Introduction
This is the second gene-focused Element of the Cambridge Elements series on Genetics in Epilepsy launched in September 2021 [Reference Poduri, George, Heinzen, Lowenstein, James and Poduri1]. The goal of this Element is to provide an in-depth, state-of-the-art review of clinical, genetic, basic science, and family perspectives on neurological and neurodevelopmental disorders (NDD) associated with pathogenic variants in SCN2A, which encodes a major voltage-gated sodium ion channel (designated NaV1.2) in the brain. The SCN2A-related disorders are clinically heterogenous with features ranging from neonatal and infantile onset epilepsy, late onset epileptic encephalopathy, autism spectrum disorder (ASD), and intellectual disability (ID). In addition to phenotype diversity, the widespread use of clinical genetic testing has resulted in more than 1,000 SCN2A variants deposited in ClinVar. With the explosion in genetic variant identification has come recognition of genotype–phenotype relationships, which when coupled with experimental demonstration of functional perturbations are guiding new therapeutic approaches. Investigations into the biology of SCN2A has led to fundamental discoveries about the physiology and pathophysiology of synaptic connections and neural circuits. As one of the earliest known epilepsy genes, SCN2A has emerged as an important genetic factor in neurodevelopment and NDD.
We hope this Element will provide opportunities for families, trainees, and health care professionals to learn about SCN2A-related disorders. Subsections of this Element offer complete discussions about clinical features, pathophysiology, genetics, model systems, and treatment. This Element begins with perspectives from parents and caregivers of children with these disorders, made possible by a parent-led advocacy group (the FamilieSCN2A Foundation). The following subsections are devoted to clinical features, genotype–phenotype correlations, basic science, and current and future therapeutic approaches. Thus, this Element on SCN2A-related disorders provides a comprehensive and in-depth review of the state of knowledge in this field, which should be valuable to scientists, clinicians, trainees, and families interested in the topic.
In addition to a thorough and informative narrative, this Element is augmented by video content, including interviews with parents of children with SCN2A-related disorders (Video 1); with Dr. Matthew State (Professor and Chair, Department of Psychiatry and Behavioral Sciences, University of California, San Francisco) on the genomics of ASD and the discovery of SCN2A as a major risk factor (Video 2); and with Dr. Steven Petrou (Professor of Neuroscience, University of Melbourne, and Chief Scientific Officer at Praxis Precision Medicines) on his career evolution from academic research to a new pharmaceutical company specializing in precision medicine for rare neurological diseases (Video 3).
This was a team effort that we hope provides inspiration to future clinicians, researchers, and patient advocates. We hope you enjoy learning about this important epilepsy gene.
Patient, Family, and Foundation Perspectives
Pathogenic variants in the SCN2A gene are associated with a broad spectrum of complex NDD that are collectively designated as SCN2A-related disorders. The primary clinical manifestations include epilepsy, ASD, movement disorders, and ID. The severity of these conditions varies among individuals, ranging from mild and well controlled to severe and treatment resistant. Even individuals with mild clinical phenotypes exhibit significant impairments compared to their age-matched peers. Those on the severe end of the spectrum are profoundly affected and heavily reliant on their caregivers for all aspects of daily life. Given the complexity of SCN2A-related disorders, clinical care teams are often multidisciplinary, emphasizing the importance of coordinated efforts to optimize care and minimize clinical risks (Figure 1).
Diagram of multispecialty care of a patient with an SCN2A-related disorder.

FamilieSCN2A Foundation
The FamilieSCN2A Foundation, founded in 2015, is the largest nonprofit patient advocacy organization representing SCN2A-related disorders and has the broadest international footprint among other SCN2A patient-advocacy groups. Focused on creating an engaged ecosystem, FamilieSCN2A Foundation acts as a central liaison between patients, caregivers, scientists, clinicians, and industry. They also work in close partnership with other large foundations such as the Simons Foundation and the Chan Zuckerberg Initiative. “Families” is part of the Foundation’s name because these rare and devastating conditions affect the entire family. The Foundation strives every day and, in every way, to improve the lives of not only the patients but also the entire family.
The missions of the Foundation are to accelerate research, foster a sense of community, and advocate for improvements in the lives of those affected. The Foundation’s vision is centered around achieving effective treatments and cures for all SCN2A-related disorders. The core values of the Foundation are urgency, integrity, collaboration, and inclusion. FamilieSCN2A strives to provide families and professionals with the information and tools needed for a rapid and accurate diagnosis as well as the resources needed to tailor treatments based on the patients’ and families’ goals informed by research knowledge.
Before 2014, there was little hope for children or families diagnosed with an SCN2A-related disorder. There were no specialists treating the condition, no support groups for families desperately seeking answers, and no researchers investigating cures. This void created a gravity that pulled together a small group of thoughtful, committed parents who sought to build a better world for their children, and thus the FamilieSCN2A Foundation was born. The Foundation grew quickly as other parents and professionals were inspired and empowered by the stated vision: a world with effective treatments and cures for all SCN2A-related disorders. As momentum built, the Foundation attracted board members that shared the core values and missions. A timeline of Foundation milestones is presented in Figure 2.
Timeline of major events involving the FamilieSCN2A Foundation.

Advocacy is critical to the mission of FamilieSCN2A. The tenets of their advocacy strategy are: awareness, empowerment, evidence-based research, and equity. Successful advocacy requires awareness within the patient/family community and beyond. Awareness of SCN2A-related disorders began with the first online support group using the Facebook platform, which launched in 2013 with just five members. Today, there are more than 1,000 participants. This private, robust group is a safe space for families, patients, and caregivers to share their journey, ask questions, learn from one another, and reduce feelings of isolation. The community of patient caregivers remains a pillar of support for individuals affected by SCN2A-related disorders.
To extend awareness beyond families, the FamilieSCN2A Foundation spearheads various awareness initiatives including state proclamations, listening sessions with the Food and Drug Administration (FDA), providing information to policymakers and drug developers, and organizing caregiver testimonies that are intended to raise awareness of the burden of disease. Education, which is vital for a community actively engaged in collaboration with clinicians, scientists, and industry, has been an important Foundation strategy. As such, the FamilieSCN2A Foundation organizes an annual family and professional conference that leaves participants feeling empowered, educated, and hopeful. Supplementing the educational initiatives are programs that support families both emotionally and financially. Initiatives include Family Meet Up Grants, a Birthday Club, and a Patient Assistance Grant program that has awarded more than $65,000 to families since 2015.
The accomplishments of the FamilieSCN2A Foundation include $4 million raised since 2015, 1,200 families supported globally, strategic partnerships with leading academic researchers, a voice with the FDA, representation at international conferences, and a growing attendance at the annual family and professional symposium. Its founders, board members, volunteers, and community (including researchers and clinicians) together have made significant progress in a thoughtful way and committed to changing the world for those affected with SCN2A-related disorders.
Family and Caregiver Perspectives
There is a dearth of literature that describes the burden of care and health-related quality of life (HRQoL) as they relate to SCN2A-related disorders. While Cohen and colleagues [Reference Cohen, Helbig, Kaufman, Schust Myers, Conway and Helbig2] summarized quality of life and its determinants in developmental and epileptic encephalopathies (DEEs), for which SCN2A pathogenic variants accounted for 24 percent of the cases (n=42/173), no peer-reviewed HRQoL studies have been published specific to SCN2A-related disorders. Given the severity of SCN2A-related disorders and their impact on individuals and their families, it is imperative that additional HRQoL studies be conducted. Further, understanding the lived experiences of patients and their caregivers facing rare neurologic diseases is critical to advancing patient-centered outcomes research and informing clinical trial design. Lived experiences also highlight unmet needs of the SCN2A-related disorders community and describe the nuances of the disease beyond clinical care. Thus, this section provides a glimpse into some of the challenges families face on a daily basis and illustrates why the patient voice is a fundamental component that complements the clinical and scientific literature on SCN2A-related disorders.
To capture the family perspective, the FamilieSCN2A Foundation interviewed caregivers of individuals with SCN2A-related disorders. Interviews were transcribed and edited into a short video that highlights various aspects of the disorder (Video 1).
I felt shattered and heartbroken. The doctors did not know anything about this diagnosis. They were new to it but luckily, they did give us all the information of the FamilieSCN2A foundation. That was helpful.
Terrible. My world just fell apart. I didn’t know what to do. Didn’t know where to look because in Belgium, there just aren’t many cases. The doctors told me not to Google it, and that was pretty scary.
I felt a mixture of emotions for the first 23 months of my son’s life. He was initially neurotypical, then seizures started. Within two months of seizures starting, we got the diagnosis. I felt overwhelmed. There was a little bit of excitement knowing that there’s something that they can possibly treat, but at the same time scary that they knew very little about it. I was scared for the future.
I was actually thrilled. It was such a relief to finally figure out what was wrong. I knew from infancy that something was wrong. I always equated it to a juggling act: she has GI reflux, and she’s got a tic-like behavior, and she’s got apraxia of speech, and she has double hip dysplasia. I kept asking, so then what is it? There has to be something underlying. I just wanted a name for it.
The following express the difficulties of having a child affected with a SCN2A-related disorder.
Charlotte was born having over 400 seizures a day and she was very prone to illness. As her life went on, she had more and more seizures, and towards the end of her life, she was in status most of the time and her brain wasn’t functioning anymore. The hardest part for me was that I am an Intensive Care Unit nurse and knew too much.
It’s kind of like stripping everything away that I had envisioned for my child. I had to re-evaluate what his future might look like. I watched him suffer and was not able to do anything about it. He still pushes through with a smile, but it’s challenging to know that there’s nothing I can do about it at this moment.
The way families feel about a cure for SCN2A-related disorders is expressed by these statements.
I believe in a cure. There are a lot of prayers as well as just the parental force of the FamilieSCN2A foundation that have come together to make me feel hopeful that there would someday be a cure.
I always believe in a cure. That’s the main reason why I attend the conferences. That why we’re not giving up, why we’re going across the globe to find people who understand and who are willing to work with us. That’s why I’m screaming from the rooftops telling everyone, “this is what my child has!” I tell every cab driver. I’m here for this, this is my daughter, she has this, I’m telling everyone.
I think there’s going to be a cure. I’m not 100 percent sure it’s going to be in her lifetime.
These quotes were in response to asking parents how they have been impacted by FamilieSCN2A Foundation.
We ask questions and we get answers that we don’t get from our doctors. We get mental as well as emotional support from each other, even coming to the conferences and learning about what the research is done or just meeting other families has been possible through the funding that is available through the FamilieSCN2A Foundation for which I am very grateful.
It’s a huge support system knowing that I’m not alone and that there are others that understand my struggle. It’s a judgment-free zone. It’s a source of hope because I see what the foundation is doing. It gives me hope that people care and they are trying to do something about SCN2A.
I believe that the foundation provides a community of support for our families. It gives families hope and I am really excited at the research that they initiated, to get doctors excited about researching this disease, and putting money towards finding a cure or at least a better quality of life for our kids.
In an effort to build upon the interview data, FamilieSCN2A disseminated a questionnaire to their community in March 2023 that asked questions related to the consequences of caregiving. Two specific questions generated data that formed visual representations of (1) how caregivers felt when their children were first diagnosed and (2) what they wished their providers knew about SCN2A-related disorders. Figure 3 illustrates word clouds representing these responses.
Parents (Sandya Crasta, Liz Hendrickx, Amy Richards, Ashley Taylor, Tracy Umezu) discussing experiences caring for their children with SCN2A-related disorders.
A transcript of this video is available in the Appendix. The video file is available at www.cambridge.org/scn2a In addition, the following paraphrased quotes describe how families felt when they received a diagnosis of SCN2A-related disorders.
Word clouds generated from caregiver responses.
Responses expressing how caregivers felt when their child was first diagnosed.

Responses expressing what caregivers wished their health care providers knew about SCN2A-related disorders.

Given the dearth of literature describing the challenges and consequences of caring for SCN2A-related disorders for patients and their families, combined with the speed at which SCN2A-related disorders are being studied and the simultaneous growth of the FamilieSCN2A Foundation, we hope this section has provided a thorough overview that highlights various aspects of the disorder and leaves families feeling empowered and health care professionals with valuable insights to navigate the complexities associated with SCN2A-related disorders.
Clinical Spectrum and Genotype–Phenotype Correlations
Pathogenic SCN2A variants are associated with a range of NDD with or without epilepsy, having symptom onset anytime between the first day of life through later childhood (Figure 4). Due to the variability in clinical presentation, data on the incidence of SCN2A-related disorders is not complete. For example, epilepsy patient cohorts exclude cases of NDD that do not have seizures. Expanded genetic testing that includes a wider spectrum of NDD will likely reveal an even greater genotype and phenotype diversity.
Clinical spectrum of SCN2A-related disorders. Black-shaded boxes indicate phenotypes associated with normal development. Unshaded boxes indicate phenotypes associated with mild or moderate to severe or profound ID. Arrows indicate predominant associated functional effects of SCN2A variants in each condition. Typical age of onset is given in months (m) and generally refers to the onset of seizures.

In a meta-analysis of genetic findings discovered using next generation sequencing-based gene panels, SCN2A was the fourth most commonly implicated gene in monogenic epilepsy (after SCN1A, KCNQ2, and CDKL5) accounting for 7 percent of cases [Reference Symonds and McTague3]. By comparison, SCN1A-associated epilepsy accounted for nearly three times as many cases (19 percent). The incidence of SCN1A-associated epilepsy was accurately estimated as 1 per 12,200 live births from a population-based cohort study in Scotland [Reference Symonds, Zuberi and Stewart4]. Based on the assumption that SCN1A-associated epilepsy is approximately three times more common than SCN2A-associated epilepsy, one might expect the incidence of SCN2A-related epilepsy to be in the range of 1 in 30,000 to 50,000 live births. An estimate of 1 per 78,608 live births for SCN2A-related epilepsy was based previously on observing seven cases of SCN2A-related epilepsy diagnosed in the single national Danish testing center between 2006 and 2014 [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5].
A comparable incidence estimate of SCN2A-related NDD comes from the UK Deciphering Developmental Disorders (DDD) study [Reference Fitzgerald, Gerety, Jones, van Kogelenberg, King and McRae6,Reference McRae, Clayton, Fitzgerald, Kaplanis, Prigmore and Rajan7], a national multicenter study in which participants with a wide range of developmental disorders underwent whole exome sequencing (WES). The DDD incidence estimate includes SCN2A-related NDD without epilepsy as a feature, but excludes self-limited and familial epilepsy cases. Analyzing the first 4,293 families in the DDD study, the authors estimated that 42 percent of the cohort carried a disease-causing de novo variant in any gene. They further estimated that a de novo variant can be expected to result in a developmental disorder between 1 in 213–448 births. Because 19 study participants with SCN2A-related NDD were among these 4,293 families, the incidence of SCN2A-related NDD can be estimated as 1 per 47,000 to 100,000 live births.
Nearly all individuals with SCN2A variants develop epilepsy at some point in their life. It is estimated that about half of individuals with SCN2A-related NDD will present with seizures in the neonatal period [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Zeng, Yang, Duan, Niu, Chen and Wang8], and 80 percent will develop seizures within the first six months of life [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Zeng, Yang, Duan, Niu, Chen and Wang8]. Focal seizures are the most common seizure type, reported in 90 percent of individuals, and epileptic spasms occur in up to 50 percent of individuals with SCN2A-related NDD [Reference Zeng, Yang, Duan, Niu, Chen and Wang8]. Nearly 80 percent of individuals with SCN2A-related epilepsy have comorbid developmental delay [Reference Zeng, Yang, Duan, Niu, Chen and Wang8]. Mosaicism is estimated to occur in 6.4 percent of pathogenic SCN2A variants, present at only 11.6 percent of variant reads (range: 11.6–39.5 percent) [Reference Stosser, Lindy, Butler, Retterer, Piccirillo-Stosser and Richard9].
Self-limited Epilepsies
Self-limited familial neonatal-infantile epilepsy (SeLFNIE), previously known as benign familial neonatal-infantile epilepsy, was the first reported SCN2A-related disorder [Reference Heron, Crossland, Andermann, Phillips, Hall and Bleasel10,Reference Berkovic, Heron, Giordano, Marini, Guerrini and Kaplan11]. Self-limited epilepsies represent approximately 20 percent of the phenotypic spectrum of reported SCN2A-related disorders [Reference Wolff, Brunklaus and Zuberi12], and include neonatal, neonatal-infantile, and infantile subtypes, which are classified by age of seizure onset along with family history [Reference Zuberi, Wirrell, Yozawitz, Wilmshurst, Specchio and Riney13]. Seizure onset ranges from 2 days to 23 months with a mean of 11.2 ± 9.2 weeks and median of 13 weeks [Reference Berkovic, Heron, Giordano, Marini, Guerrini and Kaplan11,Reference Wolff, Brunklaus and Zuberi12]. Approximately half of affected individuals have onset before age one month [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5]. Seizures are typically focal with head and eye deviation along with tonic or clonic features and possibly apnea; some evolve to bilateral tonic-clonic seizures [Reference Heron, Crossland, Andermann, Phillips, Hall and Bleasel10,Reference Berkovic, Heron, Giordano, Marini, Guerrini and Kaplan11,Reference Herlenius, Heron, Grinton, Keay, Scheffer and Mulley14,Reference Reynolds, King and Gorman15]. Seizures may occur in brief clusters (20 seconds to 3 minutes), are most often afebrile, and are usually controlled with single antiseizure medications such as phenobarbital, phenytoin, oxcarbazepine, clobazam, zonisamide, or valproic acid [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Zeng, Yang, Duan, Niu, Chen and Wang8,Reference Berkovic, Heron, Giordano, Marini, Guerrini and Kaplan11,Reference Herlenius, Heron, Grinton, Keay, Scheffer and Mulley14]. Electroencephalograms are generally normal or show focal epileptiform activity without severe encephalopathy patterns, and seizures typically resolve by age 12 months with rare recurrence later in life [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Zeng, Yang, Duan, Niu, Chen and Wang8,Reference Berkovic, Heron, Giordano, Marini, Guerrini and Kaplan11,Reference Herlenius, Heron, Grinton, Keay, Scheffer and Mulley14]. Developmental milestones and cognitive outcomes are typically normal, and SCN2A variants in SeLFNIE are frequently inherited with high penetrance. Distinguishing SeLFNIE from more severe DEEs at the time of seizure onset can be challenging. A representative clinical scenario is described in Box 1.
A three-week-old infant presented with focal motor, evolving to bilateral tonic-clonic, seizures. He had been born at term with no prenatal or perinatal complications. Interictal encephalography (EEG) demonstrated normal background rhythms. Seizures were controlled with carbamazepine. He was weaned off antiseizure medication at two years. He had recurrence of a single generalized tonic-clonic seizure following a concussion at six years but had a normal 24-hour EEG. Cognitive and developmental outcome was normal at six years.
The primary differential diagnoses for SeLFNIE, assuming structural brain imaging is normal or demonstrates nonspecific changes, include the following:
➢ KCNQ2-related epilepsy: This condition presents with a range from self-limited infantile epilepsy to DEE with or without burst suppression on electroencephalography (EEG). The seizure onset is more often neonatal and responds well to carbamazepine or other sodium channel blockers [Reference Berkovic, Heron, Giordano, Marini, Guerrini and Kaplan11,Reference Sands, Balestri, Bellini, Mulkey, Danhaive and Bakken16,Reference Weckhuysen, George, Cilio, James, Loewy, Sands, Weckhuysen and George17].
➢ PRRT2-related epilepsy: This disorder is characterized by self-limited infantile epilepsy with focal motor seizures and an excellent response to antiseizure medication, particularly sodium channel blockers [Reference Döring, Saffari and Bast18,Reference Lee, Kim, Lim and Lee19]. Seizure onset is typically at age 4–12 months with resolution by two years. Paroxysmal kinesigenic dyskinesia is also associated with pathogenic PRRT2 variants but symptoms may start after seizure onset [Reference Lee, Kim, Lim and Lee19,Reference Lee, Kim, Lim and Lee20,Reference Landolfi, Barone and Erro21].
➢ SCN8A-related epilepsy: Epilepsy associated with pathogenic SCN8A variants can manifest as self-limited infantile epilepsy or DEE. Median age of onset is approximately six months. Seizure types include focal, multifocal, or bilateral tonic-clonic. In the DEE phenotype, epileptic spasms are reported. Seizures in DEE caused by gain-of-function (GOF) SCN8A variants are reported to respond favorably to sodium channel blockers, though high doses are often required [Reference Boerma, Braun, van den Broek, van Berkestijn, Swinkels and Hagebeuk22]. Movement disorders including myoclonus, tremor, and paroxysmal dyskinesias are reported [Reference Gardella, Becker, Møller, Schubert, Lemke and Larsen23,Reference Pons, Lesca, Sanlaville, Chatron, Labalme and Manel24,Reference Johannesen, Liu, Koko, Gjerulfsen, Sonnenberg and Schubert26].
Early Infantile Developmental and Epileptic Encephalopathy
DEE represents the most frequent clinical presentation among published cases of SCN2A-related disorders. The early infantile subtype of DEE is characterized by epilepsy onset within the first three months of life, usually in the neonatal period, and represented 36 percent of participants in a study of 201 individuals with SCN2A-related disorders [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5]. In this study, 43 percent (31 of 71) of the early infantile DEE subset had an identifiable epilepsy syndrome, including early infantile DEE with burst suppression on EEG (18 of 71) and epilepsy of infancy with migrating focal seizures (EIMFS) (13 of 71), while the remaining 56 percent (40 of 71) had unclassifiable epilepsies with focal or generalized seizure types including tonic, tonic-clonic, and epileptic spasms. Initial interictal EEG often shows a burst-suppression pattern or multifocal spikes [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Olson, Kelly, LaCoursiere, Pinsky, Tambunan and Shain27,Reference Burgess, Wang, McTague, Boysen, Yang and Zeng28].
Intellectual disability, often profound or severe, is present in a majority of individuals. Comorbid movement disorders are common, including dystonia, chorea, choreoathetosis, and dyskinesia [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Burgess, Wang, McTague, Boysen, Yang and Zeng28,Reference Howell, McMahon, Carvill, Tambunan, Mackay and Rodriguez-Casero29]. Other reported comorbidities include cortical visual impairment, microcephaly, and features of dysautonomia such as temperature instability (hypo- or hyperthermia) and tachycardia [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Burgess, Wang, McTague, Boysen, Yang and Zeng28,Reference Howell, McMahon, Carvill, Tambunan, Mackay and Rodriguez-Casero29]. Brain MRI may be normal or show cerebral atrophy, hypomyelination, and T2 hyperintensities, among other abnormalities [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Olson, Kelly, LaCoursiere, Pinsky, Tambunan and Shain27,Reference Howell, McMahon, Carvill, Tambunan, Mackay and Rodriguez-Casero29,Reference Kim, Yang, Kim, Kim, Kim and Lee30]. These MRI findings are nonspecific, and cerebral atrophy specifically has been associated with other causes of early infantile developmental encephalopathy [Reference Zuberi, Wirrell, Yozawitz, Wilmshurst, Specchio and Riney13,Reference Olson, Kelly, LaCoursiere, Pinsky, Tambunan and Shain27]. Rare individuals with SCN2A-related DEE have been found to have polymicrogyria [Reference Vlachou, Larsen, Pavlidou, Ismayilova, Mazarakis and Pantazi31,32,Reference Akula, Chen, Neil, Shao, Mo and Hylton33], suggesting a potential role of sodium channels in neuronal migration during brain development. A representative clinical scenario is described in Box 2.
A female infant presented on day one of life with multiple predominantly focal tonic seizures per day. EEG initially showed burst suppression evolving to generalized slowing with multifocal or bi-frontal epileptiform activity. Phenytoin was the most effective antiseizure medication but did not achieve seizure freedom. Oxcarbazepine treatment was associated with a partial response and greater alertness but was accompanied by hyponatremia. A combination of lacosamide and lamotrigine resulted in two years of near seizure-freedom, with breakthrough only in the setting of missed or late doses. A likely pathogenic, de novo variant in SCN2A was identified by epilepsy gene panel testing.
At nine years of age, the patient exhibited global developmental delay with axial hypotonia and was unable to sit independently. She showed no purposeful hand use, smiled in response to voices but expressive communication was limited to nonspecific vocalizations. She had cortical visual impairment and made limited eye contact. Other medical issues included mild scoliosis, oral phase dysphagia with poor weight gain, and irritability.
Specific early infantile DEE phenotypes associated with pathogenic SCN2A variants are described next.
Epilepsy of Infancy with Migrating Focal Seizures (EIMFS)
EIMFS is characterized by multiple types of focal seizures that “migrate” from one hemisphere to the other, usually accompanied by severe developmental impairment [Reference Howell, McMahon, Carvill, Tambunan, Mackay and Rodriguez-Casero29]. SCN2A is second to KCNT1 as the most frequently identified gene in EIMFS [Reference Burgess, Wang, McTague, Boysen, Yang and Zeng28,Reference Howell, McMahon, Carvill, Tambunan, Mackay and Rodriguez-Casero29]. In these two cited studies of individuals with EIMFS and SCN2A variants (22 total), onset of seizures occurred between the first day of life and eight weeks for most (20/22), and less commonly after age one year (2/22). All had multifocal spikes on EEG.
Early Infantile DEE with Burst Suppression
SCN2A has been identified as a common gene associated with early infantile DEE with burst suppression on EEG formerly called Ohtahara syndrome [Reference Zeng, Yang, Duan, Niu, Chen and Wang8,Reference Olson, Kelly, LaCoursiere, Pinsky, Tambunan and Shain27,Reference Howell, McMahon, Carvill, Tambunan, Mackay and Rodriguez-Casero29,Reference Kim, Yang, Kim, Kim, Kim and Lee30,Reference Vlachou, Larsen, Pavlidou, Ismayilova, Mazarakis and Pantazi31,Reference Nakamura, Kato, Osaka, Yamashita, Nakagawa and Haginoya34,Reference Zerem, Lev, Blumkin, Goldberg-Stern, Michaeli-Yossef and Halevy35]. Seizure types typically include epileptic spasms, tonic seizures, and/or myoclonic seizures [Reference Zuberi, Wirrell, Yozawitz, Wilmshurst, Specchio and Riney13,Reference Ohtahara and Yamatogi36]. In two studies of individuals with early infantile DEE, SCN2A variants were identified in 9/67 (13.4 percent) and 2/33 (7 percent) respectively [Reference Olson, Kelly, LaCoursiere, Pinsky, Tambunan and Shain27,Reference Nakamura, Kato, Osaka, Yamashita, Nakagawa and Haginoya34]. Infantile epileptic spasms syndrome (IESS) has also been reported in individuals with SCN2A-related early infantile DEE, either independently or evolving from an initial presentation as early infantile DEE with burst suppression [Reference Zeng, Yang, Duan, Niu, Chen and Wang8].
Prognosis is variable. With regard to epilepsy, response to sodium channel blocking antiseizure medications in SCN2A-related early infantile DEE is often favorable, but high doses are often needed and there are reports of seizure relapse when plasma drug levels drop below a certain threshold [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Howell, McMahon, Carvill, Tambunan, Mackay and Rodriguez-Casero29]. There have been several reported instances of sudden unexpected death in epilepsy (SUDEP) in this subset of individuals [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Zeng, Yang, Duan, Niu, Chen and Wang8,Reference Howell, McMahon, Carvill, Tambunan, Mackay and Rodriguez-Casero29], which warrants consideration in discussing the prognosis with families. Other reported causes of early death include pneumonia (age 21 months) [Reference Howell, McMahon, Carvill, Tambunan, Mackay and Rodriguez-Casero29], respiratory failure during treatment of pneumonia [Reference Kim, Yang, Kim, Kim, Kim and Lee30], iatrogenic cardiorespiratory failure (age 19 days) [Reference Touma, Joshi, Connolly, Ellen, Hansen and Khwaja37], severe infection, and status epilepticus [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5].
The primary differential diagnoses for this group, assuming structural brain imaging is normal or demonstrates nonspecific changes, include the following:
➢ KCNT1-related disorder: As mentioned previously, KCNT1 is the gene most frequently associated with EIMFS [Reference Burgess, Wang, McTague, Boysen, Yang and Zeng28,Reference Howell, McMahon, Carvill, Tambunan, Mackay and Rodriguez-Casero29]. Differentiating features that are more associated with SCN2A include presence of severe movement disorders and seizure response to phenytoin [Reference Howell, McMahon, Carvill, Tambunan, Mackay and Rodriguez-Casero29,Reference Barcia, Fleming, Deligniere, Gazula, Brown and Langouet38]. Age of seizure onset may also be slightly later in those with KCNT1-associated EIMFS (median 3.5 weeks, compared to median of 3.5 days in those with SCN2A-associated EIMFS) [Reference Burgess, Wang, McTague, Boysen, Yang and Zeng28]. Prognosis appears to be less favorable in KCNT1-related EIMFS, with higher reported rates of severe to profound developmental impairment, refractory epilepsy, and SUDEP compared to those with SCN2A variants [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Burgess, Wang, McTague, Boysen, Yang and Zeng28,Reference Ohba, Kato, Takahashi, Osaka, Shiihara and Tohyama39].
➢ KCNQ2-related disorder: Variants in KCNQ2 are a frequently discovered genetic cause of early infantile DEE with burst suppression [Reference Olson, Kelly, LaCoursiere, Pinsky, Tambunan and Shain27,Reference Kato, Yamagata, Kubota, Arai, Yamashita and Nakagawa40], and should be considered in the differential diagnosis of a patient with this phenotype. Neonatal onset seizures associated with pathogenic KCNQ2 variant are also responsive to sodium channel blocking antiseizure medications [Reference Weckhuysen, George, Cilio, James, Loewy, Sands, Weckhuysen and George17], which may resemble the response of individuals with SCN2A GOF variants.
Later Onset Developmental and Epileptic Encephalopathies
DEEs presenting after three months of age constitute the second largest group (20–30 percent) with pathogenic SCN2A variants [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Sanders, Campbell, Cottrell, Moller, Wagner and Auldridge41]. Affected individuals typically present with generalized tonic-clonic, absence, and myoclonic seizures along with EEG features of generalized spike and wave or multifocal spikes. A number of distinct epilepsy syndrome presentations in this later onset group have been described. This includes individuals with IESS evolving to Lennox-Gastaut syndrome (LGS), myoclonic atonic epilepsy (MAE), and focal epilepsies with a condition resembling electrical status epilepticus during slow-wave sleep (ESES). Brain imaging is mostly normal with the exception of occasional nonspecific cortical atrophy. While some are cognitively normal before seizure onset, ID eventually develops in all affected individuals and the majority have severe cognitive impairment. Affected individuals have prominent comorbidities including ASD and motor symptoms such as hypotonia, choreiform, or dyskinetic movement disorders. A representative clinical description is illustrated in Box 3.
A female infant presented with epileptic spasms at age 10 months, associated with developmental regression. Prior to seizure onset, her development was mildly delayed. Prior to the genetic diagnosis, she was treated with oxcarbazepine, which led to exacerbation of seizures. At age 15 months, a heterozygous, de novo likely pathogenic variant in SCN2A was identified by WES. Treatment with ketogenic diet, vigabatrin, and ACTH improved cognition and development and temporarily resolved seizures, but epileptic spasms recurred after weaning ACTH. Additional antiseizure medications (levetiracetam, valproic acid) were not effective. At age six years, while treated with clobazam, clonazepam, and phenobarbital, she experienced up to three seizures per day. She had generalized hypotonia and did not sit independently or reach for objects. She turned her head toward sounds and made nonspecific vocalizations. Additional medical issues included intermittent choreoathetosis, autonomic dysfunction (periodic flushing with Raynaud-type phenomenon in hands and feet, constipation), mouthing, and leg-crossing stereotypes.
Later onset DEE associated with pathogenic SCN2A variants has a notable genotype–phenotype relationship [Reference Crawford, Xian, Helbig, Galer, Parthasarathy and Lewis-Smith42]. Typically, SCN2A variants in this cohort are reported to have loss-of-function (LOF) effects due to missense, protein truncating, and splice site variants or missense variants with mixed GOF/LOF effects [Reference Reynolds, King and Gorman15]. For example, the mixed function p.R853Q variant is the most common recurrent SCN2A variant [Reference Crawford, Xian, Helbig, Galer, Parthasarathy and Lewis-Smith42]. Individuals carrying this variant present with a characteristic phenotype consisting of epileptic spasms, hypsarrhythmia on EEG, and choreiform movement disorder [Reference Crawford, Xian, Helbig, Galer, Parthasarathy and Lewis-Smith42,Reference Samanta and Ramakrishnaiah43,Reference Berecki, Howell, Deerasooriya, Cilio, Oliva and Kaplan44].
In SCN2A-related disorders associated with LOF variants, seizures are typically treatment refractory and seizure freedom occurs in only 34 percent of cases [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5]. Seizures are typically unresponsive to sodium channel blockers and can worsen after the introduction of this class of antiseizure medication. Persons with later onset DEE respond better to other drug classes including benzodiazepines, levetiracetam, sodium valproate, and ACTH, compared with early onset DEE [Reference Reynolds, King and Gorman15]. Specific later onset DEE phenotypes associated with pathogenic SCN2A variants are described next.
Infantile Epileptic Spasms Syndrome (IESS)
A prominent phenotype associated with SCN2A-related later infantile onset DEE is IESS [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Crawford, Xian, Helbig, Galer, Parthasarathy and Lewis-Smith42]. Typical interictal EEG features include hypsarrhythmia or multifocal or focal epileptiform discharges, which may occur soon after the onset of epileptic spasms [Reference Zuberi, Wirrell, Yozawitz, Wilmshurst, Specchio and Riney13]. Age of onset is typically between 3 and 24 months. Developmental delay is observed after onset of epileptic spasms, but may be absent early in the course. Abnormal neurological examination findings may be present including abnormalities of posture, tone, or movement. The family, pregnancy, and birth histories are usually normal. Cognition varies from normal to severely delayed before seizure onset; however, moderate to profound developmental impairment becomes evident with time [Reference Zuberi, Wirrell, Yozawitz, Wilmshurst, Specchio and Riney13,Reference Reynolds, King and Gorman15]. Long-term prognosis is unfavorable and many subsequently evolve to LGS [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5]. SCN2A-related IESS is difficult to treat and does not respond to standard antiseizure medications, steroids, or ketogenic diet [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Nakamura, Kato, Osaka, Yamashita, Nakagawa and Haginoya34].
The main differential diagnoses for late onset IESS are other genetic etiologies including pathogenic variants in ARX, IQSEC2, TSC1, TSC2, and others [Reference Zuberi, Wirrell, Yozawitz, Wilmshurst, Specchio and Riney13]. For early onset IESS, CDKL5 and STXBP1 should be considered [Reference Zuberi, Wirrell, Yozawitz, Wilmshurst, Specchio and Riney13]. In addition, a range of chromosomal abnormalities and copy number variants has been associated with IESS [Reference Paciorkowski, Thio and Dobyns45,Reference Pavone, Polizzi, Marino, Corsello, Falsaperla and Marino46,Reference Chourasia, Yuskaitis, Libenson, Bergin, Liu and Zhang47].
SCN2A-Related Disorders without Epilepsy
Individuals with pathogenic variants in SCN2A may present with NDD without epileptic seizures. The proportion of SCN2A-related disorders without epilepsy is unclear because of an ascertainment bias arising from preferential genetic testing of children presenting with seizures as compared with children having non-syndromic ID. Perhaps the best estimate of the incidence comes from the DDD study [Reference Richardson, Baralle, Bennett, Briggs, Bijlsma and Clayton-Smith48]. Inclusion criteria for the DDD study were broad, including children with severe undiagnosed NDD and/or congenital anomalies, abnormal growth parameters, dysmorphic features, and unusual behavioral phenotypes [Reference McRae, Clayton, Fitzgerald, Kaplanis, Prigmore and Rajan7]. Richardson et al. reported the phenotypes of 22 persons including 12 presenting with neonatal or early infantile (≤3 months) onset seizures, four with ID/ASD and later onset seizures (seizure onset between two and nine years), and six with ID/ASD that had no history of epilepsy or epileptic seizures [Reference Richardson, Baralle, Bennett, Briggs, Bijlsma and Clayton-Smith48]. Thus, it could be estimated that approximately 20–30 percent of persons with NDD associated with SCN2A variants have no history of epileptic seizures.
In the large series of 201 individuals with SCN2A-associated disorders, 16 percent did not have epileptic seizures, although this sample may have ascertainment bias [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5]. In the Richardson et al. study, all participants with either later onset seizures or no history of seizures had de novo variants and most (6 of 10) were protein truncating [Reference Richardson, Baralle, Bennett, Briggs, Bijlsma and Clayton-Smith48]. In contrast, all variants associated with the early onset seizure phenotypes (≤3 months) were missense. Among the six study participants without epilepsy, the degree of ID varied from mild to profound; half were formally diagnosed with ASD and no one had ataxia. Normal or nonspecific neuroimaging findings, including cerebral atrophy and hypoplasia of the corpus callosum, were reported for this subset. EEG findings were not reported by Richardson et al. [Reference Richardson, Baralle, Bennett, Briggs, Bijlsma and Clayton-Smith48], but a case report described the EEG in the setting of later onset developmental delay without epilepsy associated with a de novo missense SCN2A variant [Reference Mangano, Fontana, Antona, Salpietro, Mangano and Giuffrè49]. Findings in this case were bilateral discharges of high amplitude sharp waves and slow waves in the parietal–temporal–occipital regions that increased during sleep and were associated with right frontal-central short sequences of 5–4 Hz spike and wave complexes.
Episodic Ataxia
Episodic ataxia is observed in a subset of individuals with pathogenic SCN2A variants [Reference Liao, Anttonen, Liukkonen, Gaily, Maljevic and Schubert50,Reference Schwarz, Hahn, Bast, Müller, Löffler and Maljevic51,Reference Gorman and King52,Reference Schwarz, Bast, Gaily, Golla, Gorman and Griffiths53]. Most have epileptic seizures beginning during the first three months of life, and all have missense variants in the gene, suggesting that episodic ataxia may be a GOF disorder related to the early onset epilepsy phenotype. Onset of ataxia ranges from 10 months to 14 years. Episodes are highly variable, ranging from brief daily events to infrequent long-lasting episodes. Acetazolamide was effective in only a minority of reported cases [Reference Schwarz, Bast, Gaily, Golla, Gorman and Griffiths53]. In a systematic review of genetic epilepsies associated with paroxysmal ataxia, potential triggers for paroxysms of ataxia in SCN2A-related disorders were cited including minor head injuries, sleep deprivation, alcohol ingestion, photic stimulation, sudden noise, vibration of the body, and menstruation [Reference Amadori, Pellino, Bansal, Mazzone, Møller and Rubboli54]. The majority (80 percent) with SCN2A-related episodic ataxia have good cognitive outcomes. A clinical vignette describing this syndrome is presented in Box 4.
A female patient with an uncomplicated prenatal and perinatal history developed focal clonic seizures beginning on the fifth day of life. These were controlled with intravenous phenytoin and did not recur. Starting at age 10 months, she developed episodes of marked truncal ataxia, lasting 12–24 hours and occurring twice per month. Introduction of carbamazepine (titrated up to 10 mg/kg twice daily) and acetazolamide (titrated up to 2 mg/kg three times daily) did not change the frequency or severity of the episodes. At age two years she had a mild global developmental delay and muscular hypotonia. She was found to have a de novo pathogenic missense variant in SCN2A.
In addition to episodic ataxia, other paroxysmal movement disorders have been anecdotally associated with SCN2A variants, including paroxysmal dyskinesia and choreoathetosis occurring in the context of both early and later onset epilepsy [Reference Liao, Anttonen, Liukkonen, Gaily, Maljevic and Schubert50,Reference Hackenberg, Baumer, Sticht, Schmitt, Kroell-Seger and Wille55,Reference Spagnoli, Fusco, Percesepe, Leuzzi and Pisani56]. Severe choreoathetosis has been reported in persons heterozygous for the recurrent SCN2A variant p.R853Q [Reference Wolff, Brunklaus and Zuberi12,Reference Samanta and Ramakrishnaiah43,Reference Berecki, Howell, Deerasooriya, Cilio, Oliva and Kaplan44]. Familial and sporadic hemiplegic migraine has also been associated with pathogenic SCN2A missense variants either with no history of epilepsy or with SeLFNIE [Reference Riant, Thompson, DeKeyser, Abramova, Gazal and Moulin57]. Alternating hemiplegia of childhood has also been associated with pathogenic SCN2A variants in cases without mutations in the main gene (ATP1A3) [Reference Panagiotakaki, Tiziano, Mikati, Vijfhuizen, Nicole and Lesca58].
There is no strong evidence supporting specific therapeutic approaches for patients with non-epileptic movement disorders associated with SCN2A variants. The majority of functionally tested variants associated with episodic ataxia exhibit GOF properties in either in silico or in vitro models. This is consistent with many cases being observed in the context of prior neonatal onset epileptic seizures. There are very few case reports of effective treatment of ataxia with sodium channel blocking medications. Treatment with acetazolamide has often been reported to be effective at reducing frequency and severity of ataxia episodes in approximately half of patients. Acetazolamide has been reported as effective in cases associated with both GOF and LOF variants in SCN2A [Reference Schwarz, Bast, Gaily, Golla, Gorman and Griffiths53].
Genotype–Phenotype Correlations
General genotype–phenotype correlations are established for SCN2A-related disorders. Individuals with self-limited epilepsy are heterozygous for mostly inherited missense variants [Reference Wolff, Brunklaus and Zuberi12]. More severe epilepsy phenotypes are mainly associated with de novo variants that confer greater electrophysiological dysfunction [Reference Wolff, Brunklaus and Zuberi12,Reference Lauxmann, Verbeek, Liu, Zaichuk, Müller and Lemke59]. There are no clear correlations between the location of the variant on the protein and phenotypic severity. Amino acid substitutions between physicochemically similar amino acids are less likely to cause severe disease and are more frequently observed in self-limited epilepsies [Reference Wolff, Brunklaus and Zuberi12,Reference Brunklaus, Ellis, Reavey, Semsarian and Zuberi60]. Functional analyses of some variants associated with self-limited epilepsy and episodic ataxia phenotypes have demonstrated GOF effects [Reference Schwarz, Hahn, Bast, Müller, Löffler and Maljevic51,Reference Schwarz, Bast, Gaily, Golla, Gorman and Griffiths53,Reference Liao, Deprez, Maljevic, Pitsch, Claes and Hristova61].
Individuals with early onset DEE mainly have missense variants (77 percent) but can also carry truncating variants (23 percent) [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Wolff, Brunklaus and Zuberi12,Reference Begemann, Acuña, Zweier, Vincent, Steindl and Bachmann-Gagescu62]. The missense variants tend to cause GOF effects; however, the functional properties are complex and do not strictly separate as GOF or LOF based on age of epilepsy onset [Reference Thompson, Potet, Abramova, DeKeyser, Ghabra and Vanoye63]. Truncating and nonsense (premature termination codon) variants present exclusively in individuals with seizure onset beyond the first year of life or without epilepsy, and those with truncating variants are reported to have seizures in 50 percent of cases [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Wolff, Brunklaus and Zuberi12]. The majority of individuals with truncating variants have features of ASD or developmental delay, compared to only 20 percent of those with missense variants. Individuals with ID/ASD and later onset epilepsy or absence of epilepsy mainly have truncating variants (75 percent) that are LOF [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Wolff, Brunklaus and Zuberi12,Reference Begemann, Acuña, Zweier, Vincent, Steindl and Bachmann-Gagescu62].
Recurrent SCN2A variants occur throughout the phenotypic spectrum (see Table 1), but individual variants may not always associate with the same phenotype, even within families [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Syrbe, Zhorov, Bertsche, Bernhard, Hornemann and Mütze64]. A computational analysis of clinical features of 413 individuals with pathogenic SCN2A variants demonstrated that only 8 of 62 individual recurrent variants were associated with similar phenotypes [Reference Crawford, Xian, Helbig, Galer, Parthasarathy and Lewis-Smith42].

* data adapted from Crawford et al. [Reference Crawford, Xian, Helbig, Galer, Parthasarathy and Lewis-Smith42]
Abbreviations: ASD = autism spectrum disorder, EI-DEE = early infantile developmental and epileptic encephalopathy, GOF = gain of function, LO-DEE = later onset developmental and epileptic encephalopathy, LOF = loss of function, NE = non-epilepsy phenotype, SLE = self-limited epilepsy
To date, the most frequently reported recurrent variants are p.R853Q (located in domain 2), p.A263V (in domain 1), and p.R1882Q (C-terminus) (Table 1). Of note, other amino acid substitutions have also been reported at the 1882 position, including p.R1882G/L/P, although these are not recurrent variants [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Schwarz, Hahn, Bast, Müller, Löffler and Maljevic51,Reference Baasch, Hüning, Gilissen, Klepper, Veltman and Gillessen-Kaesbach65]. The p.R853Q variant is associated with later onset DEE characterized by IESS, severe ID, intractable seizures, and choreoathetosis [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Wolff, Brunklaus and Zuberi12,Reference Samanta and Ramakrishnaiah43], and less commonly with ASD [Reference Crawford, Xian, Helbig, Galer, Parthasarathy and Lewis-Smith42,Reference Berecki, Howell, Heighway, Olivier, Rodda and Overmars66]. Individuals heterozygous for p.R1882Q, which causes a strong GOF, had seizure onset on day of life one and at least one died of SUDEP [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Howell, McMahon, Carvill, Tambunan, Mackay and Rodriguez-Casero29]. Phenotype is more variable for other amino acid substitutions at this position. For example, p.R1882G is associated with self-limited epilepsy with later-evolving episodic ataxia [Reference Liao, Anttonen, Liukkonen, Gaily, Maljevic and Schubert50,Reference Schwarz, Hahn, Bast, Müller, Löffler and Maljevic51,Reference Schwarz, Bast, Gaily, Golla, Gorman and Griffiths53]. The p.A263V variant is associated with early infantile DEE [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Johannesen, Miranda, Lerche and Møller67]. Two recurrent variants (p.E1211K, p.L1342P) are consistently associated with later onset DEE [Reference Berecki, Howell, Heighway, Olivier, Rodda and Overmars66]. The LOF variant p.R937C is most often reported with a phenotype of developmental delay, ID, and/or ASD without seizures [Reference Berecki, Howell, Heighway, Olivier, Rodda and Overmars66,Reference Ben-Shalom, Keeshen, Berrios, An, Sanders and Bender68].
Interpretation of SCN2A Variants
Interpretation of single nucleotide or copy number variants in SCN2A should be done in collaboration with a neurologist or geneticist with expertise in genetic variant interpretation. Features to be considered include: (1) frequency in population databases such as gnomAD,Footnote 1 (2) inheritance pattern, (3) similarity to previously described variants and/or if there are nearby pathogenic variants, (4) in silico predictions, and (5) consistency with described phenotypes. The SCN Portal,Footnote 2 ClinVar,Footnote 3 UniProt,Footnote 4 and publications provide valuable resources to determine if a variant is previously described and what clinical phenotypes are reported to determine where it is located relative to functional domains of the protein (see Figure 5B) and whether there is information on functional impact. Diagnostic laboratories will classify the variant as pathogenic, likely pathogenic, or a variant of uncertain significance based on standard guidelines [Reference Richards, Aziz, Bale, Bick, Das and Gastier-Foster69,Reference Brandt, Sack, Arjona, Tan, Mei and Cui70,Reference Riggs, Andersen, Cherry, Kantarci, Kearney and Patel71]. Functional impact of novel variants cannot be definitively determined with bioinformatics tools.
Transmembrane topology of a NaV channel.
Simplified structure of a NaV channel highlighting major functional domains.

Location by codon number of individual transmembrane segments and domains in NaV1.2.

The Biology of SCN2A
SCN2A encodes the voltage-gated sodium (NaV) channel formally named NaV1.2. The primary amino acid sequence of this protein was originally deduced from rat brain by complementary DNA (cDNA) cloning. Because it was the second sequenced brain NaV channel sequenced, it was designated as rat brain type II. Functional studies of rat NaV1.2 in Xenopus oocytes revealed a tetrodotoxin-sensitive, rapidly activating and inactivating inward current that resembled native neuron sodium currents [Reference Noda, Ikeda, Kayano, Suzuki, Takeshima and Kurasaki72,Reference Noda, Ikeda, Suzuki, Takeshima, Takahashi and Kuno73]. The primary amino acid sequence of human NaV1.2 was determined in 1992 and its functional and pharmacological properties were similar to the rat channel [Reference Ahmed, Ware, Lee, Patten, Ferrer-Montiel and Schinder74]. The human SCN2A gene resides on the long arm of chromosome 2 (2q24.3) and consists of 27 exons spanning more than 150 kilobases.
Unlike voltage-gated potassium channels, which require assembly of tetramers (4 subunits) to function, NaV channels are single polypeptides that form a pseudotetrameric arrangement of four similar domains (designated as I–IV or D1–D4) [Reference Noda, Shimizu, Tanabe, Takai, Kayano and Ikeda75]. Each of the four domains consists of six transmembrane spanning segments (S1–S6), with the fourth transmembrane segment (S4) in each domain acting as the primary voltage sensor. Segments S5 and S6, along with the intervening pore (P)-loop of each domain, interact to form a conductive pathway selective for Na+ ions (Figure 5). Sodium ion selectivity is conferred by aspartic acid (D), glutamic acid (E), lysine (K), and alanine (A) residues individually contributed by the four domains [Reference Heinemann, Terlau, Stühmer, Imoto and Numa76,Reference Schlief, Schönherr, Imoto and Heinemann77]. The III–IV cytoplasmic linker region, along with the C-terminal domain, are involved in channel inactivation [Reference Stühmer, Conti, Suzuki, Wang, Noda and Yahagi78,Reference Motoike, Liu, Glaaser, Yang, Tateyama and Kass79,Reference Capes, Goldschen-Ohm, Arcisio-Miranda, Bezanilla and Chanda80]. During the three decades after sequencing the first brain NaV channels, many research groups worked to elucidate the structural, biophysical, and pharmacological properties of NaV1.2 in great detail and investigated the underlying contribution of this channel to neuronal physiology in health and disease [Reference Catterall81,Reference Catterall, Wisedchaisri and Zheng82,Reference Catterall, Lenaeus and Gamal El-Din83]. The protein structure of NaV1.2 at atomic scale was determined using cryogenic electron microscopy [Reference Pan, Li, Huang, Huang, Gao and Shen84].
NaV1.2 Biophysical Properties
NaV channels mediate the initiation and propagation of APs [Reference Bean85]. Neuronal NaV channels, including NaV1.2, have elaborate gating mechanisms that support rapid channel activation, inactivation, and recovery from inactivation, all of which are essential to sustain high-frequency AP firing. The sequence of events occurring during a neuronal AP is well understood (Box 5). The biophysical properties of NaV1.2 that enable the AP include voltage-dependent and time-dependent (kinetic) properties. Voltage-dependence refers to the voltage range (in millivolts) over which channels activate and inactivate, while kinetic properties describe the time scales (in milliseconds) during which these events occur.
Activation of NaV channels results in a transient, rapidly inactivating current, but in some situations a small amount of non-inactivating (persistent) current remains (Figure 6). Persistent sodium current is physiologically involved in neuronal pacemaking of CA1 pyramidal neurons driven by muscarinic stimulation, as well as amplification of synaptic signals from both excitatory and inhibitory inputs [Reference Stuart and Sakmann86,Reference Stuart87,Reference Yamada-Hanff and Bean88]. Pathophysiologically, as will be discussed below, mutations that lead to abnormally large levels of persistent current can be associated with epilepsy.
Representative voltage-clamp recording of wild-type (upper trace) and pathogenic variant NaV channel with enhanced persistent sodium current (arrow).

Synaptic signals summing to produce a localized area of membrane depolarization (excitatory postsynaptic potential) of sufficient magnitude (threshold) elicits a sudden biphasic change in the membrane potential: first depolarization, then repolarization. The initial “upstroke” of the AP (depolarization phase) is caused by the opening of NaV channels. These channels are normally closed at the resting membrane potential but open (activate) upon a sufficiently strong local membrane depolarization. Activation of NaV channels is voltage-dependent owing to intrinsic voltage-sensors that are part of the protein. Opening of NaV channels allows Na+ to rush into the cell due to a favorable electrochemical gradient (high extracellular Na+ concentration and negative cell interior). The rush of Na+ into the cell causes the membrane potential to further depolarize toward the Na+ equilibrium potential (ENa), which is approximately +60 mV in mammalian neurons. Almost immediately (1–2 msec) after opening, NaV channels undergo a conformational change that closes the ion conducting pore by a process called inactivation. Sodium channels remain inactive until the membrane is repolarized. This phenomenon is a major reason for the refractory period – a short time window following an AP during which a second AP cannot be stimulated at all (absolute refractory period) or can only be elicited using a stronger second stimulus (relative refractory period).
NaV1.2 During Neurodevelopment
NaV1.2 undergoes developmentally regulated RNA splicing, resulting in two major channel isoforms with slightly different functional properties. This splicing event leads to the incorporation of an alternate exon 5 that encodes a portion of the domain I voltage sensor (S3 and S4 segments) [Reference Sarao, Gupta, Auld and Dunn89,Reference Yarowsky, Krueger, Olson, Clevinger and Koos90]. The resulting neonatal and adult splice isoforms differ by a single amino acid residue at position 209 (asparagine in neonatal NaV1.2; aspartic acid in adult NaV1.2) and in some biophysical properties [Reference Thompson, Ben-Shalom, Bender and George91]. Analysis of gene expression data from rodents and primates demonstrated that during early prenatal and immediate postnatal brain development, NaV1.2 mRNA transcripts predominantly contain exon 5N (neonatal NaV1.2) [Reference Liang, Fazel Darbandi, Pochareddy, Gulden, Gilson and Sheppard92,Reference Heighway, Sedo, Garg, Eldershaw, Perreau and Berecki93]. Progressively through the first months of postnatal development, exon 5 splice-switching leads to predominant expression of transcripts containing exon 5 A (adult NaV1.2). Neonatal NaV1.2 exhibits a depolarized voltage-dependence of activation, a hyperpolarized voltage-dependence of inactivation, and slower recovery from inactivation compared to the adult isoform. Given that immature neurons have a depolarized resting membrane potential compared with mature neurons, this combination of biophysical features would suggest that neonatal NaV1.2 may act to limit neuronal excitability during development [Reference Thompson, Ben-Shalom, Bender and George91,Reference Xu, Thomas, Jenkins, Gazina, Chiu and Heron94]. To corroborate this, a mouse model engineered to express only adult NaV1.2 throughout development showed greater neuronal excitability and increased seizure susceptibility compared to wild-type (WT) littermates [Reference Gazina, Leaw, Richards, Wimmer, Kim and Aumann95]. Thus, variants in NaV1.2 which show predominant GOF phenotypes may be particularly damaging during this critical period. Indeed, some variants exhibit exacerbated phenotypes in the neonatal isoform compared to that of the adult isoform [Reference Thompson, Ben-Shalom, Bender and George91].
Modulation and Regulation of NaV1.2
NaV1.2 activity is modulated by a number of factors, including protein–protein interactions, posttranslational modifications (e.g., phosphorylation, palmitoylation), and changes in intracellular Ca2+ concentration. Like other NaV channels, NaV1.2 interacts with members of a family of nonconducting accessory β subunits that appear to modulate channel properties and promote forward trafficking to the plasma membrane [Reference Isom, De Jongh, Patton, Reber, Offord and Charbonneau96,Reference Isom, Ragsdale, De Jongh, Westenbroek, Reber and Scheuer97,Reference Calhoun and Isom98]. Heterologous co-expression of the β1 and β2 subunits with NaV1.2 results in larger peak current amplitude, shifts in the voltage-dependence of inactivation, and faster channel activation and inactivation. In addition to β subunits, NaV1.2 also interacts with fibroblast growth homologous factors, which bind to the channel C-terminal domain. Members of this family of proteins have a range of effects on channel function. Co-expression with FGF13-1a (also known as FHF2a) results in larger current amplitude and enhanced frequency-dependent channel rundown, whereas FGF13-1b (also known as FHF2b) causes the channel to be more resistant to frequency-dependent rundown [Reference Rush, Wittmack, Tyrrell, Black, Dib-Hajj and Waxman99]. Additionally, FGF14-1b (also known as FHF4b) attenuates NaV1.2 current amplitude [Reference Laezza, Lampert, Kozel, Gerber, Rush and Nerbonne100].
Intracellular Ca2+ regulates NaV1.2 by activating Ca2+/calmodulin-dependent kinase II (CaMKII), which phosphorylates the channel. In the presence of activated CaMKII, the voltage-dependence of inactivation is significantly depolarized, and persistent current mediated by NaV1.2 is larger, suggesting that CaMKII-mediated phosphorylation enhances channel function [Reference Thompson, Hawkins, Kearney and George101]. While calmodulin was shown to interact with the C-terminus of the channel, the functional consequences of this interaction are not well established [Reference Thompson, Hawkins, Kearney and George101,Reference Wang, Chung, Yan, Wang, Lee and Pitt102].
NaV1.2 also interacts with members of the ankyrin family, cytoskeletal proteins that promote cellular localization. In neocortical pyramidal neurons, ankyrin-G scaffolds NaV1.2 to the axon initial segment (AIS) and nodes of Ranvier, whereas ankyrin-B scaffolds NaV1.2 in dendrites [Reference Nelson, Catalfio, Gupta, Min, Caballero-Floran and Dean103,Reference Gupta and Jenkins104].
NaV1.2 Function in Cells and Circuits
NaV1.2 channels are expressed throughout the central nervous system from the most ancient brainstem regions to the more recently evolved neocortex (Figure 7). There is an emerging understanding of NaV1.2 function throughout development and how dysfunction of this important channel affects neuronal excitability and integrative properties of cells and circuits. Four NaV channel subtypes are expressed in the central nervous system (NaV1.1, NaV1.2, NaV1.3, NaV1.6), which are encoded by four distinct genes (SCN1A, SCN2A, SCN3A, SCN8A, respectively). While NaV1.6 is present on the plasma membranes of a majority of neuronal classes, NaV1.1 and NaV1.2 appear to function in largely nonoverlapping groups [Reference Hu, Tian, Li, Yang, Hou and Shu105,Reference Li, Tian, Scalmani, Frassoni, Mantegazza and Wang106,Reference Tian, Wang, Ke, Guo and Shu107,Reference Yamagata, Ogiwara, Mazaki, Yanagawa and Yamakawa108,Reference Yamagata, Ogiwara, Tatsukawa, Suzuki, Otsuka and Imaeda109,Reference Yang, Xiao, Li, He, Li and Shu110]. Furthermore, NaV1.2 expression within neurons varies by neuron type and at different time points during development. In mature neurons, NaV1.2 localization is correlated with axonal myelination, with unmyelinated axons containing NaV1.2 and myelinated axons excluding NaV1.2 [Reference Yamano, Miyazaki and Nukina111]. In the latter population, emerging evidence indicates that NaV1.2 is present in dendrites [Reference Spratt, Ben-Shalom, Keeshen, Burke, Clarkson and Sanders112,Reference Spratt, Alexander, Ben-Shalom, Sahagun, Kyoung and Keeshen113]. As such, NaV1.2 function can vary markedly across the brain, affecting either input or output structures of neurons.
Localization of NaV1.2 in diverse neuronal circuits. In mature circuits from mouse and rat models, NaV1.2 expression within axons or dendrites largely corresponds to the presence of myelin. In cerebellum (left), NaV1.2 is expressed in unmyelinated granule cell axons. In hippocampus (top), NaV1.2 is expressed in unmyelinated axons of pyramidal cells. In neocortex (right), NaV1.2 is expressed in somatodendritic compartments of pyramidal cells whose axons myelinated. In striatum (bottom), current data suggests that NaV1.2 is expressed in all neuronal compartments.

NaV1.2 function, and its relationship to other ion channel classes, has been studied most extensively in neocortical structures. SCN2A is transcriptionally expressed in a range of cell classes, including excitatory and inhibitory neurons [Reference Goff and Goldberg114]; however, this expression translates into NaV1.2 predominantly in excitatory pyramidal neurons [Reference Yamagata, Ogiwara, Tatsukawa, Suzuki, Otsuka and Imaeda109]. Within pyramidal neurons, the distribution of NaV1.2 channels on the membrane changes during development. Early on, when neurons are still differentiating and elaborating neurites, NaV1.2 appears to be the sole NaV channel expressed in pyramidal neurons [Reference Gazina, Leaw, Richards, Wimmer, Kim and Aumann95]. This period spans the first year of life in humans and the first week of life in mice, a model system commonly used to study this channel. During this time period, NaV1.2 is found at highest density at the AIS, the site of AP initiation [Reference Bender and Trussell115]. Because AP initiation is supported by NaV1.2 during this developmental period, genetic variants can affect overall neuronal excitability, resulting in hyper- or hypoexcitability in developing neocortical networks. Indeed, hyperexcitability in these circuits is considered etiological to early infantile epileptic encephalopathy, which occurs within the first few months of life [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Wolff, Brunklaus and Zuberi12,Reference Sanders, Campbell, Cottrell, Moller, Wagner and Auldridge41].
After this early developmental period, NaV channels redistribute. NaV1.6 expression increases and displaces NaV1.2 in the distal AIS and at axonal nodes of Ranvier [Reference Ye, Yang, Tian, Zhu, Yin and Jiang116]. NaV1.2 expression also continues to increase [Reference Nelson, Catalfio, Gupta, Min, Caballero-Floran and Dean103], but these channels accumulate in neocortical pyramidal neuron dendrites [Reference Spratt, Ben-Shalom, Keeshen, Burke, Clarkson and Sanders112]. Here, NaV1.2 is critical for dendritic excitability and activity-dependent synaptic maturation and plasticity. Under normal conditions, APs initiated in the AIS propagate both forward to sites of neurotransmitter release and backward into the dendrite. The latter “backpropagation” is partly a passive process, limited by the electrical properties of dendritic cables, and partly supported by NaV channels. As discussed below, heterozygous loss of Scn2a, which is associated with ID/ASD, impairs this backpropagation process, with downstream effects on synaptic maturation and plasticity.
In contrast to neocortical pyramidal neurons, NaV1.2 is distributed differently in unmyelinated axons where, instead of being excluded, NaV1.2 is enriched [Reference Yamano, Miyazaki and Nukina111]. This distribution pattern is best understood in striatal medium spiny neurons [Reference Miyazaki, Oyama, Inoue, Aosaki, Abe and Kiyonari117], hippocampal pyramidal neurons [Reference Wang, Takashima, Segawa, Itoh, Shi and Hwang118,Reference Shin, Kweon, Kang, Kim, Kim and Kang119], midbrain dopaminergic neurons [Reference Yang, Xiao, Li, He, Li and Shu110], and cerebellar granule cells [Reference Wang, Derderian, Hamada, Zhou, Nelson and Kyoung120], but likely extends to other neuron classes with unmyelinated axons. Interestingly, AP threshold does not appear affected by heterozygous loss of NaV1.2 in either hippocampal pyramidal neurons or cerebellar granule cells [Reference Wang, Derderian, Hamada, Zhou, Nelson and Kyoung120], indicating that another NaV channel, presumably NaV1.6, supports AP initiation in these cells. Instead, NaV1.2 supports AP propagation, ensuring faithful transmission of those APs from the AIS to sites of neurotransmitter release.
Dysfunction of NaV1.2
Detailed analysis of NaV1.2 biophysical properties provides critical insight into how SCN2A variants can result in NDD. This work is typically done in heterologous expression systems, including human embryonic kidney cells (HEK293) that express few to no sodium channels of their own. Channel biophysical properties can then be studied using patch clamp recording. Thus, researchers have been able to quantify numerous biophysical features and assign binary classifications such as GOF or LOF to individual parameters. Data from such experiments in combination with other information such as evolutionary conservation are also useful for training and validating computational (in silico) methods to predict the general functional effects of SCN2A variants [Reference Brunklaus, Ellis, Reavey, Semsarian and Zuberi60,Reference Heyne, Baez-Nieto, Iqbal, Palmer, Brunklaus and May121,Reference Brunklaus, George, Lal, Heinzen and Goldman122].
One illustration of the GOF versus LOF concept is the effect of changes in voltage-dependence of activation on neuronal excitability (Figure 8). As stated earlier, NaV1.2 channels are activated by membrane depolarization. Any mutation that hyperpolarizes the voltage-dependence of activation would increase the number of APs fired by a given stimulus, resulting in neuronal hyperactivity. By contrast, a depolarizing shift in activation voltage-dependence would result in fewer APs and neuronal hypoactivity. One can predict that mutations that enable channel opening at weaker stimulus intensities, cause channels to remain open longer, or recover faster from inactivation will potentiate neuronal excitability. Conversely, mutations that require stronger stimuli or result in channels that prefer nonconducting states would lower neuronal excitability.
Effect of altered NaV1.2 activation voltage-dependence on neuronal excitability.
Activation curves illustrating WT channels and variants with either GOF or LOF.

Simulated APs from a layer five cortical pyramidal neuron corresponding to the three conditions. Simulations performed with NEURON as described by Thompson et al. [Reference Thompson, Ben-Shalom, Bender and George91].

Characterization of a limited number of variants has suggested that NaV1.2 variants with strong GOF properties result in early onset epileptic encephalopathies, while LOF variants are associated with later onset epilepsies, ASD, or non-syndromic ID [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Berecki, Howell, Heighway, Olivier, Rodda and Overmars66,Reference Ben-Shalom, Keeshen, Berrios, An, Sanders and Bender68,Reference Thompson, Ben-Shalom, Bender and George91]. Some variants fall neatly into these two divergent categories. For example, T236S was identified in a child diagnosed with early infantile epileptic encephalopathy (Ohtahara syndrome) and showed a large hyperpolarizing shift in the voltage-dependence of activation, consistent with GOF [Reference Thompson, Ben-Shalom, Bender and George91]. Other biophysical abnormalities interpreted to be GOF include enhanced persistent sodium current and slower onset of inactivation, which result in greater magnitudes of Na+ influx over time. By contrast, the ASD-associated variant R937H is an example of a nonconducting NaV1.2 variant [Reference Ben-Shalom, Keeshen, Berrios, An, Sanders and Bender68]. However, many NaV1.2 variants associated with neurodevelopmental diseases do not exhibit simple binary effects on channel function. Instead, many mutations exhibit a constellation of GOF and LOF properties, making the net effect on neuronal output difficult to predict [Reference Thompson, Potet, Abramova, DeKeyser, Ghabra and Vanoye63].
Additional experimental strategies such as computational modeling of APs and dynamic AP clamp can help predict the net effects of NaV1.2 dysfunction on neuronal excitability (Figure 9). In silico prediction of neuron excitability by simulating neuronal APs computationally has informed the impact of certain variants on neuronal excitability [Reference Ben-Shalom, Keeshen, Berrios, An, Sanders and Bender68,Reference Thompson, Ben-Shalom, Bender and George91], and efforts to scale this method for higher throughput are underway [Reference Ben-Shalom, Ladd, Artherya, Cross, Kim and Sanghevi123]. Dynamic AP clamp interfaces voltage-clamp recording of a heterologously expressed NaV1.2 variant with a computational model of a neuron or of an isolated neuronal domain such as the AIS. This approach has been used to predict effects of NaV1.2 variants on the propensity of an AIS model to generate APs [Reference Berecki, Howell, Deerasooriya, Cilio, Oliva and Kaplan44,Reference Berecki, Howell, Heighway, Olivier, Rodda and Overmars66]. Beyond these approaches, expressing NaV1.2 variants in native neurons may be the ultimate solution for determining pathogenic effects. The technology of using induced pluripotent stem cells (iPSC) to generate genetically defined human neurons is emerging as a new approach to assess the impact of NaV1.2 variants on neuronal firing behavior [Reference Kruth, Grisolano, Ahern and Williams124,Reference Simkin, Ambrosi, Marshall, Williams, Eisenberg and Gharib125]. To date, only a limited number of studies have demonstrated effects of either GOF or LOF variants [Reference Que, Olivero-Acosta, Zhang, Eaton, Tukker and Chen126,Reference Asadollahi, Delvendahl, Muff, Tan, Rodríguez and Turan127,Reference Mao, Mattei, Rollo, Byars, Cuddy and Berecki128]. Importantly, cultured iPSC-derived neurons exhibit features comparable to an embryonic state, which may be a limitation for modeling disorders with postnatal onset. Transgenic and knockin mouse models are covered in the next section.
Illustration of dynamic AP clamp.

SCN2A in Epilepsy
Initial reports of an association of SCN2A with human epilepsy came from family studies of benign familial neonatal-infantile seizures, which was renamed SeLFNIE in recognition that seizures are not always benign [Reference Zuberi, Wirrell, Yozawitz, Wilmshurst, Specchio and Riney13]. In 2002, SCN2A variants were found to segregate with epilepsy in two independent multigenerational pedigrees [Reference Heron, Crossland, Andermann, Phillips, Hall and Bleasel10]. The clinical spectrum expanded to include SCN2A-associated DEE with the first reported associations in 2009 [Reference Ogiwara, Ito, Sawaishi, Osaka, Mazaki and Inoue129,Reference Shi, Yasumoto, Nakagawa, Fukasawa, Uchiya and Hirose130]. SeLFNIE is a transient disorder with seizure onset in infancy that resolves after two years of age and responds well to sodium channel blockers. Inherited SeLFNIE variants are less deleterious than de novo variants associated with DEE. SCN2A-related DEE can be separated by age of seizure onset. Early seizure onset before three months of age is associated with GOF variants that respond to sodium channel blockers, while later seizure onset after three months of age is associated with partial or complete LOF variants that are exacerbated by sodium channel blockers. More details on the genotype–phenotype relationships in SCN2A-related disorders were covered the Clinical Spectrum and Genotype–Phenotype Correlations section.
The first mouse model of SCN2A-related epilepsy preceded the association with human epilepsy and provided supportive evidence for those early reports. Scn2aQ54 mice express a transgene in neurons with a GOF missense mutation that results in elevated persistent sodium current in vivo and a severe epilepsy phenotype [Reference Kearney, Plummer, Smith, Kapur, Cummins and Waxman131]. Phenotype severity in Scn2aQ54 mice is dependent on background strain, supporting a contribution of genetic modifiers even in the case of a highly penetrant driver mutation [Reference Bergren, Chen, Galecki and Kearney132]. Scn2aQ54 mice have been a useful system for identifying genetic modifiers that may contribute to variable presentation of epilepsy associated with SCN2A variants [Reference Thompson, Hawkins, Kearney and George101,Reference Jorge, Campbell, Miller, Rutter, Gurnett and Vanoye133,Reference Hawkins and Kearney134,Reference Calhoun, Hawkins, Zachwieja and Kearney135]. With the advent of genome editing technology, it has become easier to develop knockin models carrying human pathogenic variants in the corresponding position in the mouse gene. New mouse models of SCN2A-related disorders have been reported, including those with recurrent variants for early onset and later onset DEE [Reference Li, Jancovski, Jafar-Nejad, Burbano, Rollo and Richards136,Reference Jia137]. Scn2aR1882Q mice, carrying the most recurrent early onset DEE variant, exhibit a severe phenotype, with seizure onset in the neonatal period and premature lethality by postnatal day 30. Pyramidal neurons from Scn2aR1882Q mice exhibit higher evoked AP firing frequency, higher input resistance and lower rheobase, consistent with a GOF phenotype [Reference Li, Jancovski, Jafar-Nejad, Burbano, Rollo and Richards136]. In contrast, Scn2aR853Q mice, carrying the most recurrent later onset DEE variant, had no spontaneous seizures or EEG abnormalities and were less prone to seizures induced by chemoconvulsants [Reference Jia137]. Primary cortical neurons from Scn2aR853Q mice showed lower firing frequency and less network bursting compared to WT neurons, consistent with a LOF phenotype [Reference Jia137].
Pathogenic variants exhibiting mixtures of LOF and GOF properties make it challenging to translate in vitro biophysical defects into a predicted effect on neuronal excitability. One such mixed function SCN2A variant is K1422E, which exhibits altered ion selectivity allowing partial Ca2+ permeation and lower overall conductance [Reference Heinemann, Terlau, Stühmer, Imoto and Numa76,Reference Schlief, Schönherr, Imoto and Heinemann77,Reference Echevarria-Cooper, Hawkins, Misra, Huffman, Thaxton and Thompson138]. A child carrying this variant presented with developmental delay, epileptic spasms with onset at 13 months of age, and features of ASD [Reference Sundaram, Chugani, Tiwari and Huq139]. Consistent with the mixed effects on channel function, Scn2aK1422E mice exhibit a combination of GOF and LOF phenotypes, including rare spontaneous seizures, background EEG abnormalities, and altered thresholds for induced seizures, as well altered neurobehavior that overlaps features of Scn2a haploinsufficient mice [Reference Echevarria-Cooper, Hawkins, Misra, Huffman, Thaxton and Thompson138]. Recordings from layer 5b pyramidal neurons showed impaired AP initiation and larger intracellular Ca2+ transients at the AIS during the rising phase of the AP, consistent with a complex effect of the p.K1422E variant on neurons. The complex effects of mixed-effect variants suggests that therapeutic responses may be challenging to predict for these special cases. Additionally, genome background and biological sex have been shown to influence phenotype severity in Scn2aK1422E mice [Reference Echevarria-Cooper, Hawkins, Misra, Huffman, Thaxton and Thompson138,Reference Echevarria-Cooper, Hawkins and Kearney140,Reference Echevarria-Cooper and Kearney141], adding to the complexity.
SCN2A in Autism Spectrum Disorder and Intellectual Disability
SCN2A variants were first associated with ASD in 2012 in a landmark genetic analysis of exomes of 928 individuals, including 200 phenotypically discordant sibling pairs [Reference Sanders, Murtha, Gupta, Murdoch, Raubeson and Willsey142]. In this cohort, SCN2A was the only gene found to have disruptive de novo variants in two separate ASD cases, which was deemed extremely unlikely due to chance. In the decade following, the association of SCN2A with ASD strengthened, and the gene is now recognized as the leading genetic risk factor for ASD among all genes identified by exome sequencing [Reference Satterstrom, Kosmicki and Wang143,Reference Fu, Satterstrom, Peng, Brand, Collins and Dong144]. This large-scale effort was conducted by the International Autism Sequencing Consortium. One of the lead investigators was Matthew State, PhD, Professor and Chair of Psychiatry and Behavioral Sciences, University of California, San Francisco, who was interviewed for this Element by Kevin Bender, PhD (Video 2).
Two-part discussion with Matthew State, MD, PhD (Professor and Chair, Department of Psychiatry and Behavioral Sciences, University of California, San Francisco) moderated by Kevin Bender, PhD (Associate Professor of Neurology, University of California, San Francisco) on discovery of SCN2A as a risk gene for ASD, the neurobiology of this gene, and prospects for future treatments. Transcripts of these videos are available in the Appendix. The video files are available at www.cambridge.org/scn2a
Despite this strong ASD association, unraveling the contributions of SCN2A to ASD has not been straightforward. Of the 72 genes currently associated with ASD, SCN2A was initially considered an outlier, because most genes encode proteins that localized to or regulate synapses or are genes encoding chromatin modifiers in neuronal nuclei [Reference Fu, Satterstrom, Peng, Brand, Collins and Dong144]. By contrast, SCN2A encodes a NaV channel involved in AP initiation and propagation in neuronal axons, which was considered distinct from synapses and gene regulatory processes in the nucleus. It was with the discovery that NaV1.2 localizes to neocortical pyramidal cell dendrites that some level of convergence was first understood. In dendrites, NaV1.2 interacts functionally with synaptic genes, regulating dendritic excitability and synaptic plasticity. Heterozygous loss of NaV1.2 in neocortical neurons results in cell-autonomous impairments in dendritic excitability. This dendritic excitability is instructive for activity-dependent refinement of excitatory circuits. In heterozygous Scn2a knockout (Scn2a+/-) mice, excitatory synapses are less mature, with an excess of silent synapses, lower AMPA:NMDA receptor ratio, and greater proportion of NR2B-containing NMDA receptors more commonly found in developing synapses. Furthermore, impairments in NaV-supported propagation of APs into the dendritic compartment were associated with a failure to induce synaptic plasticity [Reference Nelson, Catalfio, Gupta, Min, Caballero-Floran and Dean103,Reference Spratt, Ben-Shalom, Keeshen, Burke, Clarkson and Sanders112]. Evidence has emerged that ankyrin-B, encoded by the ASD-risk gene ANK2, contributes to scaffolding NaV1.2 channels to dendritic membranes. Heterozygous loss of ankyrin-B in mice phenocopies the synaptic dysfunction discovered in Scn2a+/- mice, and this implicates dendritic dysfunction as a convergent mechanism in ASD.
Impairments in synaptic plasticity also extend to other brain regions in Scn2a haploinsufficiency. In hippocampus and cerebellum, excitatory synapses onto principal neurons (pyramidal cells and Purkinje cells, respectively) can be strengthened in response to bursts of input at high frequency. In both brain regions, the axons that convey these bursts are unmyelinated and use NaV1.2 to propagate APs from the AIS to synaptic boutons. In Scn2a+/- mice, this transmission may be less reliable, simply because there are fewer NaV channels available to initiate APs [Reference Hu and Jonas145]. As a result, synaptic plasticity that depends on burst transmission is either blunted or eliminated in both brain regions [Reference Shin, Kweon, Kang, Kim, Kim and Kang119,Reference Wang, Derderian, Hamada, Zhou, Nelson and Kyoung120].
Synaptic plasticity deficits may be linked to behavior. Although Scn2a+/- mice perform reasonably well in many behavioral assays [Reference Spratt, Ben-Shalom, Keeshen, Burke, Clarkson and Sanders112], they may have difficulties performing more complex spatial memory tasks. Unit recordings from the hippocampus revealed that these spatial memory impairments were correlated with alterations in “replay,” which is a coordinated, sequential activation of a series of hippocampal pyramidal neurons that appears associated with the animal recalling its movement through space [Reference Middleton, Kneller, Chen, Ogiwara, Montal and Yamakawa146]. Moreover, recent work examining cerebellar circuitry has revealed that Scn2a+/- mice and children heterozygous for SCN2A LOF variants exhibit hypersensitivity of the vestibulo-ocular reflex (VOR) [Reference Wang, Derderian, Hamada, Zhou, Nelson and Kyoung120]. The VOR is a conserved behavior that reflexively moves the eyes in the opposite direction of the head to help stabilize a visual scene on the retina. In mice, the combination of heightened VOR sensitivity and an inability to modulate VOR amplitude is linked specifically to heterozygous loss of Scn2a in cerebellar granule cells, which are neurons that provide excitatory input to Purkinje cells, and was found to impair synaptic plasticity between granule cells and Purkinje cells. This form of plasticity is common throughout the cerebellum [Reference Carey147]. As such, deficits in VOR behavior may serve as a bellwether of broader dysfunction in cerebellar plasticity, which could affect a range of behaviors, including motor coordination and social interaction [Reference Schmahmann148].
Mouse genetics allows for study of not only Scn2a haploinsufficiency, mimicking the condition resulting from heterozygous LOF variants, but also genetic conditions not observed in humans (Figure 10). While constitutive knockout of Scn2a is lethal [Reference Planells-Cases, Caprini, Zhang, Rockenstein, Rivera and Murre149], 75 percent loss of Scn2a throughout the brain, or conditional knockout of Scn2a in select neuron classes, is tolerated [Reference Spratt, Alexander, Ben-Shalom, Sahagun, Kyoung and Keeshen113,Reference Zhang, Chen, Eaton, Wu, Ma and Lai150]. These conditions have helped reveal phenotypes not easily observed in heterozygous Scn2a+/- mice that may nevertheless inform on the human condition. To generate 75 percent loss of Scn2a, a gene knockdown approach termed gene trap was used to block most transcription from both Scn2a alleles in mice [Reference Ma, Eaton, Liu, Zhang, Chen and Tu151]. These mice display a range of behavioral deficits not observed in heterozygous Scn2a+/- mice, including markedly elevated anxiety-related behavior, impaired nesting behavior, and fractured sleep associated with altered activity in the suprachiasmatic nucleus [Reference Ma, Eaton, Liu, Zhang, Chen and Tu151,Reference Eaton, Zhang, Ma, Park, Lietzke and Romero152]. Moreover, ex vivo recordings of principal neurons in striatum and neocortex revealed an unexpected intrinsic hyperexcitability in AP generation, which was due to lower potassium channel function that normally dampens excitability. Hyperexcitability was also observed when Scn2a expression was completely knocked out using a viral approach in adolescent mice (postnatal days 28–44). In this case, hyperexcitability was due not to a change in potassium channel expression but rather to a disrupted interaction between the loss of dendritic NaV1.2 and dendritic potassium channels, where a loss of NaV1.2-mediated depolarization led to a loss of potassium channel-mediated repolarization. Because neurons did not hyperpolarize to the same voltages between APs, it was then easier to generate subsequent APs [Reference Spratt, Alexander, Ben-Shalom, Sahagun, Kyoung and Keeshen113]. Such effects appear consistent in vivo, as conditional knockout in the same population of pyramidal cells, and only those pyramidal cells, is sufficient to promote seizures [Reference Ogiwara, Miyamoto, Tatsukawa, Yamagata, Nakayama and Atapour153].
Range of NaV1.2 dysfunction in reported mouse models of SCN2A-related disorders. Mouse models with genotypes and phenotypes associated with SCN2A-related disorders span and expand on the range of genotypes observed in the human population. GOF missense variants include those observed in children (black) and those that result in epilepsy in mice due to knockin of amino acids that would be unlikely to occur in humans (Q54). Mixed function variants, with features of both GOF and LOF, have been generated for the K1422E variant, which converts NaV1.2 from a Na+ ion selective channel to a nonselective cation channel. LOF variants include variants that truncate the NaV1.2 protein. Beyond these cases that can occur in children, mouse genetics has allowed for other approaches, including ~75 percent to 100 percent reduction of Scn2a expression, either throughout the brain or in select cells, as well as rescue alleles where full Scn2a expression can be restored in neurons using Cre recombinase.

Overall, these changes in excitability, observed when model systems are pushed beyond heterozygosity, may shed light on why some children with SCN2A protein truncating variants develop epilepsy. They suggest that Scn2a+/- mice, on the C57BL/6 background, are near a threshold for excess excitability that can be observed if the system is pushed. Of note, different strains of inbred mice display different propensity for seizures and can be either protective or more susceptible to seizure electrogenesis if converted into an epilepsy model (e.g., by providing a pro-convulsant drug or making a genetic manipulation to a key ion channel). This variability is also present in the human population and likely contributes to variability in presentation of epilepsy in those with SCN2A LOF variants. Moving forward, a deeper understanding of factors that push or pull systems from a seizure susceptibility cliff will be critical, in both mouse models and other model species.
Treatment of SCN2A-Related Disorders
Treatment of SCN2A-related disorders is primarily aimed at seizure control along with supportive therapies for symptoms related to ASD and developmental delay. When possible, treatment should to be tailored to an individual’s pathogenic variant. Certain strategies such as the use of sodium channel blocking antiseizure medications may be more effective in persons with GOF variants and less effective in cases with LOF variants. While symptomatic treatments will continue to be important, disease-modifying therapeutic strategies targeting the underlying pathophysiology are emerging. This section will initially focus on the symptomatic treatment of epilepsy followed by a discussion of emerging disease-modifying therapies for which preclinical proof-of-concept exists for SCN2A-related disorders.
Antiseizure Therapy for SCN2A-Related Disorders
Epilepsy due to GOF variants typically presents as neonatal or early infantile onset seizures. Clinical evidence supports the notion that sodium channel blockers are more often effective in this setting, but are less effective or ineffective if epilepsy begins after three months of age [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5]. The difference in epilepsy age of onset between groups with GOF versus LOF variants is a general trend but is not absolute, and some variants are difficult to categorize because they exhibit features of both GOF and LOF [Reference Thompson, Potet, Abramova, DeKeyser, Ghabra and Vanoye63].
The efficacy of specific sodium channel blockers in the setting SCN2A GOF variant is heterogeneous. In a limited number of case series, phenytoin and carbamazepine had the best response rates [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5,Reference Howell, McMahon, Carvill, Tambunan, Mackay and Rodriguez-Casero154,Reference O’Connor, Kirschenblatt, Laux, Berg, Misra and Millichap155]. High dose phenytoin may be required for efficacy [Reference Howell, McMahon, Carvill, Tambunan, Mackay and Rodriguez-Casero154,Reference Dilena, Striano and Gennaro156], but this is accompanied by greater risk of adverse effects and requires close monitoring of plasma free drug concentration. Awareness of potential drug–drug interactions and pharmacogenomic factors is also important [Reference Welzel, Ziesenitz, Waldvogel, Scheidegger, Weber and van den Anker157]. Genetic variants in CYP2C9 affecting phenytoin metabolism may predispose to toxicity [Reference Karnes, Rettie, Somogyi, Huddart, Fohner and Formea158]. While use of sodium channel blockers can produce significant seizure reduction in individuals with GOF variants, anecdotal evidence suggests developmental delays may persist [Reference Dilena, Striano and Gennaro156,Reference Adney, Millichap, DeKeyser, Abramova, Thompson and George159].
Individuals with SCN2A LOF variants may also have seizures, which tend to begin later in infancy or in early childhood and do not generally respond to sodium channel blockers [Reference Wolff, Johannesen, Hedrich, Masnada, Rubboli and Gardella5]. There is little evidence supporting specific antiseizure therapies for affected individuals with SCN2A LOF or mixed function variants. Treatment for SCN2A-related NDD is primarily supportive, with applied behavioral analysis along with physical, occupational, and speech therapies. Recognition of cortical visual impairment allows accommodations to be integrated into an individual’s care plan in a way that maximizes successful visual interaction with their environment.
Emerging Disease-Modifying Therapies for GOF SCN2A Variants
While no disease-modifying therapy is currently available for SCN2A-related disorders, many potential avenues are being investigated (Table 2), with some moving into clinical trials. The remainder of this section will focus on emerging therapeutic approaches that may have efficacy for these conditions (Figure 11). These advances are made possible by collaborations between academic laboratories and the pharmaceutical industry, as illustrated by an interview with Dr. Steven Petrou, Professor of Translational Neuroscience, University of Melbourne, and Chief Scientific Officer of Praxis Precision Medicines (Video 3).
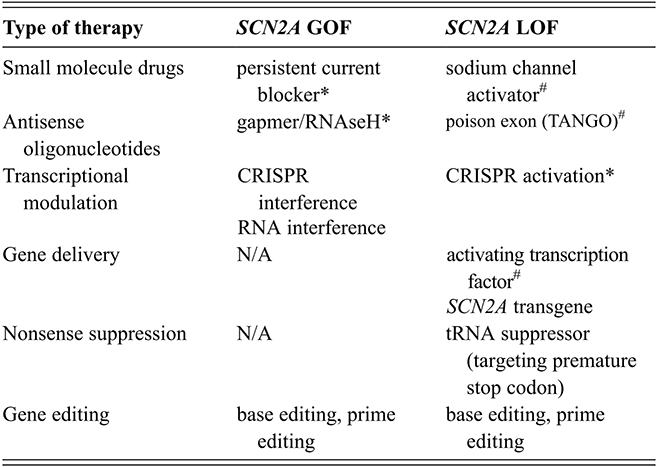
* existing preclinical proof-of-concept for SCN2A;
# preclinical proof-of-concept for another sodium channel gene;
N/A, not applicable
Overview of potential disease-modifying therapies in SCN2A-related disorders. The left side of the figure illustrates general approaches. The triangle on the right side illustrates the ease of delivery (most difficult delivery is the top). The inverted triangle on the right side illustrates the time therapy remains in the body.

Interview of Steven Petrou, PhD (Chief Scientific Officer, Praxis Precision Medicines) conducted by Alfred L. George, Jr., MD (Professor and Chair of Pharmacology, Northwestern University) explaining his transition from academic research to leading a new pharmaceutical company developing treatments for SCN2A-related disorders. A transcript of this video is available in the Appendix. The video file is available at www.cambridge.org/scn2a
Small Molecule Drugs
Conventional drug therapy, which modulates the activity of mutant sodium channels, is the mainstay of treatment for SCN2A GOF variants. However, sodium channel blocking antiseizure medications such as phenytoin do not correct the underlying functional defect and also affect normal sodium channels. New small molecule drugs in development are attempting to target channel dysfunction more precisely with the goal of being more effective with less toxicity.
One strategy targets a specific functional anomaly, enhanced persistent sodium current, exhibited by many SCN2A GOF variants. Persistent sodium current arises when channel inactivation is not complete, leading to a small level of inward current that can affect neuronal excitability. This functional anomaly occurs in many sodium channelopathies in addition to SCN2A-related disorders. The first animal model (Q54 mouse) to test the hypothesis that enhanced SCN2A persistent current can cause epilepsy was reported years before the first human mutations were discovered [Reference Kearney, Plummer, Smith, Kapur, Cummins and Waxman131]. Later investigations using this mouse model demonstrated the antiseizure efficacy of an approved antianginal drug, ranolazine [Reference Anderson, Thompson, Hawkins, Nath, Petersohn and Rajamani160], which selectively suppresses persistent current carried by cardiac sodium channels [Reference Fredj, Sampson, Liu and Kass161]. This study was motivated by in vitro evidence that ranolazine blocked persistent sodium current carried by mutant NaV1.1 channels (encoded by SCN1A) [Reference Kahlig, Lepist, Leung, Rajamani and George162,Reference Kahlig, Hirakawa, Liu, George, Belardinelli and Rajamani163]. Acute exposure of Q54 mice to ranolazine was effective at reducing seizure frequency, but poor brain penetration indicated that this drug would not be effective clinically [Reference Fredj, Sampson, Liu and Kass161]. However, a novel nonselective sodium channel blocker (GS-458967 or GS967; later named PRAX-330) was described as a potent and preferential inhibitor of persistent sodium current [Reference Koltun, Parkhill, Elzein, Kobayashi, Notte and Kalla164]. GS967 exhibits near complete seizure suppression in Q54 mice and prevented seizures following maximum electroshock [Reference Anderson, Thompson, Hawkins, Nath, Petersohn and Rajamani160]. GS967 was shown to suppress persistent current carried by recombinant human NaV1.2 [Reference Mason and Cummins165] and to prevent seizures in a mouse model of DEE caused by an SCN8A GOF variant [Reference Baker, Thompson, Hawkins, Wagnon, Wengert and Patel166,Reference Wengert, Saga, Panchal, Barker and Patel167].
Despite success in mouse models, GS967 had undesirable pharmacokinetic properties (e.g., excessive plasma and brain half-life) [Reference Koltun, Parkhill, Elzein, Kobayashi, Notte and Kalla164] and exhibits potent use-dependent block of peak sodium current [Reference Potet, Vanoye and George168]. However, the observed efficacy of preferential persistent sodium current block in mouse models of epilepsy motivated development of newer agents, including PRAX-562 [Reference Kahlig, Scott, Hatch, Griffin, Martinez Botella and Hughes169] and NBI-921352 [Reference Johnson, Focken, Khakh, Tari, Dube and Goodchild170], although neither was selective for NaV1.2. PRAX-562 is orally active and exhibits greater preference for blocking persistent current carried by mutant or neurotoxin (ATX-II)-modified NaV1.6 channels than conventional sodium channel blocking antiseizure drugs (e.g., phenytoin, carbamazepine, lamotrigine) [Reference Kahlig, Scott, Hatch, Griffin, Martinez Botella and Hughes169]. This compound has similar effects on other human sodium channels, including NaV1.2. By contrast, NBI-921352 is selective for blocking NaV1.6 channels and doesn’t specifically target persistent current [Reference Johnson, Focken, Khakh, Tari, Dube and Goodchild170]. In mice subjected to maximum electroshock, PRAX-562 exhibits dose-dependent anticonvulsant effects with less neurotoxicity than carbamazepine and lamotrigine [Reference Kahlig, Scott, Hatch, Griffin, Martinez Botella and Hughes169]. Following demonstration of tolerability in healthy volunteers [Reference Mahalingam, Oldham, Puryear, Bansal, Sriram and Patel171], PRAX-562 received orphan drug and rare pediatric disease designations by the FDA and entered Phase 2 clinical trials for SCN2A- and SCN8A-related DEE (NCT05818553, ClinicalTrials.gov).
While more targeted drugs show promise for controlling epilepsy, there remains the question of whether this type of therapy will be disease-modifying because sodium channel blockade has not been shown to rescue the neurodevelopmental phenotype. In theory, this could be related to the degree of selectivity for the mutant sodium channel and whether the small molecule corrects the specific pathophysiology of a particular pathogenic variant. It is conceivable that more precise sodium channel inhibition initiated earlier in the disease course may potentially improve seizure control and developmental outcomes.
Antisense Oligonucleotide (ASO)
Among the first disease-modifying therapies approved for NDD was an ASO (nusinersen) for spinal muscular atrophy [Reference Finkel, Mercuri, Darras, Connolly, Kuntz and Kirschner172,Reference Mercuri, Darras, Chiriboga, Day, Campbell and Connolly173]. ASOs are short stretches of synthetic nucleic acids typically comprised of deoxyribonucleic acid (DNA) or chemically modified ribonucleic acid (RNA) nucleotides. These nucleic acid drugs bind to a complementary sequence (antisense) within mRNA or pre-mRNA and act either by steric interference with enzymatic processing (e.g., splicing, translation) or by engaging ribonuclease H (RNaseH) to cleave hybrid RNA:DNA duplexes. Chemically modified RNA nucleotides (e.g., ribose-O-2-methoxyethyl or 2’-MOE) coupled with sulfur-containing (e.g., phosphorothioate) backbone linkages are employed to shield the molecule from degrading enzymes [Reference Crooke, Liang, Baker and Crooke174]. ASOs used to target mRNA or pre-mRNA for RNaseH cleavage typically use a ‘gapmer’ design consisting of a DNA core (needed to form nuclease-susceptible RNA:DNA duplexes with the target) flanked by short chemically modified RNA sequences (Figure 12). Splice-modifying ASOs can be composed of DNA or modified RNA only. These two mechanisms of action make ASOs versatile in their application (Figure 13), leading to several clinical trials for a variety of neurogenetic conditions [Reference Carvill, Matheny, Hesselberth and Demarest175,Reference Hill and Meisler176,Reference Bennett, Kordasiewicz and Cleveland177,Reference McCauley and Bennett178].
Design features and chemistry of a gapmer ASO.
Gapmer design with a 10 nucleotide DNA core and flanking five nucleotide modified RNA sequences.

Chemical structure of a 2’-MOE RNA base with phosphorothioate linkage.

Strategies for gene regulation by ASOs.
RNase H–mediated mRNA degradation.

Splice-switching ASOs that promote exon inclusion by masking silencer regions. (C) Splice-switching ASOs that promote exon skipping by masking splicing enhancer regions.

Preclinical evidence suggests that ASOs have potential therapeutic benefits for SCN2A-related disorders associated with GOF variants. Li et al. demonstrated that immediate postnatal delivery of a gapmer ASO targeting SCN2A by the intracerebroventricular route lowered spontaneous seizure frequency and prolonged survival in mice heterozygous for a GOF missense variant (equivalent to human R1882Q) [Reference Li, Jancovski, Jafar-Nejad, Burbano, Rollo and Richards136]. Administration of the ASO two weeks after birth also conferred a survival benefit, as did a second dose given approximately four weeks after birth in mice that received a first dose on day one of life. The ASO used in this study did not affect protein levels of other neuronal sodium channels, and treated mice showed no overt neurological toxicity. In 2023, a human SCN2A-selective ASO (PRAX-222, elsunersen) entered early stage clinical trials (NCT05737784, ClinicalTrials.gov).
While encouraging, these results need to be taken with tempered expectations as there are important considerations with ASO treatment in this context. A potential concern is over-reduction of NaV1.2 protein levels, which may induce a LOF phenotype, a concept referred to as the goldilocks principle [Reference Ta, Downs, Baynam, Wilson, Richmond and Leonard179]. ASOs do not allow for quick or easy dose titration given their long half-lives (more than 100 days in brain), and this creates challenges for achieving a specific range of mRNA and protein reduction. Uptake in brain is not uniform, with superficial structures and spinal cord having higher uptake than deep structures [Reference Mortberg, Gentile, Nadaf, Vanderburg, Simmons and Dubinsky180]. Another consideration is whether a simple reduction in NaV1.2 protein will result in the appropriate level of functioning protein. The ASO will down-regulate both SCN2A mRNA alleles. It is important to recognize that while we refer to variants that result in increased currents as “gain-of-function,” this is dysfunctional sodium current, and therefore reduction of both protein alleles doesn’t correct the defect specifically. The extent to which lowering the amount of NaV1.2 channels will produce a meaningful disease improvement in humans remains an open question.
Other Potential Therapeutic Strategies for SCN2A GOF
There are other emerging approaches for lowering expression of a target gene, including RNA interference [Reference Germain, Chung and Sarmiere181] and programmable transcription interference using clustered regularly interspaced short palindromic repeat (CRISPR) interference [Reference Bendixen, Jensen and Bak182] that could have value for SCN2A GOF variants [Reference Carpenter and Lignani183]. These approaches could work in a similar way as ASOs to diminish NaV1.2 channel production. In addition to suppressing expression, base editing [Reference Porto, Komor, Slaymaker and Yeo184] and prime editing [Reference Chen and Liu185] are promising technologies for correcting pathogenic variants, which have been successful for other genes and may work in mammalian brain [Reference Davis, Banskota, Levy, Newby, Wang and Anzalone186].
Emerging Disease-Modifying Therapies for LOF SCN2A Variants
CRISPR-Cas9 Activation (CRISPRa)
As mentioned earlier, CRISPR technology can enable targeted transcriptional regulation of genes. In the original embodiment, a CRISPR-associated nuclease (e.g., Cas9) cleaves genomic DNA at a position targeted by a single guide RNA (sgRNA), creating double-stranded breaks that are repaired by either nonhomologous end-joining or homology-dependent repair. An enzymatically inactive Cas9 (“dead” Cas9, dCas9) or a chimeric dCas9 fused to a transcriptional activator can similarly be positioned at a precise location in the genome, but this will not cut DNA. Rather, placing the chimeric dCas9 near the transcription start site will activate transcription for that gene. This strategy, called CRISPR activation (CRISPRa) [Reference Bendixen, Jensen and Bak182], has been demonstrated to boost SCN2A expression in heterozygous knockout mice and human stem cell-derived neurons [Reference Tamura, Nelson, Spratt, Kyoung, Zhou and Li187]. In that study, CRISPRa components were delivered in vivo using recombinant adeno-associated virus (AAV) intravenously or by stereotaxic injection directly into mouse brain. CRISPRa restored normal neuronal and dendritic excitability in heterozygous mice and did not induce seizures in WT mice. The latter observation suggests there are regulatory mechanisms to prevent SCN2A overexpression, which could convert a LOF disorder into a GOF disorder. Similar success has been demonstrated for SCN1A LOF in a mouse model of Dravet syndrome [Reference Colasante, Lignani, Brusco, Di Berardino, Carpenter and Giannelli188].
An alternative approach for transcriptional activation of brain sodium channel genes involves use of engineered transcription factors. This approach has been used successfully for boosting neuronal expression of SCN1A and improving survival of Dravet syndrome mice [Reference Tanenhaus, Stowe, Young, McLaughlin, Aeran and Lin189]. Preparations for an early phase clinical trial with the molecule used in this study (ETX101) commenced in 2022 (NCT05419492, ClinicalTrials.gov).
Other Potential Therapeutic Strategies for SCN2A LOF
Preclinical proof-of-concept exists for additional approaches for rescuing LOF variants in other sodium channel genes. Targeted augmentation of nuclear gene output (TANGO) is a strategy for suppressing unproductive mRNA splicing events arising from inclusion of “poison exons,” which evoke nonsense-mediated decay. Unproductive splicing events involve several genes associated with NDD [Reference Carvill and Mefford190], including SCN1A, which encodes the NaV1.1 sodium channel and is the major gene responsible for Dravet syndrome. A splice-switching ASO (STK-001) prevents inclusion of a poison exon in SCN1A transcripts, boosts translation of functional NaV1.1 protein, and improves survival in a Dravet syndrome mouse model [Reference Han, Chen, Christiansen, Ji, Lin and Anumonwo191,Reference Wengert, Wagley, Strohm, Reza, Wenker and Gaykema192]. Clinical trials are underway to test efficacy and safety of STK-001 in Dravet syndrome (NCT04442295, NCT04740476, ClinicalTrials.gov). Whether a similar strategy will work for SCN2A LOF variants is unknown.
Hypothetically, SCN2A-related disorders caused by LOF variants could be approached by delivering an exogenous copy of the recombinant gene or its coding region to brain using a viral vector. While conceptually appealing, this approach is not feasible with the currently available vectors that are most advanced in clinical trials (e.g., AAV), because the large size of the SCN2A coding region exceeds the capacity of these vectors. Newer vectors may overcome this size restriction as demonstrated for SCN1A [Reference Fadila, Beucher, Dopeso-Reyes, Mavashov, Brusel and Anderson193,Reference Mora-Jimenez, Valencia, Sanchez-Carpintero, Tønnesen, Fadila and Rubinstein194,Reference Chilcott, Díaz, Bertram, Berti and Karda195].
For SCN2A LOF variants that create premature termination codons (PTC, nonsense), a novel approach (nonsense suppression) utilizing modified transfer RNA (tRNA) may have value. This approach seeks to trick ribosomes into incorporating the correct amino acid at the location of the PTC by using anticodon engineered tRNA molecules [Reference Lueck, Yoon, Perales-Puchalt, Mackey, Infield and Behlke196,Reference Porter, Heil and Lueck197]. Intrinsic features of the translational machinery appear to protect native stop codons from the modified tRNA. Viral delivery of a suppressor tRNA was used successfully in vivo in mice to partially correct an engineered PTC mutation in the gene encoding α-l-iduronidase, which causes the lysosomal storage disease mucopolysaccharidosis type I in humans [Reference Wang, Zhang, Mendonca, Yukselen, Muneeruddin and Ren198]. Nonsense suppression has garnered considerable commercial interest [Reference Dolgin199] and there is interest in applying this strategy for SCN1A and SCN2A [Reference Chilcott, Díaz, Bertram, Berti and Karda195,Reference Ahern200], but major challenges must be overcome [Reference Porter, Heil and Lueck197].
Considerations for Disease-Modifying Clinical Trial Design
Bringing a disease-modifying therapy to the clinic requires not only evidence of efficacy but also a need to consider aspects of clinical trial readiness. These include precise clinical trial design (duration, most appropriate control group), accurate patient selection (age, variant type), and outcome measures that can demonstrate meaningful changes over time. Clinical trials for conventional small molecule drugs with a primary seizure endpoint are based on established designs, such as randomized, placebo-controlled trials, but these are less feasible and appropriate for disease-modifying therapies [Reference Brock, Demarest and Benke201]. Furthermore, for SCN2A-related disorders disease-modifying therapies may target specific subsets of affected individuals, which further subdivides an already rare disease and may compromise study power.
Variations from traditional clinical trial designs are required to achieve well-powered studies. One nontraditional trial design, stepped-wedge design (also known as a randomized start design) [Reference Brock, Demarest and Benke201], has specific advantages such as allowing comparison both within groups and between groups, thereby increasing the power of the study with a relatively small number of trial participants. Using this approach, all groups eventually receive treatment, which is ethically superior to trials with a placebo group. The effect of treatment duration can also be investigated because each group is introduced to the treatment at different time points. So-called “n of 1” trial designs, including individual crossover studies, may also have value in rare diseases [Reference Strupp, Kalla, Claassen, Adrion, Mansmann and Klopstock202]. Depending on the chosen outcome measures, study duration may need to be adjusted to account for developmental outcomes that require longer study times. Careful consideration of these features is needed for clinical trial designs in SCN2A-related disorders to ensure collection of the rich and diverse data necessary for successful assessment of a novel therapy.
Another challenge to clinical trial design in SCN2A-related disorders is the heterogeneous functional effects of variants. As described previously, specific strategies can be tailored to either GOF or LOF variants. While neonatal and early infantile-onset epilepsy is typically associated with GOF variants, this is not absolute. For the aforementioned clinical trial of an ASO for SCN2A GOF variants (PRAX-222), study participants must have seizure onset before three months of age (NCT05737784, ClinicalTrials.gov). However, this could potentially include some affected individuals with LOF variants unknown at the time of enrollment. For future disease-modifying therapies, especially with approaches with permanent effects such as gene delivery and gene editing, functional testing of variants prior to treatment may need to be considered [Reference Brunklaus, George, Lal, Heinzen and Goldman122]. Furthermore, individuals in this and other SCN2A-related clinical trials are enrolled beginning at age two years, which is well beyond disease onset and raises the possibility that response to therapy could be affected by prior seizure burden or pharmacological treatments.
Selection of appropriate clinical trial outcome measures is challenging for SCN2A-related disorders using existing methods. This is illustrated by a recent SCN2A clinical trial readiness study in which mean standardized domain scores on the Vineland Adaptive Behavior Scales were three standard deviations below the normative average [Reference Berg, Palac, Wilkening, Zelko and Schust Meyer203]. The Vineland has challenges assessing small changes at low scores, and therefore raw scores or growth scale values may be more effective in eliciting meaningful change without a floor effect. Seizure reduction, even greater than a 50 percent reduction, as the main outcome measure may not encapsulate the full array of disease-modifying effects families and clinicians are hoping to achieve with these therapies, which often involve more risk than traditional seizure medications. With invasive and potentially permanent types of treatment (Table 2), the benefit of such therapies needs to outweigh those risks. The hope is that the benefit from a disease-modifying therapy would entail more than reducing seizure frequency. Given that seizure control in SCN2A-related disorders does not necessarily rescue the developmental phenotype, it will be especially important to target other outcome measures, including developmental achievement, sleep, quality of life, and behavior, among others, to ensure that the full spectrum of disease-modifying effects is being assessed.
Abbreviations
- AAV
adeno-associated virus
- AIS
axon initial segment
- AP
action potential
- ASD
autism spectrum disorder
- ASO
antisense oligonucleotide
- Cas
CRISPR-associated nuclease
- CRISPR
clustered regularly interspaced short palindromic repeat
- CRISPRa
CRISPR activation
- DDD
Deciphering Developmental Disorders study
- DEE
developmental and epileptic encephalopathy
- EEG
electroencephalography
- EIMFS
epilepsy of infancy with migrating focal seizures
- ESES
electrical status epilepticus during slow-wave sleep
- FDA
Food and Drug Administration
- GOF
gain of function
- HRQoL
health-related quality of life
- ID
intellectual disability
- iPSC
induced pluripotent stem cells
- IESS
infantile epileptic spasms syndrome
- LGS
Lennox-Gastaut syndrome
- LOF
loss of function
- MAE
myoclonic atonic epilepsy
- NDD
neurodevelopmental disorders
- PTC
premature termination codons
- SeLFNIE
self-limited familial neonatal-infantile epilepsy
- sgRNA
single guide RNA
- SUDEP
sudden unexpected death in epilepsy
- TANGO
targeted augmentation of nuclear gene output
- tRNA
transfer RNA
- VOR
vestibulo-ocular reflex
- WES
whole exome sequencing
Appendix
Transcript of Video 1: Interviews of parents of children with SCN2A-related disorders
Q: How did you feel when you received the diagnosis of SCN2A-related disorder (SRD)?
Sandya Crasta, Missouri, USA: I felt shattered and heart breaking. The doctors did not know anything about this diagnosis. They were new to it, too, but luckily they do give us the information of the SCN2A Foundation, and that was helpful.
Liz Hendrickx, Belgium: My world just fell apart. I didn’t know what to do, didn’t know where to look because in Belgium no one – there just aren’t that many cases, so I couldn’t compare. And the doctors told me not to Google it [laughter], so that was pretty scary.
Ashley Taylor, Texas, USA: I felt a very mixture of emotions. For the first 23 months of my son’s life, he was very neurotypical, met his milestones and stuff; and so, seizures hit around 23 months, and within two months of those seizures hitting – and they were coming constantly – we got that diagnosis. So I felt very overwhelmed, still processing the seizures, and now we have this other diagnosis that is causing these seizures. So it was a little bit of excitement knowing that there’s something that they can possibly treat, but at the same time scary that they knew very little about it.
Amy Richards, Maryland, USA: I was actually thrilled. It was such a relief to finally figure out what was wrong. I knew from infancy that something had to have been wrong. I mean, I always equated it to a juggling act, that something would fall in our laps and, oh, she has GI reflux; oh, she’s got a tic-ing behavior; oh, she’s got apraxia of speech; oh, she has double hip dysplasia. And I kept saying, so then, what is it? There has to be something underlying. I just want a name for it.
Q: What do you feel is the hardest part about SRD?
Charlotte’s mom, Minnesota: Charlotte was very severe and was born having over 400 seizures a day, and she was very prone to illness, so spent over half of her life in the hospital and a lot of time intubated. And she suddenly declined as her life went on, and she had more and more seizures. And towards the end of it, she was in status most of the time and her brain wasn’t functioning any more. And I would say the hard part for me was that I am basically an ICU nurse, and so I knew too much, and watching her go through those things, in my heart I knew what the outcome was going to be.
Ashley Taylor, Texas, USA: Stripping everything away that I had envisioned for my child, that I had to re-evaluate, like, what his future might look like; and watching him suffer and not being able to do anything about that. And he still pushes through with a smile, and so it’s pretty – but it’s challenging to know that there’s nothing I can do about it at this moment.
Q: Do you believe in a cure for SRDs?
Sandya Crasta, Missouri, USA: I do, do strongly believe in a cure, especially because of the FamiliesSCN2A Foundation, where, you know, all the – many of the board members and the parents coming together to find a cure and fight for this cause.
Liz Hendrickx, Belgium: That’s the main reason why we’re here, why we’re not giving up, why we’re going across the globe to find people who understand, who are willing to work with us. That’s why I’m screaming it off the rooftops, telling everyone, this is what my child has. I even tell every cab driver, every Uber driver. I’m like, I’m here for this. This is my daughter, she has this. I’m telling everyone who wants to hear.
Amy Richards, Maryland, USA: I do, I think there’s going to be one. And like I said, I’m not 100 percent it’s going to be in her lifetime, but I think there’s going to be enough in her lifetime that her life gets better. But I definitely think someday, yes, there can be a cure. To think that she’s only 13, and in that short amount of time we’ve gone from me saying, there’s something wrong with this infant, just give me a name; to, we know what it is; to now, three years later, what’s happening? It blows your mind. It’s mind boggling. So yes, the amount of research that’s being done, this is – yeah, there’s a cure, it’s going to happen.
Q: How have you been impacted by FamilieSCN2A Foundation?
Ashley Taylor, Texas, USA: Huge support system, knowing that I’m not alone, and that there are others that understand my struggle, and it’s a judgment-free zone. And in the same sense, it’s a source of hope because I see what the Foundation is doing, and it gives me hope that people care and they are trying to do something about the SCN2A mutation.
Charlotte’s mom, Minnesota: I believe that the Foundation, the SCN2A Foundation provides a community of support for our families that gives families hope. And I am really excited at the research that they have initiated to start to look more into this and to get doctors, you know, excited about researching this disease; and putting money towards finding a cure, or at least a better quality of life for our kids.
Sandya Crasta, Missouri, USA: I or my family has been impacted because we’ve got this, you know, we’ve got the support of the families that are impacted through this SCN2A. We ask questions and we get answers that we don’t get from our doctors or, you know, so we get that mental as well as just the emotional support from each other. Even coming to the conferences and learning about, you know, what the research has done, or just meeting other families, has been possible through the funding that is available through the SCN2A Foundation, for which I am very grateful.
Learn more about the FamilieSCN2A Foundation at www.scn2a.org.
Transcript of Video 2 (part 1): Interview of Matthew State, MD, PhD, by Kevin Bender, PhD
Kevin Bender (KB): Hello, my name is Kevin Bender. I am a Professor in the Department of Neurology here at UCSF, and it is my honor and pleasure to be talking with Dr. Matt State, who is a human geneticist and child psychiatrist and also the Department Chair of Psychiatry and Behavioral Sciences here at UCSF. And we are hoping to have a nice chat about SCN2A and its role in autism spectrum disorder (ASD). Matt and Matt’s group have been at the forefront of human genetics in trying to understand genetic underpinnings or genetic contributions to neurodevelopmental and neuropsychiatric disorders like autism spectrum disorder, and has made some very pivotal and, I would say, foundational discoveries on the contribution of rare de novo variants in genes like SCN2A, and has really opened up this field, which I think actually recently we called it the decade of gene discovery in ASD. And it’s actually completely changed how we think about this disorder and how we can look at it and hope to develop therapeutics in the future. So, thank you, Matt, for having me today.
Matthew State (MS): It’s an honor and a pleasure right back at you.
KB: Yeah. So, how did this start? How did you guys get started with this concept? Because I think it was a controversial idea to think about de novo variants, where there is a new change in genetics in a child that wasn’t present in their parents, that is contributing to a disorder like ASD.
MS: I think, so – that’s largely the case, I think. So, you know, we began working with a particular strategy, trying to find the kinds of mutations that you’re talking about underlying common forms of psychiatric disorders. And I had a particular interest as a child psychiatrist in neurodevelopmental disorders, working with kids with autism and Tourette’s syndrome, etc. So, you know, at that time, which was really at the turn of the millennium, the conventional wisdom was that psychiatric disorders were all uniformly genetically complex and that they were going to be a consequence of common variation in the genome, sort of hundreds to thousands of different spots in the genome contributing, each one, very small, individual effects. And for a variety of reasons, I came to the study of genetics with a kind of different perspective, that was really focused on using genetics as an entrée to understand neurobiology. So as opposed to trying to figure out sort of all of autism or understand the genetic architecture of neuropsychiatric disorders, all of which are very worthwhile and interesting things to do, I had a really kind of laser-like focus on what I used to say was, if I can just find one gene that I’m absolutely confident leads to autism – because we didn’t have that in 2000, or at least not one that we were confident led to “idiopathic autism,” so, autism without other features of severe neurodevelopmental problems. So honestly, the reason that, you know, I came to it that way because I have sort of a backwards educational path. I had completed all of my clinical training, was not a scientist, and decided to go back and do a PhD, and I did it specifically because I was seeing these kids on the units and lamenting every day the fact that we had no idea what was going on at the molecular level. So, sort of this sort of reductionist idea that, if I can just find one gene, that we can then begin to pull on that thread to go from genes to understanding, you know, cellular or molecular mechanisms in biology. So, all of this was aimed at this idea that we are, you know, sort of, if we can just find …
KB: Find that one gene.
MS: Yes, just the one gene. So then we got totally lucky, and it turned out that, in fact, much more of the disorder was underlaid by these kinds of mutations, but the things we were able to look for were, for a variety of reasons – we started looking at chromosomal breaks, and particularly looking for de novo chromosomal breaks in families in which it looked like a lightning strike genetically and we had a light microscope, that was kind of the only way to look across the genome to do this; but, you know, it’s better to be lucky than to be good. I ended up at a place where we were developing tools to be able to screen the genome at much higher resolution looking for these mutations. And so by 2008 or 2009, we knew that there were changes in chromosomal structure that were contributing to idiopathic autism, that de novo rare mutations were doing that, again to a much higher rate; you know, it wasn’t like the one in a million kids, it was more like 5 to 10 percent of kids. And then the ability to sequence the coding portion of the genome came through.
So, that was kind of the evolution of the strategy. And the critical milestone in about 2012 was, you know, we were all racing, a bunch of labs, and three hit the finish line exactly at the same time in a series of papers in Nature that showed that using this approach, that you could find rare de novo mutations that were damaging to the encoded protein, easily predict that because they were stop codons or, you know, clearly things that were damaging the protein; that they were carrying very significant risks, which is, you know, one piece of the many surprises that we found. And then, as it so happened, the first gene that we identified using this approach was SCN2A.
KB: Actually, if I remember correctly in that 2012 paper, that cohort was about how large, it was about 5,000?
MS: No, no, it was much smaller.
KB: Much smaller.
MS: Right. So, I mean that was …
KB: And even within that, you identified SCN2A twice.
MS: Yes.
KB: Which seems like astronomical odds. To be able to identify one gene – you hit one gene actually two times in a very small cohort.
MS: Yeah, so, that’s exactly right. I love that that, like that was your reaction, because everyone else’s reaction was, the cohort size was only about 200 …
KB: Oh, only 200.
MS: Yes. Well, you know, there were multiple groups, and each group had about 200 families that they were doing. So the issue is that, you know, the smaller the effect, the bigger the sample size you need. So the fact that we were able to find this among 200 represents how much of an impact mutation in SCN2A have in a kid, which is like – that was our ideal, right? We wanted to find something that had big, big biological effects.
KB: This was a case where you actually caught the same lightning bolt or two bits of the same lightning bolt in the same bottle …
MS: Yes.
KB: … in this cohort of 200 kids.
MS: Yes.
KB: Since then, it’s now been 10 years, where the cohorts have gone larger and larger and larger, from 200 now I think to 60,000?
MS: Yes.
KB: Where has SCN2A fallen within this group? As the ability to actually look at this quantitatively has improved, where has SCN2A fallen?
MS: Yeah, it’s at the very – I mean it, you know, it’s turned out to be one of the most frequent among the group of genes that have rare de novo mutations, and that’s how they’re contributing to autism; and it’s turned out to have very, very large individual effects, exactly what sort of this initial hit was suggesting. And in fact, so, there were three labs that published simultaneously; three genes came out of that: CHDA, a gene called GRIN2B, and SCN2A. All three of those – so, it just goes to show you that, you know, it’s not completely random; we caught the lightning in the bottle because this is one of the more frequent with the largest effect size, so you can plausibly find it in 200 people. So, you know, again, it’s always better to be lucky than good, it’s good to be both, and in this case, you know, it was a harbinger of, you know, an overall strategy, but the gene ended up being in a whole variety of ways particularly valuable. Like, that combination of being relatively frequent and, for a rare mutation, and having such large effects; like that is absolutely the ideal …
KB: Never happens.
MS: Yeah, and you know, like I said, I came to this a kind of reformed clinician, right? With an idea that I was going to use genetics to ultimately try to get something, some knowledge that would be, that ultimately would translate into the clinic and impact. You know, again, when I started, it was like if I could know the biology in a way that would allow us to develop a treatment, even if it was for one family, two families, three families, that that would be enormous given how mysterious autism was 20 years ago and still is today. And so, you know, it’s turned out in a whole variety of ways that SCN2A really presents, I think continues to present extraordinary opportunities.
So, once we found the gene, you know, we were incredibly excited. But then there were these whole series of kind of relatively quick-in-succession discoveries about SCN2A that, you know, that made it sort of this remarkable stroke of luck. So, at one point, I can’t remember where, I called it the Rosetta stone, or potentially Rosetta stone for understanding autism.
KB: So, what do you mean by that?
MS: Yeah, yeah. So, the first is, is like, you know, it kind of would have been enough, just the fact that it was like one of the first genes, and you could think about the fact that having, you know – there were very few, in fact no psychiatric disorders at that time, apart from autism, where you could say, I have a rare mutation in coding portion of the gene and carrying very large effects that’s causing a common neuropsychiatric or neurodevelopmental disorder. We didn’t have that at the time. So, you know, like you would think, that could be enough to call it a Rosetta stone; but then it turned out, you know, that it actually really in terms of thinking about, could you elaborate biology from that finding? There are all these other characteristics that we found about the gene that really lead, I think, to extraordinary opportunities to disentangle really complicated biology. So, one of them was that, in general, as we found these things – first I mentioned we found that rare changes in chromosomal structure could lead to autism; but at the same time what we found is that the same rare chromosomal abnormality not only led to autism but could lead to a whole variety of different neurodevelopmental outcomes. And, you know, that was an extraordinary, you know, observation, kind of defied our diagnostic nosology in psychiatry; but it also kind of presents some immediate challenges to think about, okay? I thought, well, if you have this rare gene, you’re going to be able to pull on the thread; if the thread goes in 17 different directions simultaneously, you’ve got a problem, right?
KB: Yeah.
MS: So, and then with genes that we’ve identified, to some degree we found the same, actually to a large degree we found the same kind of underlying theme, which was that they were autism genes but they also were neurodevelopmental genes. But what was even kind of more complicating is that the kinds of mutations that we were finding didn’t kind of give you immediate traction on seeing whether or not you could dissect, like, why does this one gene not only lead to autism but also lead in some cases to epilepsy, for instance?
KB: Yes, very, very commonly.
MS: Yes, very, very common, so about, particularly for kids – in general in autism, about 30 percent of kids will have epilepsy, but particularly for these rare, large-effect mutations you see a lot of overlap with epilepsy. So what was immediately – well, not immediately, but pretty soon after the discovery cooled – and actually this was Stephan Sanders, who was in the lab, he was a post-doc at the time and was the lead author on the gene discovery paper, was really interested in taking a look at the range of mutations that we were finding. And what was really unusual about SNC2A is that it started to look like there might be something really specific or much more specific than any other gene that we had, about this connection between the mutation, the kind of mutation in the gene and the outcome. So, there, again, like the notion that it could be a Rosetta stone, not only to begin to understand idiopathic autism but to really get traction on the biology of how does a rare mutation that causes a range of neurodevelopmental problems in some people end up being epilepsy, in others autism, in others the overlap of epilepsy of autism? Which turns out to be an incredibly interesting story.
Transcript of Video 2 (part 2): Interview of Matthew State, MD, PhD, by Kevin Bender, PhD
Kevin Bender (KB): I still remember the first time I met you was actually down at a retreat for our neuroscience program, and you got up and gave this wonderful talk. You had just moved with Stephan [Sanders] and Jeremy Willsey, all of you moved from Yale, take over as Chair of Psychiatry and Behavioral Sciences. And it was this watershed moment for me, sitting in the back of the room where you’re displaying this huge map of genes associated with autism that you’ve just recently discovered. And there was this group that was modifying the transcription profile of neurons, and then there was this other group that was modifying synaptic transmission. And then you made this offhand comment; it was just like, oh yeah, and then we have this other gene, SCN2A, no idea what it does; it’s down here in an axon and we don’t know why it’s there, but we can’t ignore it. And I chase you down afterwards and you said, oh, hi, who are you? I said, I really want to work on this, and this was actually a moment where we got a chance to meet up, chat with Stephan, and it really started to dissect and to link the genetics to areas of biology.
And so, one of the first things we needed to do was actually characterize a lot of the variants that you had identified, and Stephan identified, within autism spectrum disorder; and they almost invariably led to loss of function, where the channel doesn’t work as well as it should. And we had a really simple idea. It was, okay, it’s a sodium channel, it’s involved in generating action potentials; so therefore, these cells are not firing as well as they should, end of story. It was only when we got a mouse that recapitulates those phenotypes that, again, we were more lucky than good. We were completely surprised by what the biology was telling us, and that was – this is biology that we’d been doing since about 2016, 2017, where we’re starting to unravel that it’s not really the initiation of an action potential; it’s actually the communication back to synapses, and those synapses that are really providing the inputs to neurons, especially in neocortex; and SCN2A is there to provide some communication, some crosstalk, some linking of activity and activity-dependent plasticity and all of these processes that are involved in learning – SCN2A is really supporting that mechanism. And so what we found invariably, both us, other labs throughout the US and abroad, we basically found that SCN2A, when you lose half of SCN2A, as happens in autism spectrum disorder, you have these very pronounced deficits of learning, and pronounced deficits in the forms of learning that we think might be closer to things that would manifest as ASD than other forms – so, things that are involved in generalization, really complex, abstract thought; things that are involved in social communication and understanding very subtle cues when you’re having a conversation with someone. And it’s really trying to understand what the biology that has been disrupted – it’s been really remarkable that SCN2A has been able to teach us this.
One of the things that we found over and over again is this situation where, when you’ve lost half of your SCN2A expression, many neurons within the brain appear to stay in this pre-critical period holding point; and it’s almost like they’re waiting for the activity-dependent refinement, the activity-dependent plasticity to occur, but it just can’t occur because you’ve lost SCN2A. And so that actually opens up an opportunity, I think, because what we’ve seen is that that holding period seems to be very, very extended in development. And it opens up the possibility that if we can restore SCN2A, in some way …
Matthew State (MS): Yes.
KB: … even late in development, we might be able to develop therapeutics that could reinvigorate plasticity mechanisms and allow for some sort of therapeutics.
MS: Yes.
KB: Because one of the problems – the thing that I struggle with as a neurophysiologist who now, who has been studying action potentials in a dark room for the last two decades, with no interaction with humans – I work with mice, I work with rats! – now meeting these families and meeting these lovely kids, these kids are only getting older …
MS: Yeah.
KB: … and we’re not helping them yet.
MS: Yes.
KB: And so, understanding the windows of opportunity for treatment of disorders like this is something that’s outrageously critical. And my question for you is, where – what should we do as biologists within the next five years or so, and where do you see the field of gene discovery and gene identification translating into therapeutic opportunities for the field?
MS: I think that there’s a possibility that SCN2A is one of those perfect targets for a whole variety of reasons. And then this, you know, the notion about this overlap in phenotype, I think is actually going to be key to progress in the clinic now. And the reason is that – it’s a long story that I won’t belabor today, but the bottom line is that doing clinical studies of core deficits in autism is extremely difficult if not impossible right now in a way that’s relevant for the FDA. That we don’t know how to measure social behavior …
KB: Yes.
MS: … in a way that is reliable enough in humans, and we can’t do it in a timeframe – I’m not sure how well we can do it ever, but we definitely can’t do it in a timeframe that might be relevant to measure in the clinic to say, oh, if we fix this gene we’re fixing social development.
KB: Um-hmm.
MS: And therefore, what we can do is, we can study epilepsy.
KB: Yes, yes.
MS: And so the opportunity, because there is overlap, strong overlap between autism and epilepsy, I think in the next five years is going to be the path towards moving from basic biology to the clinic. There are a whole variety of things that are happening around interventions that are now plausible, including targeting individual genes either with, you know, gene replacement or in some cases antisense oligonucleotides. There are a variety of ways even potentially with repurposed small molecules if you can target epilepsy. It is now, you know, that is mature enough, work in that area is mature enough.
KB: Yeah, so that’s the potential inroads to be able to …
MS: Correct. I think I’m much more optimistic that we both in the next five years are going to begin to see evidence of the kind of work that we’ve both done …
KB: Um-hmm.
MS: … and be able to really start doing bench-to-bedside work; that there’re going to be human trials that are going to give access to biology that we’ve never had before, that we’re going to have animal models we’re going to be able to really test whether or not our predictions based on animal, you know, being able to rescue in a model system, how that plays out, you know.
KB: Yeah.
MS: So, I think it’s incredibly fertile ground.
Transcript of Video 3: Interview of Steven Petrou, PhD, by Alfred L. George, Jr., MD
Al George (AG): I’m conducting an interview today with Steve Petrou, the Professor of Translational Neurosciences at University of Melbourne, and also past Director of the Florey Institute for Neurosciences there. He is also, importantly, co-founder and now Chief Scientific Officer of Praxis Precision Medicines, which has devoted a tremendous amount of energy into developing new treatments for SCN2A disorders. Steve, thank you very much for doing this interview with us.
Steve Petrou (SP): Thanks for inviting me, Al, today; delighted to be here.
AG: The first thing I think people watching this video will be interested in knowing is, what first got you interested in studying SCN2A, and what were the driving questions at that time?
SP: Great question. I mean, my background really was in hard-core ion channel biophysics, all single-channel recordings, and when I set up my laboratory, I met Sam Berkovic and got very interested in mechanisms of disease mechanisms in genetic epilepsy. And the first area I got into was GABA, and then after studying GABA, and we made a mouse model, we were thinking very much about, well, what’s the yin and yang of the brain, and the E and the I,Footnote a and obviously, you know, sodium channels and NaV1.2, 1.6 were very important channels in determining inter-neuron and pyramidal cell function. So that said, there’s a core biological interest in it. And then we started seeing information around the variants at that time responsible for a disorder called BFNIS [benign familial neonatal-infantile seizures], it’s now called self-limiting epilepsy of infancy or neonatal/infancy. And then that was interesting from a basic biology perspective because I was always very, very interested in the work on the drosophila channels and how they used the phenotype to inform the function of the Shaker, Shab, Shal, Shaw family of channels. And this sort of gave me an opportunity to say, well, you know, human biology, genetics, has done the experiment to some extent, and so how do we understand dysfunction from that perspective in really well validated mutations? So we started studying the SCN2A at that point, and that was really before all the sporadic cases and the rare epilepsies started to be revealed, of course other than Dravet, which we knew about at the time. So that was my sort of starting point.
AG: So, can you think back to when you first started studying SCN2A variants in the benign familial neonatal/infantile seizure syndrome, which was brand new, there were so few variants; did you think this was going to last?
SP: No. I just thought we’re going to just, we did a few GABAs, we’ve now done a sodium channel, there’ll be, you know, a handful – there was only three or four different genes to choose from at the time. It wasn’t a very big list. I didn’t think it was going to last. We really were thinking more about familial epilepsy, and then we also knew at the time that these, they were still very rare, these families. So the BFNIS families weren’t a common form of familial epilepsy, you know, in the overall scheme of epilepsy. So thinking a lot about, how is this going to be impactful more broadly in the disease? And so, didn’t think it was going to last, to be honest, and didn’t think 20 years later, I’d still be doing it!
AG: What was the inspiration for then deciding to start a company? And take us back to the inspiration for starting Praxis, which I recall was not originally called Praxis; that’s its newest name.Footnote b
SP: Right.
AG: Tell us that story, if you will.
SP: When I came back from my post-doc in the US into Australia and started my laboratory, I immediately got a part-time position as a VP of CNS Research at a small Australian startup called Bionomics. And that’s where all the IP that Sam Berkovic and Grant Sutherland were generating around the familial epilepsies was residing; and we were thinking there at the time, well, how do we use that knowledge to deliver better therapeutics? And so, for me, that’s where the whole concept – and that was in the early 2000s, you know, where the whole concept of around precision medicine. I think, you know, we were on, at the time, having knowledge of a mutation that was in one family in the world wasn’t very useful. So, and efforts to try and see whether they could be broadened weren’t that successful in common epilepsy. But it did teach me a lot about thinking how to turn science into impact. It wasn’t really commercialization that interested me, it never really – you know, it’s an important thing to do, but I’ve always been interested in seeing whether my research can be more impactful than just getting more h-factors and citations and papers. That was never enough of a driver for me personally. So, that was a big motivator. And that converted into consulting I did with other companies, ran a screening program for another US company for many years doing drug discovery.
AG: So, working with industry was not a foreign concept; it was natural, and it was a vehicle to maybe perhaps make more of a translational impact.
SP: It really was something that I’d always been doing, and I think, you know, later as the field matured, it became more and more apparent what you should do in that space.
AG: Steve, tell us about your interactions with SCN2A families and, in particular, the families that engaged with you to help move forward in translational activities in SCN2A?
SP: A great question, Al. It really was a convergence of several things: our continued interest in work on the biophysics of SCN2A mutations in sporadic childhood epilepsies; our beginning to make a mouse model of one of those mutations; and our interest in precision therapies. We sort of started, reached out to Ionis, thinking about how we want to apply ASOs to that arm, to the 2 A gain-of-function cases; and at the same time, really, they reached out to me, the parents reached out to me, and we actually met at an AES meeting in the US when I visited and started to talk about what we were doing. They’d just got a fairly recent diagnosis and were very interested and were very, very motivated to try and do things. And so what they did do is really provide, they’re now catalysts for really accelerating the delivery of precision medicine therapeutics. I was interested in proof-of-concept study with Ionis,Footnote c thinking, you know, how might we show that an ASO could be effective in an animal model as a potential for a future clinical program; but they said, let’s think about what it would take to do the clinical program now.
AG: Based on all this trajectory, it was interesting to those of us in the field that you made the transition almost full time to industry now as Chief Scientific Officer of Praxis. And so, a couple things: What was the motivator for doing that? And what’s your perspective on how different academic research and industry activities are?
SP: It became more and more apparent to me that my natural interest and desire is to really translate; and whilst I loved, and I still do, basic research, I still manage my laboratory, it really wasn’t enough for me to just continue along that academic path. So I did want to see what it would take, and see whether I can actually influence the rate at which we can move these programs forward. So that was really what made me – and you know, with all the knowledge I’ve had around disease modeling, you know, the new genetics coming in, there was a lot of activity and thinking around precision medicine at the time as well, and I think at the White House had an initiative on precision medicine at the time as well. So, it was sort of a perfect storm to try and launch a company like that. And so that was really the catalyst for getting the company going; and then, you know, thinking about it for four or five years really did cement the idea that I want to go full time, you know, flip my work – now I’m more or less full time at Praxis, and I have a little small secondment to keep the laboratory going at Melbourne as well.
AG: Well, as we’re drawing to the end of this interview, I want to just give you the chance to conclude by telling us what you’re most excited about for the future of SCN2A?
SP: You know, I think we’re starting to see the signs of it already, and that is a more nuanced understanding of how variants actually produce the clinical heterogeneity that we’re seeing today. And it’s obvious to me that it’s never as simple as you first realize. The fact that we’re seeing the same variant in a benign case versus a very serious case tells us that there’s not only mechanistic complexity at the neuronal and the network and the whole-brain level, but there’s always modifying effects as well. And I think what would be interesting to me is really understanding a very tight relationship between genotype and phenotype, understanding–developing a causal chain of understanding across the whole brain so we really maybe begin to understand where the pathologies that give rise to an early onset, a late onset, an autism, an autism with epilepsy – where does that pathology really emerge? And does it give us new opportunities for therapeutic intervention beyond the blocker for gain-of-function, up-regulator for loss of function; there might be new opportunities. And sort of related to that, I think, so once we develop that more nuanced understanding of the disease biology, I think there will be clear impacts beyond 2 A epilepsy, and this will probably impact other brain disorders, depression and anxiety and a whole host of other things that will gain, you know, a lot of knowledge, and the lessons learned will be able to be applied to those other areas.
Acknowledgments
We are grateful to Louise M. Tiranoff, PhD, Founder and Managing Director of Tiranoff Productions and GeneticaLens, who produced Video 3 with the editing assistance of Adam DeSantes. We acknowledge Adil Wafa for creating Figure 9, and Shaye Moore for proofreading the manuscript and transcribing the videos.
Alfred L. George, Jr., M.D. is Professor and Chair of Pharmacology at Northwestern University Feinberg School of Medicine. He has been a pioneer in elucidating the genetics and pathogenesis of channelopathies with a focus on genetic disorders caused by voltage-gated ion channel mutations including KCNQ2-associated epilepsy. He was the director of the Channelopathy-Associated Epilepsy Research Center without Walls funded by the National Institute of Neurological Disorders and Stroke.
Megan Abbott, M.D. is a pediatric neurologist at Children’s Hospital Colorado with clinical and research interests in neurogenetic conditions within epilepsy. In summer 2024, she became an assistant professor at the University of Colorado in the Department of Pediatrics, Division of Neurology. Her research is specifically looking at outcome measures that can be generalized across many developmental and epileptic encephalopathies including SCN2A.
Kevin J. Bender, Ph.D. is a Professor of Neurology at the University of California, San Francisco. He is a leader in studying the mechanisms by which neurons integrate information, and how these processes are disrupted in neuropsychiatric and neurodevelopmental disorders, including SCN2A-related disorders.
Andreas Brunklaus, M.D. is a consultant paediatric neurologist at the Royal Hospital for Children, Glasgow and honorary professor at the University of Glasgow. He is an international expert in sodium channel disorders including SCN2A-related disorders and leads international research collaborations developing diagnostic tools in epilepsy genetics.
Scott Demarest, M.D., M.S.C.S. is an Associate Professor in the Department of Pediatrics, Division of Neurology at the University of Colorado. His clinical practice and research focus on the evaluation and treatment of neurogenetic epilepsies including clinical trials for novel therapeutics, natural history studies and the development of improved outcome measures. He is the Clinical Director of Precision Medicine Institute at Children's Hospital Colorado where he supports diagnostic and therapeutic approaches to improve patient outcomes.
Shawn Egan, Ph.D. is a parent of a child affected by an SCN2A-related disorder, and he serves as Chief Scientific Officer for the FamilieSCN2A Foundation. In this role, Shawn focuses on building the necessary infrastructure to expedite the development of treatments and cures for all patients with SCN2A-related disorders. This includes empowering the SCN2A community and providing support for SCN2A-related research and drug development.
Isabel Haviland, M.D. is a research fellow in neurology at Boston Children’s Hospital and Harvard Medical School. She is focused on clinical research exploring genotype-phenotype correlations in pediatric genetic epilepsies.
Jennifer A. Kearney, Ph.D. is an Associate Professor of Pharmacology at Northwestern University Feinberg School of Medicine, the Center for Genetic Medicine and the Center for Autism and Neurodevelopment. Her research program is focused on identifying genetic factors that contribute to childhood epilepsies and neurodevelopmental disorders. Her research has contributed to understanding the genetic basis and impact of genetic modifiers on SCN2A-realted disorders using genetically engineered mouse models.
Leah Schust Myers, M.S. is the founder and Executive Director for the FamilieSCN2A Foundation. She has spent her entire career working in health care administration and never imagined she would find a use for her skills in an entirely different way. From medical secretary to hospital manager and everywhere in between, Leah learned how to manage the needs of large populations within a medical setting. When her son was diagnosed with an SCN2A-related disorder in 2012, it became abundantly clear how to leverage her experience to help others.
Heather E. Olson, M.D. is an Assistant Professor in the Department of Neurology, Division of Epilepsy and Clinical Neurophysiology at Boston Children’s Hospital. She is a pediatric neurologist with expertise in epilepsy genetics and fetal/neonatal neurology. She directs a clinical research program focusing on early-onset genetic developmental and epileptic encephalopathies, including CDKL5 Deficiency Disorder and SCN2A-related disorders.
Stephan J. Sanders, BMBS, Ph.D. is a Professor of Paediatric Neurogenetics at the University of Oxford and Associate Professor at the University of California, San Francisco. He leads a group that identifies the genetic causes of neurodevelopmental disorders and seeks to understand the consequent neurobiology and identify genome-targeted therapies. His research demonstrated that de novo SCN2A mutations are associated with autism spectrum disorders and the differing phenotypes of gain- and loss-of function variants.
Christina SanInocencio, Ph.D. is an Assistant Professor of communication at Fairfield University and adjunct professor of health communication at Stony Brook University. She has been a leader in rare epilepsy advocacy for nearly two decades and serves as a member of TASCO (team for advancing science and clinical outcomes) for FamilieSCN2A Foundation.
Joseph Symonds, M.B., Ch.B., Ph.D. is a Senior Clinical Research Fellow at the School of Health & Wellbeing at the University of Glasgow. He is a Consultant Paediatric Neurologist at the Royal Hospital for Children, Glasgow where is delivers epilepsy care for children and young people and runs a genetic epilepsy clinic. His research takes an epidemiological approach to understanding the frequency of rare genetic epilepsies, and defining the true phenotypic spectrum of each genetic epilepsy.
Christopher H. Thompson, Ph.D. is a Research Assistant Professor of Pharmacology at Northwestern Feinberg School of Medicine. His work has contributed to the elucidation of functional consequences of pathogenic variants associated with SCN2A-associated neurodevelopmental disorders, as well as pharmacology and regulation of voltage-gated sodium channels.
Annapurna H. Poduri
Boston Children’s Hospital and Harvard Medical School
Annapurna H. Poduri, MD, MPH is Associate Professor of Neurology at Harvard Medical School. She is Director of the Epilepsy Genetics Program at Boston Children’s Hospital, which focuses on the discovery of germline and mosaic variants in patients with epilepsy, and modeling epilepsy genes in zebrafish and cell-based models.
Alfred L. George, Jr.
Northwestern University Feinberg School of Medicine
Alfred L. George, Jr., MD, is Professor and Chair of Pharmacology at the Northwestern University Feinberg School of Medicine. He directs the Channelopathy-associated Epilepsy Research Center without Walls, supported by the National Institutes of Neurological Disorders and Stroke, connecting patient variants with bench science.
Erin L. Heinzen
University of North Carolina, Chapel Hill
Erin L. Heinzen, PharmD, PhD, is Associate Professor of Pharmacy and Genetics at the University of North Carolina at Chapel Hill. She has served as Principal Investigator of the Sequencing, Biostatistics and Bioinformatics Core of the Epi4 K Consortium and investigates somatic mosaic mutation in epilepsy and the mechanisms underlying SLC35A2 epilepsy.
Associate Editor
Sara James
KCNQ2 Cure Alliance and Genetic Epilepsy Team Australia
Sara James is a renowned broadcast journalist and an advocate for research in epilepsy and community–academic partnerships in genetic epilepsy. She is Vice President of the KCNQ2 Cure Alliance and a co-founder of Genetic Epilepsy Team Australia.
About the Series
Recent advances in epilepsy genetics are actively revealing numerous genetic contributors to epilepsy, both inherited and noninherited, collectively accounting for a substantial portion of otherwise unexplained epilepsies. This series integrates clinical epilepsy genetics and laboratory research, driving the field towards more precise and effective treatments.