1.1 Antibody Structure and Function 2
1.2 Tissue Preservation and Antigen Retrieval 2
1.3 Methods of Antigen Detection 3
1.4 Standardization and Quality Control 5
1.5 Immunohistochemistry in the Detection of Molecular Abnormality 6
1.6 Chromogenic In Situ Hybridization 9
1.7 Dual Chromogenic Immunohistochemistry 9
1.8 Multiplex Immunohistochemistry and Immunofluorescence 9
1.9 Future Direction 9
Introduction
Immunohistochemistry is an interdisciplinary set of scientific techniques used for the visualization of tissue-level and cellular architecture. It exists at the intersection of the fields of immunology, histology, and chemistry. Immunohistochemical staining plays a critical role in clinical diagnostic pathology. Seeing the dispersion of various proteins and other cellular components in a tissue sample allows pathologists to render or corroborate a clinical diagnosis. In order to visualize the localization of cellular components, the field exploits the natural ability of antibody proteins from the immune system to bind to a specific antigen. A parallel field, immunocytochemistry, involves the same set of techniques applied to cells, not full tissue sections. However, the two terms are sometimes used interchangeably by researchers.
In the early 1940s, scientists at Harvard Medical School, led by Dr. Albert Coons, demonstrated that antibodies that were labeled with a fluorescent tag maintained specificity for their antigen.Reference Coons, Creech and Jones1 Building on this discovery, scientists then incubated tissues with these fluorescently labeled antibodies and examined the tissues for the label under fluorescence microscopy. The localization of antibodies, and thus their tags, to the antigens allowed them to directly determine the presence and location of antigens within the tissue samples.
Over the last few decades, the techniques used in immunohistochemical staining have become exponentially more complex. Several milestones have accelerated the growth of the field of immunohistochemistry with the first one being the introduction of enzyme-labeled antibodies that allow visualization of targeted antigens by bright-field microscopy.Reference Nakane2 The production of monoclonal antibodies with greater specificity results in countless types of antibodies, tags, and reagents available for use.Reference Kohler and Milstein3 Signal strength can be improved with antigen retrieval methods that enhance the ability of antibodies to bind to epitopes in formalin-fixed paraffin-embedded tissues.Reference Shi, Key and Kalra4 Archival tissue samples can be stained, allowing for potential retrospective research. Other recent developments include sensitive secondary detection systems,Reference Hsu, Raine and Fanger5 automated staining systems,Reference Moreau, Le Neel, Joubert, Truchaud and Laboisse6 and digital imaging analysis.Reference Feldman7 The strength of the signal can be further amplified using indirect epitope detection, in which tagged secondary antibodies that are specific against the untagged primary antibody are applied to the tissue. Improvements in consistency, reproducibility, and stain quality, made possible with automated systems and image analysis technologies, allow for accurate quantification of staining in tissue samples. All of these developments in turn advance clinical diagnoses and propel research.
As the complexity of immunohistochemical techniques has progressed, so has their utility. The exploitation of the specificity of these interactions enables us to detect molecules of interest in tissue samples through immunohistochemical methods. In addition to being adjunct diagnostic tools, which are discussed in subsequent chapters, immunohistochemical markers can now be used as screening tests for hereditary syndromes (see Chapter 10). In the era of personalized medicine, biomarker expression can guide risk stratification and therapeutic decision. The association of these molecules with certain disease presentations allows us to use information gathered through staining as biomarkers for diseases. Thus, an entire line of study within the field of immunohistochemistry involves the discovery of these biomarkers and their clinical relevance.Reference Howat and Wilson8 Currently, several biomarkers are approved by the Food and Drug Administration (FDA) for diagnostic use in cancer, including Her2, estrogen receptor, epidermal growth factor receptor, and PDL1.Reference O’Hurley, Sjostedt and Rahman9 An ever-growing body of literature catalogues the relationships between expressions of various proteins and diseases. The presence of these biomarkers within a tumor can be used by the medical team to guide and personalize treatment.Reference Howat and Wilson8
Immunohistochemical techniques are growing progressively more sensitive and specific. Our understanding and manipulation of antibody-epitope interactions have allowed immunohistochemistry and other related technologies to grow into fields of their own, including immunofluorescence (IF) (see Chapter 11) and in situ hybridization (ISH) using both deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) probes. Antibodies have grown into powerful clinical and research tools. This chapter will describe the conceptual framework of immunohistochemistry, outline technical mechanisms, and explain its clinical relevance.
1.1 Antibody Structure and Function
Antibodies, or immunoglobulins, are polypeptide structures that are involved in humoral immune response. They are clonally produced in the body by B-lymphocytes. Antibodies recognize and bind with high specificity to another molecule, called an antigen. Often, this antigen is a region of a protein comprised of five to six amino acid residues, called an epitope. Antigens can be homopolymeric or multivalent with multiple identical epitopes or heteropolymeric with multiple distinct epitopes.Reference Lipman, Jackson and Trudel10 In the body, antibodies might bind to a virus to prevent it from infecting a cell. They also might form an agglutination reaction, causing a clump of antigens to form, which can then be eliminated by the immune system.
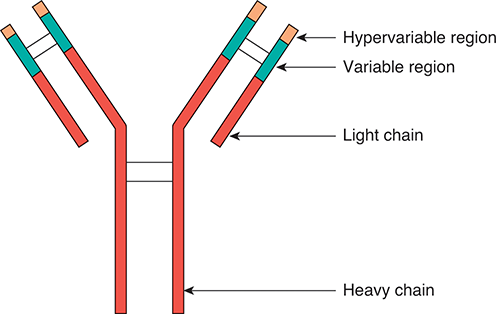
Antibodies are Y-shaped polypeptide heterotetramers composed of two “heavy” chains and two “light” chains. The quaternary structure is maintained by disulfide linkages in cysteine-rich moieties of the polypeptide chains. The structure can be further divided into the constant, or fixed, region (Fc) and the antigen-binding domain (Fab). The antigen-binding domain contains a variable region and a hypervariable region, which is on the end of the protein. It is the hypervariable region of the antigen-binding domain that determines the epitope specificity of the antibody (Figure 1.1).Reference Janeway, Travers, Walport and Shlomchik11

Figure 1.1 Structure of an antibody. An antibody protein is a heterotetramer, containing two heavy chains and two light chains. Disulfide bonds link the four polypeptides. The variable and hypervariable regions confer specificity against a certain antigen.
In the body, recombination of genomic DNA accounts for the high variability of antibody structure and, thus, the vast range of antigens against which specificity is conferred. Antibodies can be engineered to have an affinity for a countless array of antigens. Antibodies recognize epitopes based on a unique shape-size-charge profile. Antibody-antigen recognition occurs via ionic/electrostatic (non-covalent) intermolecular forces, and follows an induced-fit model.Reference Janeway, Travers, Walport and Shlomchik11 For this reason, it is critical that the structural integrity of the epitope is maintained even after the tissue has been excised. Epitopes can be linear or conformational.Reference Bogen, Vani and Sompuram12 Linear epitopes are five to seven contiguous amino acids while conformational or discontinuous epitopes are made of small groups of amino acids brought into close proximity through conformational folding or binding.Reference Janeway, Travers, Walport and Shlomchik11 While epitopes in immune responses are conformational, those in formalin-fixed paraffin-embedded tissues are invariably linear.Reference Sompuram, Vani and Hafer13 Therefore, if the antigen of interest is denatured or undergoes a conformational change, the structure of the epitope may be disrupted, and the antibody may no longer be able to bind to it.
Common sources of antibodies for clinical and research use include mouse, rat, rabbit, goat, sheep, chicken, and horse. The source of the antibody is important for indirect antigen detection. Antibodies can be produced that are specific against another species’ antibodies. This can enhance signal strength and improve technical efficiency. There are a wide variety of commercially available antibodies. Advances in molecular biology and genetic techniques have provided us with the technology necessary to engineer antibodies specific to nearly any biological macromolecule.Reference Schofield, Lewis and Austin14 Western blotting and/or immunoprecipitation are often used to determine whether or not an antibody is specific for an antigen. By engineering antibodies, specificity can be enhanced, and they can be tailored to work well in preserved tissue samples.Reference Schofield, Lewis and Austin14
Further complicating the process of antibody selection is the availability of mono- and polyclonal antibodies. Polyclonal antibodies are groups of antibodies raised against multiple epitopes of a single antigen. Monoclonal antibodies, in contrast, are raised against a single epitope of a single antigen. They are usually produced from a mouse using the technique of Kohler and Milstein.Reference Kohler and Milstein3 Rabbit monoclonal antibodies have higher affinity than mouse monoclonal antibodies, likely as a result of high glycosylation. Immunohistochemistry using polyclonal antibodies is far more sensitive. However, the use of a monoclonal antibody is a more specific process. Fittingly, different research questions can influence antibody selection. For example, if a researcher is looking simply for the presence of a protein in a cell that might exist in low concentration, it is likely advisable to use a polyclonal antibody. However, if you are looking for a specific domain of a specific protein, use of a monoclonal antibody would likely be superior. The balance between sensitivity and specificity is something that all laboratories struggle with in developing immunohistochemical assays. In addition, test volume, reagent cost, and which antibody clone to use when more than one monoclonal antibody is available are additional factors that a clinical immunohistochemical laboratory must consider.
1.2 Tissue Preservation and Antigen Retrieval
The antigenicity of a molecule, or its ability to be recognized by its antigen, is an important factor to consider in any immunohistochemical protocol. Because antigens are recognized by antibodies based on their unique properties, it is crucial that they remain structurally intact. If an antigen is present in a cell, but any of its properties (including size, shape, charge, and amino acid sequence) are compromised, the staining could fail, leading to a false negative result.Reference Khoury, Sait and Hwang15
Frequently, immunohistochemical staining is performed on archival tissue samples. Fixation stabilizes and protects the tissue from the rigorous processing and immunohistochemical staining. The most commonly used technique for tissue preservation is formalin fixation.Reference Goldstein, Hewitt and Taylor16 For over one hundred years, formalin-fixed paraffin-embedded tissues (FFPE) have been the standard tissue preparation. While the exact mechanism by which this preservation works is yet unknown, it is thought that formalin preserves tissue by inducing methylene cross-links between different protein moieties. It is very important that the preservation process is performed correctly to prevent degradation of the tissue over time. After the tissue has been fixed in formalin, it must be embedded in a wax block for storage and sectioning. The fixed tissue is first submerged in a concentrated ethanol solution. This process dehydrates the tissue. Water molecules are reactive, and their presence within a tissue could reduce the quality of preservation.Reference Xie, Chung and Ylaya17 If dehydration is incomplete, there can be a detrimental effect on the integrity of the macromolecules within the cells. Oxidation caused by any residual water in the sample can be devastating to tissue quality. Therefore, it is crucial that dehydration is completed. The ethanol is then exchanged with xylenes, then finally paraffin. The fixed tissue is then embedded in a paraffin wax and mounted onto a block. However, the addition of methylene cross-links, in conjunction with the highly hydrophobic environment created by the paraffin coating, facilitate a conformational shift in various macromolecules in the preserved tissue sample. Thus, this process maintains the integrity of the tissue structure at the cellular level over time, but it does so at the cost of molecular structural integrity. Proteins within preserved tissues can be returned to their native state, as the reactions presumed to be underlying fixation are reversible processes. Another important consideration for maintaining tissue integrity is the ischemic time, or the time elapsed between excision from the body and fixation. The longer that unpreserved tissue is cut off from the blood stream, the less likely it is that cellular architecture will be conserved, as the tissue may begin to degrade from within the paraffin block.
One of the major limitations of using IHC on preserved tissue samples is the decreased antigenicity of epitopes in fixed tissue. At some point in the fixation process, the structure of proteins within the tissue is altered. If this affects the structure of the intracellular molecular target, then the antibody will not be able to recognize and bind to it. Antibodies recognize their targets based on the unique size, shape, and charge profile of the epitope.Reference Janeway, Travers, Walport and Shlomchik11 Alteration of any of these properties diminishes the strength of the antibody-antigen interaction. In formalin-fixed, paraffin-embedded tissue, methylene cross-links induced by the fixation process and the hydrophobic environment of the wax alter protein conformation. This might mask the epitope. Alternatively, if the epitope consists of amino acids from different parts of the polypeptide that are brought together only when the protein is folded properly into its three-dimensional structure, the epitope could be completely compromised. Therefore, mitigating antigenicity loss with antigen retrieval methods is a crucial step in any immunohistochemical staining protocol.
It was demonstrated by Shi that by incubating tissue samples at high temperature prior to incubation with primary and secondary antibodies, the strength of the signal was dramatically enhanced.Reference Shi, Key and Kalra4 While the exact mechanism by which this works has yet to be determined, it is thought that antigen retrieval methods disrupt the cross-linkage between formalin and protein and restore the epitopes to their native conformations, thus enabling the antibody to bind to them with enhanced specificity. Because a macromolecule’s tertiary (three-dimensional) structure of polypeptides is environmentally sensitive, the temperature and pH at which antigen retrieval is performed will influence the efficacy of the procedure. Originally, antigen retrieval was performed by boiling formalin-fixed tissue slides in a pressure cooker prior to staining. It is believed that the high temperature of the boiling water returned epitopes to their natural state, allowing antibodies to bind in the intended manner. Later, it was demonstrated that this same enhancement of antigenicity occurred in tissues warmed in a microwave oven prior to incubation with antibodies.Reference Shi, Key and Kalra4 It is possible that the high temperature hydrolyzes the cross-links that were induced during the initial preservation process, thereby allowing the renaturation of proteins into their native states.Reference Warford, Akbar and Riberio18
Currently, there are two commonly used antigen retrieval techniques: Heat Induced Epitope Retrieval (HIER) and Protease Induced Epitope Retrieval (PIER). The latter uses an enzymatic reaction catalyzed by Proteinase K, Trypsin, Pepsin, Pronase, and others to restore antigens to their original conformations, again through an unknown mechanism. Other techniques, including incubation with peroxide, detergents, or other solvents, are less frequently used. High pH retrieval solutions are often used for antibodies with cytoplasmic or membranous staining, while low pH retrieval solutions are used for antibodies with nuclear staining. It is crucial that the conditions of antigen retrieval process be tailored to the specific antigen of interest. The variability of ideal conditions by target antigen limits the number of potential targets that can be detected on a single slide. Multiple staining is possible but must be carefully optimized.
1.3 Methods of Antigen Detection
As Coons et al. demonstrated in 1941,1 antibodies retain their properties even after the addition of a detectable tag. This allows us to directly detect the presence of antigens in tissue. However, detection of tags attached directly to antibodies binding their epitopes leads to relatively low levels of detectable staining. The staining intensity can be amplified by adding a step to the procedure. Instead of hybridizing a labeled antibody to the antigen, an unlabeled primary antibody is used. Then, after rinsing and another round of antigen retrieval, a labeled secondary antibody, specific against the primary antibody is added to the slide. Typically, these antibodies are raised in separate species. The primary antibody might be raised in a rabbit, while the secondary antibody is a mouse, anti-rabbit antibody. By adding this second layer of staining, the signal is amplified. Multiple secondary antibodies can bind to each single primary antibody. The enhanced stain quality is startlingly apparent under bright-field microscopy. Additionally, because the secondary antibody can bind to any antibody raised in the species to which it is specific, the use of this second layer can increase efficiency. When a secondary antibody is utilized, the primary antibody can be applied to the tissue in low concentrations, thereby reducing reagent consumption.Reference Brandtzaeg19
Though Coons originally labeled his antibodies with fluorescent labels, many immunohistochemical staining protocols utilize another method. The introduction of enzymes as antibody tags greatly expanded the utility of immunohistochemistry for pathology.Reference Brandtzaeg19 This method allows for the simultaneous detection of multiple cellular elements within the same tissue. In this process, the antibody is conjugated to an enzyme. The enzyme retains the ability to catalyze its reaction, while the antibody retains its specificity. Typically, the enzymes catalyze a reaction that forms a colored precipitate. The common enzymes used for this method are horseradish peroxidase and alkaline phosphatase. When staining a tissue with an enzyme-conjugated antibody, the tissue must be incubated with a chromogen reagent that reacts with the enzyme to form a colored product.
To prevent nonspecific staining and false positive results, further steps must be taken. Depending on the cell or tissue type, the enzyme that is bound to the antibody might exist in the cells. Background staining can be attributed to either nonspecific antibody staining in the setting of polyclonal antibodies or endogenous enzymes. Therefore, it is necessary to block the action of endogenous enzymes with homologous functions. Some form of peroxidase block containing hydrogen peroxide is generally used to neutralize endogenous peroxidase activity.Reference Pitts and Williams20 Depending on the tissue type of interest, there may also be endogenous immunoglobulins within the tissue. Brown et al.Reference Brown, Pemberton, Wright and Miller21 demonstrated that conjugating the antibodies that were applied experimentally to Fab fragments from antibodies of interest prevents the interaction of experimental secondary antibodies with endogenous antibodies, thereby avoiding false positive results.
Detection systems attach reporter molecules or labels to primary or secondary antibodies to visualize the antibody-antigen reaction in the tissue sections by bright-field microscopy. There have been several detection methods developed over the years with the polymer-based labeling method being currently the most widely used by the automated systems. The most common labels are enzymes such as horseradish peroxidase and alkaline phosphatase. In the presence of substrate and chromogen, enzymes produce a colored precipitate at the site of antigen-antibody reaction. 3,3’-diaminobenzidine tetrahydrochloride (DAB) is the brown chromogen for horseradish peroxidase. For alkaline phosphatase, 5-bromo-4-chloro-3-indolyl phosphate/nitro blue tetrazolium chloride (blue), fast red (red), and new fuchsin (fuchsia) are the common chromogens.
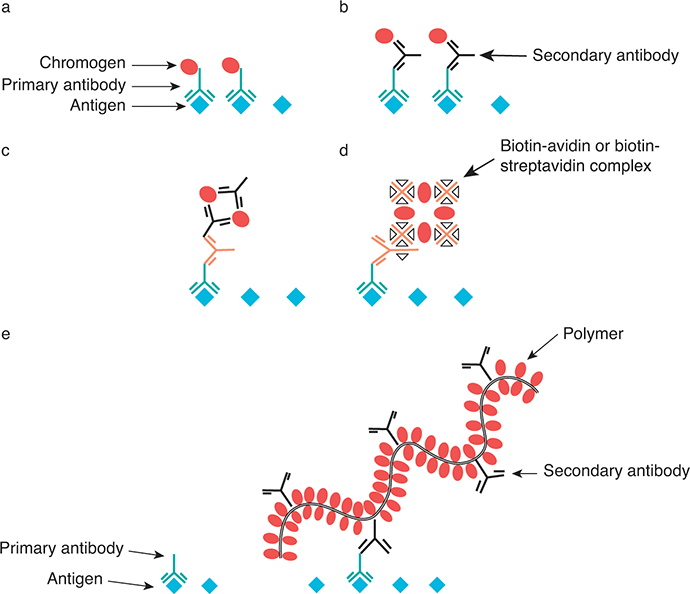
In direct method (Figure 1.2A), the label is attached directly to the antibody that targets the epitope of interest. Although rapid, this method requires a high concentration of primary antibody. The indirect or sandwich method (Figure 1.2B) allows the use of the same secondary antibody with several different primary antibodies. Since the number of labels is higher than primary antibody, this method requires less amount of primary antibody and improves the specificity of the secondary antibody. The enzyme bridge method is used mainly in research.Reference Mason, Phifer, Spicer, Swallow and Dreskin22 The peroxidase-antiperoxidase method (Figure 1.2C) is similar to the enzyme bridge method yet with hundred-fold to thousand-fold sensitivity; however, this indirect method is a laborious method.Reference Sternberger, Hardy and Cuculis23 In the biotin-avidin method (Figure 1.2D), biotin is linked to the primary antibody. Avidin is linked to peroxidase and has high affinity for biotin. The increased sensitivity is seen due to the large number of biotin molecules attached to the primary antibody.Reference Hsu, Raine and Fanger24 One disadvantage is that nonspecific or false positive staining can be seen due to endogenous biotin. Streptovidin is substituted for avidin in the biotin-streptavidin method (Figure 1.2D). The multistep includes incubation with primary antibody, reaction with biotinylated secondary antibody, binding with an enzyme (peroxidase or alkaline phosphatase) conjugated streptavidin, and detection with chromogen substrate. It has higher sensitivity than the biotin-avidin method.Reference Elias, Margiotta and Gaborc25 Alkaline phosphatase-antialkaline phosphatase method (Figure 1.2C) employs a similar principle to the peroxidase-antiperoxidase method.Reference Cordell, Falini and Erber26 This method is essential for double stain in order to avoid the cross-reactivity when two immunoperoxidase methods are used together.Reference Wagner and Worman27

Figure 1.2 Models of different detection methods in immunohistochemical staining: (A) direct method, (B) indirect method, (C) peroxidase-antiperoxidase method or alkaline phosphatase-antialkaline phosphatase method, (D) biotin-avidin method or biotin-streptavidin method, and (E) polymer labeling two-step method.
The above methods are multistep, complex, and time consuming. It is difficult to standardize and to achieve desirable sensitivity. Furthermore, recent methods including tyramine signal amplification,Reference Stack, Wang, Roman and Hoyt28 immunopolymerase chain reaction,Reference Sano, Smith and Cantor29 and end-product amplificationReference Chen, Szabolcs, Matsushima and Erlanger30 are complex and have nonspecific staining. With the availability of automated staining systems, two-step indirect polymer kits from several manufacturers including Roche/Ventana, Novovastra/Leica, Agilent/Dako, Thermo Scientific/Lab Vision, and others are widely used currently. In the polymer labeling 2-step method (Figure 1.2E) antibodies and marker systems are linked to synthetic polymer or polymerized proteins such as micropolymers, dextran, polypeptides, and DNA branches. Approximately 4–70 molecules of enzymes (peroxidase or alkaline phosphatase) and 10–20 molecules of secondary antibody are attached to a polymer.Reference Wiedorn, Goldmann, Henne, Kuhl and Vollmer31 This method increases the number of enzymes that can be coupled to the linking antibodies. The main advantages include the elimination of biotinylated secondary antibody and higher specificity.
1.4 Standardization and Quality Control
While immunostains are still done by hand in many research laboratories, automated systems are routinely used in clinical immunohistochemical laboratories. The currently available autostainers include Ventana Benchmark Ultra (Ventana Medical Systems, Tucson, Arizona), Leica Bond Max (Leica Biosystems, Buffalo Grove, Illinois), Dako Omnis (Dako North America, Carpinteria, California), and Biocare IntelliPath FLX (Biocare Medical, Concord, Massachusetts). Open systems are used in research environments since they offer flexibility in used reagents, whereas closed systems are ideal in clinical laboratories since only certain ready-to-use reagents are allowed, resulting in standardization and consistency. In an automated system with sequential processing the slides remain within the instrument throughout the staining process. On the other hand, in parallel processing, one or more steps of the staining process are performed on a separate instrument. Greater flexibility is achieved with continuous slide loading while high capacity is possible with batch slide loading. These automated systems perform antigen retrieval, antibody incubation, and chromogen application with consistency.Reference Moreau, Le Neel, Joubert, Truchaud and Laboisse6 They allow accurate, consistent, and reproducible staining by reducing human dependent errors; therefore, they have dramatically increased the efficiency in clinical and research settings and have greatly enhanced standardization.
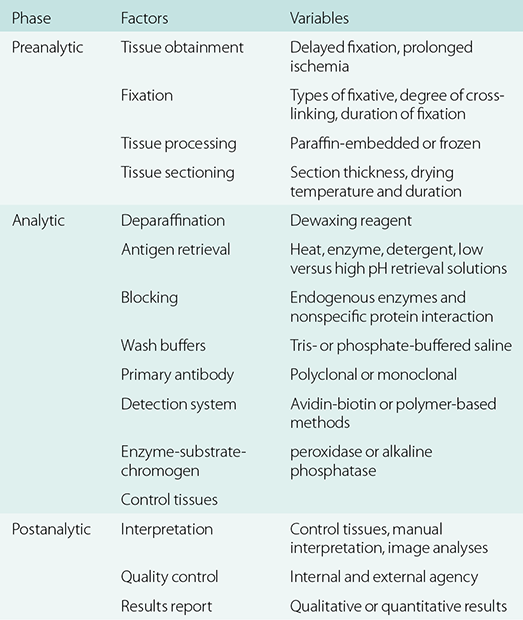
Outlined in Table 1.1 are the preanalytic, analytic, and postanalytic phases of a “total” clinical immunohistochemical test.Reference Goldstein, Hewitt and Taylor16,Reference Engel and Moore32 The fixation process is a key determinant of the outcome of immunohistochemistry with variables including types of fixative, delayed fixation, inadequate length of fixation, and over-fixation. During tissue processing, wax with a lower melting temperature (45°C) is preferred over one with a high melting temperature (65°C). Tissue sections with thickness greater than 5 micrometers often increased background staining. Clinical immunohistochemical laboratories must perform analytical validation of immunohistochemical tests prior to implementation for clinical use in accordance with the Clinical Laboratory Improvement Act of 1988.Reference Fetsch and Abati33 They must demonstrate performance characteristics of these tests including accuracy, precision/reproducibility, sensitivity, and specificity.Reference Fitzgibbons, Bradley and Fatheree34 Table 1.2 outlines the recent guidelines for validating clinical immunohistochemical assays.Reference Fitzgibbons, Bradley and Fatheree34
Table 1.1 Preanalytic, analytic, and postanalytic phases of a “total” immunohistochemical test (adapted from Engel and Moore and Goldstein et al.)Reference Goldstein, Hewitt and Taylor16, Reference Engel and Moore32
| Phase | Factors | Variables |
|---|---|---|
| Preanalytic | Tissue obtainment | Delayed fixation, prolonged ischemia |
| Fixation | Types of fixative, degree of cross-linking, duration of fixation | |
| Tissue processing | Paraffin-embedded or frozen | |
| Tissue sectioning | Section thickness, drying temperature and duration | |
| Analytic | Deparaffination | Dewaxing reagent |
| Antigen retrieval | Heat, enzyme, detergent, low versus high pH retrieval solutions | |
| Blocking | Endogenous enzymes and nonspecific protein interaction | |
| Wash buffers | Tris- or phosphate-buffered saline | |
| Primary antibody | Polyclonal or monoclonal | |
| Detection system | Avidin-biotin or polymer-based methods | |
| Enzyme-substrate-chromogen | peroxidase or alkaline phosphatase | |
| Control tissues | ||
| Postanalytic | Interpretation | Control tissues, manual interpretation, image analyses |
| Quality control | Internal and external agency | |
| Results report | Qualitative or quantitative results |
Table 1.2 Guidelines for analytic validation of clinical immunohistochemical assays (adapted from the College of American Pathologists, www.cap.org)
| Recommendation: Laboratories must validate all laboratory developed immunohistochemical tests |
| Strong recommendation: At least 90% overall concordance between the new test and expected results |
| Good practice statement: A minimum of 10 positive and 10 negative tissues for non-predictive markers |
| Good practice statement: A minimum of 20 positive and 20 negative for predictive markers |
| Good practice statement: Using validation tissues that have been fixed and processed similar to the tested clinical cases |
| Good practice statement: A minimum of 1 known positive and 1 known negative for validation of new reagent lot |
| Good practice statement: A minimum of 2 known positive and 2 known negative for validation when changing antibody dilution, antibody vendor yet same antibody clone, incubation or retrieval time yet same method |
| Good practice statement: Confirm assay performance when changing fixative type, tissue processing, antigen retrieval method, antigen detection system, and laboratory water supply |
Once an immunohistochemical test has been successfully validated, positive as well as negative control tissues must be included in every staining process, ideally on the same glass slide with the patient’s tissue sample. Nuclear staining has always been regarded as more reliable than cytoplasmic staining. Internal positive control and negative control are the ideal ways to assess for false negative staining and false positive staining, respectively.
Once the tissue samples have been stained, they can be used for diagnostic or research purposes. Depending on the stain, antigen, tissue sample, and type of information needed, the qualitative data of the stain can be interpreted differently. For certain stains and questions, knowing simply whether the sample is positive or negative for the antigen in question is sufficient. For other diagnostic purposes, a certain threshold of positive cells is necessary for definitive diagnosis. In some cases, pathologists will approximate visually the percentage of positive-staining cells. In other situations, an accurate count of positive cells is necessary. Whole slide scanning together with digital imaging analysis assist in providing consistent, reproducible, and quantitative immunohistochemical results and counting nuclear staining signals (such as Ki67, p53, estrogen receptor, and progesterone receptor).
The regulatory efforts of both the FDA and College of American Pathologists (CAP) have been focused on ensuring consistency and reliability of diagnostic tests. Participation in External Quality Assessment (EQA) and Proficiency Testing Programs is essential to maintain quality assurance. The College of American Pathologists offers a number of proficiency testing programs in the evaluation of immunohistochemical assay methodology and performance. International EQA programs designed for immunohistochemistry include Nordic Immunohistochemical Quality Control (independent scientific organization, Denmark, www.nordiqc.org) and United Kingdom National External Quality Assessment Service (UK NEQAS, Sheffield, England, www.ukneqas.org.uk).
To find the best practice in immunohistochemistry, one must know the diagnostic sensitivity and specificity of each immunostain, track the immunostain utilization data in one’s own laboratory, and replace new and better stains with the existing ones.
1.5 Immunohistochemistry in the Detection of Molecular Abnormality
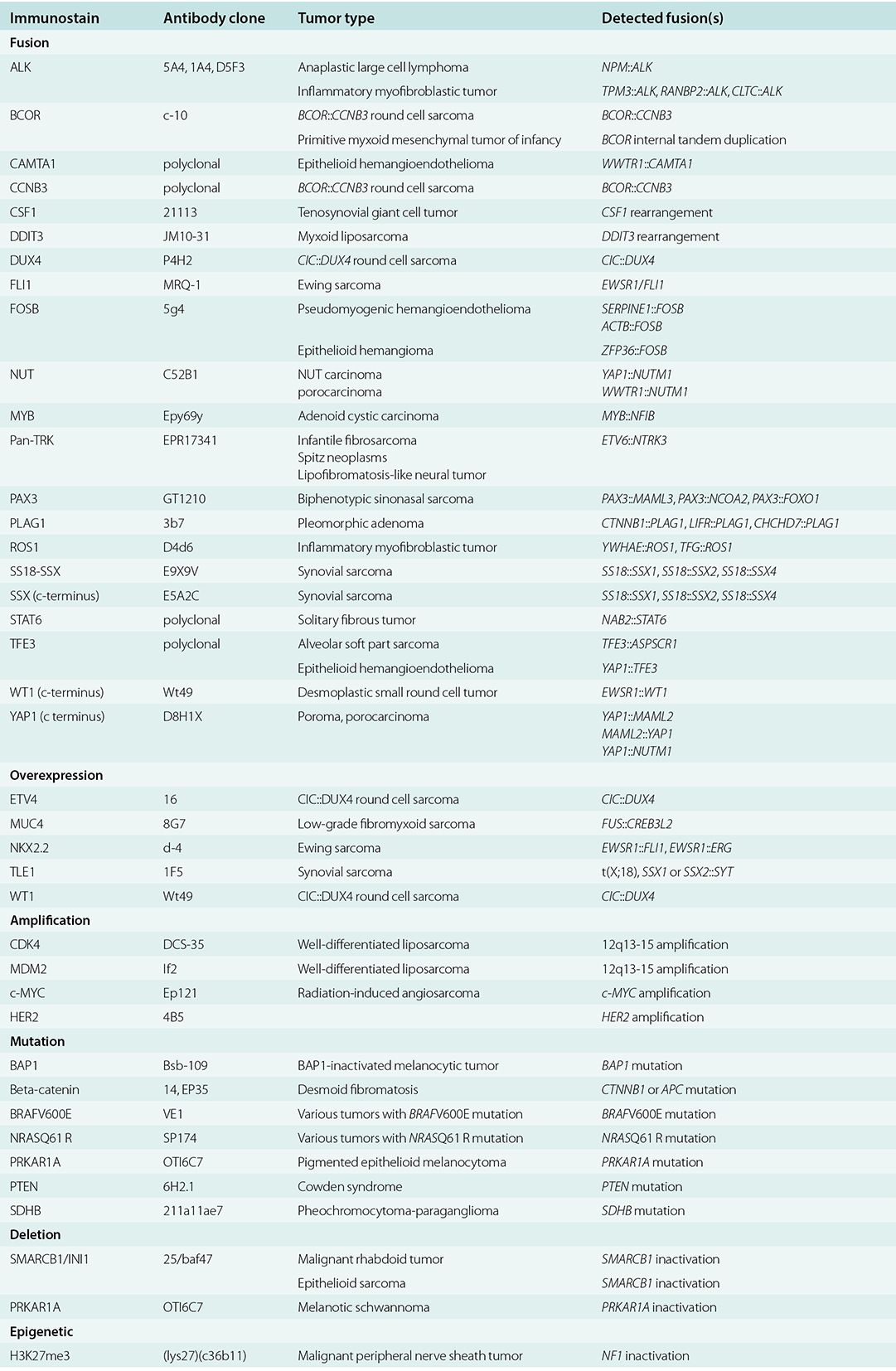
With our currently growing knowledge of tumor molecular genetics as well as the effectiveness of targeted therapy, IHC is playing an increasingly important role as diagnostic, therapeutic, and prognostic marker since it is cost-effective, rapid, and more accessible in the routine practice especially when limited tissue is available for genetic studies. IHC can currently detect the following types of molecular alterations in tumors which will be discussed in detail in the respective chapters: (1) chromosomal translocations/fusion, (2) gene overexpression, (3) gene amplification, (4) specific mutations, (5) gene deletion or loss of function, and (6) epigenetics changes (Table 1.3).
Table 1.3 Immunostains for the detection of molecular alterations
Detection of gene rearrangement can be done by different methods including IHC, fluorescence in situ hybridization (FISH), and reverse transcription-polymerase chain reaction (RT-PCR). In some cases, rearrangements involve a growing number of fusion partners, thus multiple primer sets are required for RT-PCR screening and novel rearrangements will be missed. Fluorescence in situ hybridization would require special microscope and technical expertise thus is not available in many laboratories. IHC is less laborious, cost-saving, and more widely available than FISH and RT-PCR assays. It would be a cost-effective means to screen for gene translocation which is uncommon. IHC can detect gene fusions in vascular neoplasms (CAMPTA1, FOSB, and TFE3), fibroblastic and myofibroblastic neoplasms (ALK, pan-TRK, ROS1, and STAT6), melanocytic neoplasms (ALK, pan-TRK, ROS1), and round cell tumor (BCOR, CCNB3, DUX4, and WT1). There is strong correlation between protein expression by IHC and ROS1 rearrangement detected either by FISH or next-generation sequencing (NGS).Reference Su, Goncalves, Dias-Santagata and Hoang35 Several antibody clones including ALK1 (Dako), 5A4 (Leica Microsystem), and D5F3 (Cell Signaling Technology) have been shown to exhibit high sensitivity and specificity in comparison to NGS in detecting ALK fusions.Reference Wakuda, Morita, Sekikawa, Morikawa, Miura, Doshita, Iida, Kodama, Mamesaya, Kobayashi, Ko, Ono, Kenmotsu, Naito, Murakami, Muramatsu, Kawata, Mori, Shimizu, Gon and Takahashi36
BRAFV600E and NRASQ61R immunostains can detect specific BRAF and NRAS mutations. A number of studies have demonstrated monoclonal antibody VE1 to exhibit 85–100% sensitivity and 86–100% specificity in detecting BRAFV600E mutation in melanomas.Reference Routhier, Mochel and Lynch37 Cytoplasmic expression is the typical staining pattern and background nuclear staining can be observed in some BRAF variant mutations (Figures 1.3 and 1.4).Reference Routhier, Mochel and Lynch37 In series of 171 tumors from 64 patients by Menzies et al.Reference Menzies, Lum and Wilmott38 and 230 tumors from 99 patients by Boursault et al.,Reference Boursault, Haddad and Vergier39 homogeneity of BRAFV600E expression in the primary and metastatic tumors was observed. Therefore, testing any patient’s samples with adequate viable tumor would be sufficient for determining BRAF mutation status.

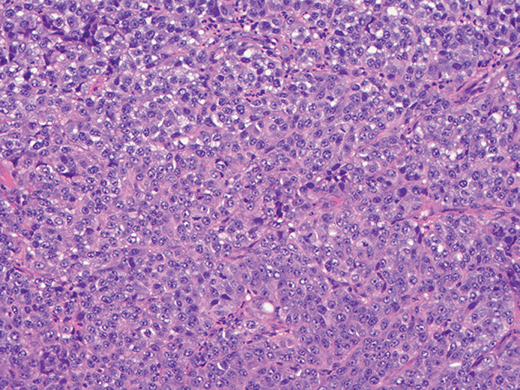
Figure 1.3 A proliferation of atypical epithelioid tumor cells characteristic of melanoma (X200).

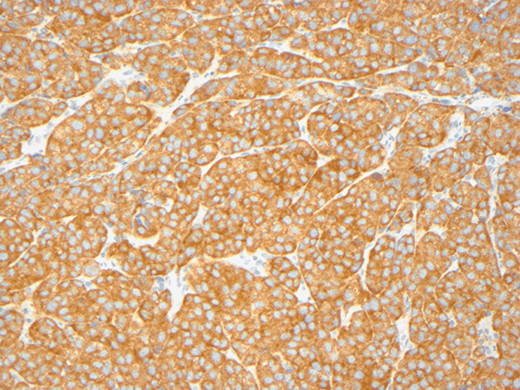
Figure 1.4 Diffuse cytoplasmic BRAFV600E expression (X200).
NRAS mutation usually occurs in codon 61 where glutamine is replaced by arginine (NRASQ61R) or leucine (NRASQ61L). Molecular detection of NRAS mutations involving codon 61, codon 12, and codon 13 can be time consuming. NRAS IHC is faster and more cost effective and has been shown to have high sensitivity and specificity.Reference Dias-Santagata, Su and Hoang40,Reference Kakavand, Walker and Lum41 The NRASQ61R immunostain (clone SP174) has 100% sensitivity and specificity, while NRASQ61L (clone 26193 from New East Bioscience) immunostain exhibits lower sensitivity (80%) and good specificity (100%).Reference Kakavand, Walker and Lum41 NRASQ61R or NRASQ61L antibody binds to a cytoplasmic protein. Positivity is considered when a strong, diffuse and homogeneous cytoplasmic staining is present (Figures 1.5 and 1.6). The positive percentage is around 80% which is easily visualized at low magnification.Reference Dias-Santagata, Su and Hoang40

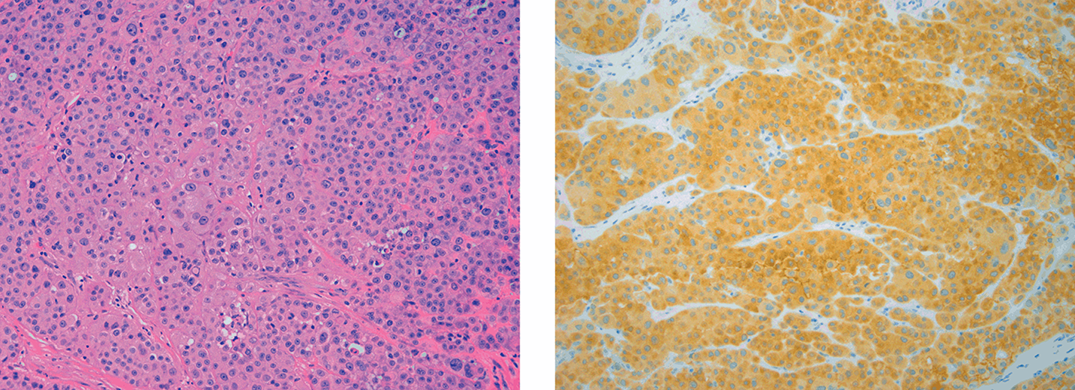
Figures 1.5 and 1.6 A melanoma with strong cytoplasmic NRASQ61R expression (X200, X200)
CDK4 and MDM2 immunostains can detect gene amplifications in well-differentiated and dedifferentiated liposarcomas. Other immunostains can detect gene deletion (SMARCB1/INI1), epigenetic alterations (H3K27me3), and increased expression discovered via gene expression profiling (ETV4, MUC4, NKX2.2, TLE1, and WT1) (Table 1.3).
1.6 Chromogenic In Situ Hybridization
RNA ISH assay (RNAscope) targeting high-risk HPV E6/E7 mRNA transcript has been reported to be a sensitive assay. HPV mRNA ish probes targeting E6 and E7 mRNA transcript of HPV6, HPV11, HPV16, HPV18 and HPV33 have shown to be helpful tests in detecting papillomavirus in formalin-fixed paraffin-embedded tissue.Reference Luk, Selinger, Cooper, Mahar and Palme42
1.7 Dual Chromogenic Immunohistochemistry
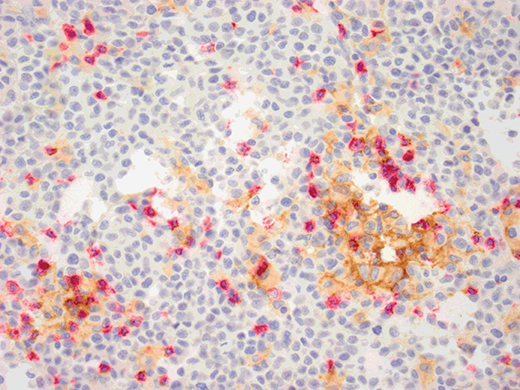
Dual chromogenic immunostain is comprised of DAB (brown) chromogen labeling one target and red chromogen for the other. Examples of clinical diagnostic stains include combined Melan-A and Ki67, combined S100 and D2-40, and combined CD4 and CD8 immunostains. In clinical practice a combined PDL1 and CD8 combined immunostain can guide treatment decision (Figure 1.7).Reference Tumeh, Harview and Yearley43 Programmed death (PD)-1 is an immunoglobulin superfamily member expressed on the surface of activated T cells and B cells, while PD-L1 is present on the surface of tumor cells. Although PD1 has two ligands, it exhibits greater affinity with PD-L1 than PD-L2. When PD-L1 expressed on tumor cells binds to PD-1 expressed on activated T-cells, an inhibition of tumor immune destruction by activated T-cells occurs.Reference Zou and Chen44 Blocking the interaction between PD1 and its ligand PD-L1 has resulted in improvements in antitumor responses.Reference Topalian, Hodi and Brahmer45 Pembrolizumab (Keytruda®) and nivolumab (Opdivo®) are two anti-PD1/PD-L1 inhibitors that targeted the PD-1 pathway and have recently been approved by the FDA (Foods and Drugs Administration) for the treatment of advanced metastatic melanoma. A landmark publication showed that patients with metastatic melanoma treated with tumor infiltrating lymphocytes (TIL) followed by high-dose IL-2 demonstrated a 34% overall response rate.Reference Rosenberg, Yannelli and Yang46 Studies on the role of tumor infiltrating lymphocytes/immune response as prognostic markers and predictor of response to immunomodulators have shown that in melanoma CD8+ T-cell density is the best predictive marker of response to PD1 blockade therapy.
1.8 Multiplex Immunohistochemistry and Immunofluorescence
Recent novel techniques including multiplexed immunohistochemistry and immunofluorescence, automated multispectral analysis, and advanced quantitation software can allow one to quantitate and assess co-expression and spatial relationship through simultaneous detection of multiple markers; whereas traditional IHC can only evaluate 1–2 proteins or 2–3 DNA/RNA molecules.Reference Stack, Wang, Roman and Hoyt47 Multiplex techniques can be cyclic immunofluorescence,Reference Zrzazhevskiy and Gao48 tyramide-base,Reference Stack, Wang, Roman and Hoyt47 epitope-targeted mass spectrometry,Reference Giesen, Wang and Schapiro49 or RNA detection.Reference Ke, Chen and Ban50 These techniques are currently utilized mainly in the research settings.
1.9 Future Direction
Bright-field RNA in situ hybridization using techniques similar to those of immunohistochemistry have allowed the detection of messenger RNA as well as long non-coding RNA with sensitivity to single RNA molecules in formalin-fixed paraffin-embedded tissues.Reference Tubbs, Wang and Wang51,Reference Ricciuti, Mencaroni, Paglialunga, Paciullo, Crino, Chiari and Metro52 Recent studies have reported the potential of formalin-free, non-crosslinking fixatives to replace formalin in diagnostic pathology and research.Reference Kothmaier, Rohrer, Stacher, Quehenberger, Becker and Popper53 Quantum dots are semiconductor nanoparticles that emit different wavelengths depending on their size, shape, and composition.Reference Jin and Hildebrandt54 Due to their narrow emission spectra and higher fluorescent signal per unit of light absorbed, quantum dots has been reported to be more accurate and precise at low concentration than traditional IHC.Reference Chen, Sun and Gong55 In addition to detecting proteins, new immunostains will continue to play an important role in diagnostic pathology with an increased availability of novel techniques, sensitive and specific biomarkers, predictive markers, and mutation-specific antibodies.