The acronym GWAS stands for genome-wide association study and is pronounced to sound like ‘gee-wass’. This term refers to a new large-scale molecular genetic research approach that has, over the past 5 years, made major contributions to advancing our understanding of many common human diseases, including diabetes, heart disease, inflammatory bowel disease, various cancers and rheumatoid arthritis. The studies are starting to provide insights into the aetiology and pathogenesis of major psychiatric illness, including the biological relationships between the traditional clinical diagnostic categories. This article outlines the key take-home messages emerging from the research and the implications for clinical psychiatry.
Some useful terms to remember
As in many research fields, some technical words crop up repeatedly and can sound confusing. Some readers may therefore find it useful to refresh acquaintance with several common terms and basic genetic concepts (a brief glossary is provided in Table 1). Phenotype refers to the observable characteristics (or symptoms of illness) under consideration. Genotype refers to the set of genes an individual possesses that are relevant to the phenotype being considered. All human phenotypes are the end products of the interplay between genes and environment. This applies to normal human traits, such as height, weight and blood lipid levels, as well as personality and behavioural traits. It also applies to illnesses such as heart disease, diabetes, asthma, cancers and all of the major psychiatric illnesses.
TABLE 1 Terms commonly used in psychiatric genetics

At a conceptual level, genes are the basic units of inheritance that are passed from parent to child. At a molecular level, a gene is that part of a deoxyribonucleic acid (DNA) molecule containing the information that allows cells to make proteins. The genetic information is coded in the sequence of the nucleotide bases that make up the DNA molecule. Ribonucleic acid (RNA) is an intermediate molecule, chemically rather similar to DNA, that transfers genetic information between DNA and proteins. Proteins are the basic building blocks for all cells and tissues: they allow cells to ‘work’.
Genetic variation
Within each gene there may be DNA sequence differences between individuals that make the effects of that gene (and the corresponding protein) different for any given person. Any difference is usually only slight, but sometimes the difference in protein function can be dramatic. Different forms of a given variation in DNA sequence are called alleles. There are many different sorts of genetic variation. From the viewpoint of how common a variant is within the population, some variants are very rare (these are usually called mutations) and some are common (usually called polymorphisms). Regarding the length of a sequence of DNA involved in the variant, the smallest type of genetic variant is a difference at a single nucleotide base. A common single base-pair variant is called a single nucleotide polymorphism (SNP; pronounced ‘snip’); this is the main variant studied within GWAS. Typically, there is about one such variant every 1000 bases. Genetic differences may be larger, going from differences at a small DNA sequence (e.g. two, or a few base pairs in a row), through differences at thousands of base pairs in a row, right up to differences in whole chromosomes. Over the past few years there has been a great deal of interest in the intermediate size of genetic variants (thousands up to a few millions of base pairs). They have been found to be more common than had previously been thought and they can be important in disease susceptibility. Such variants are known as structural genomic variants or copy number variants (CNVs) and will be mentioned again later because they can be studied using GWAS. Box 1 contains a little more information about genetic variation.
BOX 1 A little more about genetic variation
Since each individual has two copies of each autosomal gene (derived from each parental chromosome) they also possess two alleles at any given locus (the location of a gene on a chromosome), the combination of which makes up the genotype. Individuals in the population differ in the specific DNA base sequence at many locations in their genomes. Much of this variation is ‘silent’ and has no effect but some of the variation influences the expression or function of proteins and thereby influences normal human traits and/or disease susceptibility.
What is a GWAS?
In the GWAS approach, a very large number of SNPs (usually hundreds of thousands or millions) is examined in a large number of individuals (typically many thousands) in order to provide an acceptable level of genetic information across all the chromosomes (the whole ‘genome’) (Reference Corvin, Craddock and SullivanCorvin 2010). The technical advance that has made this possible is the availability of genotyping ‘chips’ which can characterise DNA sequence variation at many hundreds of thousands of SNPs for modest cost. Owing to an important biological property of chromosomes (known as linkage disequilibrium, the correlation of genetic variants that are located close together), GWAS provide excellent information for a substantial proportion of the common DNA variants that occur in humans. In other words, directly genotyping, for example, 1 million SNPs can provide information about many other SNPs (perhaps another 4 million) that were not directly genotyped. Genome-wide association studies can also provide information about rare CNVs. However, there are some very important types of rare genetic variant for which GWAS provide no good information. Specifically, rare variants that are a change at a single base of the DNA sequence (so-called point mutations) are not detected in GWAS. (Recall that it is common single base variants that GWAS are designed to detect – any single base variant that has a population frequency of, for example, only 1 in 1000 is undetectable by GWAS). However, we know that such rare variants can have an impact on biological function and some can influence disease risk. It is therefore extremely important to recognise that GWAS cannot detect much of the rare variation that may influence disease susceptibility, even if the rare variants had a very large effect. Such rare variants require approaches based on sequencing (see section later).
Genome-wide association studies are used with either a large sample of unrelated cases and unrelated controls, or with a large family-based sample in which many of the individuals are related. In practice, most GWAS are of the unrelated case–control design. One reason is that adequately powered GWAS for common diseases require very large sample sizes, and unrelated case–control samples are usually much easier and cheaper to collect than are family-based samples. Another important practical consideration is that case–control designs can use a single large common set of controls, the allele frequencies in which can be contrasted with many different disorders. This is more economically attractive than family designs in which the controls are unique to that study. This ‘shared controls’ design was pioneered in the Wellcome Trust Case Control Consortium (2007) study of 2000 cases for each of the 7 common diseases, which were compared with 3000 shared ‘controls’ for 500 000 common DNA variants. Bipolar disorder was one of the seven diseases studied; the others were: coronary artery disease, Crohn's disease, hypertension, rheumatoid arthritis, and types 1 and 2 diabetes.
The (not very distant) future: whole genome sequencing
An important limitation of GWAS has been stressed above – namely that, although providing good information about common genetic variation (polymorphisms), they do not provide information about the vast majority of the rare variation (e.g. point mutations). The perfect genetic analysis would provide complete information at every variable point in the genome. We are not quite there yet with the technology but this will become a realistic possibility over the next few years with so-called ‘next generation sequencing’, whereby it will be feasible to determine the full DNA sequence for each person for an acceptable cost (Reference MardisMardis 2008).
Key messages for clinical psychiatrists about findings from GWAS
The following sections outline major points that will help psychiatrists to understand the direction of research and to answer patients' questions.
GWAS is a powerful method for studying common diseases
The original proof of principle for GWAS in human disease was provided by the identification of the gene encoding complement factor H as a risk locus for age-related macular degeneration (Reference Klein, Zeiss and ChewKlein 2005). This study was highly atypical in that the risk variant identified had a relatively large effect size that was detectable in a mere 96 cases and 50 controls typed for only about 116 000 SNPs. Subsequently, GWAS have resulted in the identification of a large number of alleles which have been confidently associated with common diseases. These include susceptibility alleles for non-psychiatric diseases such as asthma, coronary artery disease, atrial fibrillation, Crohn's disease, rheumatoid arthritis, type 1 and type 2 diabetes, obesity, prostate cancer, breast cancer and coeliac disease (Reference Petretto, Liu and AitmanPetretto 2007; Reference Corvin, Craddock and SullivanCorvin 2010). It also includes susceptibility alleles for psychiatric disorders, including schizophrenia, bipolar disorder, Alzheimer's disease and autism.
GWAS are delivering robust findings for psychiatric disorders
As mentioned in the preceding section, GWAS have already contributed to the identification of susceptibility alleles for major psychiatric disorders (Reference Owen, Craddock and O'DonovanOwen 2010). Among the earliest convincing GWAS findings, reported in 2008, was a bipolar disorder study of approximately 10 000 individuals that showed strong (genome-wide significant) evidence for association with susceptibility to bipolar disorder at variants within two genes involved in ion channel function: ANK3 (encoding the protein Ankyrin G) and CACNA1C (encoding the alpha-1C subunit of the L-type voltage-gated calcium channel) (Reference Ferreira, O'Donovan and MengFerreira 2008). A similar study in nearly 20 000 individuals showed strong evidence for association with susceptibility to schizophrenia at a variant within ZNF804A (encoding a zinc finger transcription factor) (Reference O'Donovan, Craddock and NortonO'Donovan 2008). As with all research, further study and replication of results is important, and subsequent work has provided support for these findings, as well as highlighting further loci of interest (some of which are shown in Table 2) (Reference Ripke, Sanders and KendlerRipke 2011; Reference Sklar, Ripke and ScottSklar 2011). It is important to stress that the molecular mechanisms that influence risk are not yet understood. The field is moving rapidly. At the time of writing, approximately six loci have been reported at genome-wide levels of statistical significance for bipolar disorder and about ten loci for schizophrenia.
TABLE 2 Some loci that have genome-wide significant statistical support in genome-wide associated studies in psychosis

Effect sizes are usually small, so large samples are needed
A very clear message from the many GWAS of common diseases so far is the importance of large samples powered to detect small effect sizes. This is consistent with theoretical predictions and, with few exceptions, the effect sizes that have been identified in studies of common diseases have been in the small range. For example, in the Wellcome Trust Case Control Consortium GWAS of seven common diseases (2007), per allele odds ratios of identified loci were in the range of 1.2–1.5 (meaning the risk allele increases susceptibility by about 20–50% at most). To have reasonable power to detect such loci requires samples of the order of 2000 cases and 2000 controls or larger. Many of the loci identified more recently have smaller effect sizes, which require substantially larger samples, in the tens of thousands.
For any given disease there are many susceptibility alleles and genes
For each of the common familial diseases that have been studied using GWAS, it is clear that there are many susceptibility loci and that they have a range of allele frequencies and effect sizes. Thus, at any point in time, the available data (usually summarised via a global meta-analysis) provide only a partial picture of the full genetic variation that influences susceptibility to that illness. Larger samples allow more loci to be discovered. For example, in Crohn's disease more than 80 susceptibility loci have been robustly implicated by GWAS. To date, studies of schizophrenia and bipolar disorder have robustly demonstrated fewer than 20 susceptibility loci, but the pattern of association within the data shows that there are probably many hundreds or thousands of common alleles that influence susceptibility to these disorders (Reference Purcell, Wray and StonePurcell 2009). An increasing number can be robustly identified as the number of individuals investigated increases.
Genetics will not form the basis for classification or diagnosis, but will help in moving towards more useful nosological entities
Molecular genetics will never provide a simple, gene-based classification of psychiatric illness (as it will not for other common familial illnesses) (Reference KendlerKendler 2006; Reference Craddock, Kendler and NealeCraddock 2009a). The notion that there is a ‘gene for …’ one or more psychiatric disorders is inappropriate and unhelpful. Rather, there is a complex relationship between genotype and phenotype that involves multiple genes and environmental factors, together with random variation. Nonetheless, molecular genetic findings can be expected to help delineate the relationship between specific biological pathways/systems and broad patterns, or domains, of psychopathology (Reference Craddock and OwenCraddock 2010). A precedent for such insights from genetic studies is already emerging from GWAS in other areas of medicine that have revealed unforeseen biological relationships among different autoimmune diseases (Reference Lettre and RiouxLettre 2008). It is clear that genetic findings will not map cleanly onto the existing diagnostic categories and we can expect that genetic associations may assist us in finding more useful and valid nosological entities.
At a genetic level, psychiatric disorders are not fundamentally different from non-psychiatric disorders
Although the phenotype issues provide a particular challenge for psychiatric genetics, findings from genetic epidemiology, such as familial recurrence risks and estimates of heritability, show that many types of major psychiatric illness are among the most genetically influenced of human traits and diseases (Reference McGuffin, Owen and GottesmanMcGuffin 2002). As for other disorders, it is likely that a range of mechanisms influence genetic risk, including common polymorphisms, rare mutations and structural rearrangements. There are no strong theoretical reasons to expect that the genetic mechanisms underlying major psychiatric illness will be qualitatively different from those underlying non-psychiatric disorders. The findings being delivered by GWAS are consistent with this. This helps remind us that psychiatric disorders are not, in principle, different from non-psychiatric disorders.
Rare CNVs have been robustly associated with risk of schizophrenia and other psychiatric and non-psychiatric disorders
As mentioned earlier, GWAS data-sets can be used to identify CNVs within individuals and test for association of one or more such variants with susceptibility to illness. Using this approach, it has been shown that some rare, large CNVs increase the risk for schizophrenia. Typically, the effect size is substantially larger than for the common SNP susceptibility alleles. For example, a CNV that disrupts the gene NRXN1 increases the risk for schizophrenia about eightfold (Reference Kirov, Rujescu and IngasonKirov 2009). NRXN1 encodes the protein neurexin that acts at synapses and is involved in the development and maintenance of normal brain functioning by mediating signalling across the synapse and affecting the properties of neural networks by specifying synaptic functions. As for the common SNP susceptibility alleles mentioned earlier, the risk CNVs are not disease/disorder-specific. For example, the NRXN1 CNV also increases the risk of autism and intellectual disability. Typically, the CNVs shown to be associated with risk of schizophrenia are very large, disrupt multiple genes and are also associated with a range of possible neuropsychiatric and non-psychiatric phenotypes. Such findings serve as a reminder that psychiatry is very much part of medicine and that brain dysfunction expresses itself in clinical pictures that can cut across current psychiatric subspecialties.
There is overlap in genetic susceptibility across traditional psychiatric diagnostic categories
One of the most striking and interesting early observations from GWAS has been the lack of diagnostic specificity for some of the best-supported susceptibility loci for schizophrenia and bipolar disorder (Reference Williams, Craddock and RussoWilliams 2011). For example, the risk allele at CACNA1C, identified originally in studies of bipolar disorder, has been shown also to increase the risk for schizophrenia and for (non-bipolar) recurrent major depression (Reference Green, Grozeva and JonesGreen et al 2010). This suggests that the same underlying mechanism may play a role in multiple traditional diagnostic categories. Although this does not, of itself, invalidate the traditional diagnostic groups, it strongly suggests the possibility of more biologically valid diagnostic entities that are based on the underlying pathogenesis and which may cut across existing descriptive categories. Identification of such categories would be good for patients and good for psychiatry.
Identifying susceptibility loci helps to pinpoint biological systems involved in illness
In the earlier history of psychiatric genetic association studies, it was usual to study specific variants within candidate genes – that is, genes that were suspected a priori to be involved in illness susceptibility. Examples were genes encoding dopamine receptors or the serotonin transporter; the selection of plausible candidates was based on a specific hypothesis about disease pathogenesis, often based on extrapolation from knowledge of the action of a drug that is effective in treatment. The enormous limitation of such an approach was (and remains) the lack of understanding of pathogenesis of psychiatric illness – which is one of the major rationales for using a systematic genetic approach such as GWAS to identify the mechanisms of pathogenesis. What has been striking from the results of GWAS is that the previously suspected candidates have not been implicated. Rather, unsuspected and novel genes, and hence, proteins and potential pathways, have been implicated (Table 2). This will, of course, open up entirely new avenues for study and development of therapeutic and preventative approaches. Here it is worth noting explicitly that understanding biological mechanisms does not mean that the therapeutic and preventative approaches that follow will all be drug based; psychological and lifestyle interventions are also likely to flow from improved understanding.
The simplest type of genetic analysis of GWAS data considers one SNP at a time. However, methods of analysis of GWAS data exist that seek to identify patterns of association across the whole genome that delineate biological pathways involved in susceptibility to illness, rather than just focusing on single loci or even single genes. Perhaps the most consistent and interesting finding to emerge from such an approach to date is the involvement of L-type voltage-gated calcium channels in susceptibility to bipolar disorder (Reference Sklar, Ripke and ScottSklar 2011), with evidence accumulating of a wider involvement in other psychiatric phenotypes.
No finding yet warrants clinical genetic testing but some may do so in the near future
Consider the susceptibility allele for bipolar disorder and other psychiatric phenotypes that has been robustly identified within CACNA1C (Box 2). It is common in the general population (allele frequency 30%) and is associated with a very small increase in risk to those carrying it: the risk is increased by about 18%. Clearly, most of those with the risk allele do not develop bipolar disorder and this highlights that many other factors (genetic and non-genetic) must be involved in influencing whether a particular person becomes ill and when. It should be intuitively obvious that this would not be helpful in predictive testing, at least if used on its own. In contrast, knowledge that CACNA1C is involved in the pathogenesis of bipolar disorder provides new avenues for research using a whole host of research approaches to better understand illness (Reference Craddock and SklarCraddock 2009b).
BOX 2 Findings at CACNA1C: a glimpse of things ahead?
As mentioned in the main text, the association signal within the CACNA1C gene with susceptibility to bipolar disorder occurs with an allele that is present in the normal population with a frequency of about 30%. The association is statistically highly significant but the effect size is very small.
A fascinating observation is that a rare mutation in the coding sequence of CACNA1C (i.e. a variant that changes the amino acid sequence of the calcium channel protein) causes a multisystem developmental disorder, Timothy syndrome, which affects many tissues, including heart tissue, causing cardiac conduction defects (perhaps not surprisingly for a calcium channel protein). Of enormous interest for us in psychiatry is that 80% of adults with Timothy syndrome have autism spectrum diagnoses. Thus, a dramatic change of the protein sequence can manifest in a high proportion of individuals as autism spectrum disorder, whereas a common variation outside the coding sequence exerts a modest effect on risk of bipolar disorder and other psychiatric illness. This tantalising observation gives hope that, as we delineate the pathways and mechanisms that contribute to disease, we may better understand why psychiatric phenotypes are so variable and commonly co-occur (e.g. mood disorder and autism spectrum disorder) – and, perhaps, grasp some of the associations seen between psychiatric and non-psychiatric illness (such as mood disorder and heart disease).
As just explained, although a common allele within the gene CACNA1C is robustly associated with susceptibility to bipolar disorder and other psychiatric phenotypes, there is no immediate clinical utility in testing a person for the presence or absence of the common risk variant. The same reasoning applies to the other common variants that have been robustly associated with risk of psychiatric illness.
In contrast, because of the larger effect sizes and also the potential for increasing the risk of physical disorders, it is possible that testing for rare CNVs that are associated with disease risk could have clinical benefits in the foreseeable future. For example, testing a person with a diagnosis of schizophrenia for the presence of a schizophrenia-associated CNV that is known also to increase the risk of congenital heart disease might be beneficial in targeting further cardiac investigation that could bring overall benefits for the patient's healthcare and quality of life. However, as with any clinical test, substantial work is required to determine potential clinical benefits and potential disadvantages before the test enters routine services. Those working in the psychiatry of intellectual disability are, of course, already very familiar with including genetic investigation, when appropriate, within the clinical assessment. As knowledge develops, other parts of psychiatry will need to be willing to embrace new technologies if they are shown to be clinically beneficial.
Conclusions
A range of genetic and non-genetic research approaches is required to help us better understand the major biological, psychological and social processes that contribute to psychiatric illness.
Together with complementary research approaches, the ongoing major investments of time and money in GWAS for psychiatric disorders has the potential to identify pathways involved in illness and help psychiatry move towards approaches to diagnosis and treatment that are grounded in a better understanding of pathogenesis (Reference Craddock and OwenCraddock 2010). This would be of great benefit to patients.
MCQs
Select the single best option for each question stem
-
1 Which of the following genes has been implicated in both bipolar disorder and Timothy syndrome:
-
a MAOA
-
b CACNA1C
-
c TF4
-
d COMT
-
e NRXN1.
-
-
2 Which of the following is not true about GWAS:
-
a GWAS are useful for detecting common genetic variants that influence susceptibility of illness
-
b GWAS are useful for detecting rare CNVs that influence susceptibility to illness
-
c GWAS are useful for detecting rare single base mutations that influence susceptibility to illness
-
d GWAS approaches have already identified more than 80 susceptibility loci for Crohn's disease
-
e GWAS approaches have already identified more than ten susceptibility loci for mood and psychotic disorders.
-
-
3 Which of the following is true:
-
a most genetic variants cause illness
-
b most people with a diagnosis of schizophrenia have large CNVs that are thought to cause the illness
-
c genetic variants that have been found to be associated with psychiatric illness are usually highly diagnosis-specific
-
d recent genetic findings suggest that environmental factors are largely unimportant in the development of psychiatric illnesses
-
e it is likely that there are hundreds or thousands of genetic loci that influence susceptibility to major psychiatric disorders.
-
-
4 Which of the following is true:
-
a GWAS typically analyse about 25 000 000 SNPs on a genotyping chip
-
b GWAS are so powerful that they can show clear-cut positive findings with about 50 cases and 50 controls for psychiatric disorders
-
c GWAS were originally developed in the 1990s
-
d sequencing of the whole genome will be more useful than GWAS because it provides information that GWAS cannot
-
e it is very unlikely that whole genome sequencing will be widely available within the next decade.
-
-
5 Which of the following genes or genetic loci has not been strongly associated in GWAS with psychiatric illness (at least to date):
-
a MHC
-
b ZNF804A
-
c MIR137
-
d CACNA1C
-
e COMT.
-
MCQ answers
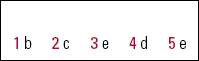
| 1 b | 2 c | 3 e | 4 d | 5 e |
Acknowledgements
I am grateful to all colleagues and collaborators in the Wellcome Trust Case Control Consortium and am indebted to the many individuals who have participated in, and helped with, research, particularly those involved with the Bipolar Disorder Research Network and the National Centre for Mental Health.





eLetters
No eLetters have been published for this article.