Introduction
Limpets are ubiquitous and abundant invertebrates in aquatic environments, yet they remain comparatively understudied for parasitological research relative to other gastropod taxa. As molluscs, limpets play important roles in the transmission of digenean trematodes, serving either as first intermediate hosts harbouring sporocysts, rediae, and cercariae, or as second intermediate hosts harbouring metacercariae (Koppel et al. Reference Koppel, Leung and Poulin2011). Given their wide distribution and potential role in parasite transmission, investigating the diversity of trematodes infecting limpets can help resolve partially understood life cycles and provide predictive insights into the parasite diversity of local vertebrate hosts.
In New Zealand, fewer than 0.5% of marine gastropod taxa have documented helminth parasites (Bennett et al. Reference Bennett, Presswell and Poulin2022b). Although knowledge of marine helminth diversity has expanded in recent years (Bennett et al. Reference Bennett, Poulin and Presswell2022a), many life cycles remain incomplete (Bennett et al. Reference Bennett, Presswell and Poulin2023). The Otago Harbour mudflats, in New Zealand’s South Island, are known to support a rich diversity of trematode parasites (Leung et al. Reference Leung, Donald, Keeney, Koehler, Peoples and Poulin2009b). While previous biodiversity surveys in Otago’s coastal marine ecosystem have successfully matched various larval stages to adult parasites (Bennett et al. Reference Bennett, Presswell and Poulin2023), limpets (here including limpet-shaped gastropods) have been overlooked. Koppel et al. (Reference Koppel, Leung and Poulin2011) examined only one limpet species (Notoacmea scapha) and recovered metacercariae from three trematode species in Otago Harbour, highlighting limpets’ potential role as viable intermediate hosts and transmitters of parasites.
Of particular interest is the role of siphonariids (false limpets), which are known to host marine schistosomes (Digenea: Schistosomatidae) in Argentina, Australia, and Kuwait (Abdul-Salam and Al-Khedery Reference Abdul-Salam and Al-Khedery1992; Brant et al. Reference Brant, Loker, Casalins and Flores2017; Ewers Reference Ewers1961; Lorenti et al. Reference Lorenti, Brant, Gilardoni, Diaz and Cremonte2022). Schistosomes are dioecious blood flukes of birds and mammals. Unlike most trematodes, they lack a metacercarial stage, and cercariae leave the molluscan host and penetrate the definitive host (Gabrielli and Garba Djirmay, Reference Gabrielli and Garba Djirmay2023). Some species cause schistosomiasis in humans, while others can induce human cercarial dermatitis (HCD, or swimmer’s itch) when non-human schistosome cercariae accidentally penetrate human skin (Gabrielli and Garba Djirmay Reference Gabrielli and Garba Djirmay2023; Horak et al. Reference Horak, Mikes, Lichtenbergova, Skala, Soldanova and Brant2015). In New Zealand, outbreaks of HCD have been reported from freshwater avian schistosomes in the genus of Trichobilharzia (Davis et al. Reference Davis, Blair and Brant2022), but the diversity and zoonotic potential of marine schistosomes remain poorly understood. Recently, a schistosome (Schistosomatidae gen. sp. 1) was reported from the limpet Patelloida corticate in New Zealand (Bennett et al. Reference Bennett, Poulin and Presswell2022a). However, this host may have been misidentified and is likely Siphonaria australis. This represents only the second marine schistosome species recorded in New Zealand, the first being Ornithobilharzia canaliculata from the gull Larus dominicanus (Bennett et al. Reference Bennett, Presswell and Poulin2023; Rind Reference Rind1984). Therefore, examining these gastropods in greater depth could reveal the diversity, distribution, and public health potential of marine schistosomes in New Zealand.
In this study, we adopt a morphological definition of ‘limpet’, encompassing gastropods with a patelliform (limpet-shaped) shell regardless of taxonomic lineage. Our aims are to investigate the limpet species of Otago Harbour and molecularly identify the trematode parasites they harbour. We also conducted a comprehensive review of available information on trematode parasites infecting limpets in both marine and freshwater ecosystems worldwide, to provide an up-to-date synthesis of the trematode taxa using these molluscs as hosts.
Methods
Field collection and dissection
Limpets were collected by hand at low tide from five sites along Otago Harbour, South Island, New Zealand, between December 2024 and March 2025. These sampling sites include Lower Portobello Bay (latitude, longitude: −45.832203, 170.672795), Wellers Rock (−45.797962, 170.715378), Macandrew Bay (−45.868997, 170.596806), Broad Bay (−45.848534, 170.619328), and Company Bay (−45.856851, 170.598893). Sample sizes vary across sampling sites (Table 1). Prior to dissection, limpets were maintained in aerated containers with fresh seawater. Each limpet was removed from its shell using forceps, and internal organs were examined under a dissection microscope. Muscle tissue was firmly pressed between two glass plates and examined under the dissection microscope for parasites. All parasites recovered were preserved in 70% ethanol for subsequent DNA sequencing. Limpets were identified to species or genus using published keys (Carson and Morris Reference Carson and Morris2017; Jones et al. Reference Jones, Marsden, Holdaway and Jones2005), and through personal communication with expert molluscan taxonomist Prof. Hamish Spencer (Zoology Department, University of Otago, Dunedin, New Zealand).
Table 1. Parasite prevalence from the limpets collected at five Otago Peninsula sites facing the Otago Harbour

Parasite identification and haplotype network
Genomic DNA was extracted from individual cercariae or metacercariae using the DNeasy® Blood &Tissue Kit (Qiagen, Hilden, Germany) following the manufacturer’s protocol. Two genetic markers were targeted for amplification: a partial region of the 28S rRNA gene and the cytochrome c oxidase subunit I (cox1) gene. For the 28S rRNA gene, primers T16 and T30 (Harper and Saunders 2001) were used under the following thermal cycling conditions: initial denaturation at 94°C for 5 min; 38 cycles of denaturation at 94°C for 30 s, annealing at 45°C for 30 s, and extension at 72°C for 2 min; followed by a final extension at 72°C for 7 min. For the cox1 gene, primers JB3 (Bowles 1993) and Trem.cox1.rrnl (Kralova-Hromadova 2008) were used with the following protocol: initial denaturation at 95°C for 2 min; 40 cycles of denaturation at 95°C for 30 s, annealing at 48°C for 40 s, and extension at 72°C for 1 min; followed by a final extension at 72°C for 10 min. PCR products were cleaned using ExoSAP-ITTM Express PCR Product Cleanup Reagent (USB Corporation, Cleveland, OH, USA) following the manufacturer’s instructions. Sanger sequencing by capillary electrophoresis was performed by the Genetic Analysis Service, Department of Anatomy, University of Otago, New Zealand.
Raw sequences were trimmed using default parameters and manually curated to correct ambiguous bases in Geneious Prime® version 2025.2.2. Depending on the parasite, we either used the 28S gene or a combination of 28S and cox1 genes to identify species to the lowest taxonomic level possible through BLASTn (Basic Local Alignment Search Tool for nucleotides) searches against the NCBI GenBank database (https://blast.ncbi.nlm.nih.gov/). To examine the genetic structure of the schistosome, we constructed a haplotype network from aligned cox1 sequence data. Approximately 33 bp downstream of the cox1 stop codon were retained in the alignment due to their high sequence quality. Some sequences were shorter in length compared to others, but still included in the haplotype network with the assumption that they matched 100% to longer haplotype representatives. Sequences were aligned using the ClustalW algorithm implemented in the msa package (version 1.38.0) (Bodenhofer et al. Reference Bodenhofer, Bonatesta, Horejs-Kainrath and Hochreiter2015) within R Statistical Software (version 4.2.2; R Core Team, 2022). The haplotype network was constructed using the pegas package (version 1.3) (Paradis Reference Paradis2010) to visualize genetic relationships among haplotypes.
Literature review
To survey trematodes parasitising limpets, a topic search was conducted in Web of Science in July, 2025 using the following query: ‘(Trematod* OR Digenea OR Fluke OR Redia* OR Cercaria* OR Sporocyst* OR Metacercaria*) AND (Limpet* OR Patellogastropod* OR Patellid* OR Fissurellid* OR Lepetelloid* OR Siphonariid* OR Phenacolepadid* OR Tylodinid* OR Latiid* OR Latia OR Trimuscul* OR Ancylid*)’. For completeness, besides the ‘true limpets’ (family Patellidae), we also included in our review all major gastropod groups commonly referred to as ‘limpets’ in the literature (e.g., family Fissurellidae – keyhole limpets; family Siphonariidae – false limpets; family Calyptraeidae – slipper limpets). The title and abstract of all publications retrieved by the search were checked, and all relevant publications were retained. From each publication, we extracted the host species name, the trematode family and (if given) species name, the sampling locality, and (if available) the prevalence of infection.
Results
Trematode survey and molecular identification
We collected and examined four limpet taxa (Siphonaria australis Quoy & Gaimard, 1833, Cellana ornata Dillwyn, 1817, Cellana strigilis Hombron & Jacquinot, 1841, Notoacmea sp.) from five intertidal sites in Otago Harbour and identified two trematode species (Table 1). Siphonaria australis was the most common and abundant species, occurring at all five sites. Schistosome infections were detected in S. australis at every site, with a prevalence of sporocysts ranging from 2.6% to 100%, though the 100% prevalence was based on a sample size of only two limpets from Lower Portobello Bay. Notoacmea sp. were found at four sites, but only individuals from Lower Protobello Bay hosted metacercaeriae of Acanthoparyphium sp. A. (11.4% prevalence). No parasites were detected in any of the Cellana ornata and C. strigilis examined.
Cercariae from the false limpet S. australis were identified as Gigantobilharzia cf. patagonense (Figure 1). The 28S rRNA sequence obtained (~560 bp, GenBank accession PX833732) differed by a single nucleotide (99.83% identity) from Gigantobilharzia patagonense (syn. Marinabilharzia patagonense) described from the kelp gull Larus dominicanus in Argentina (GenBank accessions OK338634-OK338636) (Lorenti et al. Reference Lorenti, Brant, Gilardoni, Diaz and Cremonte2022). It also shared a 99.83% identity with Schistosomatidae sp. recorded from L. dominicanus (KX302891) and Siphonaria lessonii (KX302889) in Argentina (Brant et al. Reference Brant, Loker, Casalins and Flores2017), and a 99.81% identity (89% query coverage) with a sequence from Patelloida corticata in New Zealand (ON661331) (Bennett et al. Reference Bennett, Poulin and Presswell2022a). Additionally, the sequence shared 98.91% identity with Schistosomatidae sp. from penguin Spheniscus demersus from South Africa (KM023789) (Aldhoun and Horne Reference Aldhoun and Horne2015). For the cox1 gene, our consensus sequence shared 96.69% identity (44% query coverage) with G. patagonensis from the brown-hooded gull Chroicocephalus maculipennis (OK338770) and 95.83% identity (55% query coverage) with G. patagonense from L. dominicanus (OK338769) (Lorenti et al. Reference Lorenti, Brant, Gilardoni, Diaz and Cremonte2022).

Figure 1. Photomicrographs of the cercaria of Gigantobilharzia cf. patagonensis showing (A) dorsal view and (B) lateral view.
Metacercariae found infecting Notoacmea sp. at Lower Portobello (Table 1) were identified as Acanthoparyphium sp. A. Our 28S rRNA sequence was identical (100% match) to various representatives of Acanthoparyphium sp. A of (Leung et al. Reference Leung, Keeney and Poulin2009a) recorded from New Zealand intertidal species, including cockle Austrovenus stutchburyi (OQ407748), the green chiton Chiton glaucus (OQ407746), and the limpet Notoacmea scapha (OQ407744) (Bennett et al. Reference Bennett, Presswell and Poulin2023). It also shared 98.72% identity with isolates of another unidentified Acanthoparyphium sp. from N. scapha (ON661320) (Bennett et al. Reference Bennett, Poulin and Presswell2022a).
Haplotype network analysis of Gigantobilharzia cf. patagonensis
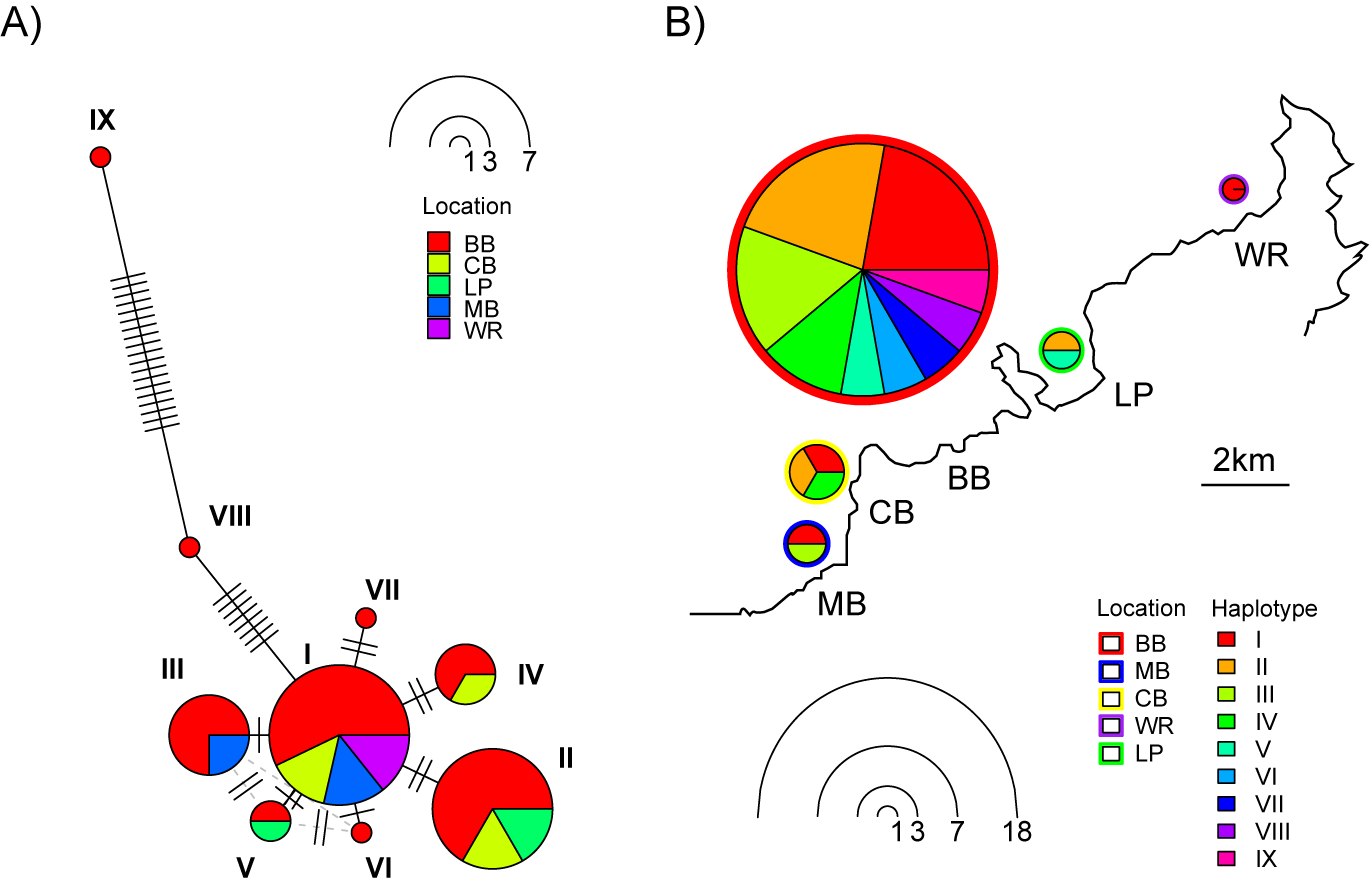
We identified nine unique G. cf. patagonensis haplotypes based on cox1 sequences (Figure 2A, accessions PX910024-PX910034). The network displayed a star-like topology dominated by haplotype I, which was the most common lineage and present in four of the five sampling locations (absent only from Lower Portobello) across Otago Harbour, New Zealand. Haplotypes III, V, and VI differed from haplotype I by 1bp, while haplotypes II, IV, and VII differed by 2 bp. Broad Bay exhibited the highest genetic diversity, containing all nine identified haplotypes. This included two divergent lineages found exclusively at this site: Haplotype VIII (8 mutational steps from haplotype I) and haplotype IX (a further 19 mutational steps away). Despite the variation, haplotypes were mixed among sites (Figure 2B), showing random sorting with no clear geographic structuring across the Otago Peninsula.

Figure 2. Haplotype network of Gigantobilharzia cf. patagonensis recovered from Siphonaria australis, based on cox1 sequences from five Otago Harbour sites (A) and the distribution of haplotypes by location (B). BB, Broad Bay; CB, Company Bay; LP, Lower Portobello; MB, Macandrew Bay; WR, Wellers Rock.
Literature survey of limpet-associated trematodes
The bibliographical survey recovered trematodes from 12 superfamilies and 23 families utilising marine or freshwater limpets as first or second intermediate hosts (Table 2). In the freshwater environment, trematode superfamilies included the Paramphistomoidea, Diplostomoidea, Schistosomatoidea, Echinostomatoidea, Monorchioidea, and Gorgoderoidea. In the marine environment, trematodes belonged to the superfamilies Gymnophalloidea, Hemiuroidea, Lepocreadioidea, Pronocephaloidea, Microphalloidea, and Opecoeloidea, as well as the Schistosomatoidea and Echinostomatoidea. Although the Schistosomatoidea and Echinostomatoidea superfamilies were found in both environments, no trematode families were shared between freshwater and marine systems. These trematodes utilised limpet-shaped hosts from nine families: three freshwater families (Planorbidae, Bumupiidae, Acroloxidae), and six marine families (Siphonariidae, Lottidae, Patellidae, Fissurellidae, Nacellidae, Calyptraeidae).
Table 2. Summary of trematode taxa reported from limpet hosts from previous publications

* IH-1 = First intermediate host; IH-2 = second intermediate host; DH = definitive host.
** FW, Freshwater; MR, marine.
The prevalence of trematode infection in freshwater environments ranged from 0 to 20%, with all recorded trematodes using limpets as first intermediate hosts. In marine environments, prevalence ranged from 0.1 to 100%, and many trematode species also use limpets as second intermediate hosts. Trematodes of the genus Proctoeces (family Fellodistomidae) can mature into egg-producing adult worms in their molluscan host, and thus also use limpets as their definitive host. Records of limpet-associated trematodes are most numerous in the United States for freshwater systems and in Argentina and Ireland for marine systems.
Discussion
The role of limpets (in the broad sense) in trematode life cycles remains poorly documented. Here, we identified two trematode species in intertidal limpets from Otago Harbour. The first is the avian schistosome, Gigantobilharzia cf. patagonensis, which uses Siphonaria australis as its first intermediate host. The second is Acanthoparyphium sp. A metacercaria, recovered from Notoacmea sp., used as a second intermediate host. Gigantobilharzia cf. patagonensis was recovered from all five sampling sites, with prevalence reaching up to 100%. Genetic analyses revealed considerable diversity of cox1 sequences within the Otago Harbour population. In addition, our literature review highlights the broad diversity of trematodes infecting limpets in both marine and freshwater systems around the world.
We report Siphonaria australis as a new first intermediate host record for Gigantobilharzia cf. patagonensis. Previous studies in Otago recovered matching sequences (Bennett et al. Reference Bennett, Poulin and Presswell2022a), but the host was identified as the true limpet Patelloida corticata. Given that schistosomes within the Gigantobilharzia clade are known to infect pulmonate gastropods (Siphonariidae) (Brant et al. Reference Brant, Loker, Casalins and Flores2017; Ewers Reference Ewers1961; Lorenti et al. Reference Lorenti, Brant, Gilardoni, Diaz and Cremonte2022), the earlier record likely reflects host misidentification, with S. australis representing the intermediate host in this region.
The Otago Harbour isolate is closely related to Gigantobilharzia patagonensis, which infects S. lessoni in Argentina (Brant et al. Reference Brant, Loker, Casalins and Flores2017; Lorenti et al. Reference Lorenti, Brant, Gilardoni, Diaz and Cremonte2022). Definitive hosts of G. patagonensis include the kelp gull Larus dominicanus in Argentina and the African penguin Spheniscus demersus in South Africa (Aldhoun and Horne Reference Aldhoun and Horne2015). In New Zealand, the life cycle involves the black-backed gull L. dominicanus (Bennett et al. Reference Bennett, Presswell and Poulin2023). As these gulls are non-migratory, introduction via bird migration is unlikely.
Analysis of the mitochondrial cox1 gene of G. cf. patagonensis revealed high haplotype diversity within the Otago Harbour population. The haplotype network displayed a star-like topology dominated by a single common lineage (haplotype I), suggesting a potential recent population expansion. Broad Bay exhibited the highest diversity, containing unique, divergent lineages. This genetic diversity suggests a large, stable parasite population within the harbour, likely maintained by the constant movement of avian definitive hosts dropping schistosome eggs across different coastal sites. Prevalence in S. australis (2.6–100%) was notably higher than that reported for G. patagonensis in S. lessoni from Argentina (0.9–6.1%) (Alda and Martorelli Reference Alda and Martorelli2009; Brant et al. Reference Brant, Loker, Casalins and Flores2017). However, extreme values in our dataset may result from small sample sizes (e.g., 100% prevalence at Lower Portobello Bay was based on only two limpets, and our highest number of haplotypes came from Broad Bay, which has the highest number of limpets examined).
Avian schistosomes can cause HCD when their cercariae penetrate human skin. While freshwater outbreaks associated with Trichobilharzia are well documented (Davis et al. Reference Davis, Blair and Brant2022; Horak et al. Reference Horak, Mikes, Lichtenbergova, Skala, Soldanova and Brant2015), marine species also pose risks to humans (Ewers Reference Ewers1961). Compared with freshwater schistosomes, marine taxa remain less studied worldwide (Brant and Loker Reference Brant and Loker2013; Khosravi et al. Reference Khosravi, Thieltges, Shamseddin and Georgieva2022); however, genera such as Ornithobilharzia have previously been reported in New Zealand (Bennett et al. Reference Bennett, Presswell and Poulin2023; Rind Reference Rind1984). The molecular confirmation of Gigantobilharzia in five bays within Otago Harbour expands the known diversity of marine schistosomes in the region. Its presence in intertidal zones highlights a potential public health concern if this species has zoonotic potential, and underscores the need for monitoring schistosome populations in recreational coastal environments. In the context of climate change, swimmer’s itch is likely to become increasingly prevalent in cool, temperate areas like New Zealand (Soleng et al. Reference Soleng, Gundersen and Lindstedt2025), reinforcing the importance of clinical awareness and improved knowledge of parasite distribution and host knowledge.
Metacercariae recovered from Notoacmea sp. were identified as Acanthoparyphium sp. A of Leung et al. (Reference Leung, Keeney and Poulin2009a) (Echinostomatoidea: Himasthlidae). The 28S rRNA sequences matched 100% with isolates previously recorded from the New Zealand cockle Austrovenus stutchburyi and the chiton Chiton glaucusi (Bennett et al. Reference Bennett, Presswell and Poulin2023). Acanthoparyphium sp. A was also reported in Notoacmea scapha from Lower Protobello Bay in 2010, where limpets were suggested to act as dead-end hosts unlikely to transmit metacercariae to suitable definitive hosts, such as oystercatchers, due to their small body size (Koppel et al. Reference Koppel, Leung and Poulin2011). Finally, we found no trematode infection in either of the other two limpet species examined from Otago Harbour, Cellana ornata and C. strigilis; however, the numbers of specimens of these species that could be collected and examined were too low to conclude that they are not used as hosts by trematodes.
Our literature survey revealed that limpets play important roles in trematode life cycles across 12 superfamilies and 23 families. Ecological roles of limpets differ between freshwater and marine environments. Freshwater limpets (Planorbidae, Burnupiidae, Acroloxidae) serve exclusively as first intermediate hosts, whereas marine limpets (Siphonariidae, Lottidae, Patellidae, Fissurellidae, Nacellidae, Calyptraeidae) can serve as both first and second intermediate hosts. No trematode families exploit both marine and freshwater limpets, although members of superfamilies Echinostomatoidea and Schistosomatoidea occur in both environments. Prevalence tends to be higher in limpets serving as second intermediate hosts, likely resulting from accumulating metacercarial encystment, whereas first intermediate host infections, which often result in castration and increased mortality due to cercariae asexual amplification, occur at much lower prevalences.
Different limpet families exhibit distinct parasitism profiles. For example, Siphonariidae (air-breathing false limpets) are key first intermediate hosts for marine Schistosomatidae and Hemiuridae, and second intermediate hosts for Microphallidae (Gilardoni et al. Reference Gilardoni, Etchegoin, Diaz, Ituarte and Cremonte2011). In contrast, the Fissurellidae (keyhole limpets) are often exploited by Fellodistomidae, specifically the genus Proctoeces. Notably, Proctoeces lintoni is reported to exhibit progenesis (precocious maturity) in Fissurellidae in Chile, whereby limpets act as alternative definitive hosts alongside clingfish (George-Nascimento et al. Reference George-Nascimento, Balboa, Aldana and Olmos1998).
Although sample size varied among limpet species and sampling sites and may influence prevalence estimates or the likelihood of detecting rare haplotypes, this study provides important insights into the diversity and ecology of marine schistosomes in New Zealand. We documented high G. cf. patagonensis prevalence and genetic diversity in Otago Harbour, thus highlighting potential public health risks of the zoonotic disease HCD. Future work can expand geographic sampling and incorporate definitive host screening to resolve life cycles fully and assess the distribution of G. cf. patagonensis and the risk of HCD in recreational coastal environments.
Acknowledgements
We would like to acknowledge Priscila Salloum and Lilou Mayeur for their assistance with field collections. We also thank Hamish Spencer for sharing his expertise in limpet identification.
Financial support
This work was supported by a Ministry of Business, Innovation and Employment Endeavour Fund Emerging Aquatic Diseases: A novel diagnostic pipeline and management framework, Award no: CAWX2207, recipient: Cawthron Institute.
Competing interests
The authors declare no conflict of interest.