Introduction
Selecting elite genotypes based on morphological features from naturally growing seedling populations forms the foundation of present-day fruit culture. This stands in contrast to the relatively recent practice of developing varieties through hybridization, which only emerged in the 1900s. Venturing into the evolution of varieties in perennial tree fruits presents evident concerns, notably heterozygosity and lengthy gestation cycles. Furthermore, the limited number of breeding programmes with enduring financial and institutional support adds to these challenges. These challenges become more pressing when considering certain nutraceutically rich fruit crops like jamun, renowned as a health tonic in various medical systems including Ayurveda, Unani, and traditional therapies. Jamun (Syzygium cumini L. Skeels), also referred to as Indian blackberry, jambu, java plum, black plum and jambul Portuguese plum (Sharma et al., Reference Sharma, Mehta, Mehta, Nagar and Mishra2012), is a wild indigenous minor fruit tree of India (Singh, Reference Singh1969) belonging to the Myrtaceae family with a chromosome number of 2n = 40 (Tewari et al., Reference Tewari, Singh, Nainwal and Nair2017; Singh et al., Reference Singh, Singh, Saroj and Mishra2019a). Its distribution spans the Western Ghats and Indo-Gangetic regions of the Indian subcontinent. The Indian subcontinent earned the name ‘Jambudeep’ due to the profusion of jamun trees (Karmarkar, Reference Karmarkar1955). Beyond India, it is also cultivated in various Asian, African and American countries (Singh et al., Reference Singh, Singh, Saroj and Mishra2019a). Although organized plantations of this crop are scarce, jamun trees can be observed growing sporadically in patches across different villages in various Indian states.
Jamun's significance extends beyond its fresh consumption. Its fruit, leaves, stem bark and seeds find application in traditional medicine. The fruit's pulp and seeds contain anthocyanins, fibres, jamboline, glucoside, ellagitannins and resveratrol, which aid in reducing diseases induced by oxidative stress (Kumar et al., Reference Kumar, Misra and Mishra2011). These components exhibit robust antioxidant, chemoprotective, functional and nutraceutical properties (Kapoor et al., Reference Kapoor, Ranote and Sharma2015; Hameed et al., Reference Hameed, Gupta, Rahman, Anjum, Nayik, Nayik and Gul2020). Jamun fruit extract demonstrates anticancer activity against leukaemia cancer cell lines, attributed to compounds like kaempferol 7-O-methyl ether and sterols such as γ-sitosterol (Afify et al., Reference Afify, Fayed, Shalaby and El-Shemy2011). The fruit and seeds are widely used for their anti-glycaemic and anti-hypercholesterolaemic effects (Raza et al., Reference Raza, Butt and Suleria2017). Additionally, its leaves and bark are employed in treating conditions such as hypertension, gingivitis, diuretic effects, antiscorbutic properties, chronic diarrhoea and spleen enlargement (Achrekar et al., Reference Achrekar, Kakliji, Pote and Kelkar1991; Maran et al., Reference Maran, Sivakumar, Thirugnanasambandham and Sridhar2014).
Despite advancements in producing high-yield varieties and improved agricultural techniques (Singh et al., Reference Singh, Bajpai, Singh, Ravishankar, Tandon and Reddy2010), the growth of jamun cultivation remains restricted to local markets, prompting the need to address existing concerns (Singh et al., Reference Singh, Singh, Saroj, Rao and Mishra2019b). This is highlighted by the short harvest season (lasting about two weeks) in various Indian jamun-growing regions (Singh et al., Reference Singh, Bajpai, Singh, Ravishankar, Tandon and Reddy2010; Mishra et al., Reference Mishra, Singh, Kumar, Singh, Singh, Swamy and Ghosh2014), requiring the development of early, medium and late maturing varieties to extend harvest duration and overcome seasonal limitations (Roy, Reference Roy2014). Furthermore, ripe jamun fruits are susceptible to spoilage and possess a limited shelf life (Madani et al., Reference Madani, Mirshekari, Yahia, Golding, Hajivand and Dastjerdy2021), necessitating the creation of new varieties with firmer texture and prolonged shelf life (Singh et al., Reference Singh, Bajpai, Singh, Ravishankar, Tandon and Reddy2010). Syzygium plants exhibit polyembryony and exhibit consistent traits. Natural variations occur in fruit attributes such as shape, size, total soluble solids (TSS), acidity and bearing precocity. Opting for selection as the preferred breeding approach allows the choice of desired genotypes in their native environments. The inherent variability provides an opportunity to develop varieties with reduced seed size, increased pulp, lower astringency and an extended shelf life. Gepts (Reference Gepts1993) proposes that understanding phenotypic traits, coupled with genetic analysis, serves as the initial step in conserving, sustaining and capitalizing existing genetic diversity. Prioritizing the genetic enhancement of jamun includes the creation of compact, early-bearing cultivars with sizable fruits, high pulp content and resilience against pests and diseases (Mishra et al., Reference Mishra, Singh, Kumar, Singh, Singh, Swamy and Ghosh2014; Singh et al., Reference Singh, Singh, Saroj and Mishra2019a).
While molecular markers enhance precision and efficiency in crop improvement, jamun genetic resources are largely characterized through morphological traits (Ghojage et al., Reference Ghojage, Swamy, Kanamadi, Jagdeesh, Kumar, Patil and Reddy2011; Plathia et al., Reference Plathia, Wali, Bakshi and Sharma2018; Singh et al., Reference Singh, Singh, Saroj, Rao and Mishra2019b; Bhatnagar et al., Reference Bhatnagar, Sharma and Singh2020; Ud Din et al., Reference Ud Din, Jaskani, Naqvi and Awan2020; Singh et al., Reference Singh, Verma, Prasad, Kumar, Sharma and Singh2022). However, morphological selection of certain jamun cultivars possess significant drawbacks (Madani et al., Reference Madani, Mirshekari, Yahia, Golding, Hajivand and Dastjerdy2021), raising concerns about genetic diversity loss (Singh et al., Reference Singh, Singh, Saroj and Mishra2019a). This underscores the importance of employing molecular and genomic tools to enhance germplasm management and conduct association mapping for the development of core collections in jamun (Jamnadass et al., Reference Jamnadass, Lowe and Dawson2009; Patzak et al., Reference Patzak, Paprštein, Henychová and Sedlák2012). While some studies indicate the potential of molecular markers to differentiate jamun accessions and assess genetic relationships (Shakya et al., Reference Shakya, Siddiqui, Srivatawa and Bajpai2010; Khan et al., Reference Khan, Vaishali and Sharma2011; Ahmad et al., Reference Ahmad, Bhagat, Simachalam and Srivastava2012; Singh et al., Reference Singh, Singh, Bajpai and Ahmad2014b; Gajera et al., Reference Gajera, Gevariya, Patel and Golakiya2018), most of these investigations were limited to specific geographic regions and used a small marker set. It is demonstrated that using sufficient, frequently occurring and easily reproducible polymorphic markers (Belaj et al., Reference Belaj, De la Rosa, Lorite, Mariotti, Cultrera, Beuzón, González-Plaza, Munoz-Mérida, Trelles and Baldoni2018) across a substantial number of accessions from diverse environments (Singh et al., Reference Singh, Rana, Singh, Kumar, Kumar and Singh2014a; Luo et al., Reference Luo, Brock, Dyer, Kutchan, Schachtman, Augustin, Ge, Fahlgren and Abdel-Haleem2019) provides a more accurate estimate of genetic diversity and relationships. Furthermore, it is worth noting that earlier molecular investigations predominantly utilized markers such as random amplified polymorphic DNA (RAPD) and inter simple sequence repeats (ISSR), as highlighted in studies conducted by Ahmad et al. (Reference Ahmad, Bhagat, Simachalam and Srivastava2012), Gajera et al. (Reference Gajera, Gevariya, Patel and Golakiya2018) and (Khodaee et al., Reference Khodaee, Azizinezhad, Etminan and Khosroshahi2021). More robust markers like simple sequence repeats have not been explored for the specific jamun species or cross-species amplification. However, in a more recent development, Singh et al. (Reference Singh, Verma, Prasad, Kumar, Sharma and Singh2022) introduced a more robust and advanced approach known as the CAAT box-derived polymorphism marker technique for assessment of genetic diversity and population structure within the jamun species.
Despite the absence of DNA sequence information, molecular markers like RAPD and ISSR are employed to assess the efficiency of these markers to analyse species diversity. Thus, this study's focus is to assess the variability of S. cumini populations in the North Western Himalayan region using both morphological descriptors and RAPD and ISSR molecular markers. This study's novelty lies in providing insights into the evaluation of genetic diversity among different S. cumini accessions, a potential fruit crop with valuable nutraceutical properties and economic potential. The identified elite genotypes hold promise for release as varieties after evaluation or for utilization in future breeding programmes.
Materials and methods
Geographical site
At the beginning of the experiment, the sites containing abundant populations of natural S. cumini across three districts in the Himachal province of the North Western Himalayas were pinpointed. All three of these districts are situated in a sub-tropical climate. Providing geographical and climatic insights into specific study regions or districts, the focus extends to Mandi, Hamirpur and Kangra (online Supplementary Fig. S1). The Mandi district is identified as having a sub-humid and wet temperate climate, encompassing an elevation range from 550 to 3960 m above mean sea level. Its geographical coordinates are latitude 31°35′21.12″ N and longitude 76°55′05.55″ E, with an annual precipitation of 1338.9 mm. In the Hamirpur district, the classification aligns with sub-mountain, low hills and subtropical characteristics. The elevation spans from 470 to 1235 m above mean sea level, and its geographic position is latitude 31°41′10.32″ N, longitude 76°31′16.71″ E. The district receives an annual rainfall of 1328.5 mm. District Kangra stands out for its sub-mountain, low hills and subtropical/alpine attributes. The elevation range covers 350–4880 m above mean sea level, with geographic coordinates of 31°05′59.29″ N and longitude 76°16′08.76″ E. Notably, Kangra records an annual precipitation of 1949.4 mm. These insights contribute to an understanding of the diverse climates, elevations and geographic locations of the respective districts.
Sample collection and screening of genotypes
A preliminary survey was conducted on the seedling populations of S. cumini within their natural habitats with the aim of identifying a viable population for more in-depth investigation by assessing their fruiting traits. The initial data gathered from the nearby residents played a crucial role in guiding this selection process.
For screening studies, a numerical scoring system was developed on the basis of seven characters/traits of horticultural importance and consumer preference. The numerical scoring system adopted for selection is given below:

Here, we evaluated a population of 82 trees surveyed during fruiting season of jamun during 2019 and 2020 (refer to online Supplementary Table S1 and Fig. S1). Fresh ripe fruits were collected from marked plants during the fruiting season to record data on fruit size, colour, TSS, profuse bearing and regular bearing character of the fruit. Trees obtaining high scores in aggregate were taken as elite trees and considered as selected genotypes.
Morphological and biochemical analysis
For the purpose of morpho-molecular analysis, we examined 15 specific genotypes chosen from a larger population of 82 trees that had been surveyed for their horticulturally significant traits (online Supplementary Fig. S2). The assessments regarding tree, foliage and fruit characteristics were documented following the guidelines outlined in the descriptor provided by the National Bureau of Plant Genetic Resources (NBPGR) (Anonymous, 2002). Quantitative attributes were measured and weighed using precise equipment: an electronic balance (Kern, ABJ-NM, Powai, Mumbai, Maharashtra (India)) for weight measurements and a digital Vernier caliper (Mitutoyo, ASX) for measurements in millimetres. The measurements were expressed in grams (g) and millimetres (mm), respectively. Colour-related attributes were visually evaluated by referencing colour charts. Data pertaining to quantitative traits, such as tree height, tree canopy area, tree girth, leaf dimensions along with their ratios, petiole length, fruit dimensions, fruit weight, pulp weight, seed weight, seed dimensions, pulp-to-seed ratio, pulp percentage, fruit size and seed size, were recorded in accordance with the crop descriptor. Additionally, qualitative characteristics including tree habit, tree foliage type, bark colour, leaf colour, leaf shape, leaf apex, leaf base, fruit shape, fruit apex, fruit base, fruit colour and pulp colour were noted.
For the assessment of fruit sweetness, a small quantity of fresh fruit juice was placed on the prism of an Erma-hand refractometer. The sweetness, measured in terms of TSS, was determined using the refractometer (0–32° Brix). The juice's titrable acidity, reducing sugar, total sugar and ascorbic acid content were determined following the method described by Ranganna (Reference Ranganna1986). Non-reducing sugar content was evaluated using the procedure outlined by Sadasivam and Manickam (Reference Sadasivam and Manickam1996). All measurements and observations were conducted in triplicate, with each replicate consisting of five samples.
Isolation of genomic DNA
Different procedures were utilized to standardize the DNA extraction protocol and amplification procedure for jamun accessions. Disease-free, young, fresh green leaves were individually collected from superior selected plants originating from various regions including Mandi, Hamirpur and Kangra (HP). The leaves were carefully cleaned to remove soil particles, wrapped in aluminium foil and transported to the laboratory in an icebox. Subsequently, they were stored in a deep freezer until further processing.
The genomic DNA was isolated using a modified version of the CTAB method, based on the approach outlined by Doyle (Reference Doyle, Hewitt, Johnson and Young1991). While Doyle (Reference Doyle, Hewitt, Johnson and Young1991) employed a 2% CTAB solution along with 1.4 M NaCl, 20 mM EDTA and 0.2% β-mercaptoethanol for the extraction buffer, the current protocol introduced several adjustments. The extraction buffer composition was modified to include 6% CTAB, 5 M NaCl, 0.5 M EDTA and 2% polyvinylpyrrolidone. The incubation time was prolonged to 60 min, and the temperature during the water bath treatment was raised to 65°C, in contrast to the 30 min treatment at 60°C described by Doyle. Furthermore, the cold isopropanol precipitation step was extended to an overnight duration. The incorporation of higher concentrations of NaCl and EDTA in the extraction buffer was intended to facilitate DNA extraction while minimizing DNA degradation, aided by the presence of EDTA which assists in protein denaturation. The longer incubation time and higher temperature in the water bath were implemented to optimize the extraction efficiency. These modifications to the protocol led to the successful extraction of a substantial amount of DNA from all jamun genotypes. The DNA concentration and purity were assessed using a UV-vis spectrophotometer (Thermo Scientific Evolution 200) and gel electrophoresis. The DNA was dissolved in 50 μl TE buffer and subsequently stored at −20°C.
Molecular analysis of genomic DNA
DNA samples from 15 carefully selected jamun genotypes were subjected to amplification using a total of 16 RAPD and 10 ISSR markers. Following the initial screening process, primers that demonstrated both polymorphic and consistent band patterns, referred to as informative primers, were chosen for further amplification targeting S. cumini accessions. The amplification procedure was executed in a reaction volume of 10 μl, composed of 0.5 μl Taq DNA polymerase, 1 μl Taq Buffer, 0.5 μl dNTP mix, 0.5 μl of the selected primer, 4 μl genomic DNA and 3.5 μl autoclaved distilled water. All reagents utilized in the PCR reaction mixture were procured from Genei Private Limited, located in Bangalore, India.
The amplification process comprised a total of 35 cycles, performed using a Mini AMP Plus thermocycler. Each cycle consisted of denaturation, annealing and initial extension stages. The specific thermal profile employed for both RAPD and ISSR amplification begins with initial denaturation at 94.0°C for 5.0 min, followed by final denaturation at the same temperature for 1 min. This initial denaturation aims to denature the DNA template by disrupting hydrogen bonds, readying it for amplification. Subsequently, at 35.0°C (for RAPD) or 55.0°C (for ISSR), the primer annealing phase lasts for 1.0 min. The goal is to facilitate the binding of primers to complementary DNA sequences, aiding targeted amplification. The subsequent initial extension occurs at a temperature of 72.0°C for 1.0 min (for RAPD) or 2.0 min (for ISSR), succeeded by final extension at 72.0°C, maintained for 5.0 min (for RAPD) or 7.0 min (for ISSR). Finally, the temperature is lowered to 4.0°C, and this indefinite ‘hold’ step preserves DNA at a low temperature, effectively preventing degradation or further reactions. The RAPD and ISSR techniques share a comparable PCR process while diverging in annealing temperature and time, reflecting differences in their primers and target regions. RAPD employs random primers and employs a lower annealing temperature (35.0°C) for shorter sequences, while ISSR uses primers designed for specific microsatellite regions, necessitating a higher annealing temperature (55.0°C) for enhanced specificity.
The amplified PCR product bands were examined on a 1.8% agarose gel, subjected to an electric field of 75 V until the loading dye migrated to the leading edge of the gel. Subsequently, the gel was visualized using a UV trans-illuminator, and the image was captured using a gel documentation unit. TBE buffer (1×) was employed both as the gel matrix and tray buffer. To analyse the correct banding pattern, a 1 kb DNA ladder marker was utilized as a reference standard.
Statistical analysis
The data obtained from these investigations were subjected to statistical analysis as procedures described by Gomez and Gomez (Reference Gomez and Gomez1984). Statistical analysis was performed for each observed character by using MS-Excel (version 2019) and OPSTAT. The critical difference was calculated at 5% level of significance. Principal component analysis (PCA) was carried out to standardize the data set and to observe interrelationships of samples and analysed using Microsoft Excel 2019 described by Zielinski et al. (Reference Zielinski, Haminiuk, Alberti, Nogueira, Demiate and Granato2014) with the trial version of XLSTAT (version 2020; Addinsoft). Agglomerative hierarchical clustering (AHC) has been performed by using the trial version of XLSTAT (version 2020; Addinsoft) and Euclidean used as a distance measure for dendrogram construction.
For molecular data analysis, each amplified product was scored across all genotypes for molecular analysis with respect to all informative primers. Presence of an amplified product was designated as ‘1’ and absence was marked as ‘0’. Numerical Taxonomic and Multivariate Analysis System (NTSYS-pc) version 2.02 (Rohlf, Reference Rohlf1998) software program was used to analyse molecular data matrix. Similarity index was estimated using SIMQUAL function of NTSYS, which computes a variety of similarity coefficient for qualitative data (nominal data). Dendrogram was constructed using UPGMA available in NTSYS software.
Results
Morphological characterization
Tree and foliage characters
Considerable variation was evident in terms of tree characteristics, with significant differences observed in tree height (ranging from 516.87 to 2052.62 cm), trunk girth (ranging from 96.52 to 274.86 cm), tree canopy area for the east–west direction (ranging from 373.38 to 1325.88 cm) and tree canopy area for the north–south direction (ranging from 497.84 to 2024.38 cm) (online Supplementary Table S2). Among the recorded measurements, tree 39 exhibited the maximum tree height and trunk girth, while the most extensive tree canopy was found in tree 43 for the east–west direction and tree 44 for the north–south direction. A majority of the genotypes displayed a spreading tree habit and dense foliage. Among the genotypes, nine had a grey tree bark colour, whereas three genotypes each exhibited brown and light grey bark colours. Leaf colour predominantly ranged from green to dark green, and the leaf shape was predominantly elliptic among 11 genotypes, while the remaining exhibited lanceolate, elliptic oblong and broadly ovate shapes. Acute and round leaf bases were prevalent, and leaf apices were mainly acute and acuminated across all screened genotypes.
In terms of leaf quantitative attributes, considerable variability was observed in leaf length (ranging from 12.50 to 18.66 cm), leaf width (ranging from 5.10 to 7.36 cm), leaf petiole length (ranging from 1.53 to 3.43 cm) and the ratio of leaf length to width (ranging from 1.88 to 3.18) (online Supplementary Table S3).
Fruit physicochemical characters
A significant degree of variation was observed in the fruit characteristics of S. cumini (online Supplementary Table S4). Noteworthy variations encompassed attributes such as fruit weight (ranging from 3.24 to 6.43 g), fruit length (ranging from 19.90 to 28.03 mm), fruit width (ranging from 15.63 to 20.34 mm), fruit size (ranging from 3.44 to 5.54 cm2), pulp weight (ranging from 2.34 to 5.87 g), seed weight (ranging from 0.47 to 1.40 g), seed length (ranging from 12.33 to 18.23 mm), seed width (ranging from 5.92 to 10.27 mm), pulp-to-seed ratio (ranging from 2.00 to 10.72) and pulp percentage (ranging from 66.51 to 91.30%). Most of the fruit-related parameters, including fruit weight, fruit length, fruit size, pulp weight, seed length, seed size, pulp percentage and pulp-to-seed ratio, reached their maximum values in tree 43. Conversely, fruit width, seed weight and seed width exhibited their highest measurements in tree 5 (online Supplementary Table S5). In terms of fruit shapes, oblong configurations were predominant across the genotypes, followed by ellipsoid, oval and round shapes. The fruit apex was round in 10 genotypes and flat in the remaining five, while the fruit base was flat in 11 genotypes and depressed in the other four (Fig. 1).
Fruit characteristics of superior selected jamun genotypes.

Regarding biochemical attributes, a wide range of variation was observed among the superior selected trees studied (online Supplementary Table S6). Notable variability extended to TSS levels (ranging from 9.93 to 22.27°B), acidity (ranging from 0.36 to 0.73), the TSS-to-acidity ratio (ranging from 17.68 to 43.37), total sugar content (ranging from 9.51 to 29.25%), reducing sugar content (ranging from 8.29 to 18.41%), non-reducing sugar content (ranging from 0.52 to 2.47%), the sugar-to-acid ratio (ranging from 13.11 to 39.64) and ascorbic acid content (ranging from 13.60 to 34.45 mg/100 g). The highest TSS, total sugar content and reducing sugar content were observed in tree 49. Tree 54 exhibited the highest acidity, tree 13 had the highest ascorbic acid content, tree 5 showcased the highest non-reducing sugar content, tree 23 presented the highest TSS-to-acidity ratio and tree 23 had the highest sugar-to-acid ratio.
Principal component analysis (PCA)
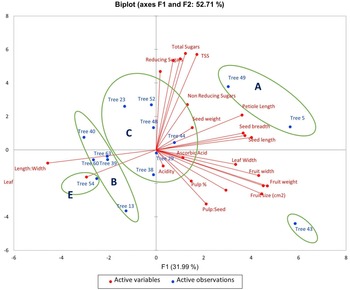
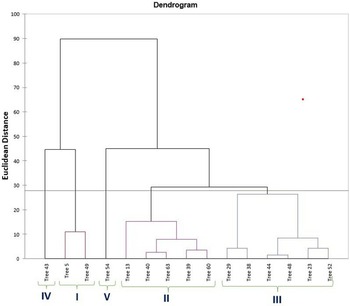
PCA was utilized to elucidate the variations among distinct genotypes based on their characteristics. A comprehensive examination of six principal components, namely PC1, PC2, PC3, PC4, PC5 and PC6, was undertaken. These components collectively accounted for variations of 31.99, 20.71, 18.95, 7.16, 5.97 and 5.08, respectively (Table 1). The orientation of the vector assigned to a variable reflects its relative contribution to the initial two PCs. Variables placed at narrow angles denote positive correlations, whereas those at wider angles indicate negative correlations. Mean PC scores for replicates of reference accessions are illustrated in red. The 2D biplot of PCA effectively illustrates the combined contribution of PC1 and PC2, amounting to 52.71% of the total variance (Figs 2 and 3). PC1 exhibited an eigenvalue of 7.358, elucidating 31.99% of the overall variability. The variance encompassed by PC1 predominantly stemmed from attributes such as fruit width (0.302), fruit weight (0.327), petiole length (0.253), leaf length-to-width ratio (−0.319), pulp weight (0.311) and fruit size (0.315). In parallel, PC2 demonstrated an eigenvalue of 4.765, contributing 20.71% to the total variability and was chiefly characterized by attributes like TSS (0.399), reducing sugars (0.380), total sugars (0.402), TSS-to-acidity ratio (0.374) and sugar-to-acid ratio (0.327). The AHC analysis was performed on quantitative data derived from 23 attributes, utilizing the Euclidean dissimilarity matrix and the Ward's minimum variance clustering method (online Supplementary Fig. S3). The threshold dissimilarity value for clustering was set at 28, represented by a dotted line along the Y-axis. Among the 15 jamun genotypes, the AHC yielded five distinct clusters. Cluster I encompassed two genotypes (trees 5 and 49), while cluster II comprised five genotypes (trees 13, 40, 63, 39, 60) (online Supplementary Table S7). Cluster III, the largest group, comprised six genotypes (trees 29, 38, 44, 48, 23, 52), and clusters IV and V consisted of a single genotype each, specifically trees 43 and 54, respectively.
Eigen value and percentage of variation for corresponding first 3 principal components based on morphological characters

Biplot of loading and score values of PC1 and PC2 for combined morphological and biochemical characters.

Dendrogram based on combined morphological and biochemical characters using AHC.

Molecular characterization
RAPD and ISSR analysis of genomic DNA
For the RAPD analysis, a total of 16 primers were examined, with 15 of them yielding robust amplification (online Supplementary Fig. S3). In contrast, when considering ISSR primers, all 10 screened primers exhibited successful amplification (online Supplementary Fig. S4). The analysis generated a total of 956 bands, out of which 911 displayed polymorphism, accounting for 95.29% of the bands, while the remaining 45 bands were monomorphic. The size of all bands fell within the range of 100–1000 bp (Table 2). A similarity matrix was constructed using Jaccard's coefficient based on the complete set of screened primers. The computed similarity coefficient values spanned from 0.37 to 0.77. Notably, the lowest similarity coefficient (0.37) was observed between trees 29 and 43, highlighting their distinctiveness. On the other end of the spectrum, the highest similarity coefficient (0.77) was noted between trees 49 and 52.
Scorable bands/amplified band and polymorphic bands generated by PCR using RAPD and ISSR primers

Cluster analysis based on RAPD and ISSR profile
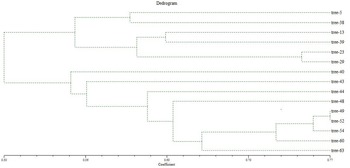
During the cluster analysis, the dendrogram underwent a partition, resulting in two major clusters identified at a similarity index value of 0.47 (Fig. 4). Cluster 1 encompassed six jamun genotypes, branching apart at a similarity index of 0.554. Within this cluster, two genotypes, namely trees 5 and 38, shared a resemblance, while four other genotypes (trees 13, 39, 23, 29) were situated within the similarity index range of 0.602–0.742. As indicated by the dendrogram, tree 40 (0.548) exhibited dissimilarity, followed by trees 43 (0.56), 44 (0.616) and 48 (0.637); so, these genotypes stood apart as distinct entities with the lowest similarity index values, highlighting their diversity.
Dendrogram of 15 selected trees of jamun based on RAPD and ISSR analysis using UPGMA.

Moving to cluster 2, the highest similarity index value of 0.77 was observed between genotypes trees 49 and 52, signifying their strong similarity and positioning them as the most closely related cultivars. The remaining three genotypes trees 54 (0.756), 60 (0.728) and 63 (0.66) were isolated as individual branches, indicating their dissimilarity from the other genotypes.
Discussion
Selection based on characterization is a crucial technique within crop improvement initiatives, aimed at acquiring desirable physical and fruit-related traits. Despite its status as a native fruit crop, the history of genetic enhancement for jamun in India has been brief (Singh et al., Reference Singh, Bajpai, Singh, Ravishankar, Tandon and Reddy2010). Even though a few improved cultivars have recently been introduced (Singh et al., Reference Singh, Singh, Saroj and Mishra2019a), the increasing medicinal potential of jamun underscores the necessity for intensified cultivar development strategies employing molecular and genomic resources. Prior attempts to assess jamun's genetic diversity employed either physical (Kaur and Singh, Reference Kaur and Singh2017; Anushma and Sane, Reference Anushma and Sane2018; Singh et al., Reference Singh, Singh, Saroj, Rao and Mishra2019b; Ud Din et al., Reference Ud Din, Jaskani, Naqvi and Awan2020) or molecular (Shakya et al., Reference Shakya, Siddiqui, Srivatawa and Bajpai2010; Khan et al., Reference Khan, Vaishali and Sharma2011; Ahmad et al., Reference Ahmad, Bhagat, Simachalam and Srivastava2012) markers. Some studies simultaneously explored both aspects (Shakya et al., Reference Shakya, Siddiqui, Srivatawa and Bajpai2010; Khan et al., Reference Khan, Vaishali and Sharma2011; Cheong and Sanmukhiya, Reference Cheong and Sanmukhiya2013; Singh et al., Reference Singh, Singh, Bajpai and Ahmad2014b; Gajera et al., Reference Gajera, Gevariya, Patel and Golakiya2018), but they were limited to a few locally accessible Syzygium species/accessions and utilized a small number of RAPD and IISR markers, thus hindering meaningful conclusions. In contrast, our study investigated a considerable number of wild genotypes collected from three distinct districts in Himachal Pradesh, examining morphological and molecular variations to assess genetic diversity. While previous research acknowledged genetic variability in jamun, the exploration of genetic diversity within the wild populations of the North-West Himalayas remains scarce.
Our findings in morphological characteristics unveiled substantial genetic variability in superior S. cumini genotypes in terms of plant growth and fruit physicochemical attributes (Anushma and Sane, Reference Anushma and Sane2018; Khadivi et al., Reference Khadivi, Mirheidari, Moradi and Paryan2020; Guo et al., Reference Guo, Liu, Li, Cao, Zhang, Zhang, Zhang, Deng, Niu, Su and Li2022; Singh et al., Reference Singh, Verma, Prasad, Kumar, Sharma and Singh2022). These variations can be attributed to diverse edaphic, climatic and genetic factors (Agrawal et al., Reference Agrawal, Rangare and Nair2017; Kaur and Singh, Reference Kaur and Singh2017; Plathia et al., Reference Plathia, Wali, Bakshi and Sharma2018), influenced by spatial and temporal differences in sample collection sites (Devi et al., Reference Devi, Swamy and Naik2016; Ningot et al., Reference Ningot, Dahale, Bharad and Nagre2017; Akhila and Hiremath, Reference Akhila and Hiremath2018; Babu et al., Reference Babu, Raghavendra and Basavaraj2019). PCA demonstrated a significant amount of variation (90.34%) for morphological traits, akin to findings in other crops like mulberry (Natić et al., Reference Natić, Dabić, Papetti, Akšić, Ognjanov, Ljubojević and Tešić2015), hog plum (Silva et al., Reference Silva, Rossi, Dardengo, Tiago, Silveira and Souza2017) and rice (Bollinedi et al., Reference Bollinedi, Vinod, Bisht, Chauhan, Krishnan, Bhowmick, Nagarajan, Rao, Ellur and Singh2020) regarding physicochemical attributes. Traits displaying higher variability are prime targets for enhancement and contribute to a robust gene transfer during breeding programmes (Aliyu et al., Reference Aliyu, Akoroda and Padulosi2000; Aliyu and Awopetu, Reference Aliyu and Awopetu2007). However, clustering patterns revealed that fruits collected from the same districts or geographical areas did not consistently cluster together. The clustering of jamun genotypes based on morphological traits did not align well with their geographical origins, analogous to findings in apple (Gasi et al., Reference Gasi, Simon, Pojskic, Kurtovic and Pejic2010) and olive (Hagidimitriou et al., Reference Hagidimitriou, Katsiotis, Menexes, Pontikis and Loukas2005). Although morphological traits are useful for distinguishing crop genotypes, their applicability in deducing precise genetic relationships in a diverse array of accessions remains limited (Hagidimitriou et al., Reference Hagidimitriou, Katsiotis, Menexes, Pontikis and Loukas2005). Distinct geographical origins do not necessarily imply genetic disparities; the grouping of accessions from various populations within the same clusters suggests shared ancestry or frequent inter-regional seed exchanges (Norouzi et al., Reference Norouzi, Erfani-Moghadam, Fazeli and Khadivi2017; Wever et al., Reference Wever, Holler, Becker, Biertümpfel, Kohler, van Inghelandt, Westhoff, Pude and Pestsova2019). Bird-mediated seed dispersal, attracted to jamun fruits (Singh et al., Reference Singh, Bajpai, Singh, Ravishankar, Tandon and Reddy2010), potentially contributes to seed dispersal over extensive distances (Wadl et al., Reference Wadl, Rinehart, Olsen, Waldo and Kirkbride2022). Euclidean dissimilarity distance unveiled close similarity among genotypes from the same districts. For instance, genotypes from Hamirpur (trees 44 and 48) exhibited the shortest Euclidean distances, possibly due to shared gene pools as chance seedlings (Rohini et al., Reference Rohini, Sankaran, Rajkumar, Prakash, Gaikwad, Chaudhury and Malik2020). The existence of nucellar embryony in jamun (Sivasubramaniam and Selvarani, Reference Sivasubramaniam and Selvarani2012) might also play a role in the minimal morphological differences observed among certain accessions (Omar et al., Reference Omar, El Sayed, Mohamed, Ramadan, Ayoub and Rohn2021).
Conversely, molecular analysis indicated a high level of diversity, evidenced by the broad range of similarity coefficient values (Jaccard's coefficient) derived from RAPD and ISSR, showing 95.29% polymorphism with similarity coefficients spanning from 0.37 to 0.77 (Khan et al., Reference Khan, Vaishali and Sharma2010, Reference Khan, Vaishali and Sharma2011; Shakya et al., Reference Shakya, Siddiqui, Srivatawa and Bajpai2010; Ahmad et al., Reference Ahmad, Bhagat, Simachalam and Srivastava2012; Singh et al., Reference Singh, Singh, Bajpai and Ahmad2014b; Gajera et al., Reference Gajera, Gevariya, Patel and Golakiya2018; Khodaee et al., Reference Khodaee, Azizinezhad, Etminan and Khosroshahi2021). Through the analysis of both primers, all genotypes were divided into two main clusters, with trees 40, 43, 44 and 48 standing apart from others due to their distinctiveness at minimum similarity index values, signifying their diverse nature. However, molecular data-based UPGMA clustering, though superior to morphological traits in terms of region-wide grouping of accessions, still placed genotypes from diverse regions into the same cluster. Animal-mediated seed dispersal (Krishnamurthy et al., Reference Krishnamurthy, Shaanker and Ganeshaiah1997) could contribute to inter-regional genetic similarities in jamun. While cross-pollination and ongoing seed propagation seem pivotal reasons for heightened molecular variance within jamun populations (Keneni et al., Reference Keneni, Bekele, Imtiaz, Dagne, Getu and Assefa2012; Liu et al., Reference Liu, Ge, Wang, Li, Yang, Cui and Qu2013), animal-mediated seed dispersal (Krishnamurthy et al., Reference Krishnamurthy, Shaanker and Ganeshaiah1997) might also influence within-population gene flow. These variations could stem from genetically distinct parentage (Shekhawat et al., Reference Shekhawat, Rai, Shekhawat and Kataria2018) and a prolonged history of micro-climatic adaptability. Moreover, genetic differences might arise from continuous seed propagation combined with human selection for various physicochemical traits (Singh et al., Reference Singh, Bajpai, Singh, Ravishankar, Tandon and Reddy2010) and farmer-to-farmer seed exchanges (Liu et al., Reference Liu, Ge, Wang, Li, Yang, Cui and Qu2013). Given the considerable value of this underexplored species, the chosen genotypes will undergo further testing and preservation as invaluable genetic resources.
This study represents the first comprehensive exploration of wild jamun genotypes in the North-Western Himalayas. By combining robust RAPD and ISSR marker systems with morpho-physiological traits, we have unveiled valuable insights into the genetic relationships among wild population of Indian jamun across this region. The observed genetic diversity serves as a significant resource that holds great potential for enhancing various breeding programmes. The notable genetic distances found in this study highlight the substantial diversity present within the population under investigation. This diverse genotypic spectrum presents a valuable resource for targeted future endeavours.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S147926212300076X.
Acknowledgements
The authors would like to thank College of Horticulture and Forestry, University of Horticulture and Forestry, Nauni, Solan, India for the financial support granted for this research, the laboratory of Fruit Science and Molecular Biology and Biotechnology for providing equipment and reagents for this research.