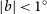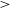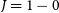Carlosbarbosaite, ideally (UO2)2Nb2O6(OH)2·2H2O, is a new mineral which occurs as a late cavity filling in albite in the Jaguaraçu pegmatite, Jaguaraçu municipality, Minas Gerais, Brazil. The name honours Carlos do Prado Barbosa (1917–2003). Carlosbarbosaite forms long flattened lath-like crystals with a very simple orthorhombic morphology. The crystals are elongated along [001] and flattened on (100); they are up to 120 μm long and 2–5 μm thick. The colour is cream to pale yellow, the streak yellowish white and the lustre vitreous. The mineral is transparent (as individual crystals) to translucent (massive). It is not fluorescent under either long-wave or short-wave ultraviolet radiation. Carlosbarbosaite is biaxial(+) with α = 1.760(5), β = 1.775(5), γ = 1.795(5), 2Vmeas. = 70(1)º, 2Vcalc. = 83º. The orientation is X || a, Y || b, Z || c. Pleochroism is weak, in yellowish green shades, which are most intense in the Z direction. Two samples were analysed. For sample 1, the composition is: UO3 54.52, CaO 2.07, Ce2O3 0.33, Nd2O3 0.49, Nb2O5 14.11, Ta2O5 15.25, TiO2 2.20, SiO2 2.14, Fe2O3 1.08, Al2O3 0.73, H2O (calc.) 11.49, total 104.41 wt.%; the empirical formula is (□0.68Ca0.28Nd0.02Ce0.02)Σ=1.00[U1.44□0.56O2.88(H2O)1.12](Nb0.80Ta0.52Si0.27Ti0.21Al0.11Fe0.10)Σ=2.01 O4.72(OH)3.20(H2O)2.08. For sample 2, the composition is: UO3 41.83, CaO 2.10, Ce2O3 0.31, Nd2O3 1.12, Nb2 O5 14.64, Ta2O5 16.34, TiO2 0.95, SiO2 3.55, Fe2O3 0.89, Al2O3 0.71, H2O (calc.) 14.99, total 97.43 wt.%; the empirical formula is (□0.67Ca0.27Nd0.05Ce0.01)Σ=1.00[U1.04□0.96O2.08(H2O)1.92] (Nb0.79Ta0.53Si0.42Ti0.08Al0.10Fe0.08)Σ=2.00O4.00(OH)3.96(H2O)2.04. The ideal endmember formula is (UO2)2Nb2O6(OH)2·2H2O. Calculated densities are 4.713 g cm-3 (sample 1) and 4.172 g cm-3 (sample 2). Infrared spectra show that both (OH) and H2O are present. The strongest eight X-ray powder-diffraction lines [listed as d in Å (I)(hkl)] are: 8.405(8)(110), 7.081(10)(200), 4.201(9)(220), 3.333(6)(202), 3.053(8)(022), 2.931(7)(420), 2.803(6)(222) and 2.589(5)(040,402). The crystal structure was solved using single-crystal X-ray diffraction (R = 0.037) which gave the following data: orthorhombic, Cmcm, a = 14.150(6), b = 10.395(4), c = 7.529(3) Å, V = 1107(1) Å3, Z = 4. The crystal structure contains a single U site with an appreciable deficiency in electron scattering, which is populated by U atoms and vacancies. The U site is surrounded by seven O atoms in a pentagonal bipyramidal arrangement. The Nb site is coordinated by four O atoms and two OH groups in an octahedral arrangement. The half-occupied tunnel Ca site is coordinated by four O atoms and four H2O groups. Octahedrally coordinated Nb polyhedra share edges and corners to form Nb2O6(OH)2 double chains, and edge-sharing pentagonal bipyramidal U polyhedra form UO5 chains. The Nb2O6(OH)2 and UO5 chains share edges to form an open U—Nb—φ framework with tunnels along [001] that contain Ca(H2O)4 clusters. Carlosbarbosaite is closely related to a family of synthetic U–Nb–ϕ framework tunnel structures, it differs in that is has an (OH)-bearing framework and Ca(H2O)4 tunnel occupant. The structure of carlosbarbosaite resembles that of holfertite.