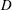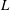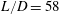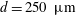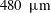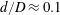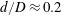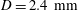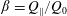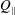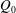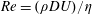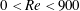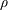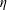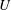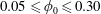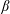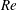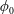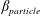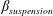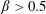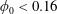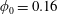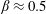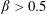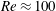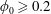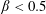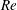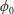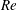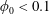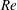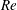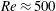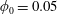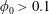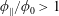12 results
Nomenclature for Pediatric and Congenital Cardiac Care: Unification of Clinical and Administrative Nomenclature – The 2021 International Paediatric and Congenital Cardiac Code (IPCCC) and the Eleventh Revision of the International Classification of Diseases (ICD-11)
- Part of
- Jeffrey P. Jacobs, Rodney C. G. Franklin, Marie J. Béland, Diane E. Spicer, Steven D. Colan, Henry L. Walters III, Frédérique Bailliard, Lucile Houyel, James D. St. Louis, Leo Lopez, Vera D. Aiello, J. William Gaynor, Otto N. Krogmann, Hiromi Kurosawa, Bohdan J. Maruszewski, Giovanni Stellin, Paul Morris Weinberg, Marshall Lewis Jacobs, Jeffrey R. Boris, Meryl S. Cohen, Allen D. Everett, Jorge M. Giroud, Kristine J. Guleserian, Marina L. Hughes, Amy L. Juraszek, Stephen P. Seslar, Charles W. Shepard, Shubhika Srivastava, Andrew C. Cook, Adrian Crucean, Lazaro E. Hernandez, Rohit S. Loomba, Lindsay S. Rogers, Stephen P. Sanders, Jill J. Savla, Elif Seda Selamet Tierney, Justin T. Tretter, Lianyi Wang, Martin J. Elliott, Constantine Mavroudis, Christo I. Tchervenkov
-
- Journal:
- Cardiology in the Young / Volume 31 / Issue 7 / July 2021
- Published online by Cambridge University Press:
- 29 July 2021, pp. 1057-1188
-
- Article
-
- You have access Access
- Open access
- HTML
- Export citation
-
Substantial progress has been made in the standardization of nomenclature for paediatric and congenital cardiac care. In 1936, Maude Abbott published her Atlas of Congenital Cardiac Disease, which was the first formal attempt to classify congenital heart disease. The International Paediatric and Congenital Cardiac Code (IPCCC) is now utilized worldwide and has most recently become the paediatric and congenital cardiac component of the Eleventh Revision of the International Classification of Diseases (ICD-11). The most recent publication of the IPCCC was in 2017. This manuscript provides an updated 2021 version of the IPCCC.
The International Society for Nomenclature of Paediatric and Congenital Heart Disease (ISNPCHD), in collaboration with the World Health Organization (WHO), developed the paediatric and congenital cardiac nomenclature that is now within the eleventh version of the International Classification of Diseases (ICD-11). This unification of IPCCC and ICD-11 is the IPCCC ICD-11 Nomenclature and is the first time that the clinical nomenclature for paediatric and congenital cardiac care and the administrative nomenclature for paediatric and congenital cardiac care are harmonized. The resultant congenital cardiac component of ICD-11 was increased from 29 congenital cardiac codes in ICD-9 and 73 congenital cardiac codes in ICD-10 to 318 codes submitted by ISNPCHD through 2018 for incorporation into ICD-11. After these 318 terms were incorporated into ICD-11 in 2018, the WHO ICD-11 team added an additional 49 terms, some of which are acceptable legacy terms from ICD-10, while others provide greater granularity than the ISNPCHD thought was originally acceptable. Thus, the total number of paediatric and congenital cardiac terms in ICD-11 is 367. In this manuscript, we describe and review the terminology, hierarchy, and definitions of the IPCCC ICD-11 Nomenclature. This article, therefore, presents a global system of nomenclature for paediatric and congenital cardiac care that unifies clinical and administrative nomenclature.
The members of ISNPCHD realize that the nomenclature published in this manuscript will continue to evolve. The version of the IPCCC that was published in 2017 has evolved and changed, and it is now replaced by this 2021 version. In the future, ISNPCHD will again publish updated versions of IPCCC, as IPCCC continues to evolve.
Suspension flow through an asymmetric T-junction
- Sojwal Manoorkar, Sreenath Krishnan, Omer Sedes, Eric S. G. Shaqfeh, Gianluca Iaccarino, Jeffrey F. Morris
-
- Journal:
- Journal of Fluid Mechanics / Volume 844 / 10 June 2018
- Published online by Cambridge University Press:
- 04 April 2018, pp. 247-273
-
- Article
- Export citation
-
The flow of a suspension through a bifurcating channel is studied experimentally and by computational methods. The geometry considered is an ‘asymmetric T’, as flow in the entering branch divides to either continue straight or to make a right angle turn. All branches are of the same square cross-section of side length
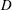 $D$, with inlet and outlet section lengths
$D$, with inlet and outlet section lengths 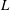 $L$ yielding
$L$ yielding 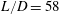 $L/D=58$ in the experiments. The suspensions are composed of neutrally buoyant spherical particles in a Newtonian liquid, with mean particle diameters of
$L/D=58$ in the experiments. The suspensions are composed of neutrally buoyant spherical particles in a Newtonian liquid, with mean particle diameters of 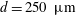 $d=250~\unicode[STIX]{x03BC}\text{m}$ and
$d=250~\unicode[STIX]{x03BC}\text{m}$ and 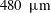 $480~\unicode[STIX]{x03BC}\text{m}$ resulting in
$480~\unicode[STIX]{x03BC}\text{m}$ resulting in 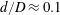 $d/D\approx 0.1$ to
$d/D\approx 0.1$ to 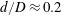 $d/D\approx 0.2$ for
$d/D\approx 0.2$ for 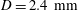 $D=2.4~\text{mm}$. The flow rate ratio
$D=2.4~\text{mm}$. The flow rate ratio 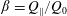 $\unicode[STIX]{x1D6FD}=Q_{\Vert }/Q_{0}$, defined for the bulk, fluid and particles, is used to characterize the flow behaviour; here
$\unicode[STIX]{x1D6FD}=Q_{\Vert }/Q_{0}$, defined for the bulk, fluid and particles, is used to characterize the flow behaviour; here 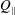 $Q_{\Vert }$ and
$Q_{\Vert }$ and 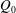 $Q_{0}$ are volumetric flow rates in the straight outlet branch and inlet branch, respectively. The channel Reynolds number
$Q_{0}$ are volumetric flow rates in the straight outlet branch and inlet branch, respectively. The channel Reynolds number 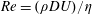 $Re=(\unicode[STIX]{x1D70C}DU)/\unicode[STIX]{x1D702}$ was varied over
$Re=(\unicode[STIX]{x1D70C}DU)/\unicode[STIX]{x1D702}$ was varied over 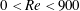 $0<Re<900$, with
$0<Re<900$, with 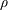 $\unicode[STIX]{x1D70C}$ and
$\unicode[STIX]{x1D70C}$ and 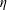 $\unicode[STIX]{x1D702}$ the fluid density and viscosity, respectively, and
$\unicode[STIX]{x1D702}$ the fluid density and viscosity, respectively, and 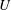 $U$ the mean velocity in the inlet channel; the inlet particle volume fraction was
$U$ the mean velocity in the inlet channel; the inlet particle volume fraction was 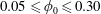 $0.05\leqslant \unicode[STIX]{x1D719}_{0}\leqslant 0.30$. Experimental and numerical results for single-phase Newtonian fluid both show
$0.05\leqslant \unicode[STIX]{x1D719}_{0}\leqslant 0.30$. Experimental and numerical results for single-phase Newtonian fluid both show 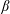 $\unicode[STIX]{x1D6FD}$ increasing with
$\unicode[STIX]{x1D6FD}$ increasing with 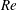 $Re$, implying more material tending toward the straight branch as the inertia of the flow increases. In suspension flow at small
$Re$, implying more material tending toward the straight branch as the inertia of the flow increases. In suspension flow at small 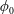 $\unicode[STIX]{x1D719}_{0}$, inertial migration of particles in the inlet branch affects the flow rate ratio for particles (
$\unicode[STIX]{x1D719}_{0}$, inertial migration of particles in the inlet branch affects the flow rate ratio for particles (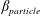 $\unicode[STIX]{x1D6FD}_{\mathit{particle}}$) and suspension (
$\unicode[STIX]{x1D6FD}_{\mathit{particle}}$) and suspension (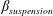 $\unicode[STIX]{x1D6FD}_{\mathit{suspension}}$). The flow split for the bulk suspension satisfies
$\unicode[STIX]{x1D6FD}_{\mathit{suspension}}$). The flow split for the bulk suspension satisfies 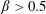 $\unicode[STIX]{x1D6FD}>0.5$ for
$\unicode[STIX]{x1D6FD}>0.5$ for 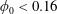 $\unicode[STIX]{x1D719}_{0}<0.16$ while
$\unicode[STIX]{x1D719}_{0}<0.16$ while 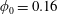 $\unicode[STIX]{x1D719}_{0}=0.16$ crosses from
$\unicode[STIX]{x1D719}_{0}=0.16$ crosses from 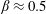 $\unicode[STIX]{x1D6FD}\approx 0.5$ to
$\unicode[STIX]{x1D6FD}\approx 0.5$ to 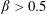 $\unicode[STIX]{x1D6FD}>0.5$ at
$\unicode[STIX]{x1D6FD}>0.5$ at 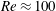 $Re\approx 100$. For
$Re\approx 100$. For 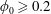 $\unicode[STIX]{x1D719}_{0}\geqslant 0.2$,
$\unicode[STIX]{x1D719}_{0}\geqslant 0.2$, 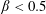 $\unicode[STIX]{x1D6FD}<0.5$ at all
$\unicode[STIX]{x1D6FD}<0.5$ at all 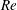 $Re$ studied. A complex dependence of the mean solid fraction in the downstream branches upon inlet fraction
$Re$ studied. A complex dependence of the mean solid fraction in the downstream branches upon inlet fraction 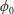 $\unicode[STIX]{x1D719}_{0}$ and
$\unicode[STIX]{x1D719}_{0}$ and 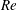 $Re$ is observed: for
$Re$ is observed: for 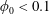 $\unicode[STIX]{x1D719}_{0}<0.1$, the solid fraction in the straight downstream branch initially decreases with
$\unicode[STIX]{x1D719}_{0}<0.1$, the solid fraction in the straight downstream branch initially decreases with 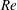 $Re$, before increasing to surpass the inlet fraction at large
$Re$, before increasing to surpass the inlet fraction at large 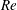 $Re$ (
$Re$ (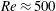 $Re\approx 500$ for
$Re\approx 500$ for 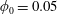 $\unicode[STIX]{x1D719}_{0}=0.05$). At
$\unicode[STIX]{x1D719}_{0}=0.05$). At 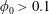 $\unicode[STIX]{x1D719}_{0}>0.1$, the solid fraction in the straight branch satisfies
$\unicode[STIX]{x1D719}_{0}>0.1$, the solid fraction in the straight branch satisfies 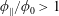 $\unicode[STIX]{x1D719}_{\Vert }/\unicode[STIX]{x1D719}_{0}>1$, and this ratio grows with
$\unicode[STIX]{x1D719}_{\Vert }/\unicode[STIX]{x1D719}_{0}>1$, and this ratio grows with 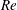 $Re$. Discrete-particle simulations employing immersed boundary and lattice-Boltzmann techniques are used to analyse these phenomena, allowing rationalization of aspects of this complex behaviour as being due to particle migration in the inlet branch.
$Re$. Discrete-particle simulations employing immersed boundary and lattice-Boltzmann techniques are used to analyse these phenomena, allowing rationalization of aspects of this complex behaviour as being due to particle migration in the inlet branch.
10 - Bayesian Models for Flexible Integrative Analysis of Multi-Platform Genomics Data
- from Part B - Vertical Integrative Analysis (General Methods)
-
- By Elizabeth J. McGuffey, United States Naval Academy, Annapolis, MD, Jeffrey S. Morris, UT MD Anderson Cancer Center, Houston, TX, Ganiraju C. Manyam, UT MD Anderson Cancer Center, Houston, TX, Raymond J. Carroll, Texas A&M University, College Station, TX, Veerabhadran Baladandayuthapani, UT MD Anderson Cancer Center, Houston, TX
- George Tseng, University of Pittsburgh, Debashis Ghosh, Pennsylvania State University, Xianghong Jasmine Zhou, University of Southern California
-
- Book:
- Integrating Omics Data
- Published online:
- 05 September 2015
- Print publication:
- 23 September 2015, pp 221-241
-
- Chapter
- Export citation
Contributors
-
- By Ghazi Al-Rawas, Vazken Andréassian, Tianqi Ao, Stacey A. Archfield, Berit Arheimer, András Bárdossy, Trent Biggs, Günter Blöschl, Theresa Blume, Marco Borga, Helge Bormann, Gianluca Botter, Tom Brown, Donald H. Burn, Sean K. Carey, Attilio Castellarin, Francis Chiew, François Colin, Paulin Coulibaly, Armand Crabit, Barry Croke, Siegfried Demuth, Qingyun Duan, Giuliano Di Baldassarre, Thomas Dunne, Ying Fan, Xing Fang, Boris Gartsman, Alexander Gelfan, Mikhail Georgievski, Nick van de Giesen, David C. Goodrich, Hoshin V. Gupta, Khaled Haddad, David M. Hannah, H. A. P. Hapuarachchi, Hege Hisdal, Kamila Hlavčová, Markus Hrachowitz, Denis A. Hughes, Günter Humer, Ruud Hurkmans, Vito Iacobellis, Elena Ilyichyova, Hiroshi Ishidaira, Graham Jewitt, Shaofeng Jia, Jeffrey R. Kennedy, Anthony S. Kiem, Robert Kirnbauer, Thomas R. Kjeldsen, Jürgen Komma, Leonid M. Korytny, Charles N. Kroll, George Kuczera, Gregor Laaha, Henny A. J. van Lanen, Hjalmar Laudon, Jens Liebe, Shijun Lin, Göran Lindström, Suxia Liu, Jun Magome, Danny G. Marks, Dominic Mazvimavi, Jeffrey J. McDonnell, Brian L. McGlynn, Kevin J. McGuire, Neil McIntyre, Thomas A. McMahon, Ralf Merz, Robert A. Metcalfe, Alberto Montanari, David Morris, Roger Moussa, Lakshman Nandagiri, Thomas Nester, Taha B. M. J. Ouarda, Ludovic Oudin, Juraj Parajka, Charles S. Pearson, Murray C. Peel, Charles Perrin, John W. Pomeroy, David A. Post, Ataur Rahman, Liliang Ren, Magdalena Rogger, Dan Rosbjerg, José Luis Salinas, Jos Samuel, Eric Sauquet, Hubert H. G. Savenije, Takahiro Sayama, John C. Schaake, Kevin Shook, Murugesu Sivapalan, Jon Olav Skøien, Chris Soulsby, Christopher Spence, R. ‘Sri’ Srikanthan, Tammo S. Steenhuis, Jan Szolgay, Yasuto Tachikawa, Kuniyoshi Takeuchi, Lena M. Tallaksen, Dörthe Tetzlaff, Sally E. Thompson, Elena Toth, Peter A. Troch, Remko Uijlenhoet, Carl L. Unkrich, Alberto Viglione, Neil R. Viney, Richard M. Vogel, Thorsten Wagener, M. Todd Walter, Guoqiang Wang, Markus Weiler, Rolf Weingartner, Erwin Weinmann, Hessel Winsemius, Ross A. Woods, Dawen Yang, Chihiro Yoshimura, Andy Young, Gordon Young, Erwin Zehe, Yongqiang Zhang, Maichun C. Zhou
- Edited by Günter Blöschl, Technische Universität Wien, Austria, Murugesu Sivapalan, University of Illinois, Urbana-Champaign, Thorsten Wagener, University of Bristol, Alberto Viglione, Technische Universität Wien, Austria, Hubert Savenije, Technische Universiteit Delft, The Netherlands
-
- Book:
- Runoff Prediction in Ungauged Basins
- Published online:
- 05 April 2013
- Print publication:
- 18 April 2013, pp ix-xiv
-
- Chapter
- Export citation
Contributors
-
- By Charles E. Argoff, Gerard A. Banez, Samantha Boris-Karpel, Barbara K. Bruce, Alexandra S. Bullough, Annmarie Cano, Victor T. Chang, Elizabeth A. Clark, Daniel J. Clauw, June L. Dahl, Tam K. Dao, Amber M. Davis, Courtney L. Dixon, Michael H. Ebert, Robin M. Gallagher, Gerald W. Grass, Carmen R. Green, Jay Gunkelman, Bradford D. Hare, Jennifer A. Haythornthwaite, Jaclyn Heller Issner, W. Michael Hooten, Mark P. Jensen, Mark E. Jones, Robert D. Kerns, Raphael J. Leo, Morris Maizels, Mary E. Murawski, Brooke Myers-Sorger, Akiko Okifuji, Renata Okonkwo, John D. Otis, Stacy C. Parenteau, Laura E. Pence, Donald B. Penzien, Donna B. Pincus, Ellyn Poltrock Stein, Wendy J. Quinton, Jeanetta C. Rains, M. Carrington Reid, Thomas J. Romano, Jeffrey D. Rome, Robert L. Ruff, Suzanne S. Ruff, Steven H. Sanders, Ingra Schellenberg, John J. Sellinger, Howard S. Smith, Brenda Stoelb, Jon Streltzer, Mark D. Sullivan, Kimberly S. Swanson, Gabriel Tan, Stephen Thielke, Beverly E. Thorn, Cynthia O. Townsend, Dennis C. Turk, Stephanie C. Wallio, Lawrence J. Weinberger, David A. Williams, Hilary Wilson
- Edited by Michael H. Ebert, Yale University, Connecticut, Robert D. Kerns, Yale University, Connecticut
-
- Book:
- Behavioral and Psychopharmacologic Pain Management
- Published online:
- 10 January 2011
- Print publication:
- 25 November 2010, pp ix-xii
-
- Chapter
- Export citation
Contributors
-
- By Claude Alain, Amy F. T. Arnsten, Lars Bäckman, Malcolm A. Binns, Sandra E. Black, S. Thomas Carmichael, Keith D. Cicerone, Maurizio Corbetta, Bruce Crosson, Jeffrey L. Cummings, Deirdre R. Dawson, Michael deRiesthal, Roger A. Dixon, Laura Eggermont, Kirk I. Erickson, Anthony Feinstein, Susan M. Fitzpatrick, Fu Qiang Gao, Douglas D. Garrett, Omar Ghaffar, Robbin Gibb, Elizabeth L. Glisky, Martha L. Glisky, Leslie J. Gonzalez Rothi, Cheryl L. Grady, Carol Greenwood, Gerri Hanten, Richard G. Hunter, Masud Husain, Narinder Kapur, Bryan Kolb, Arthur F. Kramer, Susan A. Leon, Harvey S. Levin, Brian Levine, Nadina Lincoln, Thomas W. McAllister, Edward McAuley, Bruce S. McEwen, David M. Morris, Stephen E. Nadeau, Roshan das Nair, Matthew Parrott, Jennie Ponsford, George P. Prigatano, Joel Ramirez, John M. Ringman, Ian H. Robertson, Amy D. Rodriguez, John C. Rosenbek, Bernhard Ross, Erik Scherder, Victoria Singh-Curry, Trudi Stickland, Donald T. Stuss, Edward Taub, Gary R. Turner, Harry V. Vinters, Samuel Weiss, John Whyte, Barbara A. Wilson, Gordon Winocur, J. Martin Wojtowicz
- Edited by Donald T. Stuss, University of Toronto, Gordon Winocur, University of Toronto, Ian H. Robertson, Trinity College, Dublin
-
- Book:
- Cognitive Neurorehabilitation
- Published online:
- 05 September 2015
- Print publication:
- 11 September 2008, pp ix-xiv
-
- Chapter
- Export citation
1 - An Introduction to High-Throughput Bioinformatics Data
- Edited by Kim-Anh Do, University of Texas, MD Anderson Cancer Center, Peter Müller, Swiss Federal Institute of Technology, Zürich, Marina Vannucci, Rice University, Houston
-
- Book:
- Bayesian Inference for Gene Expression and Proteomics
- Published online:
- 23 November 2009
- Print publication:
- 24 July 2006, pp 1-39
-
- Chapter
- Export citation
-
Summary
Abstract
High throughput biological assays supply thousands of measurements per sample, and the sheer amount of related data increases the need for better models to enhance inference. Such models, however, are more effective if they take into account the idiosyncracies associated with the specific methods of measurement: where the numbers come from. We illustrate this point by describing three different measurement platforms: microarrays, serial analysis of gene expression (SAGE), and proteomic mass spectrometry.
Introduction
In our view, high-throughput biological experiments involve three phases: experimental design, measurement and preprocessing, and postprocessing. These phases are otherwise known as deciding what you want to measure, getting the right numbers and assembling them in a matrix, and mining the matrix for information. Of these, it is primarily the middle step that is unique to the particular measurement technology employed, and it is there that we shall focus our attention. This is not meant to imply that the other steps are less important! It is still a truism that the best analysis may not be able to save you if your experimental design is poor.
We simply wish to emphasize that each type of data has its own quirks associated with the methods of measurement, and understanding these quirks allows us to craft ever more sophisticated probability models to improve our analyses. These probability models should ideally also let us exploit information across measurements made in parallel, and across samples. Crafting these models leads to the development of brand-new statistical methods, many of which are discussed in this volume.
In this chapter, we address the importance of measurement-specific methodology by discussing several approaches in detail. We cannot be all-inclusive, so we shall focus on three.
12 - Bayesian Mixture Models for Gene Expression and Protein Profiles
- Edited by Kim-Anh Do, University of Texas, MD Anderson Cancer Center, Peter Müller, Swiss Federal Institute of Technology, Zürich, Marina Vannucci, Rice University, Houston
-
- Book:
- Bayesian Inference for Gene Expression and Proteomics
- Published online:
- 23 November 2009
- Print publication:
- 24 July 2006, pp 238-253
-
- Chapter
- Export citation
-
Summary
Abstract
We review the use of semiparametric mixture models for Bayesian inference in high-throughput genomic data. We discuss three specific approaches for microarray data, for protein mass spectrometry experiments, and for serial analysis of gene expression (SAGE) data. For the microarray data and the protein mass spectrometry we assume group comparison experiments, that is, experiments that seek to identify genes and proteins that are differentially expressed across two biologic conditions of interest. For the SAGE data example we consider inference for a single biologic sample. For all three applications we use flexible mixture models to implement inference. For the microarray data we define a Dirichlet process mixture of normal model. For the mass spectrometry data we introduce a mixture of Beta model. The proposed inference for SAGE data is based on a semiparametric mixture of Poisson distributions.
Introduction
We discuss semiparametric Bayesian data analysis for high-throughput genomic data. We introduce suitable semiparametric mixture models to implement inference for microarray data, mass spectrometry data, and SAGE data. The proposed models include a Dirichlet process mixture of normals for microarray data, a mixture of Beta distributions with a random number of terms for mass spectrometry data, and a Dirichlet process mixture of Poisson model for SAGE data. For the microarray data and the protein mass spectrometry data we consider experiments that compare two biologic conditions of interest. We assume that the aim of the experiment is to find genes and proteins, respectively, that are differentially expressed under the two conditions. For the SAGE example, we propose data analysis for a single biologic sample.
14 - Analysis of Mass Spectrometry Data Using Bayesian Wavelet-Based Functional Mixed Models
- Edited by Kim-Anh Do, University of Texas, MD Anderson Cancer Center, Peter Müller, Swiss Federal Institute of Technology, Zürich, Marina Vannucci, Rice University, Houston
-
- Book:
- Bayesian Inference for Gene Expression and Proteomics
- Published online:
- 23 November 2009
- Print publication:
- 24 July 2006, pp 269-292
-
- Chapter
- Export citation
-
Summary
Abstract
In this chapter, we demonstrate how to analyze MALDI-TOF/SELDI-TOF mass spectrometry data using the wavelet-based functional mixed model introduced by J. S. Morris and R. J. Carroll (wavelet-based functional mixed models. Journal of the Royal Statistical Society, Series B, in 2006, which generalizes the linear mixed model to the case of functional data. This approach models each spectrum as a function, and is very general, accommodating a broad class of experimental designs and allowing one to model nonparametric functional effects for various factors, which can be conditions of interest (e.g., cancer/normal) or experimental factors (blocking factors). Inference on these functional effects allows us to identify protein peaks related to various outcomes of interest, including dichotomous outcomes, categorical outcomes, continuous outcomes, and any interactions among factors. Functional random effects make it possible to account for correlation between spectra from the same individual or block in a flexible manner. After fitting this model using Markov chain Monte Carlo, the output can be used to perform peak detection and identify the peaks that are related to factors of interest, while automatically adjusting for nonlinear block effects that are characteristic of these data. We apply this method to mass spectrometry data from a University of Texas M.D. Anderson Cancer Center experiment studying the serum proteome of mice injected with one of two cell lines in one of two organs. This methodology appears promising for the analysis of mass spectrometry proteomics data, and may have application for other types of proteomics data as well.
Introduction
MALDI-TOF is a mass-spectrometry-based proteomics method that yields spiky functional data, with peaks corresponding to proteins present in the biological sample.
13 - Shrinkage Estimation for SAGE Data Using a Mixture Dirichlet Prior
- Edited by Kim-Anh Do, University of Texas, MD Anderson Cancer Center, Peter Müller, Swiss Federal Institute of Technology, Zürich, Marina Vannucci, Rice University, Houston
-
- Book:
- Bayesian Inference for Gene Expression and Proteomics
- Published online:
- 23 November 2009
- Print publication:
- 24 July 2006, pp 254-268
-
- Chapter
- Export citation
-
Summary
Abstract
Serial analysis of gene expression (SAGE) is a technique for estimating the gene expression profile of a biological sample. Any efficient inference in SAGE must be based upon efficient estimates of these gene expression profiles, which consist of the estimated relative abundances for each mRNA species present in the sample. The data from SAGE experiments are counts for each observed mRNA species, and can be modeled using a multinomial distribution with two characteristics: skewness in the distribution of relative abundances and small sample size relative to the dimension. As a result of these characteristics, a given SAGE sample will fail to capture a large number of expressed mRNA species present in the tissue. Standard empirical estimates of the relative abundances effectively ignore these missing, unobserved species, and consequently tend to also overestimate the abundance of the scarce observed species comprising a vast majority of the total. In this chapter, we review a new Bayesian procedure that yields improved estimates for the missing and scarce species without trading off much efficiency for the abundant species. The key to the procedure is the mixture Dirichlet prior, which stochastically partitions the mRNA species into abundant and scarce strata, with each stratum modeled with its own multivariate prior, a scalar multiple of a symmetric Dirichlet. Simulation studies demonstrate that the resulting shrinkage estimators have efficiency advantages over the maximum likelihood estimator for SAGE scenarios simulated.
Introduction
Serial analysis of gene expression (SAGE) is a method for estimating the gene expression profile of a biological sample of interest. In this chapter, we review a method introduced in Morris, Baggerly, and Coombes (2003) for obtaining Bayesian shrinkage estimates of these profiles using a fully specified probability model.
Looking Backward, Looking Forward: MLA Members Speak
- April Alliston, Elizabeth Ammons, Jean Arnold, Nina Baym, Sandra L. Beckett, Peter G. Beidler, Roger A. Berger, Sandra Bermann, J.J. Wilson, Troy Boone, Alison Booth, Wayne C. Booth, James Phelan, Marie Borroff, Ihab Hassan, Ulrich Weisstein, Zack Bowen, Jill Campbell, Dan Campion, Jay Caplan, Maurice Charney, Beverly Lyon Clark, Robert A. Colby, Thomas C. Coleman III, Nicole Cooley, Richard Dellamora, Morris Dickstein, Terrell Dixon, Emory Elliott, Caryl Emerson, Ann W. Engar, Lars Engle, Kai Hammermeister, N. N. Feltes, Mary Anne Ferguson, Annie Finch, Shelley Fisher Fishkin, Jerry Aline Flieger, Norman Friedman, Rosemarie Garland-Thomson, Sandra M. Gilbert, Laurie Grobman, George Guida, Liselotte Gumpel, R. K. Gupta, Florence Howe, Cathy L. Jrade, Richard A. Kaye, Calhoun Winton, Murray Krieger, Robert Langbaum, Richard A. Lanham, Marilee Lindemann, Paul Michael Lützeler, Thomas J. Lynn, Juliet Flower MacCannell, Michelle A. Massé, Irving Massey, Georges May, Christian W. Hallstein, Gita May, Lucy McDiarmid, Ellen Messer-Davidow, Koritha Mitchell, Robin Smiles, Kenyatta Albeny, George Monteiro, Joel Myerson, Alan Nadel, Ashton Nichols, Jeffrey Nishimura, Neal Oxenhandler, David Palumbo-Liu, Vincent P. Pecora, David Porter, Nancy Potter, Ronald C. Rosbottom, Elias L. Rivers, Gerhard F. Strasser, J. L. Styan, Marianna De Marco Torgovnick, Gary Totten, David van Leer, Asha Varadharajan, Orrin N. C. Wang, Sharon Willis, Louise E. Wright, Donald A. Yates, Takayuki Yokota-Murakami, Richard E. Zeikowitz, Angelika Bammer, Dale Bauer, Karl Beckson, Betsy A. Bowen, Stacey Donohue, Sheila Emerson, Gwendolyn Audrey Foster, Jay L. Halio, Karl Kroeber, Terence Hawkes, William B. Hunter, Mary Jambus, Willard F. King, Nancy K. Miller, Jody Norton, Ann Pellegrini, S. P. Rosenbaum, Lorie Roth, Robert Scholes, Joanne Shattock, Rosemary T. VanArsdel, Alfred Bendixen, Alarma Kathleen Brown, Michael J. Kiskis, Debra A. Castillo, Rey Chow, John F. Crossen, Robert F. Fleissner, Regenia Gagnier, Nicholas Howe, M. Thomas Inge, Frank Mehring, Hyungji Park, Jahan Ramazani, Kenneth M. Roemer, Deborah D. Rogers, A. LaVonne Brown Ruoff, Regina M. Schwartz, John T. Shawcross, Brenda R. Silver, Andrew von Hendy, Virginia Wright Wexman, Britta Zangen, A. Owen Aldridge, Paula R. Backscheider, Roland Bartel, E. M. Forster, Milton Birnbaum, Jonathan Bishop, Crystal Downing, Frank H. Ellis, Roberto Forns-Broggi, James R. Giles, Mary E. Giles, Susan Blair Green, Madelyn Gutwirth, Constance B. Hieatt, Titi Adepitan, Edgar C. Knowlton, Jr., Emanuel Mussman, Sally Todd Nelson, Robert O. Preyer, David Diego Rodriguez, Guy Stern, James Thorpe, Robert J. Wilson, Rebecca S. Beal, Joyce Simutis, Betsy Bowden, Sara Cooper, Wheeler Winston Dixon, Tarek el Ariss, Richard Jewell, John W. Kronik, Wendy Martin, Stuart Y. McDougal, Hugo Méndez-Ramírez, Ivy Schweitzer, Armand E. Singer, G. Thomas Tanselle, Tom Bishop, Mary Ann Caws, Marcel Gutwirth, Christophe Ippolito, Lawrence D. Kritzman, James Longenbach, Tim McCracken, Wolfe S. Molitor, Diane Quantic, Gregory Rabassa, Ellen M. Tsagaris, Anthony C. Yu, Betty Jean Craige, Wendell V. Harris, J. Hillis Miller, Jesse G. Swan, Helene Zimmer-Loew, Peter Berek, James Chandler, Hanna K. Charney, Philip Cohen, Judith Fetterley, Herbert Lindenberger, Julia Reinhard Lupton, Maximillian E. Novak, Richard Ohmann, Marjorie Perloff, Mark Reynolds, James Sledd, Harriet Turner, Marie Umeh, Flavia Aloya, Regina Barreca, Konrad Bieber, Ellis Hanson, William J. Hyde, Holly A. Laird, David Leverenz, Allen Michie, J. Wesley Miller, Marvin Rosenberg, Daniel R. Schwarz, Elizabeth Welt Trahan, Jean Fagan Yellin
-
- Journal:
- PMLA / Publications of the Modern Language Association of America / Volume 115 / Issue 7 / December 2000
- Published online by Cambridge University Press:
- 23 October 2020, pp. 1986-2078
- Print publication:
- December 2000
-
- Article
- Export citation
Homage to Rudolf Carnap
- Herbert Feigl, Carl G. Hempel, Richard C. Jeffrey, W. V. Quine, A. Shimony, Yehoshua Bar-Hillel, Herbert G. Bohnert, Robert S. Cohen, Charles Hartshorne, David Kaplan, Charles Morris, Maria Reichenbach, Wolfgang Stegmüller
-
- Journal:
- PSA: Proceedings of the Biennial Meeting of the Philosophy of Science Association / Volume 1970 / 1970
- Published online by Cambridge University Press:
- 28 February 2022, pp. XI-LXVI
- Print publication:
- 1970
-
- Article
- Export citation