Burkholderia pseudomallei is a Gram-negative bacterium naturally occurring in rice-farming fields, rubber plantations, agricultural sites and water in endemic regions, particularly Southeast Asia and northern Australia [Reference Raja, Ahmed and Singh1]. It is the causative agent of meliodosis, a potentially acute fulminating disease in animals and humans with the clinical syndromes varying from bacteraemia, pneumonia, skin or soft tissue infection, and brain, splenic and liver abscesses; pneumonia is the most common presentation and is involved in about half of all cases [Reference Cheng and Currie2]. Most cases of melioidosis occur within latitudes 20° N and 20° S [Reference Dance3], and may account for 20% of all community-acquired septicaemias and 20–40% of sepsis-related mortality in northeast Thailand and Australia [Reference Wiersinga4, Reference Currie5].
The first human case of melioidosis in China was reported in Hong Kong in 1983 [Reference So6], followed by five cases a year later [Reference So7]. A seroprevalence survey in Hong Kong showed 14% seroconversion in a tuberculosis sanatorium in 1987 [Reference So8], and since these cases had never travelled outside Hong Kong the infections were regarded as being acquired locally. On mainland China, probably due to unawareness of melioidosis, it was not until 1990 that the first human case was diagnosed in Hainan, a southernmost province [Reference Li9], and subsequently some animal and human cases were reported in Guangdong, Guangxi and Hainan [Reference Wiersinga4].
A seroprevalence study of antibody responses to B. pseudomallei in humans and animals undertaken in 1979 in Nanning, Guangxi showed that the seropositive rate in humans was lower than that in animals (9·7% for humans vs. 34% in cattle and 15% in pigs). B. pseudomallei has routinely been isolated since 1975 from clinical samples in Guangxi, but there is limited information on the geographic distribution of this species in the region. To this end we undertook a survey of the distribution of B. pseudomallei in soil and water sites throughout this region.
In total, 274 soil and water samples were collected from 77 sites mainly in the rice field regions in Guangxi in the winter of 2007 (Fig. 1). About 10 g soil was sampled at a depth of 30–60 cm and placed into a sterile plastic bag; each site was sampled from two separate holes at the same time. Water samples were 10 ml volumes of pond surface water collected in sterile plastic tubes. All samples were stored in an ice box for transportation to the laboratory. About 5 g soil was shaken vigorously in 2 ml distilled water in sterile screw-capped glass vials; 1 ml water sample was added to 9 ml selective enrichment broth consisting of threonine-basal salt plus colistin (TBSS-C50 broth) [Reference Galimand and Dodin10] and incubated at 42°C for 48 h. One drop of soil suspension or 10 μl of the 48-h water enrichment culture were spread on Ashdown's agar. The plates were incubated at 42°C for 6 days and isolates with dry, wrinkled, violet to purple colonies were regarded as presumptive B. pseudomallei, and were stored in Luria–Bertani broth containing 15% glycerol at −70°C for further identification. For quantitative culture, 5 g soil was shaken in 5 ml water and 10- and 100-μl aliquots were plated on Ashdown's agar as above. All presumptive B. pseudomallei isolates were identified by phenotypic tests [Reference Walsh and Wuthiekanun11], and confirmed by PCR amplification of the fliC and 16S rDNA genes as described by Su et al. [Reference Su12]. The 16S rDNA amplicon was sequenced and the phylogenetic relatedness to the reference strain B. pseudomallei strain (CMCC 53001) was displayed by the Neighbour-Joining arithmetic method.
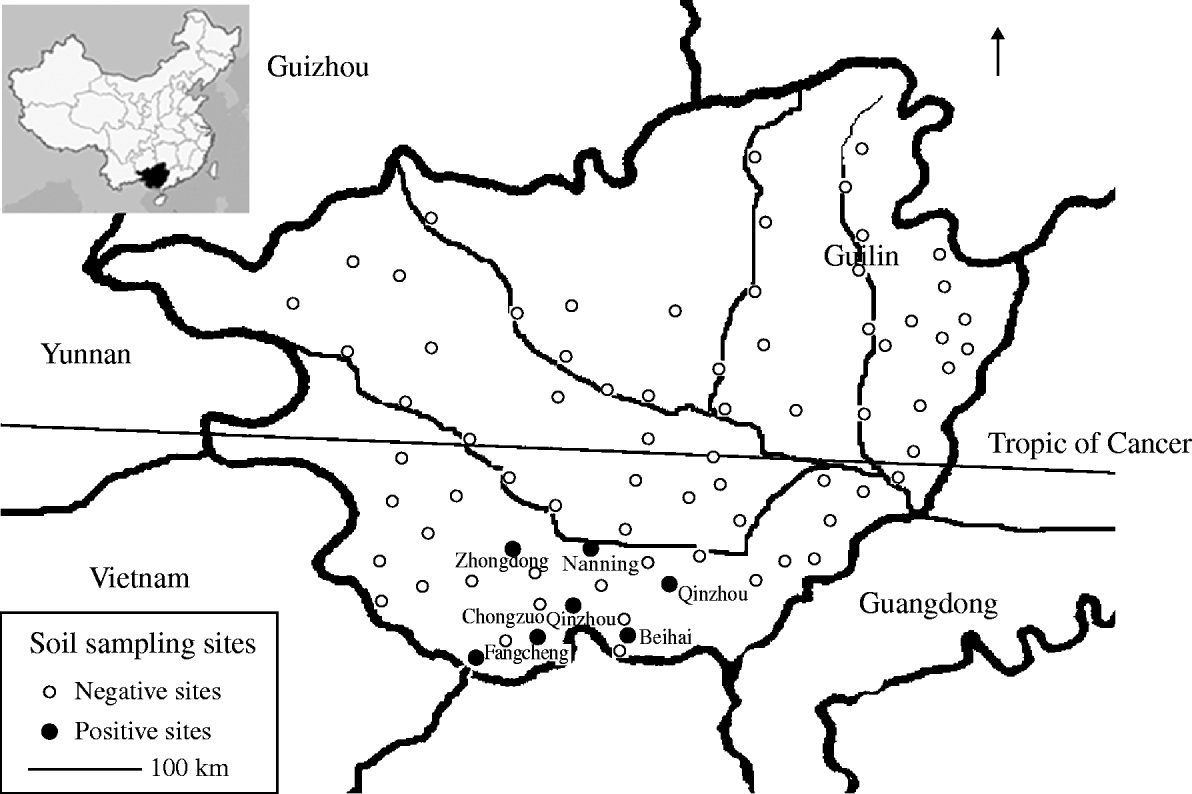
Fig. 1. Map illustrating soil sampling sites in Guangxi, China.
B. pseudomallei was detected in 13/154 (8·4%) soil samples, and seven sampling sites (9·1%) (Fig. 1). It was not found in any of the surface water samples. Counts of B. pseudomallei in the soil samples ranged from 23 to 521 c.f.u./g. Isolates identified by phenotypic tests were positive for fliC and 16S rDNA genes. Sequencing of the latter revealed high similarity to the reference strain.
There was an uneven distribution of B. pseudomallei in Guangxi. Four of the seven positive sites were located in coastal areas (Qinzhou, Beihai, Fangcheng) and the remainder to the south of Nanning city, a plain of rice fields <200 m above sea level (latitude 23·5° N). No positive sample was identified in mountainous regions in the north and west (Fig. 1) and no B. pseudomallei were detected in samples from Chongzuo where cases of melioidosis have been reported. The geographical distribution of B. pseudomallei is possibly affected by several factors including altitude, rainfall, and temperature and local ecological conditions; indeed 11 negative sampling sites were located close to seven positive sites.
The bacterial counts in the soil are probably related to the risk of developing melioidosis. The concentrations of B. pseudomallei in soil (23–521 c.f.u./g) were comparable to levels found in studies in Thailand (1–17 000) and Laos (10–1200) [Reference Smith13, Reference Lee, Wang and Yap14]. Previous antibody prevalence surveys to B. pseudomallei in animals and humans reported rates of 9·7% in Nanning and 1·8% in Guilin [Reference Li15]. This is consistent with the fact that one-third of the sites around Nanning cultured positive for B. pseudomallei while Guilin region yielded no positive isolations.
Recreational or occupational exposure to soil and water contaminated with B. pseudomallei are recognized risk factors for contracting melioidosis [Reference Suputtamongkol16]. Although no cases have yet been reported in Guangxi, physicians in this area should consider melioidosis when patients present with an unknown fever or community-acquired pneumonia. Special attention should be given to seropositive individuals as the pathogen may persist in humans for very long periods and be reactivated leading to relapse, especially among immunocompromised individuals, in whom the disease can be fatal [Reference Su12, Reference Currie17].
ACKNOWLEDGEMENTS
We thank Mr Yan Jianping and Wang Jianzhou for technical assistance. The project was supported by a grant (24GFCX-YJ-2306) from the Chinese Academy of Sciences.
DECLARATION OF INTEREST
None.



