Introduction
Vancomycin-resistant enterococci (VRE) is a generic term for enterococci that have acquired resistance to vancomycin. The first reported case of VRE was in the United Kingdom in 1988 [Reference Uttley1]. VRE causes various healthcare-associated infections, including urinary tract infections, cholangitis, cholecystitis, and infective endocarditis, despite their relatively low virulence [Reference O’Driscoll and Crank2]. Around 10% of colonized patients develop a clinical infection within 30 days [Reference Willems3]. VRE bacteraemia increases mortality 1.8–2.5 times compared to vancomycin-susceptible strains [Reference DiazGranados4, Reference Prematunge5]. Outbreaks of VRE are often protracted and require intensive, multifaceted infection control interventions, resulting in a substantial economic burden [Reference van der Pol6]. A Canadian study reported that VRE infections incurred an average additional cost of $17949 and prolonged hospital stays by 13.8 days per patient [Reference Lloyd-Smith7].
Given these important clinical and economic consequences, preventing nosocomial transmission of VRE is of paramount importance. Effective control strategies require identification of patients with active VRE infection and those who are colonized with VRE. Screening and isolating VRE carriers is a cost-effective infection prevention and control intervention [Reference Mac8–Reference Weterings11]; however, the optimal timing, target population, and duration of VRE screening remain unclear. This study aimed to evaluate the effectiveness of multiple screening strategies for VRE outbreak control and to identify risk factors for VRE acquisition during the initial hospital-wide screening.
Methods
Study setting and periods
In this retrospective study, we analysed the overall course of a VRE outbreak and subsequent ongoing screening tests conducted at Fujisawa City Hospital, a 536-bed tertiary care centre in Japan, between May 2010 and March 2021. The hospital features six dedicated infectious disease beds, with average patient length of stay of 13.2 days.
Infection control measures
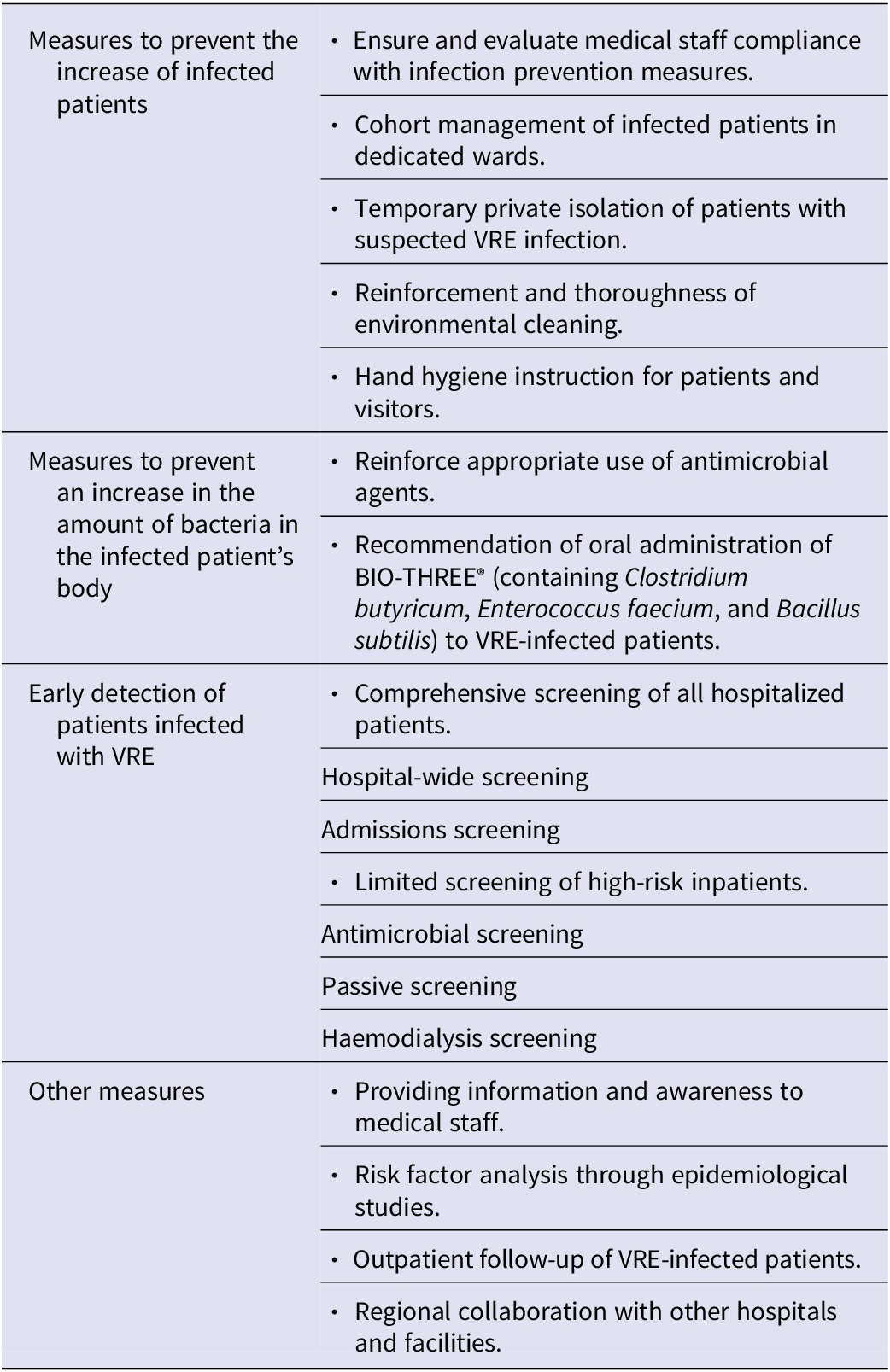
To implement VRE infection control measures, we referenced several guidelines developed in other countries [Reference Muto12–Reference Siegel14], which include antimicrobial stewardship, screening tests, patient isolation, environmental cleaning, and adherence to contact infection control measures. Based on these references, we implemented various measures to prevent the spread of VRE (Table 1). These measures aimed to prevent an increase in the number of infected patients, reduce the bacterial burden via antimicrobial stewardship, and promote early detection of VRE colonization or infection using various screening tests. We conducted a case – control study to analyse the risk factors approximately 7 months after we identified the first case. We primarily followed five screening strategies: hospital-wide, admission, antimicrobial, passive, and haemodialysis screening. Admission screening was performed for all patients upon admission. Antimicrobial screening was performed among patients who had received glycopeptides, third- and fourth-generation cephalosporins, beta-lactamase inhibitor combinations, carbapenems, aminoglycosides, or clindamycin for ≥7 days [Reference Farhadi15, Reference Gouliouris16]. Passive screening is an automated screening method to detect VRE when routine stool culture is submitted to identify patients not captured in antimicrobial screening. Haemodialysis screening was performed every 2 weeks in patients undergoing haemodialysis during hospitalization. In addition to the five primary screening tests, we performed routine screening of wards with VRE-positive patients and screening upon transfer of these patients to other wards. Furthermore, to promptly and effectively control the outbreak, we organized meetings to discuss countermeasures that included external experts in microbiology, infection control, infectious disease treatment, medical epidemiology, and public health.
Measures to control the spread of VRE infection

Abbreviation: VRE, vancomycin-resistant enterococci.
Microbiologic testing
Rectal swabs were collected from patients by nurses using BD BBL CultureSwab Plus™ (Nippon Becton Dickinson Co. Ltd., Tokyo, Japan) for VRE screening. The swabs were directly inoculated onto BD VRE Selective Agar (Nippon Becton Dickinson Co. Ltd., Tokyo, Japan). The agar plates were divided into two sections. A seed swab was directly applied to one section and streaked using a 1-μL inoculating loop. The agar plates were incubated aerobically at 35 C for 48 h, with colony observation performed every 24 h. If suspected colonies were observed, Gram staining was performed. Gram-positive cocci in short chains were considered presumptive VRE and were sub-cultured onto blood agar for pure culture isolation. Colonies grown in pure culture were subjected to catalase and PYR tests (SwabColor PYR; Iwaki & Co., Ltd., Tokyo, Japan). Isolates exhibiting a negative catalase test and a positive PYR test were presumptively identified as Enterococcus species. Species identification and antimicrobial susceptibility testing were then performed using the MicroScan WalkAway 40 system (Siemens Healthineers, Erlangen, Germany) with the Prompt method for bacterial suspension preparation. The MicroScan WalkAway Pos Combo 3.1 J Panel was used to determine minimum inhibitory concentration values. VRE was confirmed if the minimum inhibitory concentration for vancomycin was >16 μg/mL for Enterococcus species.
Detection of vancomycin resistance genes was performed using primers described by Dutka-Malen et al. [Reference Dutka-Malen, Evers and Courvalin17].
Molecular typing of VRE isolates
To investigate the genetic relatedness among the VRE isolates, molecular typing was performed using pulsed-field gel electrophoresis (PFGE). Agarose gel plugs containing bacterial cells were prepared and digested with the restriction enzyme SmaI (BioLabs, Ipswich, MA, USA). Electrophoresis was conducted using the CHEF DRII system (Bio-Rad Laboratories, Hercules, CA, USA) under the following conditions: 6.0 V/cm at 14 C. Phylogenetic trees were constructed based on the banding patterns using Fingerprinting II software (Bio-Rad Laboratories). Isolates showing PFGE patterns with a difference of up to three-fragment difference were defined as the same clone [Reference Tenover18].
Case–control study
To identify risk factors associated with VRE acquisition, a retrospective case–control study was conducted using VRE-positive cases identified during the initial hospital-wide screening conducted in December 2010 (n = 23). We retrospectively collected patient data from electronic medical records, including age, sex, pre-admission residence, activities of daily living, presence or absence of enteral feeding, dependency on assistance with hygiene and toileting, presence of an indwelling urinary catheter, diaper use, oral care practices, airway suctioning requirements, history of intensive care unit admission, medical procedures performed, use of medical devices, and prior exposure to antibiotics. Age was analysed as a continuous variable. Prior antibiotic exposure was treated as a categorical variable, defined as exposure to specific antimicrobial agents within the preceding 3 months. Sensitivity analyses were not performed because all eligible cases and controls from the initial hospital-wide screening were included, leaving no scope for alternative analytic scenarios.
This retrospective case–control cohort was defined by the initial hospital-wide screening at the start of the outbreak. Patients for whom consent was not obtained were excluded. We included every VRE-positive patient detected at that screening (n = 23) and all other inpatients across all wards who underwent the same screening round and tested negative for VRE (n = 376). All inpatients were screened during this round, and no patients had missing core clinical data. Because the cohort was fixed by this screening round rather than by a target sample size, no a priori sample size calculation was performed. This approach ensured that all available cases were included, thereby maximizing statistical power and minimizing selection bias.
Statistical analysis
All statistical analyses were conducted using IBM SPSS Statistics 29.0 software (IBM Corp., Armonk, NY, USA). Risk factor analysis was performed using the chi-square test, and a p-value of <0.05 was considered statistically significant. In the case–control study, univariate analysis was performed to statistically assess various potential risk factors hypothesized associated with VRE acquisition. Multivariable logistic regression was not performed due to the limited number of cases (n = 23).
Ethical considerations
Management and infection control interventions for outbreaks were conducted as part of normal infection control programs. This study was approved by the ethical review committee of Fujisawa City Hospital (approval number F2023003). To ensure participant autonomy, an opt-out option was provided.
Results
Outbreak overview
Table 2 summarizes the timeline of implemented infection control measures and important events. In May 2010, VRE was detected in the bile of a 72-year-old male patient who was hospitalized with acute cholangitis, marking the index case in this outbreak. Standard precautions, including hand hygiene, were promptly reinforced, and thorough environmental cleaning was implemented. Specifically, to interrupt indirect contact transmission via contaminated environmental surfaces, we focused on disinfecting frequently touched surfaces, such as shared toilets, bed rails, bedside tables, and handrails. To ensure the quality of cleaning, a head nurse with infection control nurse certification conducted checks on cleaning procedures and provided feedback to cleaning staff and/or contractors. Simultaneously, screening of all patients in the same ward revealed VRE carriage in an additional 13 patients, all of whom were promptly placed in single rooms or cohort isolation. Given the continued increase in VRE-positive cases, a task force meeting with external experts was convened in December 2010. Following this meeting, multiple screening tests were sequentially introduced. By October 2011, all VRE-positive inpatients were discharged.
Overview of events related to the VRE outbreak and the implemented infection control measures and the number of VRE positive patients

Abbreviation: VRE, vancomycin-resistant enterococci.
Over the approximate 10-year period from May 2010 to March 2021, a total of 169 cases of VRE were detected, comprising seven symptomatic patients and 162 asymptomatic carriers identified through screening. Figure 1 displays the epidemiologic curve alongside screening tests performed. The study period was divided into four distinct phases: May 2010–October 2011 (phase 1), October 2011–September 2013 (phase 2), September 2013–December 2017 (phase 3), and January 2018–March 2021 (phase 4). Phase 1 was defined as the period from initial outbreak recognition to the point when all VRE-positive inpatients were discharged. Phase 2 was defined as the period during which efforts were made to identify potential VRE carriers by continuing various screening procedures initiated in phase 1. Phase 3 was defined as the period when admission screening was limited to high-risk patients. High-risk patients were defined as those with a history of VRE detection, diaper use, inter-facility transfer, or pre-admission antibiotic use for ≥7 days. Phase 4 was defined as the period between the reduction in hospital-wide screening owing to two consecutive instances with no newly detected cases, and March 2021 when all VRE screening tests were discontinued owing to no positive results in the remaining admission, antimicrobial, and passive screening.
Epidemiologic curve showing the number of patients with newly detected vancomycin-resistant enterococci (VRE; solid line) and number of screening tests performed (dashed line). The transient sharp increases observed in the number of screenings correspond to the periodic implementation of hospital-wide screening strategies. No additional VRE cases were detected after March 2017, and hospital-wide screening was discontinued in January 2018 following a 9-month period with no newly identified cases.

Risk factor analysis
Table 3 presents the results of a case–control study conducted during the initial hospital-wide screening (VRE positive n = 23, VRE negative n = 376). Oral care practices, haemodialysis, and prior exposure to carbapenems, glycopeptides, and piperacillin/tazobactam were identified as significant risk factors for VRE acquisition.
Comparison of risk factors between patients with VRE and a control group

Abbreviations: ABPC, ampicillin; ABPC/SBT, ampicillin/sulbactam; AG, aminoglycosides; CEZ, cefazolin; CFPM, cefepime; CLDM, clindamycin; CMZ, cefmetazole; CPFX, ciprofloxacin; CTM, cefotiam; CTRX, ceftriaxone; CTX, cefotaxime; 95%CI, 95% confidence interval; GP, glycopeptides; ICU, intensive care unit; LVFX, levofloxacin; MEPM, meropenem; PIPC/TAZ, piperacillin/tazobactam; RR, relative risk; VRE, vancomycin-resistant enterococci.
Screening tests
Table 4 shows positivity rates for the five screening strategies implemented in each phase of the study period. Hospital-wide screening was performed a total of 32 times until Phase 3, with a positivity rate of 43/12,534 (0.34%). Admission screening was narrowed to high-risk individuals from Phase 3 onward, with a final positivity rate of 46/46,516 (0.10%). The positivity rates for antibiotic screening, passive screening, and dialysis patient screening were 31/9,230 (0.34%), 5/5,846 (0.09%), and 1/566 (0.18%), respectively. Hospital-wide screening and antibiotic screening had the highest detection rates. Of the 162 patients with positive results on screening, 46 (28.4%) were detected in admission screening, 43 (26.5%) in hospital-wide screening, and 31 (19.1%) in antimicrobial screening.
Trends in new VRE detection rates by screening type for each of the four phases

Abbreviation: VRE, vancomycin-resistant enterococci.
Phylogenetic tree analysis
To understand the relatedness of Enterococcus isolates, we performed PFGE and phylogenetic analysis on 106 VRE isolates from Phase 1 to 3 from the index case’s ward toilets (Figure 2). All isolates were identified as Enterococcus faecium. The results showed that 49 of 106 patient isolates (46.2%) shared the same PFGE pattern (group a), suggesting a high likelihood of horizontal transmission. Twelve additional minor groups (b through m) likely represented single strains. Notably, three group c isolates had a PFGE pattern identical to that of isolates from the toilet environment, indicating potential indirect transmission via the toilet environment. In contrast, 26 isolates (24.5%) displayed unique PFGE patterns. Overall, 39 distinct VRE lineages were identified during Phase 1. The vanA and vanB genes were detected in 96 isolates (90.6%) and nine isolates (8.5%), respectively, while one isolate could not be genotyped. All isolates belonging to clonal groups a through m, which represented multiple identical strains, carried the vanA gene.
Molecular phylogenetic tree constructed from 109 vancomycin-resistant enterococci (106 clinical, three from toilet facilities) using pulsed-field gel electrophoresis (PFGE). Isolates with identical PFGE banding patterns were grouped and designated as groups a through m. Within group c (n = 6), three isolates originated from the toilet environment. In contrast, 26 isolates exhibited unique PFGE patterns, indicating that a total of 39 distinct PFGE types were identified across the isolates analysed.

Discussion
We reported the characteristics of a large VRE outbreak involving over 100 cases, which was among the largest outbreaks reported in Japan at the time of its occurrence. VRE is recognized globally as a major healthcare threat, listed alongside methicillin-resistant Staphylococcus aureus, carbapenem-resistant Enterobacterales, and multidrug-resistant Pseudomonas aeruginosa by the United States (U.S.) Centers for Disease Control and Prevention [19]. In the U.S., VRE accounts for an estimated 5400 deaths annually and ranks among the leading pathogens responsible for healthcare-associated infections [20]. In Europe, surveillance data from the European Centre for Disease Prevention and Control indicate that the proportion of VRE has increased in several EU/EEA countries in recent years, although substantial inter-country variability persists [21, 22]. In 2010, resistance rates were 4.9% in China [Reference Zhao23] and 24.9% in Taiwan [Reference Wang24]; recent data (2023) from the national Japan Nosocomial Infections Surveillance database showed a relatively low rate of 1.9% for E. faecium in Japan [25]. Despite this low prevalence, outbreaks such as that reported herein can place a large burden on healthcare systems and raise serious public health concerns.
Our retrospective case–control study, based on hospital-wide screening approximately 7 months after the index case, identified several risk factors for VRE colonization, including haemodialysis, oral care practices, and prior antimicrobial exposure. Consistent with U.S. studies [Reference Fridkin26, Reference Bae27], we found strong associations between VRE carriage and exposure to vancomycin, carbapenems, glycopeptides, and piperacillin/tazobactam, likely owing to selective pressure and disruption of intestinal microbiota [Reference Donskey28–Reference Karki31]. Patients with such antimicrobial histories consistently showed higher positivity rates (0.34%) across all outbreak phases, supporting their inclusion as a high-risk screening target.
Hospital-wide screening had a high positivity rate in Phase 1 (0.91%), which declined to 0.26% in Phase 2 and 0.10% in Phase 3, indicating reduced cost-effectiveness. Accordingly, this broad strategy was discontinued after two consecutive rounds without new VRE detections. Haemodialysis patients had a five-fold increased risk of VRE carriage, likely due to immunosuppression and frequent healthcare contact [Reference Shaghaghian32, Reference Zacharioudakis33]. Positivity remained low (0.18%) whilst intensive interventions were in place, including retraining in personal protective equipment use, enhanced disinfection, and biweekly screening. Nonetheless, the vulnerability of this population justified their continued inclusion in targeted screening.
Early detection of asymptomatic carriers is crucial because VRE can be acquired during hospitalization owing to antibiotic pressure and nosocomial transmission. Although admission screening is ideal, it has limitations; only 28.4% (46/162) of VRE-positive individuals in our study were identified at admission (Table 4). Most were detected through subsequent in-hospital screening. Given the limited sensitivity of a single rectal swab (56–70%) [Reference D’Agata34–Reference Brasg36], repeated and multifaceted screening—particularly in high-risk patients – is essential to avoid missed carriers.
Screening methodology should also be considered when interpreting the declining positivity. Rectal swabs were cultured by direct plating onto selective agar, reflecting routine microbiological practice during the early outbreak period. Culture-based screening may be less sensitive than liquid-based or molecular methods, particularly for detecting low-density gastrointestinal colonization [Reference Ledeboer and Dallas37]. However, the same methodology was applied consistently throughout all outbreak phases, allowing internally consistent longitudinal comparisons. Moreover, the decline in screening positivity coincided with the disappearance of new clinical VRE infections and de-escalation of outbreak control measures, suggesting genuine epidemiological improvement rather than solely methodological under-detection. Several international frameworks, including guidance from the European Centre for Disease Prevention and Control, recommend risk-based screening and intensified surveillance during VRE outbreaks [38–40]. However, the optimal screening intensity and surveillance duration remain uncertain [Reference Isenman, Michaels and Fisher41]. A meta-analysis suggested that cost-effectiveness depends on local variables such as prevalence, resource utilization, and infection control practices [Reference Macdougall42]. In our setting, new VRE cases emerged intermittently over 7 years, likely influenced by community reservoirs. Even a single case in low-prevalence countries like Japan warrants robust control measures. Given the potential for long-term intestinal carriage and ongoing community transmission, extended surveillance is advisable.
Unlike typical VRE outbreaks involving clonal expansion, we identified 39 genetically distinct strains via PFGE analysis of 106 isolates from infected patients. Although these strains may have originated from a common ancestor, subsequent genetic divergence likely contributed to the observed heterogeneity. Supporting this, a study at a healthcare facility in London reported the co-circulation of 45 distinct VRE strains within a single institution [Reference Uttley43]. These findings should also be interpreted in light of the molecular typing method used in this study. Molecular typing was performed using PFGE; however, PFGE has lower discriminatory resolution than whole-genome sequencing (WGS) and cannot fully resolve fine-scale transmission events. Future studies using WGS may therefore provide more detailed insights into the genetic relatedness and transmission dynamics of VRE strains. Given that enterococci can asymptomatically colonize the gastrointestinal tract, it is plausible that multiple VRE strains were already present in the community before the outbreak was recognized. Supporting this possibility, a comparative analysis of antimicrobial-resistant bacteria in hospital versus municipal wastewater demonstrated higher concentrations of extended-spectrum β-lactamase-producing organisms and carbapenemase-producing Enterobacteriaceae in hospital effluent, whereas VRE levels were similar between hospital and municipal wastewater [Reference Hassoun-Kheir44]. These findings suggest the potential for widespread VRE dissemination in the community. Although wastewater sampling was not performed in this study, previous wastewater surveillance studies suggest that VRE may circulate beyond healthcare settings [Reference Geissler45]. E. faecium carrying vanA or vanB determinants has been detected in municipal wastewater and surface water, indicating environmental reservoirs of glycopeptide resistance. Genetic relatedness between wastewater and clinical isolates has also been reported [Reference Zhang46], raising the possibility of bidirectional transmission between healthcare facilities and the community [Reference Furukawa47]. Consequently, a comprehensive strategy involving universal screening of all admitted patients and enhanced environmental cleaning and disinfection – rather than targeted screening and infection control alone – proved effective in mitigating VRE acquisition within our facility. Early implementation of broad-based screening during VRE outbreaks may therefore help identify previously unrecognized carriers.
The study period ended in early 2021 during the early phase of the COVID-19 pandemic. Recent surveillance reports suggest increasing VRE incidence in some regions in the post-pandemic period [Reference Lagadinou48]. Changes in antimicrobial use, infection control practices, hospital occupancy, and patient movement during the pandemic may have influenced VRE epidemiology. Conversely, reductions in elective admissions and strengthened infection control measures may have temporarily reduced transmission opportunities. Therefore, the epidemiology observed in this study may not fully reflect post-pandemic trends.
This study has several limitations. First, due to the limited number of VRE-positive cases in our case–control study, we were unable to perform a multivariable analysis to adjust for potential confounders. Future research with larger sample sizes is needed to explore these relationships more thoroughly. Second, we relied on culture-based detection methods, which may be less sensitive than emerging molecular assays. Future studies should evaluate the cost-effectiveness of molecular techniques. Third, optimal screening strategies vary by region; our findings are most applicable to settings with low VRE prevalence and should be interpreted accordingly.
Conclusion
Our findings demonstrate that no single screening strategy is universally optimal for VRE outbreak control. Instead, early implementation of broad-based screening is crucial to identify the true extent of transmission, particularly in low-prevalence settings where undetected colonization may sustain prolonged outbreaks. Subsequent dynamic, data-driven refinement of screening strategies based on positivity rates and local epidemiology enables more efficient allocation of infection control resources while maintaining effective containment. This adaptive approach provides a practical framework for the sustainable management of prolonged VRE outbreaks in healthcare settings.
Data availability statement
The data presented are available upon reasonable request from the corresponding author after publication.
Acknowledgements
We would like to extend our thanks to Shuhei Fujimoto, Norio Omagari, Rumiko Komaba, Makiko Sugawa, Hanako Misao, and Toshihiro Mitsuda for their contribution as external experts. We thank Analisa Avila, MPH, ELS, of Edanz (https://jp.edanz.com/ac) for editing a draft of this manuscript.
Author contribution
Conceptualization: H.S., T.K., T.O., M.S., N.Y., A.S., M.N.; Data Curation: H.S.; Formal Analysis: H.S.; Investigation: H.S., T.K., T.O., M.S., N.Y., A.S., M.N.; Methodology: H.S., T.K., T.O.; Supervision: M.S., N.Y., A.S., M.N.; Validation: T.K., T.O.; Writing – Original Draft: H.S.; Writing – Review and Editing: H.S.
All authors approved the final manuscript as submitted and agree to be accountable for all aspects of the work.
Competing interests
The authors declare no conflicts of interest.