recorded. With very intense beams (approximately 1 MHz count rate per pixel), the electron arrival rate can exceed the counting rate of the detector; this does no harm to the detector, but electrons are missed and the counts no longer represent an accurate reflection of arrival rates (and even a little below this level, counting linearity is lost). In this data, however, the beam current was not high enough to cause such an effect and the slight dips in intensity in the center of Figures 1h and 1i are due to the real internal structure of the brightest portion of the diffraction pattern being resolved at the highest bit depth.
The fastest frame rate demonstrated above, corresponding to 80 µs per scan pixel, is still substantially slower than that of commonly used scintillator or photomultiplier tube-based STEM detectors. While switching between these detectors is a common procedure with little overhead, pixelated STEM detectors with higher speeds would be beneficial for the efficiency of live-imaging and for low-dose imaging of beam-sensitive materials without having to reduce the beam current. The limiting factor in the Medipix3 setup we used is the 120 MHz clock of the Merlin readout system (Plackett et al., Reference Plackett, Horswell, Gimenez, Marchal, Omar and Tartoni2013). The Medipix3 chip itself can be clocked to 200–250 MHz, potential allowing a doubling of acquisition speed. Additionally, the Medipix3 chip allows readout of regions of interest (ROI), potentially allowing much higher sub-frame readout rates, but readout systems with this capability are not yet commonly available.
The pnCCD detector (Ryll et al., Reference Ryll, Simson, Hartmann, Holl, Huth, Ihle, Kondo, Kotula, Liebel, Müller-Caspary, Rosenauer, Sagawa, Schmidt, Soltau and Strüder2016) is a radiation hard ![]() $264 \times 264$ pixel CCD-based sensor with a full frame rate of 1000 fps. The chip supports binning along one axis, allowing speeds of up to 4,000 fps with
$264 \times 264$ pixel CCD-based sensor with a full frame rate of 1000 fps. The chip supports binning along one axis, allowing speeds of up to 4,000 fps with ![]() $4\times$ binning (
$4\times$ binning (![]() $264\times 66$ pixels). However, it was recently reported that these speeds may be doubled with optimization of the operation conditions and timing coordination of the readout ASICs (Huth et al., Reference Huth, Ritz, O'Leary, Griffiths, Nellist and Soltau2019). By windowing to a 24-pixel wide strip, the detector is reported to operated at 10,000 fps, which is approaching that of the 12,500 fps
$264\times 66$ pixels). However, it was recently reported that these speeds may be doubled with optimization of the operation conditions and timing coordination of the readout ASICs (Huth et al., Reference Huth, Ritz, O'Leary, Griffiths, Nellist and Soltau2019). By windowing to a 24-pixel wide strip, the detector is reported to operated at 10,000 fps, which is approaching that of the 12,500 fps ![]() $256\times 256$ 1-bit data shown in this work. At the maximum SNR mode of operation and imaging 200 keV electrons, the maximum number of primary electrons that may be measured per pixel is about 1 in the pnCCD detector. This number rises to about 7 in the highest capacity mode of operation, but with a reduction in the SNR. This detector does not benefit from the gapless or noise-free readout of the Medipix detector used here, but can still produce excellent results at low doses (Huth et al., Reference Huth, Ritz, O'Leary, Griffiths, Nellist and Soltau2019).
$256\times 256$ 1-bit data shown in this work. At the maximum SNR mode of operation and imaging 200 keV electrons, the maximum number of primary electrons that may be measured per pixel is about 1 in the pnCCD detector. This number rises to about 7 in the highest capacity mode of operation, but with a reduction in the SNR. This detector does not benefit from the gapless or noise-free readout of the Medipix detector used here, but can still produce excellent results at low doses (Huth et al., Reference Huth, Ritz, O'Leary, Griffiths, Nellist and Soltau2019).
Alternative modes of operation can yield even faster data readouts from current generation detectors. The Timepix3 chip (Poikela et al., Reference Poikela, Plosila, Westerlund, Campbell, Gaspari, Llopart, Gromov, Kluit, van Beuzekom, Zappon, Zivkovic, Brezina, Desch, Fu and Kruth2014) operates with a 640 MHz clock, giving a timestamp resolution of 1.56 ns, and supports a data-driven mode of acquisition. In this mode, only data from events are read out rather than the full array of pixel counts. The data packet itself includes additional information such as time over threshold and time of arrival and, consequently, additional or alternative strategies are required to process this type of data (with the potential benefit of more advanced signal processing). The maximum hit rate is 40 M hits/s/cm2 (approximately 80 M hits/s for a single 1.98 cm2 chip) and provides faster readout in a event driven mode than in a frame mode for less than 50% occupancy. While this approach allows very short effective exposures, due to the increased size of the packet, the overall counting rate is reduced compared with the Medipix3 detector. However, unlike in the 1-bit Medipix3 data, the intensity distribution of the image signal would be accurately recorded without saturation in event driven modes of acquisition.
Beyond this, the collaboration behind the next-generation Medipix4 detector is targeting imaging rates which are compatible with human CT imaging (Campbell et al., Reference Campbell, Alozy, Ballabriga, Frojdh, Heijne, Llopart, Poikela, Tlustos, Valerio and Wong2016). The typical detector dose varies in CT imaging, but can be of the order of 103–104 M hits/s/cm2 (Taguchi & Iwanczyk, Reference Taguchi and Iwanczyk2013), significantly higher than is possible in current-generation Medipix detectors.
Regardless of the imaging mode used for the collection of the source data, smaller bit depths also make both the file storage and the data processing more efficient; a 6-bit dataset is about four times smaller than a 24-bit one of the same scan area, making it much more convenient to store and transfer. This advantage also extends to the data processing, since loading and processing data files which are four times smaller will be much quicker.
Medipix3 Data Acquisition
Data from the Medipix3 detector were acquired through the Merlin readout system (Plackett et al., Reference Plackett, Horswell, Gimenez, Marchal, Omar and Tartoni2013). This allows setting of the acquisition parameters, either through a graphical user interface (GUI) or over TCP/IP, and reads and processes the raw data through a field-programmable gate array (FPGA), returning the data to the acquisition computer. The FPGA processing can be bypassed to some extent by operating the system in a “raw” mode. This enables larger scan sizes at high frame rates, with the requirement that the data must be reshaped post-acquisition, and with no live visualization of the acquired images directly in the Merlin software. However, the Merlin TCP/IP data API remains functional, so it is possible to get live imaging through other means (see the section "Live Data Processing").
The Merlin system can be triggered by software over TCP/IP or by hardware (TTL) input. We typically use the latter approach and couple to the TTL signals produced by a Gatan DigiScan system, as shown schematically in Figure 2a. This produces extra acquisitions due to triggers sent during the flyback time, and the handling of these is discussed in the section "Data Storage". The main advantage of this approach is that Gatan Digital Micrograph (DM), in addition to allowing access to microscope control, can be used for setting scan parameters in one of several ways discussed below, and additional STEM detector signals may be acquired simultaneously.

Fig. 2. (a) Schematic of the Merlin Medipix3 readout control architecture and screenshots of (b) low-level (Merlin Control) and (c) optimized continuous read-write mode (MERLIN_PixSTEM) plugins for Gatan DM. Note that the DM plugin in (c) is not related to the similarly named Python pixStem library.
The simultaneously acquired DM datasets also serve to document the microscope and scan parameters in the data tags, which can then be used in data conversion (discussed in the section "Data Storage"), abstracting away the differences in how various microscope manufacturers provide microscope configuration information.
When regular STEM detectors can be used to navigate, the images produced from them may be used to set ROI scans using an image produced by a prior “survey” scan, following the spectrum imaging methodology, or regular STEM scans may be used to maximize read rates. When these approaches are used, a scripted DM plugin may be used for setting low-level Merlin parameters (Merlin DM Plugin devs, 2017), as shown in Figure 2b. Alternatively, real-space pixel sizes and scan ranges may be set and low-level DM commands used to configure and enable the scan. A scripted DM GUI has been developed, MERLIN_PixSTEM, to coordinate this with configuring the Merlin system to acquire data in the optimized continuous read–write mode and is shown in Figure 2c. Among the other features implemented, this plugin also allows different projection system settings to be saved and restored, enabling efficient switching between different detectors. The Merlin system includes two TCP/IP servers, shown in red in Figure 2a, one for setting and reading acquisition parameters and the second for image data transfer. The DM-scripted GUIs, indicated in cyan in Figure 2a, interface with the Merlin communication server through a separate TCP/IP C++ plugin (Merlin DM Plugin devs, 2017), shown in red. The TCP/IP plugin may be installed alone, allowing it to be used for many other communication purposes. For more advanced control of the Merlin system over TCP/IP, a Python implementation of Merlin TCP/IP commands has been developed (Merlin Interface devs, 2016).
An example of an additional STEM signal we collect is the noise correction (NC) signal, which may be used for gun-noise correction of the 4D dataset. The NC signal is produced by a current pickup attached to the condenser aperture and gives a measure of the gun emission. A similar approach to gun signal measurement was recently reported and shown to have good linearity to the probe current (House et al., Reference House, Tom Schamp and Yang2018). Correction of gun noise is particularly useful in intensity-based low contrast imaging modes, as shown in the bright field (BF) images of a mouse liver microtomed thin section in Figure 3. The image in Figure 3a is produced by summing the entire diffraction pattern (an example is shown in the inset) at each scan position. Sample contrast primarily arises due to incoherent Rutherford scattering of electrons to angles beyond the detector. The large circular feature in the bottom left corner is part of a mitochondrion organelle, while the darker spotted stripe structures are endoplasmic reticula studded with ribosomes.

Fig. 3. Gun noise correction in a BF STEM image produced from a 4D dataset from a thin fixed but unstained section of mouse liver, showing part of one cell including the end of a mitochondrion organelle and part of the rough endoplasmic reticulum. Measured (a) BF image and (b) the same image after gun noise correction using the fpd.tem_tools.nc_correct function. The inset in (a) shows the summed diffraction pattern on a logarithmic scale, while that in (b) shows the recorded gun noise. The acceleration voltage was 200 kV, the objective lens was off, the condenser aperture was 30 µm, the camera length was 600 cm, the convergence semi-angle was 13.1 mrad, and the pixel spacing was 3.7 nm.
The horizontal stripes in the as-measured image in Figure 3a are from short-period variations in the cold-FEG emission. The gun signal measured by the NC detector is shown in the inset of Figure 3b. Figure 3b itself shows the corrected image produced by minimizing the contrast introduced by these gun emission current variations using a linear gun-noise model. This reveals much more detail in the BF image than previously seen in Figure 3a. The residual horizontal features in the corrected image are most likely a result of variations in linearity of the NC detector signal from things like amplifier drift or external noise. Taking the corrected image as a reference, the power SNR of the uncorrected image, calculated using the implementation of the two-image method (Frank, Reference Frank1980) in the fpd.utils module, is 11 dB, giving a measure of the improvement in the image quality by applying gun-noise correction.
The Medipix3 data acquired following the above methodologies can be saved to disk on the acquisition computer in a flat binary format and may also be sent over the network using TCP/IP. The conversion of the binary data to more appropriate formats is discussed in the section "Data Storage". The network transfer of data has many potential uses, and we discuss these in the context of live data processing in the next section.
Live Data Processing
Live feedback from the data collected by a fast pixelated detector in a STEM acquisition is crucial for both optimizing imaging conditions and navigating to ROI in a sample. This is especially true for some modes of imaging where traditional STEM detectors may not produce useful contrast, such as when imaging magnetic features which are typically not visible in STEM without a custom segmented detector and a readout system (McGrouther et al., Reference McGrouther, Benitez Romero, McFadzean and McVitie2014). To facilitate real-time feedback, we developed the Python library fpd_live_imaging (FPD Live Imaging devs, 2015), which implements multiple common analysis routines and wraps processing routines from other libraries (fpd devs, 2015). Although the fpd_live_imaging package was developed for use with the Medipix3 detector and the Merlin readout system, its design is modular and can easily be extended to work with any detector. Our implementation takes advantage of the many cores available in modern CPUs by employing Python's multiprocessing library. Shared parameters and data are passed between the separate processes through “queue” objects or other shared memory. This approach enables good performance, even at very high data rates.
The internal workings of the package are outlined in Figure 4. The Medipix3 1R insertion and retraction mechanism (shown colored in purple, i) is controllable through a serial interface (shown in green) and is made possible through library function calls. As discussed in the previous section, the Merlin system (drawn in yellow, ii) can be interfaced with via two TCP/IP servers (shown in red), which are utilized by the fpd_live_imaging package (white, iii) to get data from the detector and to control the acquisition of data. The first step in the visualization is receiving the raw binary data from the Merlin TCP/IP data interface using the receive_data_medipix function (iv). This function runs a TCP/IP socket which gets the raw binary data and passes it along to a parser function. The function has its own CPU process to be able to handle the very high framerate of the Medipix3 detector. Due to the nature of the TCP/IP protocol, the raw binary images can be split into different fragments. These fragments are pieced together in the parse function, which results in the image in the form of a NumPy array (Oliphant, Reference Oliphant2006). The function also handles the bit depth of the data and the number of pixels in the detector, and also runs in its own separate CPU process.

Fig. 4. Schematic of the fast pixelated detector live visualization library architecture, showing the relationship between the Medipix3 detector and the retraction mechanism (purple, i), the Merlin readout system (yellow, ii), and the fpd_live_imaging library (white, iii). Multiple processing classes (blue, v) are implemented for scanning and imaging modes.
After having constructed the image in the form of the NumPy array, a copy is sent to any number of data processing classes. These data processing classes are shown in blue (v) in Figure 4 and can be separated into two categories based on the imaging mode: scanning and parallel. The scanning data classes include things like virtual bright field and annular darkfield, where the input detector image is reduced to a single output value. In the parallel data classes, the output image is the same size as the input one, and the processing methods include passing through the input image, a thresholded version of the input image, or a Fourier transformed image. All these run in separate CPU processes. In addition to the aforementioned processing classes, are ones for single pixel extraction, center of mass, and phase-correlation for electro- or magneto-static field imaging, and routines for HOLZ analysis. Virtual detector imaging and HOLZ data processing are covered in Part II of this work (Paterson et al., Reference Paterson, Webster, Ross, Paton, Macgregor, McGrouther, MacLaren and Nord2020b), while field mapping will be covered in Part III.
The processing time varies greatly, depending on the computational complexity of the routine (Nord et al., Reference Nord, Krajnak, Bali, Hlawacek, Liersch, Fassbender, McVitie, Paterson, Maclaren and McGrouther2016), and the choice of routine depends upon the nature of the sample. For example, in magnetic imaging, the integrated induction components perpendicular to the electron path can be determined from deflections in the position of the bright field disc (Chapman & Scheinfein, Reference Chapman and Scheinfein1999). The center of mass calculation provides good contrast in many cases but can be affected by the crystallinity of the sample due to intensity diffracted from the bright field disc to angles either outside or inside the detector collection angle (Chapman et al., Reference Chapman, McFadyen and McVitie1990). Phase- or cross-correlation (Krajnak et al., Reference Krajnak, McGrouther, Maneuski, O'Shea and McVitie2016) approaches can greatly improve upon this at the expense of computation time and can be crucial to detecting magnetic contrast in highly diffracting samples. On the other hand, single pixel extraction, where a single pixel on the edge of the disc is used as a measure of up to around pixel-level disc shifts, requires the minimum of processing and is orders of magnitude faster, taking approximately 2 µs when the ![]() $256\times 256$ scan position 12-bit dataset is in memory (Nord et al., Reference Nord, Krajnak, Bali, Hlawacek, Liersch, Fassbender, McVitie, Paterson, Maclaren and McGrouther2016). As each selected pixel gives a measure of a component of the integrated induction in a direction tangential to the disc, the use of only two pixels out of each diffraction image is sufficient to form a qualitative 2D vector map, which allows the user to at least navigate to an appropriate position, and set magnification and focus. Multiple processes may be run sequentially or simultaneously, allowing the tradeoff between runtime and sensitivity to be seen in real time.
$256\times 256$ scan position 12-bit dataset is in memory (Nord et al., Reference Nord, Krajnak, Bali, Hlawacek, Liersch, Fassbender, McVitie, Paterson, Maclaren and McGrouther2016). As each selected pixel gives a measure of a component of the integrated induction in a direction tangential to the disc, the use of only two pixels out of each diffraction image is sufficient to form a qualitative 2D vector map, which allows the user to at least navigate to an appropriate position, and set magnification and focus. Multiple processes may be run sequentially or simultaneously, allowing the tradeoff between runtime and sensitivity to be seen in real time.
The output data from any kind of processing is sent to a visualization class, which shows the result of the processing on the computer running the fpd_live_imaging package. Due to rescaling of the intensity to optimize the contrast, this visualization is qualitative, while the calculations themselves can be quantitative. This computer may be anywhere on the network. The visualization is separated into parallel and scanning modes, as shown in pink (vi) in Figure 4, and they also run in separate CPU processes. An example of the visualization GUI is shown in Figure 5b. In this case, the image is from thresholded center of mass analysis of data from a patterned DC sputtered 8 nm permalloy film capped with 4 nm of copper. The 2 µm discs were patterned with a Ga focused ion beam, and the contrast in the resulting structures shows they support magnetic vortices. A detailed study of the sample will be published elsewhere. The GUI has buttons for setting the brightness and contrast during the acquisition, and the analysis parameters can be tuned during imaging, allowing for live optimization of the required contrast. Alternatively, the processed data can be sent over TCP/IP to any computer on the network, for example, directly into Digital Micrograph.

Fig. 5. fpd_live_imaging's graphical user interface showing (a) the control window for the visualization and (b) a thresholded center of mass contrast of a patterned 8-nm permalloy film capped with 4 nm of copper. The contrast in the 2 µm discs represents the beam deflection along a single axis and shows that the discs support magnetic vortices. The inset in (b) shows the thresholded detector image.
All the above processes are orchestrated from the “Acquisition Control” class (shown in brown, vii in Figure 4), which handles the initialization and connection of all of these separate functions. For ease of use, the Acquisition Control class can be accessed through a GUI, as shown in Figure 5a. This allows for starting and stopping of the acquisition, modification of the scan parameters, the addition and removal of processing classes, modification of their parameters, and insertion and retraction of the detector itself.
The three separate stages described above, reading data from the detector, processing the images, and visualizing or sending the result over TCP/IP, are implemented in modular design, making simple the addition of new detector data sources, image processing classes, and visualization.
Data Storage
The principal issues when choosing a file format for fast pixelated detector data are common across data from all detectors: the ability to store the data with the dimensionality of the scan, store metadata along with the detector data, allow access to subsets of the data without reading the entire and often very large dataset into memory, support compression, and be an open format with read and write support across a variety of programming languages. An HDF5 (The HDF Group,1997–2018)-based format was chosen for our use since it meets all of the above requirements.
The HDF format has long been widely used in the synchrotron community and is increasingly being used in electron microscopy (de la Peña et al., Reference de la Peña, Ostasevicius, Fauske, Burdet, Prestat, Jokubauskas, Nord, Sarahan, MacArthur, Johnstone, Taillon, Caron, Migunov, Furnival, Eljarrat, Mazzucco, Aarholt, Walls, Slater, Winkler, Martineau, Donval, McLeod, Hoglund, Alxneit, Hjorth, Henninen, Zagonel and Garmannslund2018; EMD authors, 2019; Somnath et al., Reference Somnath, Smith, Laanait, Vasudevan, Ievlev, Belianinov, Lupini, Shankar, Kalinin and Jesse2019). It can be both read and written in a number of programming languages, including MatLab, C++, Python, Java, R, and Gatan Digital Micrograph through a third party plugin (Niermann, Reference Niermann2016). The HDF5 format consists of an arbitrary structure of hierarchies of groups containing further groups or datasets, enabling the relationship between data to be indicated by the file structure. For datasets, the data-type definitions are stored with the data, making it self-describing and ensuring maximum portability. Additionally, all groups and datasets can have attributes, allowing user and acquisition metadata to be stored along with the detector data in appropriate locations. The datasets may be of any number of dimensions, and so, it is ideal for multidimensional data from fast pixelated detectors when used in STEM or other acquisition modes.
HDF5 has in-built support for a variety of compression algorithms and other so-called “filters,” all providing transparent read and write access to the data. To allow access to subsets of the data without having to decompress the entire dataset, the dataset can be divided into smaller pieces and stored in a B-tree, a balanced hierarchical data structure, by enabling “chunking.” Figure 6a shows an example of the potential chunking of a one-dimensional (1D) scan dataset. The stack of images (shown on the left) occupy a 3D data “cube” (middle), with one axis being the scan dimension. On the right of panel (a), we show the same dataset with two chunks along each dimension, with each chunk in a different color. The dataset access sequence is summarized in Figure 6b. When indexing a chunked dataset, the B-tree is navigated, each chunk containing the required data is decompressed, and only the selected components are returned. For example, when reading the image slice shown by the blue dashed line in the right of Figure 6a, each of the top four chunks must be read.

Fig. 6. (a) Example of potential dataset chunking for data from a 1D scan stored in an HDF5 file. (b) Data indexing sequence for chunked data. (c–f) HDF5 chunk performance metrics for the ![]() $256 \times 256$ probe position STEM dataset from Figure 3 with a
$256 \times 256$ probe position STEM dataset from Figure 3 with a ![]() $256\times 256$ Medipix3 detector in a 12-bit mode. Level 4 GZIP compression was used. All panels show metrics for hypercube chunks (solid lines) and hyperrectangle chunks (dashed lines) with dimensions matching those of the detector (detY, detX). The inset in (c) shows a diffraction image on a logarithmic scale. The (e,f) read and (c) write times are the ratios of the values to those for the hypercube chunk length of 16 (marked by symbols). The compression ratios (d) are of the entire HDF5 file relative to only the raw Merlin binary file. The read times in (e) and (f) are those required to load a 128-sided hypercube or single slices of the dataset, respectively, into an in-memory NumPy array using h5py.
$256\times 256$ Medipix3 detector in a 12-bit mode. Level 4 GZIP compression was used. All panels show metrics for hypercube chunks (solid lines) and hyperrectangle chunks (dashed lines) with dimensions matching those of the detector (detY, detX). The inset in (c) shows a diffraction image on a logarithmic scale. The (e,f) read and (c) write times are the ratios of the values to those for the hypercube chunk length of 16 (marked by symbols). The compression ratios (d) are of the entire HDF5 file relative to only the raw Merlin binary file. The read times in (e) and (f) are those required to load a 128-sided hypercube or single slices of the dataset, respectively, into an in-memory NumPy array using h5py.
Chunk Size
When choosing a chunk size, a compromise is made between the cost of B-tree navigation, compression level, and data reading speed, with the optimum choice ultimately depending on the intended data access pattern. For STEM data, the diffraction pattern can be sparse and compression can be optimized by chunking in both the scan and image dimensions. Although most of the issues discussed in this section will be common to all detectors, we note that the Medipix detector has three relevant features which separate it from most others. The first two are that the detector can unambiguously detect single electron impacts, and that the data are read out free of addition noise, meaning that dark regions of images can be truly filled with zeros rather than noise. The third point is that data from the Medipix3 detector are zero padded to align it with common data types (e.g. 12-bit data are stored as 16-bit), allowing compression to achieve significant reductions in file size. For example, with a chunking of 16 along each axis of a 4D dataset, the in-built lossless GZIP compression at level 4 typically reduces the data size of a scanning acquisition of ![]() $256\times 256$ probe positions in a 12-bit mode from 8.6 to 2.7 GB.
$256\times 256$ probe positions in a 12-bit mode from 8.6 to 2.7 GB.
Many chunking strategies are possible, and here, we explore two of them: a hypercube with equal chunk sizes across all axes; and a hyperrectangle with equal chunk sizes along the scan axes, and detector axes chunk sizes matching the detector dimensions. The first approach gives the most uniform data access properties across different axes and, importantly, can improve data processing efficiency by allowing reduction of the volumes of data that must be read for some analyses. For example, with a chunk size of 16 along each axis, getting the direct beam in a dataset where it resides in four chunks would require loading into memory only those chunks, corresponding to only 1.6% of the total file. The second approach of hyperrectangles is more suited to applications which only ever access full images, since requesting even a single pixel from an image would require the entire image to be read by the HDF5 library, and, similarly, indexing a 2D slice perpendicular to the detector axis would require the entire dataset to be read, resulting in significant overheads.
Figures 6c–6f show three HDF5 performance metrics as a function of the chunk edge length for the two chunking approaches (hypercube: solid lines; hyperrectangle: dashed lines) using the liver sample data in Figure 3. The three metrics are normalized write time (Fig. 6c), compression ratio (Fig. 6d), and normalized read time (Figs. 6e, 6f). The write and read times were normalized with respect to the values for the hypercube chunk length of 16 (marked by symbols), while the compression ratio is with respect to the size of the raw Merlin data file. Hypercube chunks of length 16 are used as the reference because they are the default values in our implementation, as a result of it being a reasonable compromise for most applications.
For hyperrectangle chunks, the write time (Fig. 6c), compression ratio (Fig. 6d), and read time for a 128-side length hypercube (Fig. 6e) are mostly independent of chunk size, whereas there are significant variations for hypercubes. As the chunk size is increased, all three metrics start very high (poor) due to the very large number of chunks, and then either go through a minimum at a chunk length around 16–32 (write time and compression) before increasing slightly, or plateau (hypercube read time). With all else being equal, the hypercube read time for the largest chunk sizes in Figure 6e should be ![]() $4\times$ worse for the hyperrectangle than for the hypercube, due to the overhead from the HDF5 library having to read the entire dataset, and the actual value of 5.1 is close to this. Similarly, reading a subset of any of the chunk sizes shown in Figure 6e will result in performance reduction, with smaller chunk sizes being less affected.
$4\times$ worse for the hyperrectangle than for the hypercube, due to the overhead from the HDF5 library having to read the entire dataset, and the actual value of 5.1 is close to this. Similarly, reading a subset of any of the chunk sizes shown in Figure 6e will result in performance reduction, with smaller chunk sizes being less affected.
In Figure 6f we show the read times for indexing a single slice of the dataset, with the slice creating a detector image (thick lines) or a scan image (thin lines). As explained above, when indexing across a non-detector axis, the entire dataset must be read and this is the reason for the high and chunk size independent read times for the hyperrectangle approach (thick dashed line). Above a chunk length of around 16, all other datasets lie on top of one another and follow a linear relationship. Below this point, the hypercube read times begin to plateau and then increase, while the hyperrectangle chunked read times maintain the linear trend all the way to chunk lengths of 1. At this chunk size, reading a single image is ![]() $141\times$ faster than for our default setting of length 16 hyperchunks, which is not as fast as the
$141\times$ faster than for our default setting of length 16 hyperchunks, which is not as fast as the ![]() $256\times$ smaller data would predict, due to additional overheads. Thus, for reading single images, having chunks of single images is a simple strategy to maximize performance, at the expense of flexibility in reading the data in other ways. However, a very simple method that often allows for similar read speed while maintaining flexibility is to read images (when they are needed as full images) from a hypercube chunked dataset in numbers that are aligned to a chunk size. This is the approach of data processing using the fpd package, which is discussed in more detail in Part II (Paterson et al., Reference Paterson, Webster, Ross, Paton, Macgregor, McGrouther, MacLaren and Nord2020b). For the example in Figure 6f, taking length 16 hypercubes as the reference, reading
$256\times$ smaller data would predict, due to additional overheads. Thus, for reading single images, having chunks of single images is a simple strategy to maximize performance, at the expense of flexibility in reading the data in other ways. However, a very simple method that often allows for similar read speed while maintaining flexibility is to read images (when they are needed as full images) from a hypercube chunked dataset in numbers that are aligned to a chunk size. This is the approach of data processing using the fpd package, which is discussed in more detail in Part II (Paterson et al., Reference Paterson, Webster, Ross, Paton, Macgregor, McGrouther, MacLaren and Nord2020b). For the example in Figure 6f, taking length 16 hypercubes as the reference, reading ![]() $16\times 16$ images with hyperrectangles of length 1 takes
$16\times 16$ images with hyperrectangles of length 1 takes ![]() $0.95\times$ the reference time, while reading the same with hyperrectangles of length 16 takes
$0.95\times$ the reference time, while reading the same with hyperrectangles of length 16 takes ![]() $1.12\times$ the reference time. For markedly different datasets or where the data access pattern is known in advance, the optimum chunking may be somewhat different, and this can be set by the user at the point of conversion.
$1.12\times$ the reference time. For markedly different datasets or where the data access pattern is known in advance, the optimum chunking may be somewhat different, and this can be set by the user at the point of conversion.
Merlin Data
The Merlin readout software stores the detector and readout system parameters in a separate header file, and the detector data as a stream of uncompressed binary data, with each image containing a variable length header of acquisition parameters specific to that image. The MerlinBinary class from the fpd_file module of the fpd library (fpd devs, 2015; fpd demos devs, 2018) allows parsing of data files and array access to raw data using memory mapping and conversion to the HDF5 format. The scan parameters and metadata can be extracted from Digital Micrograph files acquired simultaneously with the diffraction patterns, or may be supplied separately. For the former case, the DM files are accessed through the HyperSpy library (de la Peña et al., Reference de la Peña, Ostasevicius, Fauske, Burdet, Prestat, Jokubauskas, Nord, Sarahan, MacArthur, Johnstone, Taillon, Caron, Migunov, Furnival, Eljarrat, Mazzucco, Aarholt, Walls, Slater, Winkler, Martineau, Donval, McLeod, Hoglund, Alxneit, Hjorth, Henninen, Zagonel and Garmannslund2018) and are also embedded in the HDF5 file as raw binary blobs for reuse in the proprietary DM software. All DM files are also stored in the HDF5 file in the open EMD format (EMD authors, 2019) (discussed in the next section). Examples of Merlin data converted to HDF5 format using the MerlinBinary class are available in the open data deposit for this work (Nord et al., Reference Nord, Webster, Paton, McVitie, McGrouther, MacLaren and Paterson2019b).
Figure 7a shows an example of a 2D scan when the acquisition is being triggered by the microscope scanning system. Images, indicated in green, are acquired on a regular scan grid. As discussed in the section "Medipix3 Data Acquisition", during the time when the beam is being moved from the end of one row to the start of the next, the “flyback” time, the DigiScan system continues to send triggers, causing additional images to be acquired. These are shown in red and may be excluded during data access and conversion to the HDF5 format with appropriate parameter settings.
The image data in the Merlin binary file is in C-order, that is, with the fastest moving index being in the last dimension, as depicted in Figure 7a. The HDF5 library is a self-describing one and returns datasets stored within it in the form appropriate for the library being used, but stores the data internally in C-order. C-order is also the default ordering in NumPy, and thus, we naturally store the multidimensional pixelated STEM datasets in C-order, with the first axes being the scan ones and the last two being the detector ones. Most pixelated STEM datasets are 4D and of the type described. However, the Medipix3 detector can be operated in color mode, where an additional axis representing multiple thresholds exists between the scan and detector axes. This axis, while not generally used in STEM acquisitions at present, is used for spectroscopic X-ray imaging, is useful for characterizing the detector performance using X-rays, and is supported by the fpd library.
While the HDF5 conversion is most appropriate for data archival and later processing of data acquired under all modes of operation, the MerlinBinary class also provides a memory mapped array interface to the data on disk for most, but not all, acquisitions. For example, 1-bit data acquired in raw mode is stored as 1-bit by the Merlin system and has the image segments out of order, and cannot currently be easily memory mapped. However, in most cases, this mode of access allows the dataset to be visualized or processed without conversion of the data on disk to the HDF5 format, which is particularly useful for checking datasets immediately after acquisition.
HDF5 File Structure
Figure 7b shows an overview of one of the HDF5 files read in HDFView (The HDF Group, 2017), a Java GUI program that allows, among many other things, quick inspection of HDF file contents. Information from the binary headers such as the DAC values, the image exposure, the comparator threshold values, and the acquisition time are automatically extracted and included as datasets in the HDF5 file. During conversion to the HDF5 format, a sum of both the image and diffraction dimensions are generated and stored in the HDF5 file as separate datasets, resulting in bright field and diffraction sum images. These images may be used for data inspection and navigation without having to process the entire dataset in order to re-render them every time the file is loaded. Also during conversion, any bad pixels in the detector (hot, noisy, or dead) may be replaced by interpolated values when the user supplies a mask image, which can be important for optimization of some forms of data analysis. Additionally, an arbitrary function may be applied to each image during conversion. This has many possible uses, including to correct for unwanted image shifts due to descan and to apply image transforms to correct image distortion.
The detector data, including images created from it, is stored in the EMD format, a simple open subset of the HDF5 format, created by a specific collection of datasets and attributes (EMD authors, 2019). The EMD datasets may be read in software such as EMD viewer (EMDViewer devs, 2015), HyperSpy (de la Peña et al., Reference de la Peña, Ostasevicius, Fauske, Burdet, Prestat, Jokubauskas, Nord, Sarahan, MacArthur, Johnstone, Taillon, Caron, Migunov, Furnival, Eljarrat, Mazzucco, Aarholt, Walls, Slater, Winkler, Martineau, Donval, McLeod, Hoglund, Alxneit, Hjorth, Henninen, Zagonel and Garmannslund2018) and, of course, any HDF5 reader. Many utility functions are also provided in the fpd.fpd_file module, allowing conversion to other formats and extraction of data. These include ones to access datasets as a namedtuple (fpd_to_tuple) which allows for indexing or tab completion and one for accessing the same as HyperSpy objects (fpd_to_hyperspy).
Other projects have also adopted or are compatible with the EMD format (de la Peña et al., Reference de la Peña, Ostasevicius, Fauske, Burdet, Prestat, Jokubauskas, Nord, Sarahan, MacArthur, Johnstone, Taillon, Caron, Migunov, Furnival, Eljarrat, Mazzucco, Aarholt, Walls, Slater, Winkler, Martineau, Donval, McLeod, Hoglund, Alxneit, Hjorth, Henninen, Zagonel and Garmannslund2018; Savitzky et al., Reference Savitzky, Zeltmann, Barnard, Brown, Henderson and Ginsburg2019), and we note that work is currently underway by the LiberTEM project (LiberTEM devs, 2018) to include (transmission) electron microscopy data in the NeXus data format (Könnecke et al., Reference Könnecke, Akeroyd, Bernstein, Brewster, Campbell, Clausen, Cottrell, Hoffmann, Jemian, Männicke, Osborn, Peterson, Richter, Suzuki, Watts, Wintersberger and Wuttke2015). The Nexus format is another open subset of the HDF5 file format, originally developed to improve the data exchange within the fields of neutron, X-ray, and muon experiments. Having a common data format across all of these fields would be beneficial, since it would make the sharing of data and of processing routines that rely on metadata easier than is currently the case. This format could clearly be used within the suite of tools described in this paper.
Merlin Equipped Scanning Precession Electron Diffraction Systems
A recent development in precession electron diffraction (PED) is the use of fast pixelated detectors (Midgley & Eggeman, Reference Midgley and Eggeman2015). One such example is the work by NanoMEGAS to incorporate a Medipix3 DED into their DigiSTAR precession system in order to enable high fidelity recording of diffraction patterns in scanning PED (SPED) applications, as has been demostrated in recent work (MacLaren et al., Reference MacLaren, Frutos-Myro, McGrouther, McFadzean, Weiss, Cosart, Portillo, Robins, Nicolopoulos, del Busto and Skogeby2020). Additional benefits brought about by the use of DEDs in SPED will be discussed in Part II of this work (Paterson et al., Reference Paterson, Webster, Ross, Paton, Macgregor, McGrouther, MacLaren and Nord2020b). We note here, however, that the properties of the 4D dataset obtained by such a system are in many ways equivalent to those of 4D non-SPED datasets, and so many of the same issues of data access, storage, and processing apply here too. To enable these datasets to be more easily used, the topspin_app5_to_hdf5 function of the fpd.fpd_io module allows the conversion of data originally recorded in the native NanoMEGAS TopSpin app5 format to one almost identical to the HDF5 format outlined above. Alternatively, the Merlin acquisition software can be programmed to output the data directly to a raw file, whilst the acquisition is being performed and controlled by the TopSpin software. The main differences between the converted files is the inclusion of precession metadata instead of Medipix3 metadata in SPED datasets, and the absence of simultaneously acquired DM datasets.
Summary
The use of fast pixelated detectors for electron imaging is a burgeoning field with the prospect of revolutionizing many aspects of TEM and, in particular, scanning TEM. We have presented many of the key tools needed to (i) acquire data from fast pixelated detectors, (ii) analyze in real time the data from one and visualize the results, and (iii) store data from them in an optimized way.
The software packages presented are hosted in public repositories (fpd devs, 2015; FPD Live Imaging devs, 2015; pixStem devs, 2015; Merlin Interface devs, 2016; Merlin DM Plugin devs, 2017; fpd demos devs, 2018), are under active development, and contain many more features than are covered in this short publication. Many of the data analysis algorithms in these libraries are applicable to data from any detector. Most of these packages are provided under an open source licence, allowing transparency of the algorithms implemented and for them to be continually improved upon by the community.
Part II of this paper will cover post-acquisition processing and visualization of data from fast pixelated detectors, with examples of their application to the study of the structure of materials studied using scanning TEM (Paterson et al., Reference Paterson, Webster, Ross, Paton, Macgregor, McGrouther, MacLaren and Nord2020b). A final part III will cover aspects related to differential phase contrast analysis.
Availability of data and materials
The source data and scripts to analyze the data and produce the results presented here are publicly available at https://doi.org/10.5281/zenodo.3479124 (Nord et al., Reference Nord, Webster, Paton, McVitie, McGrouther, MacLaren and Paterson2019b). The software presented in this work is available in public Git repositories (fpd devs, 2015; FPD Live Imaging devs, 2015; pixStem devs, 2015; Merlin Interface devs, 2016; Merlin DM Plugin devs, 2017; fpd demos devs, 2018), which also document the source of contributions to the code. Enquiries about the MERLIN_PixSTEM DM plugin should be directed to Damien.McGrouther@glasgow.ac.uk.
Acknowledgments
We thank Diamond Light Source, especially Dr. Chris Allen and Prof. Angus Kirkland, for access and support in the use of the electron Physical Science Imaging Centre E02 microscope (proposal no. EM16952) that provided the atomic resolution SrTiO3 data presented here; Quantum Detectors Ltd. for Medipix3 detector support; Dr. Bruno Humbel from Okinawa Institute of Science and Technology and Dr. Caroline Kizilyaprak from the University of Lausanne for providing the liver sample; Dr. Kayla Fallon from the University of Glasgow for the sample data shown in Figure 5b, and Dr. Sinan Azzawi and Prof. Del Atkinson from University of Durham for providing the sample; Dr. Ingrid Hallsteinsen and Prof. Thomas Tybell from the Norwegian University of Science and Technology (NTNU) for providing the SrTiO3 sample shown in Figure 3; and NanoMEGAS for the loan of the DigiSTAR precession system and TopSpin acquisition software.
Author contributions statement
G.W.P. and M.N. were the principal authors of the libraries reported herein (details of all contributions are documented in the repositories), and have made almost all of these available under open source licence GPLv3 for the benefit of the community. R.W.H.W., S.McV., I.M., K.A.P., and D.McG. have all contributed through acquisition and analysis of data and through participation in the revision of the manuscript. R.W.H.W. and K.A.P. have also made contributions to the source codes in these libraries. G.W.P and M.N. have led the drafting of this manuscript.
Financial support
The performance of this work was mainly supported by the Engineering and Physical Sciences Research Council (EPSRC) of the UK via the project “Fast Pixel Detectors: a paradigm shift in STEM imaging” (grant no. EP/M009963/1). G.W.P. received additional support from the EPSRC under grant no. EP/M024423/1. M.N. received additional support for this work from the European Union's Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement no. 838001. The studentship of R.W.H.W. was supported by the EPSRC Doctoral Training Partnership grant no. EP/N509668/1. S.McV. was supported by EPSRC grant no. EP/M024423/1. I.M. was supported by EPSRC grant no. EP/M009963/1. The studentship of K.A.P. was funded entirely by the UK Science and Technology Facilities Council (STFC) Industrial CASE studentship “Next2 TEM Detection” (no. ST/P002471/1) with Quantum Detectors Ltd. as the industrial partner. D.McG. was also supported by EPSRC grant no. EP/M009963/1. As an inventor of intellectual property related to the MERLIN detector hardware, he is a beneficiary of the license agreement between the University of Glasgow and Quantum Detectors Ltd. The development of the integration of TopSpin with the Merlin readout of the Medipix3 camera has been performed with the aid of financial assistance from the EPSRC under grant no. EP/R511705/1 and through direct collaboration between NanoMEGAS and Quantum Detectors Ltd.