Introduction
Ticks (Ixodida) are comprised of 3 extant families, Argasidae (soft ticks), Ixodidae (hard ticks) and Nuttalliellidae and 2 extinct families, Deinocrotonidae and Khimairidae (Mans, Reference Mans2023). The Argasidae is divided into the Argasinae and Ornithodorinae subfamilies (Mans, Reference Mans2023). The Ornithodorinae currently comprise the genera Alectorobius, Antricola (including Parantricola), Carios, Chiropterargas, Nothoaspis, Ornithodoros (including Microargas, Ornamentum, Ornithodoros and Pavlovskyella), Otobius, Reticulinasus and Subparmatus (Mans et al., Reference Mans, Kelava, Pienaar, Featherston, de Castro, Quetglas, Reeves, Durden, Miller, Laverty, Shao, Takano, Kawabata, Moustafa, Nakao, Matsuno, Greay, Evasco, Barker and Barker2021; Mans, Reference Mans2023). The Argasinae currently comprise the genera Alveonasus, Argas (including Argas and Persicargas), Navis, Ogadenus, Proknekalia and Secretargas (Mans et al., Reference Mans, Kelava, Pienaar, Featherston, de Castro, Quetglas, Reeves, Durden, Miller, Laverty, Shao, Takano, Kawabata, Moustafa, Nakao, Matsuno, Greay, Evasco, Barker and Barker2021; Mans, Reference Mans2023). Otobius lagophilus Cooley and Kohls, 1940 has recently been shown to group in the Argasinae even though Otobius sensu stricto group within the Ornithodorinae (Kneubehl et al., Reference Kneubehl, Muñoz-Leal, Filatov, de Klerk, Pienaar, Lohmeyer, Bermúdez, Suriyamongkol, Mali, Kanduma, Latif, Sarih, Bouattour, de León, Teel, Labruna, Mans and Lopez2022). Understanding the relationships between the families and the relationships between genera within the families is important for an understanding of tick evolution (Mans et al., Reference Mans, de Castro, Pienaar, de Klerk, Gaven, Genu and Latif2016; Mans, Reference Mans2023). Generic relationships in the Argasidae in particular is to a large extent unresolved, while many genera seem to be paraphyletic or polyphyletic, leading to the possibility that argasid genera may increase in the future. A number of genera and species have been shown to group outside of their traditional placement in the argasid subfamilies, notably Proknekalia and Otobius lagophilus in the Argasinae and Carios and Chiropterargas in the Ornithodorinae (Mans et al., Reference Mans, Featherston, Kvas, Pillay, de Klerk, Pienaar, de Castro, Schwan, Lopez, Teel, Pérez de León, Sonenshine, Egekwu, Bakkes, Heyne, Kanduma, Nyangiwe, Bouattour and Latif2019, Reference Mans, Kelava, Pienaar, Featherston, de Castro, Quetglas, Reeves, Durden, Miller, Laverty, Shao, Takano, Kawabata, Moustafa, Nakao, Matsuno, Greay, Evasco, Barker and Barker2021; Kneubehl et al., Reference Kneubehl, Muñoz-Leal, Filatov, de Klerk, Pienaar, Lohmeyer, Bermúdez, Suriyamongkol, Mali, Kanduma, Latif, Sarih, Bouattour, de León, Teel, Labruna, Mans and Lopez2022). Placement of argasid genera and species within the subfamilies is therefore important and has not been confirmed for the genera Alveonasus or Microargas. The genus Alveonasus is of particular interest since the major systematic schools have differed regarding its affinity to the various subfamilies (Pospelova-Shtrom, Reference Pospelova-Shtrom1969; Keirans et al., Reference Keirans, Hoogstraal and Clifford1977; Klompen and Oliver, Reference Klompen and Oliver1993).
The genus Alveonasus (Schulze, Reference Schulze1941) is characterized by a non-mammilated body integument with numerous depressions around which wrinkled ridges radiate to give a madreporean sculpturing effect (Clifford et al., Reference Clifford, Kohls and Sonenshine1964). The type species is Alveonasus lahorensis (Neumann, 1908) (Schulze, Reference Schulze1941). The genus is composed of 8 species and is mainly found in the Afrotropic and Palearctic regions, although Alveonasus cooleyi (McIvor, Reference McIvor1941) derives from the Nearctic (Table 1). Alveonasus lahorensis is unique in the Argasidae in that it is a 2-host tick species with larvae feeding and detaching as engorged third-instar nymphs that then moult to adults (Hoogstraal, Reference Hoogstraal1985). Hosts include cattle, sheep, camels and wild ungulates. Its geographic distribution covers a wide range of the Palearctic region including Central Asia (southern former USSR), East Asia (China and Tibet), South Asia (Kashmir and Pakistan), Southwest Asia (Iran to Saudi Arabia) and Southeast Europe (Turkey, Bulgaria, Yugoslavia and Greece) (Hoogstraal, Reference Hoogstraal1985).
Current species placed in the genus Alveonasus and their geographic distribution. Specific type localities are also indicated.
Alveonasus was placed within the Ornithodorinae by the Russian (Pospelova-Shtrom, Reference Pospelova-Shtrom1946, Reference Pospelova-Shtrom1969), American (Clifford et al., Reference Clifford, Kohls and Sonenshine1964; Hoogstraal, Reference Hoogstraal1985; Guglielmone et al., Reference Guglielmone, Robbins, Apanaskevich, Petney, Estrada-Peña, Horak, Shao and Barker2010) and French (Camicas and Morel, Reference Camicas and Morel1977; Camicas et al., Reference Camicas, Hervy, Adam and Morel1998) schools, mostly based on the fact that its body margin is rounded and lack a sutural groove. However, cladistic analysis based on 83 biological and morphological characters placed Alveonasus in the Argasinae (Klompen and Oliver, Reference Klompen and Oliver1993). Phylogenetic analysis of the nuclear 18S ribosomal RNA gene (Black et al., Reference Black, Klompen and Keirans1997; Mans et al., Reference Mans, Featherston, Kvas, Pillay, de Klerk, Pienaar, de Castro, Schwan, Lopez, Teel, Pérez de León, Sonenshine, Egekwu, Bakkes, Heyne, Kanduma, Nyangiwe, Bouattour and Latif2019, Reference Mans, Kelava, Pienaar, Featherston, de Castro, Quetglas, Reeves, Durden, Miller, Laverty, Shao, Takano, Kawabata, Moustafa, Nakao, Matsuno, Greay, Evasco, Barker and Barker2021) and the mitochondrial 12S and 16S ribosomal RNA genes (Zhao et al., Reference Zhao, Lin, Li, Li, He, Zhang, Pan, Wang, Gao, Johnson, Yuan, Lv, Wu and Liu2018), also placed Alveonasus within the Argasinae. Based on these considerations, Alveonasus was placed within the Argasinae even though analysis of its full mitochondrial genome was not presented (Mans et al., Reference Mans, Featherston, Kvas, Pillay, de Klerk, Pienaar, de Castro, Schwan, Lopez, Teel, Pérez de León, Sonenshine, Egekwu, Bakkes, Heyne, Kanduma, Nyangiwe, Bouattour and Latif2019, Reference Mans, Kelava, Pienaar, Featherston, de Castro, Quetglas, Reeves, Durden, Miller, Laverty, Shao, Takano, Kawabata, Moustafa, Nakao, Matsuno, Greay, Evasco, Barker and Barker2021). The current study sequenced the mitochondrial genome and full-length 18S and 28S ribosomal RNA of A. lahorensis, the type species of this genus from Pakistan. The results support the placement of Alveonasus within the Argasinae based on both nuclear and mitochondrial gene analysis. In addition, we report the mitochondrial genomes of the type species for Argas: the pigeon tick, Argas reflexus Fabricius, 1794 from Germany and the bat tick, Alectorobius kelleyi (Cooley and Kohls, 1941) from the USA.
Materials and methods
Ticks and datasets
The ticks analyzed in the current study include A. lahorensis collected from sheep in Khyber Pakhtunkhwa, Pakistan (collected and identified by Abid Ali in 2022) and A. reflexus, historical collection and identification by Hans Dautel in Berlin, Germany. In both cases, voucher specimens have been deposited in the Gertrud Theiler National Tick Collection (Agricultural Research Council – Onderstepoort Veterinary Research). The data for A. kelleyi was obtained from the NCBI SRA database (SRR23908069) (Occi et al., Reference Occi, Price, Hall, Campbell, Stronsick, Sullivan, Pesapane, Gonzalez, Toledo and Fonseca2023). Tick nomenclature is used according to the proposals by Mans et al. (Reference Mans, Featherston, Kvas, Pillay, de Klerk, Pienaar, de Castro, Schwan, Lopez, Teel, Pérez de León, Sonenshine, Egekwu, Bakkes, Heyne, Kanduma, Nyangiwe, Bouattour and Latif2019) and Mans et al. (Reference Mans, Kelava, Pienaar, Featherston, de Castro, Quetglas, Reeves, Durden, Miller, Laverty, Shao, Takano, Kawabata, Moustafa, Nakao, Matsuno, Greay, Evasco, Barker and Barker2021).
Next-generation sequencing, assembly and mitochondrial genome annotation
Genomic DNA was extracted using the QIAamp DNA Blood Mini Kit (Qiagen), processed using the MGIEasy Universal DNA Library Prep kit (MGI, Shenzhen, China) and sequenced on the MGI DNBSEQ-G400 sequencing instrument using the PE150 (paired-end 2 × 150 bp) format (Agricultural Research Council-Biotechnology Platform, South Africa). Paired-end sequence data were quality trimmed (0.001 quality limit) and MGI adapters were removed using CLC Genomics Workbench v.20.1 software (Qiagen). Standard assembly parameters (mismatch cost-2, insertion cost-3, deletion cost-3, length fraction-0.9, similarity-0.9, minimum contig length-200 and automatic bubble size) were used and assembly was performed using a word size of 49 in CLC Genomics Workbench v.20.1 software (Qiagen). Contigs were identified as mitochondrial, 18S or 28S rRNA using BLASTN analysis (Altschul et al., Reference Altschul, Gish, Miller, Myers and Lipman1990). Final contigs were obtained by mapping data back to the contigs using CLC Genomics Workbench v.20.1 (mismatch cost-2, insertion cost-3, deletion cost-3, length fraction-0.5 and similarity-0.9), to obtain consensus sequences and final coverage values. The mitochondrial genome was annotated using the MITOS and ARWEN servers to identify tRNA genes (Laslett and Canbäck, Reference Laslett and Canbäck2008; Bernt et al., Reference Bernt, Donath, Jühling, Externbrink, Florentz, Fritzsch, Pütz, Middendorf and Stadler2013). Protein-coding genes were identified using the Expasy Translation Server (https://web.expasy.org/translate/) and BLASTP analysis (Altschul et al., Reference Altschul, Gish, Miller, Myers and Lipman1990).
16S rRNA phylogenetic analysis
Sequences of A. lahorensis, A. reflexus and A. kelleyi were used to download the top 100 most closely related sequences from GenBank using BLASTN analysis (Altschul et al., Reference Altschul, Gish, Miller, Myers and Lipman1990). Sequences from the BLASTN analysis were combined to produce a non-redundant dataset with sequences that represent unique species. This yielded a final dataset of 107 sequences that was aligned using MAFFT taking rRNA secondary structure into account (Q-INS-i) (1PAM/k = 2 scoring matrix) (Katoh and Standley, Reference Katoh and Standley2013). Maximum likelihood analysis was performed using IQ-Tree2 v 2.2.0 (Minh et al., Reference Minh, Schmidt, Chernomor, Schrempf, Woodhams, von Haeseler and Lanfear2020) with an alignment size of 336 bp. The most optimal substitution model used was GTR + F + I + G4. Nodal support was estimated using ultrafast bootstrap (n = 10 000) and the 50% consensus tree was reported.
Cytochrome oxidase I phylogenetic analysis
Sequences of A. lahorensis, A. reflexus and A. kelleyi were used to download the top 100 most closely related sequences from GenBank using BLASTN analysis (Altschul et al., Reference Altschul, Gish, Miller, Myers and Lipman1990). Sequences from the BLASTN analysis were combined to produce a non-redundant dataset with sequences that represented unique species. This yielded a final dataset of 107 sequences that was aligned using MAFFT taking rRNA secondary structure into account (Q-INS-i) (1PAM/k = 2 scoring matrix) (Katoh and Standley, Reference Katoh and Standley2013). A neighbour-joining analysis was performed in Mega 5 (Tamura et al., Reference Tamura, Peterson, Peterson, Stecher, Nei and Kumar2011), using the amino acid matrix derived from the nucleotide sequence with the Jones–Taylor–Thornton amino acid substitution model with uniform rates across sites. Nodal support was estimated using 10 000 bootstraps.
Mitochondrial genome phylogenetic analysis
Translated protein sequences for the ATP synthase 6 (ATP6), ATP synthase 8 (ATP8), cytochrome oxidase I (COX1), cytochrome oxidase II (COX2), cytochrome oxidase III (COX3), cytochrome b (Cytb), NADH dehydrogenase subunit 1 (ND1), NADH dehydrogenase subunit 2 (ND2), NADH dehydrogenase subunit 3 (ND3), NADH dehydrogenase subunit 4 (ND4), NADH dehydrogenase subunit 4L (ND4L), NADH dehydrogenase subunit 5 (ND5) and NADH dehydrogenase subunit 6 (ND6) genes were used for phylogenetic analysis (Mans et al., Reference Mans, de Klerk, Pienaar, de Castro and Latif2012). Multiple sequence alignments for each protein were performed separately using MAFFT with iterative alignment (FFT-NS-i) and the BLOSUM62 amino acid scoring matrix (Katoh and Standley, Reference Katoh and Standley2013). Maximum likelihood analysis was performed in IQ-Tree2 IQ-Tree2 v 2.2.0 (Minh et al., Reference Minh, Schmidt, Chernomor, Schrempf, Woodhams, von Haeseler and Lanfear2020). An optimal substitution model was calculated for each protein partition: ATP6 (mtMet + F + R5), ATP8 (mtVer + F + G4), COX1 (mtART + R5), COX2 (mtMAM + F + I + I + R4), COX3 (mtART + F + I + I + R5), CYTB (mtMet + I + G4), NAD1 (mtZOA + F + I + I + R5), NAD2 (mtMAM + F + I + I + R6), NAD3 (mtMet + I + G4), NAD4 (mtMet + F + I + I + R5), NAD4L (mtMet + F + G4), NAD5 (mtMet + F + I + I + R6) and NAD6 (mtVer + F + G4). Absent protein genes were treated as missing data. An edge-proportional partition model with proportional branch lengths (-spp) was used to allow each partition its own specific rate to accommodate different evolutionary rates between partitions. Nodal support was estimated using ultrafast bootstrap (n = 1 000 000) and the 50% consensus tree was reported.
18S-28S Ribosomal RNA phylogenetic analysis
The 18S and 28S rRNA genes from the Ixodida were downloaded from GenBank and only included in the analysis where both sequences were available for a species. Sequences were cleaned up to include a single representative for each species, except for A. lahorensis where the 18S rRNA gene from Black et al. (Reference Black, Klompen and Keirans1997) was also included. The 18S and 28S rRNA genes were aligned separately with MAFFT taking rRNA secondary structure into account (Q-INS-i) (1PAM/k = 2 scoring matrix) (Katoh and Standley, Reference Katoh and Standley2013). GBLOCKS was used to remove columns with less than 50% coverage (Castresana, Reference Castresana2000) resulting in alignments of 1014 bp for the 18S and 540 bp for the 28S rRNA genes. Maximum likelihood analysis was performed using IQ-Tree2 IQ-Tree2 v 2.2.0 (Minh et al., Reference Minh, Schmidt, Chernomor, Schrempf, Woodhams, von Haeseler and Lanfear2020). The most optimal substitution model (18S: TIM3e + I + I + R2; 28S: TPM3 + I + I + R2) for each alignment was automatically selected. Absent genes were treated as missing data. An edge-proportional partition model with proportional branch lengths (-spp) was used, to allow different rate parameters for each partition to accommodate different evolutionary rates between partitions. Nodal support was estimated using ultrafast bootstrap (n = 1 000 000) and the 50% consensus tree was reported. For Bayesian analysis, alignments were concatenated to produce a matrix with 1831 positions.
Results
Mitochondrial gene structure
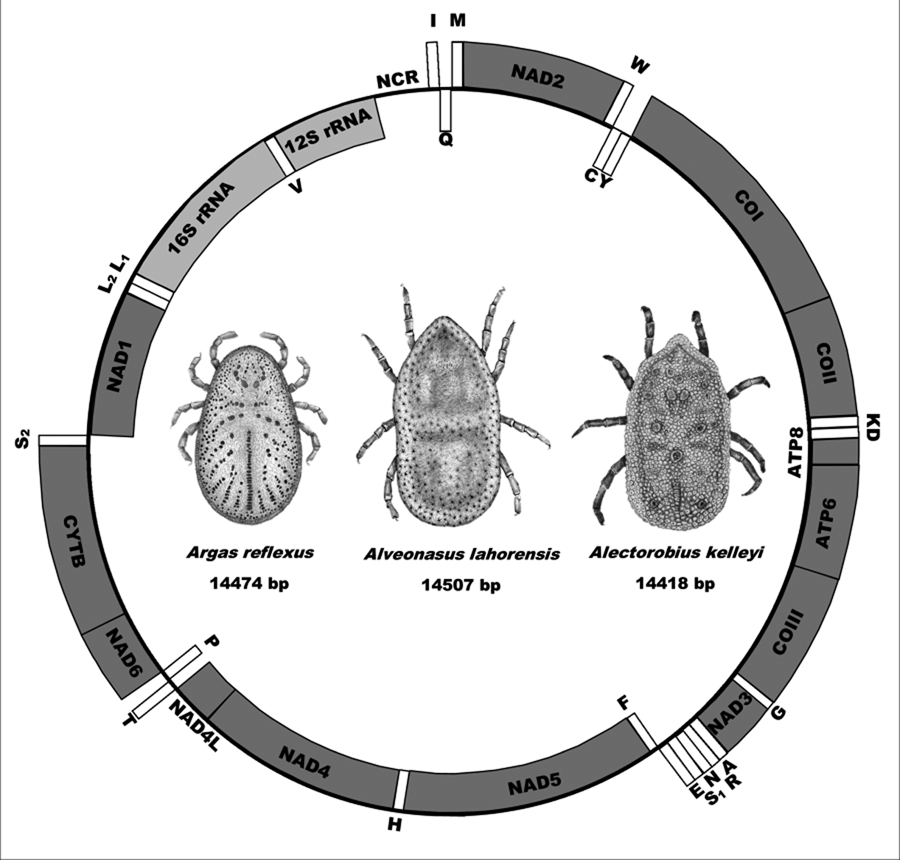
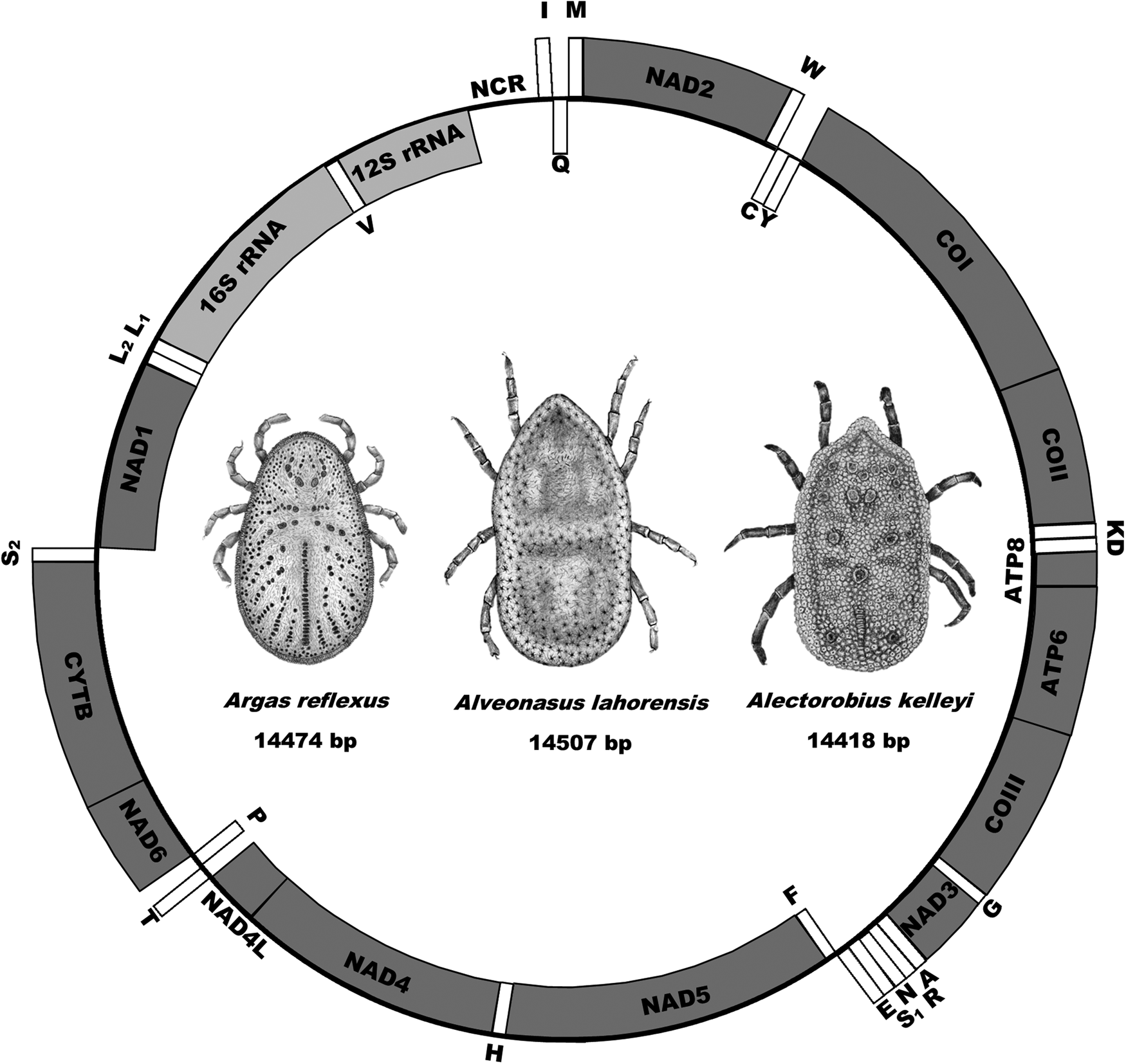
The mitochondrial genomes of A. lahorensis (PP072240-coverage 273; PP072241-coverage 784), A. reflexus (PP072242 – coverage 832) and A. kelleyi (SRR23908069 – coverage 167) all possess the standard mitochondrial gene structure and number of genes observed in argasid species (Shao et al., Reference Shao, Aoki, Mitani, Tabuchi, Barker and Fukunaga2004; Mans et al., Reference Mans, de Klerk, Pienaar, de Castro and Latif2012;, Reference Mans, Featherston, Kvas, Pillay, de Klerk, Pienaar, de Castro, Schwan, Lopez, Teel, Pérez de León, Sonenshine, Egekwu, Bakkes, Heyne, Kanduma, Nyangiwe, Bouattour and Latif2019, Reference Mans, Kelava, Pienaar, Featherston, de Castro, Quetglas, Reeves, Durden, Miller, Laverty, Shao, Takano, Kawabata, Moustafa, Nakao, Matsuno, Greay, Evasco, Barker and Barker2021; Burger et al., Reference Burger, Shao, Labruna and Barker2014; Kneubehl et al., Reference Kneubehl, Muñoz-Leal, Filatov, de Klerk, Pienaar, Lohmeyer, Bermúdez, Suriyamongkol, Mali, Kanduma, Latif, Sarih, Bouattour, de León, Teel, Labruna, Mans and Lopez2022) that include the 16S and 12S ribosomal RNA genes, the 13 protein coding genes and the 22 tRNA genes (Fig. 1).
Mitochondrial genome arrangement for Alveonasus lahorensis, Argas reflexus and Alectorobius kelleyi. Genes on the outside are on the forward strand whereas genes on the inside are on the reverse or complementary strand. Protein-encoding genes are in dark grey, ribosomal genes in light grey whereas transfer RNA genes are in white boxes. Also indicated are mitochondrial genome sizes.
16S Ribosomal gene analysis
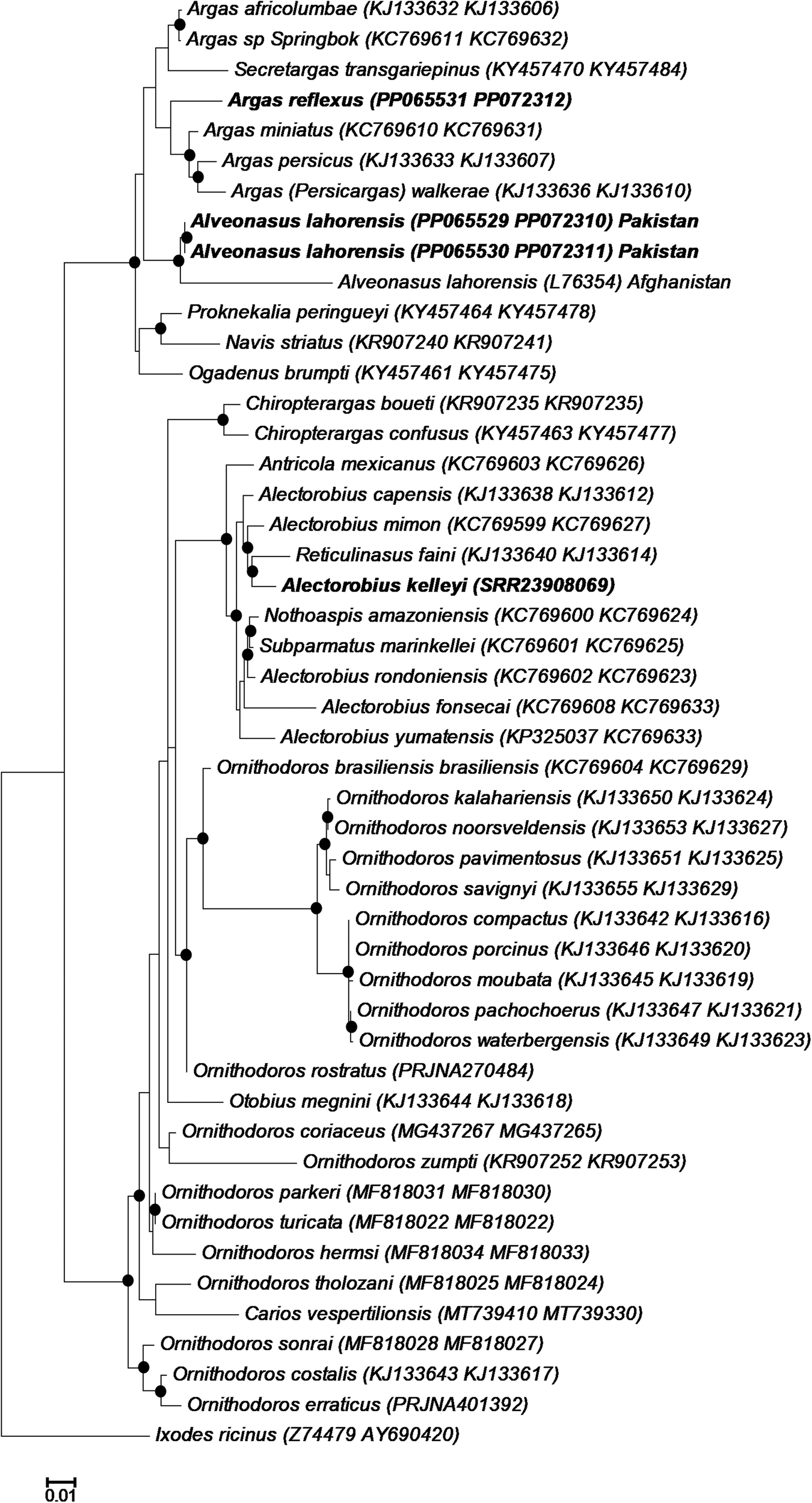
A large number of 16S rRNA sequences were previously deposited in GenBank for A. lahorensis, but none from the type locality (Pakistan). To confirm the relationship of A. lahorensis from Pakistan to published sequences, a BLASTN analysis was performed to determine all closely related sequences, followed by a multiple sequence alignment of all sequences annotated as A. lahorensis. All sequences in GenBank (n = 41) derive from the Tarim basin in China (Zhao et al., Reference Zhao, Lin, Li, Li, He, Zhang, Pan, Wang, Gao, Johnson, Yuan, Lv, Wu and Liu2018), and showed 96.15–99.78% (median 99.35%) sequence identity over a 461 bp region with those from Pakistan. For A. kelleyi no 16S rRNA data has yet been deposited in GenBank, while A. reflexus shows 99–100% sequence identity to 4 sequences in Genbank that derive from Germany (ON366980), Spain (MW289075, MW289076) and Poland (AF001401). Phylogenetic analysis showed that A. lahorensis grouped within a monophyletic clade within the Argasinae (Fig. 2). Within this clade A. lahorensis from Pakistan grouped within other sequences from China. Argas refelxus grouped within the genus Argas with a number of sequences annotated as A. reflexus, while A. kelleyi grouped within the Alectorobius genus (Fig. 2).
Phylogenetic analysis of the 16S rRNA gene. Indicated are selected members of various genera and subgenera as well as those sequences available in GenBank for Argas reflexus, Alveonasus lahorensis and Alectorobius kelleyi. The current specimens are indicated in bold and GenBank accession numbers in brackets. Bootstrap values are indicated.

Cytochrome oxidase I gene analysis
Similar to the 16S rRNA gene, a number of A. lahorensis COI genes have been deposited in GenBank. A BLASTN analysis was performed to determine all closely related sequences, followed by a multiple sequence alignment of all sequences annotated as A. lahorensis. The sequences in GenBank (n = 8) derive from the Tarim basin in China (unpublished) and Iran (Hosseini-Chegeni et al., Reference Hosseini-Chegeni, Tavakoli, Koshki and Khedri2019), and showed 99.08–99.54% sequence identity over a ~650 bp region. There are no COI genes available for A. reflexus, while A. kelleyi retrieved 15 hits annotated as A. kelleyi, all from the USA, with sequence identities that ranged from 96.20% to 100%. The COI gene for A. canestrinii is also available in GenBank (MH673048) and the hypothesis that Alveonasus is a monophyletic lineage was tested by performing a phylogenetic analysis of the Argasidae (Fig. 3). All A. lahorensis sequences grouped within a well-supported monophyletic clade within the Argasinae. The tree also included A. canestrinii, but this sequence did not group in a monophyletic clade with A. lahorensis, also previously observed (Hosseini-Chegeni et al., Reference Hosseini-Chegeni, Tavakoli, Koshki and Khedri2019), suggesting that Alveonasus may be paraphyletic. Alectorobius kelleyi grouped within Alectorobius in a well-supported clade with other A. kelleyi sequences, while A. reflexus grouped in a clade with other Argas spp.
Phylogenetic analysis of the cytochrome oxidase I gene. Indicated are selected members of various genera and subgenera as well as those sequences available in GenBank for Argas reflexus, Alveonasus lahorensis and Alectorobius kelleyi. The current specimens are indicated in bold and GenBank accession numbers in brackets. The tree was rooted using Ixodes anatis. Bootstrap values above 70% are indicated with black dots and above 90% with white dots.

Mitochondrial genome analysis
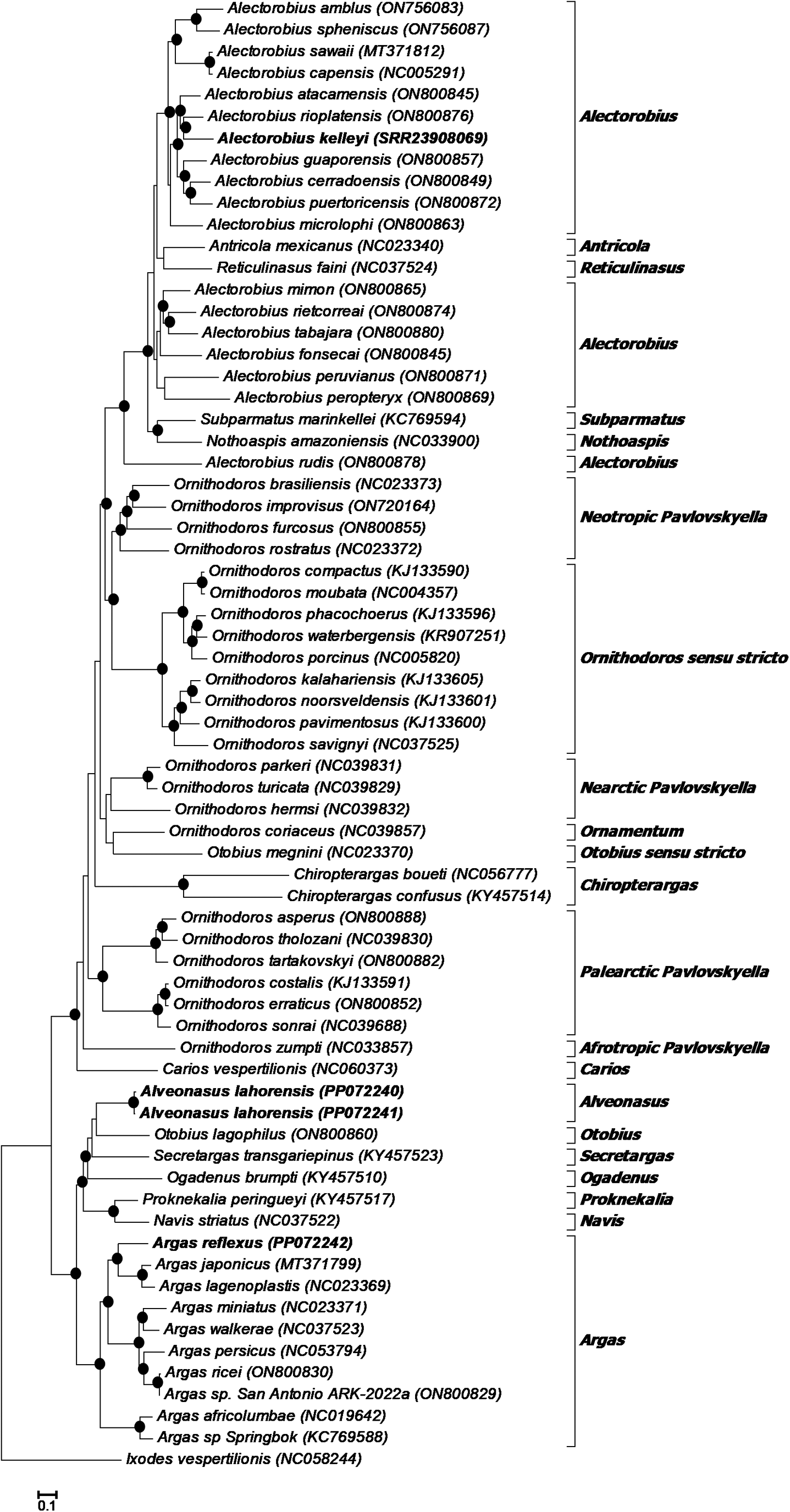
Phylogenetic analysis using the 13 protein coding genes indicates that A. lahorensis group with good support within the Argasinae in a clade formed by Alveonasus, Navis, Ogadenus, Otobius lagophilus, Proknekalia and Secretargas (Fig. 4). Argas reflexus group within the Argas clade, while A. kelleyi group within the Alectorobius clade.
Phylogenetic analysis of the 13 mitochondrial protein coding genes. Indicated are species with their mitochondrial genome accession numbers. The current specimens are indicated in bold. Also indicated are the genera or subgenera of various clades. The tree was rooted using Ixodes vespertilionis. Bootstrap values above 90% are indicated with black dots.

18S-28S rRNA gene analysis
To confirm that the sequences group within the various subfamilies as indicated for mitogenome analysis, an analysis of the nuclear 18S and 28S ribosomal RNA genes was also performed (Fig. 5). Alveonasus lahorensis grouped in the Argasinae with good support. The sequences from Pakistan grouped as sister group to an A. lahorensis sequence from Afghanistan (L76354). The sequences from Pakistan showed 100% sequence identity, however, pairwise comparison showed only 95% sequence identity to A. lahorensis (Afghanistan). This translates to 42 differences that include 4 gapped positions and would suggest different species (Mans et al., Reference Mans, de Klerk, Pienaar, de Castro and Latif2015). Given the wide geographic distribution of this species, the possibility exist that it is composed of a species complex. Wider geographic sampling is needed to investigate this possibility. Argas reflexus grouped within a clade with other Argas members, while A. kelleyi grouped within a clade formed by members of the Alectorobius, Antricola, Reticulinasus and Subparmatus genera.
Phylogenetic analysis of the 18S-28S ribosomal RNA genes. Indicated are species with their 18S and 28S rRNA accession numbers in brackets. The current specimens are indicated in bold. The tree was rooted using Ixodes ricinus. Bootstrap values above 90% are indicated with black dots.
Discussion
Relationship of Alveonasus to other argasid genera
Alveonasus was considered a connecting intermediate lineage between Argas and Ornithodoros (Clifford et al., Reference Clifford, Kohls and Sonenshine1964). The latter study included Proknekalia peringueyi (Bedford and Hewitt, 1925) and Proknekalia peusi (Schulze, 1943) in Alveonasus based on their wrinkled integument and absence of the preanal and tranverse anal groove in adults and a large dorsal plate and respiratory apparatus in larvae. Proknekalia was raised to its own genus (Keirans et al., Reference Keirans, Hoogstraal and Clifford1977) as supported by the placement of Proknekalia in its own clade in the Argasinae by mitochondrial and 18S-28S rRNA analysis (Mans et al., Reference Mans, Featherston, Kvas, Pillay, de Klerk, Pienaar, de Castro, Schwan, Lopez, Teel, Pérez de León, Sonenshine, Egekwu, Bakkes, Heyne, Kanduma, Nyangiwe, Bouattour and Latif2019; Mans et al., Reference Mans, Kelava, Pienaar, Featherston, de Castro, Quetglas, Reeves, Durden, Miller, Laverty, Shao, Takano, Kawabata, Moustafa, Nakao, Matsuno, Greay, Evasco, Barker and Barker2021; Current study). Both Alveonasus and Proknekalia were considered special branches within the genus Ornithodoros and attempts to relate these lineages to Argas was considered to be based on biological artefacts (Keirans et al., Reference Keirans, Hoogstraal and Clifford1977). However, the current study shows that both mitochondrial and nuclear genetic data support an association within the Argasinae. Together with the cladistic data from Klompen and Oliver (Reference Klompen and Oliver1993), the evidence that these genera belong within the Argasinae is convincing. It is of interest that Alveonasus clusters within a clade formed by Ogadenus, Otobius lagophilus, Navis, Proknekalia and Secretargas. Hoogstraal (Reference Hoogstraal1985) placed Alveonasus within the Ornithodorinae, but did recognize a resemblance between Alveonasus, Ogadenus and Secretargas. Pospelova-Shtrom (Reference Pospelova-Shtrom1969) also placed Alveonasus (including Proknekalia and Ogadenus) and Otobius in the Otobiini tribe, although this tribe was placed within the Ornithodorinae. There was therefore recognition of relationships between these genera, perhaps obscured by the conviction that Alveonasus had to belong to the Ornithodorinae (Keirans et al., Reference Keirans, Hoogstraal and Clifford1977). Cladistic analysis placed Alveonasus, Ogadenus, Proknekalia and Secretargas into a larger clade of related lineages termed the Alveonasus group (Klompen, Reference Klompen1992).
Given that O. lagophilus also group within the Argasinae, it is evident that there is an evolutionary relationship with other Argasinae genera. Conversely, the surprising finding that O. megnini group within the Ornithodorinae needs to be addressed. The placement of O. lagophilus within Otobius seemed fairly straightforward. Adults from both species presents a panduriform shape and adults do not feed (Clifford et al., Reference Clifford, Kohls and Sonenshine1964; Herring and Beck, Reference Herring and Beck1965). Nymphs present posterodorsal spines, although these differ in number and size between O. megnini and O. lagophilus, with the latter's spines more slender (Herring and Beck, Reference Herring and Beck1965). Conversely, larvae from O. lagophilus lack eyes while O. megnini possess 2 pairs of eyes (Herring and Beck, Reference Herring and Beck1965). Otobius lagophilus parasitize rabbits exclusively and is found on the face, while O. megnini parasitizes a range of domestic animals and is usually found in the ears. Otobius lagophilus also completes 1 larval and 1 nymphal moult, while O. megnini completes 2 nymphal moults (Herring and Beck, Reference Herring and Beck1965). There is therefore significant differences that may have indicated that these belong to different genera. Even so, both species lack Pd1 palpal setae, a characteristic used to differentiate Argasinae and Ornithodorinae, the former possessing palpal setae Pd1 (Klompen, Reference Klompen1992; Klompen and Oliver, Reference Klompen and Oliver1993). In the case of O. lagophilus these setae would have been lost. Loss of morphological characters within lineages may be more probable than gain of a character via independent convergent evolution (homoplasy). Some Argasinae may therefore lack Pd1 palpal setae and this character may not be stable for all members, especially since its functional significance is not apparent for the lineage.
Paraphyly of the genus Alveonasus: larger implications
It may be expected that members from the same genus and geographic region should share a recent common ancestor and therefore group as sister clades. However, even though A. canestrinii and A. lahorensis occur in the same region (Hosseini-Chegeni et al., Reference Hosseini-Chegeni, Tavakoli, Koshki and Khedri2019), they do not group as sister clades or even as a monophyletic clade to the exclusion of other genera (Fig. 3). Cladistic analysis of larval characters also failed to provide support for a monophyletic Alveonasus when A. canestrinii, A. eboris and A. lahorensis was compared (Klompen, Reference Klompen1992). Additional support for paraphyly may be found in the unique biology of A. lahorensis which is the only argasid species to display 2-host behaviour with the larvae and nymphs feeding on the same host for up to 3–6 weeks before the engorged third-instar nymph drops and moults to an adult (Hoogstraal, Reference Hoogstraal1985). This suggests that Alveonasus may be comprised of different independent lineages (i.e. genera). In fact, given the wide geographic distribution observed for members of Alveonasus, it may be expected that Alveonasus may likely be comprised of a large number of independent lineages, probably linked to their diverse geographic distribution (Table 1). Indeed, a similar phenomenon is seen for the subgenus Pavlovskyella, where different geographic lineages group as unique clades in the Ornithodorinae (Mans et al., Reference Mans, Featherston, Kvas, Pillay, de Klerk, Pienaar, de Castro, Schwan, Lopez, Teel, Pérez de León, Sonenshine, Egekwu, Bakkes, Heyne, Kanduma, Nyangiwe, Bouattour and Latif2019). It also implies that the eponymous feature of Alveonasus, the madreporian sculpturing of the integument (Clifford et al., Reference Clifford, Kohls and Sonenshine1964), evolved independently or derive from similar, but independent developmental pathways (Mans, Reference Mans2023). If an ancestral developmental pathway for the Alveonasus sensu Klompen (Reference Klompen1992) group was present, perhaps characterized by polygonal depressions surrounded by ridges, formed by integumental folds from the centre of these depressions (Klompen and Oliver, Reference Klompen and Oliver1993), it resulted in numerous integumental variations, some madreporean and others more accentuated by the ridges. Depending on how the integumental development plan unfolds in each species, the specific patterns observed may not be stable enough for genus level classification.
Given the recent surprising placement of O. lagophilus in the Argasinae (Kneubehl et al., Reference Kneubehl, Muñoz-Leal, Filatov, de Klerk, Pienaar, Lohmeyer, Bermúdez, Suriyamongkol, Mali, Kanduma, Latif, Sarih, Bouattour, de León, Teel, Labruna, Mans and Lopez2022), the placement of other members of Alveonasus in the Argasinae (and in Alveonasus) should be empirically confirmed. While it is likely that all members would eventually group within a larger clade formed by Alveonasus, Navis, Ogadenus, Proknekalia, Secretargas and O. lagophilus (the Alveonasus group sensu Klompen, Reference Klompen1992), this cannot be taken for granted. A concerted effort to place the remaining Alveonasus species will be important, since it is likely that the unexpected grouping of the remaining lineages may profoundly impact our understanding of the evolution of this group.
Conclusion
The current study showed that the type species for Alveonasus group within the Argasinae and not the Ornithodorinae as suggested by the American (Clifford et al., Reference Clifford, Kohls and Sonenshine1964; Hoogstraal, Reference Hoogstraal1985; Guglielmone et al., Reference Guglielmone, Robbins, Apanaskevich, Petney, Estrada-Peña, Horak, Shao and Barker2010), Russian (Pospelova-Shtrom, Reference Pospelova-Shtrom1946, Reference Pospelova-Shtrom1969) and French (Camicas and Morel, Reference Camicas and Morel1977; Camicas et al., Reference Camicas, Hervy, Adam and Morel1998) schools. Inclusion of Alveonasus in the Argasinae has been indicated by previous molecular studies using nuclear 18S rRNA and mitochondrial 16S rRNA (Black et al., Reference Black, Klompen and Keirans1997; Zhao et al., Reference Zhao, Lin, Li, Li, He, Zhang, Pan, Wang, Gao, Johnson, Yuan, Lv, Wu and Liu2018). However, the inclusion of Alveonasus in the Argasinae was not specifically discussed nor proposed in these studies, nor recognized by the tick community (except for Mans et al., Reference Mans, Featherston, Kvas, Pillay, de Klerk, Pienaar, de Castro, Schwan, Lopez, Teel, Pérez de León, Sonenshine, Egekwu, Bakkes, Heyne, Kanduma, Nyangiwe, Bouattour and Latif2019 and Mans et al., Reference Mans, Kelava, Pienaar, Featherston, de Castro, Quetglas, Reeves, Durden, Miller, Laverty, Shao, Takano, Kawabata, Moustafa, Nakao, Matsuno, Greay, Evasco, Barker and Barker2021). The current study is therefore the first to provide evidence that unambiguously places Alveonasus in the Argasinae and recognize its placement in this subfamily, after the proposal by Klompen and Oliver (Reference Klompen and Oliver1993), that Alveonasus should be placed in the Argasinae based on cladistic analysis. This is also one of the last major genera for which mitochondrial genome data has been reported. The only formally recognized genus or lineage for which mitochondrial genome data has not yet been reported is the subgenus Microargas (monotypic species Ornithodoros transversus Banks (1902)). It is becoming clear that many more evolutionary independent lineages may exist in the Argasidae that may warrant generic status that include members from Alectorobius, Pavlovskyella, Otobius and now Alveonasus (Mans, Reference Mans2023).
Data availability statement
All data in the study is available in the public databases as listed by their accession numbers.
Acknowledgements
All data in the study is available in the public databases as listed by their accession numbers.
Author contributions
BJM: Study design, data analysis, writing and revision; LCD: writing and revision, RP: data analysis, writing and revision; MdC: data analysis, writing and revision; MK: sample collection, writing and revision; MMA: sample collection, writing and revision; AA: sample collection, writing and revision; AA: Study design, data analysis, writing and revision.
Financial support
This study was supported by the National Research Foundation of South Africa (Grant Number: 137966). The researchers supporting project number (RSP2024R494), King Saud University, Riyadh, Saudi Arabia.
Competing interests
The authors declare there are no conflicts of interest.
Ethical standards
Not applicable