INTRODUCTION
An essential assumption in modelling the spread of infectious diseases is that the force of infection, which is the probability for a susceptible to acquire the infection, varies over time as a function of the level of infectivity in the population [Reference Anderson and May1]. For many infectious diseases, the force of infection is also known to depend on age. The equation describing the dependence of the force of infection on age and time is given by
The coefficients β(a, a′) are called the transmission coefficients and I(a′, t) is the number of infectious individuals at age a′ and time t. These transmission coefficients combine epidemiological, environmental and social factors affecting the transmission rate between an infective of age a′ and a susceptible of age a [Reference Anderson and May1, Reference Halloran, Watelet and Struchiner2]. For the discrete case with a population divided into a finite number, say n, of age groups, Anderson & May [Reference Anderson and May3] introduced the WAIFW (Who Acquires Infection From Whom) matrix in which the ij th entry of the matrix, βij, is the transmission coefficient from an infective in age group j to a susceptible in age group i. Let Ī i be the total number of infectious individuals in the ith age group at time t, i=1, …, n, then the age- and time-dependent force of infection can be approximated by the matrix product
Here, Ī=(Ī 1, …, Ī n) is the vector in which the ith element is the number of infectious individuals (prevalence of infectivity) in age group i, λ=(λ1, …, λn) is the vector in which the ith element is the force of infection specific to age group i and W is a known WAIFW matrix. The configuration of the WAIFW matrix represents a priori knowledge (or assumptions) about the mixing patterns in the population. Several configurations are discussed in the literature (see e.g. [Reference Anderson and May1, Reference Halloran, Watelet and Struchiner2, Reference Farrington, Kanaan and Gay4–Reference Wallinga6]). For example, for a model with five age groups the WAIFW matrix W 1 in equation (3) represents a mixing pattern for which individuals are mixing only with individuals from their own age group (assortative mixing [Reference Greenhalgh and Dietz5]) with a specific age-dependent transmission coefficient while W 2 represents a mixing pattern similar to W 1, also accounting for an additional mixing of individuals with individuals of other age groups with a ‘background’ transmission coefficient:

Note that both matrices have five unknown parameters and both are symmetric. For each of these contact structures, the number of parameters is equal to the number of age groups. This is a condition to have a solution [Reference Anderson and May3]. Suppose that the population is divided into n age groups and let ![]() =(
=(![]() 1, …,
1, …, ![]() n) be the estimated vector of age-specific force of infection in each age group. If the structure of the WAIFW matrix is known and consists of n unknown parameters, the WAIFW matrix can be estimated using the equality
n) be the estimated vector of age-specific force of infection in each age group. If the structure of the WAIFW matrix is known and consists of n unknown parameters, the WAIFW matrix can be estimated using the equality

with N the total population size, D the mean duration of infectiousness, L the life-expectancy at birth, and
Here, a i−a i−1 is the width of the ith age group. Hence, as long as the WAIFW matrix has a known configuration with n unknown parameters, the parameter vector β=(β1, …, βn) is identifiable. Note that we expect that βi⩾0, i=1, 2, … , n.
The basic reproductive number R 0 can be computed as the dominant eigenvalue of a matrix for which the ijth entry is the basic reproductive number ![]() , specific to the transmission from an infective in age group j to a susceptible in age group i. More precisely,
, specific to the transmission from an infective in age group j to a susceptible in age group i. More precisely, ![]() where D is the duration of infectiousness assumed independent of age and N i is the size of the population in age group i. Therefore, the estimator for R 0 depends on the configuration of the WAIFW matrix. Farrington et al. [Reference Farrington, Kanaan and Gay4] showed that different configurations of the WAIFW matrix can lead to quite different estimates for R 0. For example, Farrington et al. [Reference Farrington, Kanaan and Gay4] estimated R 0 for mumps to be equal to 25·5, 8·0 and 3·3 for the configuration of W 2, W 3 and W 4, respectively:
where D is the duration of infectiousness assumed independent of age and N i is the size of the population in age group i. Therefore, the estimator for R 0 depends on the configuration of the WAIFW matrix. Farrington et al. [Reference Farrington, Kanaan and Gay4] showed that different configurations of the WAIFW matrix can lead to quite different estimates for R 0. For example, Farrington et al. [Reference Farrington, Kanaan and Gay4] estimated R 0 for mumps to be equal to 25·5, 8·0 and 3·3 for the configuration of W 2, W 3 and W 4, respectively:

Hence, the uncertainty related to the WAIFW matrix is coming from two different sources: (1) the uncertainty about the unknown transmission coefficients βi and (2) the uncertainty about the configuration of the WAIFW matrix. Furthermore, Wallinga et al. [Reference Wallinga6] showed that the basic reproductive number for measles ranges between 770·38 when assortative mixing pattern is assumed and 1·43 when infant mixing is assumed, i.e. infants are assumed to be the source of all infection [Reference Wallinga6].
In this paper, we present an investigation of the estimation of the basic reproductive number, R 0, and p c, the minimal proportion of the population that needs to be vaccinated to eliminate the infection, for varicella in Belgium for which, currently, there is no vaccination programme. Following the approach of Greenhalgh & Dietz [Reference Greenhalgh and Dietz5] we show that, depending on our assumption about the contact patterns, R 0 ranges between 3·12 and 68·57, and p c ranges between 67·9% and 98·5%.
This paper is organized as follow. In the next section, we present six possible configurations for the WAIFW matrix for varicella and discuss the estimation of the age-dependent force of infection from serological data using fractional polynomials. In the ‘Estimation of the transmission coefficients’ section, the WAIFW matrices are estimated using the integrated force of infection in the relevant age groups. Parametric and non-parametric bootstrap is used to calculate confidence intervals for the transmission coefficients. The estimation of R 0 and p c is discussed in the ‘Estimation of R 0 from the WAIFW matrices’ section.
Estimation of R 0 and the WAIFW matrix for varicella based on serological data
Age-dependent transmission coefficients
Several authors [Reference Farrington, Kanaan and Gay4, Reference Greenhalgh and Dietz5, Reference Greenhalgh7] illustrated how the estimation of R 0 is influenced by the configuration of the WAIFW matrix. A specific configuration of the WAIFW matrix represents specific assumptions about age-dependent transmission coefficients in the population which in turn represents prior assumptions about the mixing patterns in the population.
We illustrate these concepts for varicella in Belgium. The population was divided into the following six age groups, taking into account the schooling system in Belgium, 6 months–1 year, 2–5 years, 6–11 years, 12–18 years, 19–30 years and 31–44 years. Several configurations were discussed by Anderson & May [Reference Anderson and May1] and Greenhalgh & Dietz [Reference Greenhalgh and Dietz5]. Assortative mixing assumes that all contacts occurs within the age groups [Reference Greenhalgh and Dietz5]. The matrix W V1 has specific transmission coefficients within each age group, i.e. for the transmission among hosts belonging to the same age group on the diagonal, and of a common ‘background’ transmission coefficient between any two different age groups. This ‘background’ transmission coefficient is assumed equal to the transmission coefficients in the oldest age group (β6). Note that the transmission coefficient in the oldest age group and the ‘background’ transmission coefficient between different age groups are both expected to be smaller than the transmission coefficients in younger age groups. The second configuration, W V2, assumes that the main route of transmission for a directly transmitted viral infection like varicella is in kindergarten children or in the classroom. This is expressed by a unique coefficient β2 for the (presumed high) transmission between infectious and susceptible hosts in the range 2–5 years and two other specific coefficients (β1 and β3) for transmission amongst other hosts aged <12 years. The third configuration of the WAIFW matrix, W V3 is a minor variation of the structure W V2 with a common transmission coefficient β1 between hosts in the range 6 months–1 year and hosts aged <12 years. For the fourth configuration, W V4, the transmission coefficient depends only on the age group of the susceptible host. The susceptible hosts of a given age group are assumed to be as likely to acquire infection from infectious hosts of any age. Note that this structure is not symmetric. The fifth matrix structure, W V5, is a minor variation of the assortative structure W V1 in which there is a common transmission coefficient in the two older age groups and a distinct ‘background’ coefficient for the transmission between any other two different age groups. Finally, the sixth configuration, W V6, is the extreme case of assortative mixing for which there is only transmission assumed between hosts belonging to the same age group. Of these proposed configurations, the latter structure is most obviously unrealistic, it is nevertheless useful as it provides an upper bound to the basic reproductive number R 0 [Reference Greenhalgh and Dietz5].



Estimation of the age-dependent force of infection for varicella
The estimation of the WAIFW matrix requires the estimation of the force of infection from pre-vaccination data. Following the methodology proposed by Anderson & May [Reference Anderson and May1] (see also [Reference Halloran, Watelet and Struchiner2, Reference Greenhalgh7]) we assume that an age-specific serological profile can be estimated from pre-vaccination data. This can be done by modelling pre-vaccination seroprevalence data. The age-dependent force of infection can be derived from the estimated model for the prevalence of seropositive hosts by using non-parametric methods (discussed in [Reference Keiding8, Reference Shkedy9]), or parametric methods (discussed in [Reference Farrington, Kanaan and Gay4, Reference Grenfell and Anderson10–Reference Shkedy13]).
For varicella, we estimated the force of infection by using a seroprevalence dataset consisting of 1673 individuals aged between 1 and 44 years that was sampled in Antwerp (Belgium) between October 1999 and April 2000 and reported by Thiry et al. [Reference Thiry14]. The sera were residual specimens submitted to medical laboratories for diagnostic purposes. Sera for the 1–11 years age group were collected from outpatients hospitals in Antwerp, sera for the 12–18 years age group were collected from volunteers in vaccine trials and sera for age groups >16 years were provided by a medical laboratory in Antwerp. The population was stratified by age in order to sample about 100 observations per age group. The force of infection can be estimated from this serological sample under the assumption that the disease is in a steady state.
For the analysis presented in this paper, fractional polynomial [Reference Royston and Altman15] models were used to describe the dependency of the force of infection on age, as discussed in Shkedy et al. [Reference Shkedy13]. Briefly, a generalized linear model for the binary data with logit link was used to estimate the force of infection. The linear predictor for that model is given by
where m is an integer, p 1<p 2<…<pm is a sequence of powers and H i(a) is a transformation function given by
with p 0=0 and H 0=1. As shown in Shkedy et al. [Reference Shkedy13] the force of infection in this model can be expressed as
where ηm(a, β, p 1, p 2, …, p m)′ denotes the partial derivative of ηm(a, β, p 1, p 2, …, p m) with respect to age a. Figure 1 shows, for the varicella dataset, the estimated model for the prevalence of seropositive hosts (Fig. 1a) and the force of infection (Fig. 1b). Constrained fractional polynomials were fitted to ensure that the estimated force of infection will be non-negative. The model fit was based on the value of the Akaike Information Criterion (AIC) and the selected model has exponents p 1=−0·4 and p 2=−0·3 with an AIC equal to 125·769.

Fig. 1. Estimated prevalence (a) and force of infection (b) for varicella in Belgium. – – –, Integrated force of infection.
The estimated force of infection is given by equation (9), with m=2 and η2(a)=−40·231 a −0·3+28·153 a −0·4+11·303. According to this model, the force of infection for varicella in Belgium peaks at 2 years of age with a value of ℓ(2)=0·3111 and drops monotonically for older susceptibles. At 44 years of age the force of infection is estimated to be 0·0315. The mean force of infection for each of the six age groups can be estimated by integrating from the parametric model of the force of infection in equation (9). Hence, we first use a flexible parametric model to estimate the force of infection and integrate over the age groups thereafter. The advantage of using fractional polynomials is that this integration can be performed analytically and the force of infection in age group i (i=1, …, 6) is given by
where a i−1 and a i are the lower and upper bounds of age group i, respectively. The estimates for the six age groups were ![]() 1=0·254,
1=0·254, ![]() 2=0·267,
2=0·267, ![]() 3=0·160,
3=0·160, ![]() 4=0·096,
4=0·096, ![]() 5=0·059 and
5=0·059 and ![]() 6=0·037 for age groups 6 months–1 year, 2–5 years, 6–11 years, 12–18 years, 19–30 years and 31–44 years, respectively. Figure 1b shows the force of infection estimated by the fractional polynomial (solid line) and the integrated force of infection (dashed line) for the six age groups.
6=0·037 for age groups 6 months–1 year, 2–5 years, 6–11 years, 12–18 years, 19–30 years and 31–44 years, respectively. Figure 1b shows the force of infection estimated by the fractional polynomial (solid line) and the integrated force of infection (dashed line) for the six age groups.
Estimation of the transmission coefficients
Once the estimates of the force of infection are obtained, the elements of the WAIFW matrix can be computed by
where L is the life-expectancy at birth, D is the mean duration of infectiousness, N is the total population size and Ψj is given by equation (5). Substituting the ![]() is by their expression, we have
is by their expression, we have
It is easy to show that for i=2, …, 6:
Hence, in our model for varicella using six age groups, the expressions for the Ψi s are
for i=1, …, 6, where a 0=0·5, a 1=2, a 2=6, a 3=12, a 4=19, a 5=31, a 6=45, N=10237988, D=7/365 years and L=78 years.
System (4) (i=1, …, n) is a linear system of n equations in n 2 unknowns: the elements of the WAIFW matrix βij (i, j=1, …, n). Since this system is underdetermined, we need to impose a structure upon the WAIFW matrix, limiting the number of unknowns to n (n=6 for our model of varicella). We have estimated the elements of the WAIFW matrix for the six types of matrix structure described above. For example, the estimate of β6 for the WAIFW matrix structure W V3 is given by
The expressions for the other βis are given in the Appendix for matrix W V3. Similar expressions can be derived for each of the five other matrix structures.
The estimates of the βis are given in Table 1 for the six WAIFW matrix structures described above (see also Fig. 2), together with two types of confidence intervals (CIs): the non-parametric bootstrap percentile 95% CIs and the parametric bootstrap percentile 95% CIs. We now provide details on the computations.
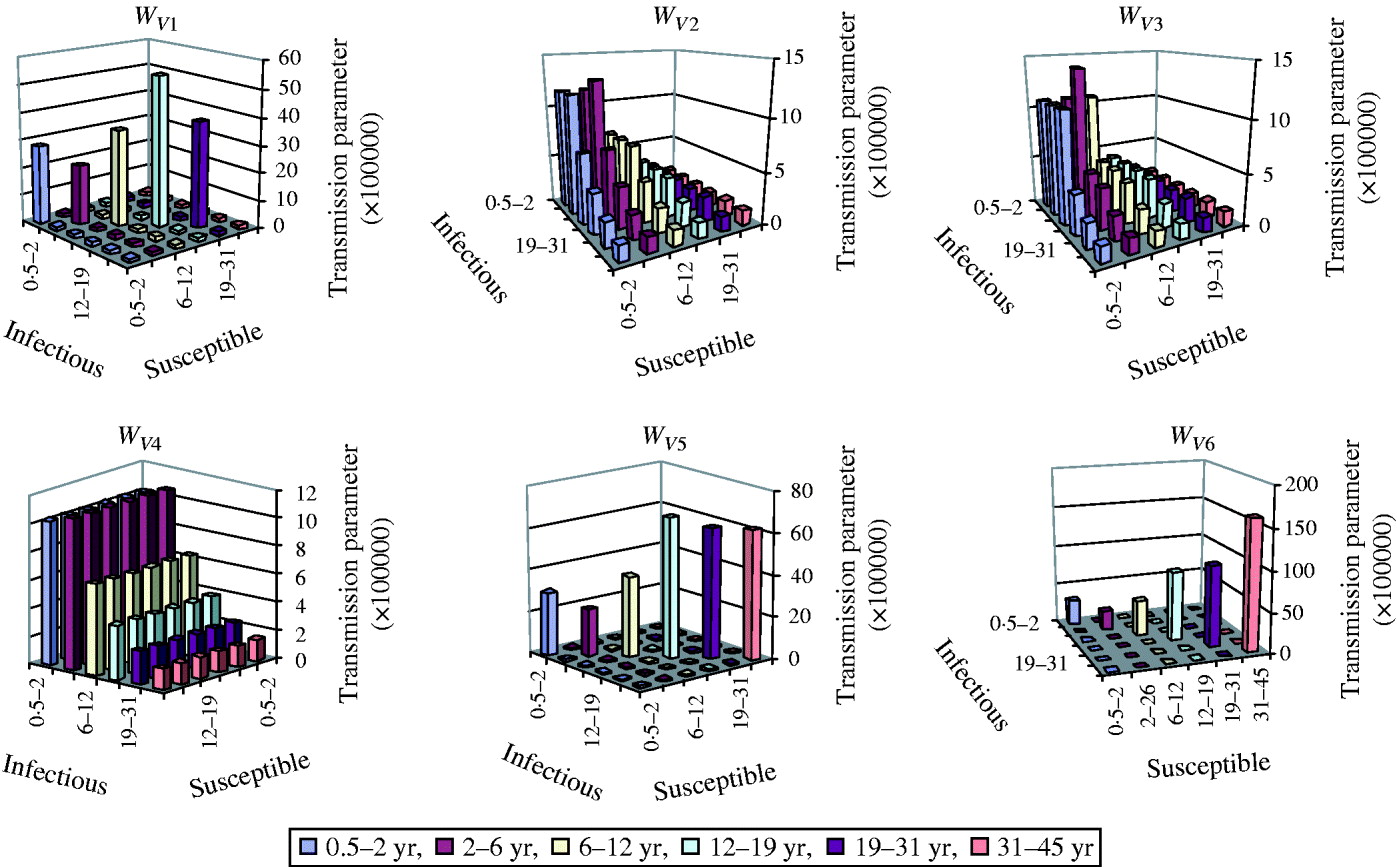
Fig. 2. WAIFW matrices for varicella in Belgium. Configurations 1–6. The upper limit is not included in the category age.
Table 1. Parameter estimates for the transmission coefficients. Decimal points are shown to illustrate 74differences in the confidence intervals

Bootstrap confidence intervals
In the seroprevalence sample, we have 44 samples by age of size N i (i=1, …, 44) and let p i be the proportion of subjects of age i who are seropositive for varicella-zoster virus antibodies. The total number of subjects in the seroprevalence sample is ![]() . A non-parametric bootstrap sample is a sample of size N obtained by drawing a random sample (with replacement) for each age i. The sample for age i is a sample of size N i drawn from a Bernoulli distribution with probability p i. Since we are interested in the number of subjects seropositive for varicella-zoster virus in the sample, we can equivalently generate a random value from the binomial distribution (N i, p i) for each age i. We have used 1000 bootstrap samples in the computation since this is deemed necessary for the estimation of confidence intervals. For each bootstrap sample, we can then estimate the parameters of the model. For these estimates, we can estimate the mean force of infection λ1, λ2, …, λ6, in each of the six age groups. In this way we obtain 1000 bootstrap values for each λi (i=1, …, 6). The non-parametric percentile bootstrap 95% CI for a parameter (for example λ1), is obtained from the bootstrap distribution of λ1 by taking the 25th and the 975th value in the sequentially ordered set of 1000 bootstrap values of this parameter.
. A non-parametric bootstrap sample is a sample of size N obtained by drawing a random sample (with replacement) for each age i. The sample for age i is a sample of size N i drawn from a Bernoulli distribution with probability p i. Since we are interested in the number of subjects seropositive for varicella-zoster virus in the sample, we can equivalently generate a random value from the binomial distribution (N i, p i) for each age i. We have used 1000 bootstrap samples in the computation since this is deemed necessary for the estimation of confidence intervals. For each bootstrap sample, we can then estimate the parameters of the model. For these estimates, we can estimate the mean force of infection λ1, λ2, …, λ6, in each of the six age groups. In this way we obtain 1000 bootstrap values for each λi (i=1, …, 6). The non-parametric percentile bootstrap 95% CI for a parameter (for example λ1), is obtained from the bootstrap distribution of λ1 by taking the 25th and the 975th value in the sequentially ordered set of 1000 bootstrap values of this parameter.
The parametric bootstrap and the computation of the parametric percentile bootstrap 95% CIs is performed in the same way as described above for the non-parametric bootstrap except that instead of drawing for each age i a random sample with replacement from the Bernoulli distribution with probability p i, i.e. from the data, we can use instead the parametric model for the cumulative distribution of the age at infection π(a) and draw the random sample from the Bernoulli distribution with probability f i, where f i is the fitted value for age i.
With the matrix structure W V1, the transmission is the highest between hosts aged 12–18 years (54·5×10−5, see also Fig. 2). This implies that the highest probability of infection is between an infective aged 12–18 years and a susceptible belonging to the same age group, presumably because of the high frequency of close contacts. In the other age groups, the transmission increases almost monotonically with age for hosts aged ⩽18 years and decreases monotonically with age for hosts aged >19 years. The transmission coefficient among hosts aged 31–44 years, which is also the ‘background’ transmission coefficient between hosts belonging to different age groups is very low (1·5×10−5). The two highest transmission coefficients with the matrix structure W V2 are among hosts aged 2–5 years (12·7×10−5) and between hosts aged 6 months–1 year and all hosts up to 6 years of age (11·6×10−5). The mixing pattern with matrix W V3 is quite similar to the pattern with W V2. The two highest transmission coefficients with W V3 is among hosts aged 2–5 years (14×10−5) and between hosts aged 6 months–1 year and all hosts up to 12 years of age (10·8×10−5). With matrix structure W V4, the two highest transmission coefficients are for susceptible hosts aged 2–5 years (10·7×10−5) and hosts aged 6 months–1 year (10·2×10−5). The transmission coefficient decreases monotonically with increasing age for susceptible hosts aged >2 years. Below 18 years of age, W V5 and W V1 have similar mixing patterns. Transmission is highest between hosts aged 12–18 years (66·7×10−5) and increases almost monotonically with age for hosts aged ⩽18 years and decreases monotonically with age for hosts aged >19 years. However, the transmission coefficient in the 19–30 years age group, which is constrained to be equal to the parameter in the 31–44 years group is much higher than with matrix W V1. Just as with W V1, the ‘background’ transmission coefficient between hosts belonging to different age groups is very low (0·9×10−5). With matrix W V6 with no transmission at all between hosts belonging to different age groups, the transmission coefficient increases almost monotonically with age and is the highest in hosts aged 31–44 years (162·4×10−5).
Estimation of R 0 from the WAIFW matrices
The global basic reproductive number R 0 for the population is the dominant eigenvalue of the ‘next generation matrix’ whose elements are the individual basic reproductive numbers ![]() (i=1, 2, …, 6, j=1, 2, …, 6) for the transmission of the infection from an infectious person in the age group j to a susceptible person in the age group i. By definition of the basic reproductive number, each
(i=1, 2, …, 6, j=1, 2, …, 6) for the transmission of the infection from an infectious person in the age group j to a susceptible person in the age group i. By definition of the basic reproductive number, each ![]() , where βij is the ijth element of the WAIFW matrix, D is the mean duration of infectiousness and N i is the total population in age group i. For varicella, the mean duration of infectiousness is 7 days=7/365 years. Like the elements of the WAIFW matrix, the 95% CIs for R 0 are computed using two different methods: percentile non-parametric bootstrap and percentile parametric bootstrap.
, where βij is the ijth element of the WAIFW matrix, D is the mean duration of infectiousness and N i is the total population in age group i. For varicella, the mean duration of infectiousness is 7 days=7/365 years. Like the elements of the WAIFW matrix, the 95% CIs for R 0 are computed using two different methods: percentile non-parametric bootstrap and percentile parametric bootstrap.
The minimal immunization coverage needed for elimination, i.e. the proportion of the total population to be immunized immediately after waning of maternal antibodies in order to eliminate varicella, p c, is obtained by the relationship:
Bootstrap confidence interval for estimation of R0 and pc
Table 2 and Figure 3 show the parameter estimates for R 0 and p c. Depending on the configuration of the WAIFW matrix, the basic reproductive number ranges between 3·12 (95% non-parametric CI 2·78–3·50) for W V3 to 68·57 (95% non-parametric CI 43·64–111·20) for the assortative mixing pattern W V6. This implies that across the different configurations of the WAIFW matrices p c ranges from 63·99% (lower limit for W V3) to 99·10% (upper limit for the assortative mixing pattern).

Fig. 3. Estimates for (a) p c and (b) R 0 and non-parametric bootstrap confidence intervals.
Table 2. Estimates of the basic reproductive number R0 and pc for different matrix structures for the ‘Who Acquires Infection From Whom’ (WAIFW) matrix

DISCUSSION
When the force of infection is both time and age dependent, the WAIFW matrix is a central parameter in modelling the spread of the infection in the population. A structure has to be assumed for the WAIFW matrix in order to be able to estimate the transmission coefficients and different structures lead to different estimates for the basic reproductive number R 0 and the minimal immunization coverage needed for elimination of the infection in the population p c. In this paper, we have estimated R 0 and p c for varicella in Belgium for different configurations of the WAIFW matrix. First, the force of infection has been estimated from seroprevalence data stratified by age, using a parametric model with fractional polynomials. The estimates of the mean force of infection over six age groups has then given the means to estimate the transmission coefficients, R 0 and p c for six different configurations of the WAIFW matrix. The variability of these parameters has been estimated through the computation of bootstrap confidence intervals. The results show that the values of R 0 and p c are sensitive to the structure of the WAIFW matrix, with the estimates of R 0 ranging between 3·12 and 68·57 and those of p c ranging between 67·92% and 98·54% for the six configurations chosen. Preliminary empirical data gathered by surveys about the mixing patterns for directly transmitted infections like varicella tend to show that people mostly mix with other people of the same age (configurations W V1, W V5, and W V6). However, although it has the advantage of providing an upper bound on the values of R 0 and p c, configuration W V6 is not realistic since people do not mix exclusively with people of the same age group. Moreover, the assumption of W V4, that only the age of the susceptible hosts matters and not the age of the infectious hosts seems a priori unrealistic. On the other hand, mixing patterns like W V2 and W V3 are probably realistic for a childhood disease like varicella for which it can reasonably be assumed that transmission takes place mainly amongst groups of young children. Hence, configurations W V1, W V2, W V3, and W V5 are probably the most relevant for varicella, which tends to support a value of R 0 between 3·12 and 26·33 and a value of p c between 67·92% and 96·20%. Although using a different model with a force of infection that varies over time and only five age groups, Whitaker & Farrington [Reference Whitaker and Farrington16] obtained similar values for R 0 for varicella in the United Kingdom. The estimates they obtained with a WAIFW matrix with a configuration similar to our W V2 and W V3 matrices were 3·02 and 3·14 in 1970 and 1998, respectively, while their estimate was 11·79 at both time-points for a WAIFW configuration similar to our assortative W V1 matrix. For elimination to be possible at a critical uptake, u c, the vaccine's lifelong efficacy, e, should be between 64·0% and 97·4%, respectively, in order for elimination of varicella by vaccination to be theoretically possible (as u c=p c/e). Different means can be investigated to determine more precisely the value of R 0 and p c. Gathering empirical data about mixing patterns [Reference Edmunds, O'Callahgan and Nokes17, Reference Beutels18] should help us to better determine which is the most plausible configuration for a given infection in a population. Another possible avenue is to estimate R 0 and p c using seroprevalence data from different infections that have a similar type of transmission, e.g. varicella, parvovirus, measles, mumps and rubella. Assuming a symmetric WAIFW matrix, seroprevalence data from three or four different infections would be enough to estimate the 15 (or 21), transmission coefficients with five (or six), age groups without additional assumptions about the mixing pattern. In any case, it seems clear that further studies on the most appropriate configuration of the WAIFW matrix are necessary to reduce variation in estimated R 0 and associated parameters.
APPENDIX
Expression of the βis for the structure W V3:

ACKNOWLEDGEMENTS
This paper is based on T.v.E.'s M.Sc. thesis in Biostatistics, at Hasselt University. Z.S., M.A., P.V.D. and P.B. acknowledge funding from ‘SIMID’, a strategic basic research project of the Institute for the Promotion of Innovation by Science and Technology in Flanders (IWT), project no. 060081 and POLYMOD, a European Commission project funded within the Sixth Framework Programme, Contract no.: SSP22-CT-2004-502084. Z.S., M.A. and G.M. gratefully acknowledge the financial support from the IAP research network no. P5/24 of the Belgian Government (Belgian Science Policy).
DECLARATION OF INTEREST
This work is unrelated to T.v.E.'s work at GlaxoSmithKline Biologicals where he is employed as a mathematical modeller.







