Embracing diverse genetic data plays an important role in advancing our understanding of the genetic contribution to the development of diseases and ensuring health equity when translating genetic findings into practice. The presence of evolutionary driving forces, including genetic drift, mutation, migration, natural selection, and recombination, has led to genetic structure differences between populations, which might create disparities in genetic association findings. Including genetic data from diverse backgrounds, therefore, can help improve our knowledge of genetic contributions to diseases and inferences across populations. Furthermore, this can avoid potential bias resulting from using single-population genetic findings and prevent health disparities for underrepresented populations.
Many public genetic variation catalogues have attempted to capture genetic diversity across various geographic areas and/or ethnic groups (Fairley et al., Reference Fairley, Lowy-Gallego, Perry and Flicek2020; International HapMap Consortium, 2003; 1000 Genomes Project Consortium et al., Reference Auton, Brooks, Durbin, Garrison, Kang, Korbel, Marchini, McCarthy, McVean and Abecasis2015). However, no single reference panel covers all potential populations of research interest and simultaneously provides genetic data with large sample sizes. In addition, while the majority of current ancestry inference tools have been designed and validated to work on estimating continental or broad geographical populations (Alexander et al., Reference Alexander, Novembre and Lange2009; Price et al., Reference Price, Patterson, Plenge, Weinblatt, Shadick and Reich2006; Pritchard et al., Reference Pritchard, Stephens and Donnelly2000), studies on the transferability of genetic findings such as polygenic risk scores (PRSs) have shown inconsistent results that occur not only in different superpopulations but also at the subcontinental level (Gola et al., Reference Gola, Erdmann, Läll, Mägi, Müller-Myhsok, Schunkert and König2020; Kerminen et al., Reference Kerminen, Martin, Koskela, Ruotsalainen, Havulinna, Surakka, Palotie, Perola, Salomaa, Daly, Ripatti and Pirinen2019). This impedes the adoption of genetic research in clinical practice, especially in ethnically diverse countries. Furthermore, some of these groups cannot be fully captured by population labels from existing reference panels such as 1000 Genomes. Hence, it may be advantageous to include a larger number of subcontinental populations for ancestry estimation.
The UK Biobank is a vast resource of genetic and phenotypic data that has made significant contributions to health studies (Sudlow et al., Reference Sudlow, Gallacher, Allen, Beral, Burton, Danesh, Downey, Elliott, Green, Landray, Liu, Matthews, Ong, Pell, Silman, Young, Sprosen, Peakman and Collins2015). The cohort contains participants from various ethnic backgrounds (self-reported) and countries of origin, and as such represents a potential repository of population diversity across the world. Moreover, the UK Biobank population clusters defined in a previous study have been observed to correlate with regions of birth (Constantinescu et al., Reference Constantinescu, Mitchell, Zheng, Bull, Timpson, Amulic, Vincent and Hughes2022), which is also in alignment with the correspondence of genetic variations and geographical locations in general. This underpins the potential of the UK Biobank as a reference dataset for ancestry estimation on a global scale. There are several studies that used unsupervised algorithms (Constantinescu et al., Reference Constantinescu, Mitchell, Zheng, Bull, Timpson, Amulic, Vincent and Hughes2022; Diaz-Papkovich et al., Reference Diaz-Papkovich, Zabad, Ben-Eghan, Anderson-Trocmé, Femerling, Nathan, Patel and Gravel2023) and identity-by-descent (IBD)-based haplotype clustering (Gilbert et al., Reference Gilbert, Shanmugam and Cavalleri2022) to capture the population structure of the UKB data at the subcontinental level. The limitation of these approaches is their exploratory nature, meaning that the clusters formed are not predetermined and are significantly dependent on tuning parameters (e.g., the chosen number of clusters, distance metric, and length of IBD segments) and the patterns of genetic data. Although additional information can be used to identify the common patterns of resulting clusters, formed clusters may only present the specific patterns of the studied dataset and not align with predefined population categories. This unpredicted discrepancy makes it challenging to compare and/or integrate results from different studies and datasets, limiting the practical significance of these findings (Constantinescu et al., Reference Constantinescu, Mitchell, Zheng, Bull, Timpson, Amulic, Vincent and Hughes2022; Diaz-Papkovich et al., Reference Diaz-Papkovich, Zabad, Ben-Eghan, Anderson-Trocmé, Femerling, Nathan, Patel and Gravel2023; Gilbert et al., Reference Gilbert, Shanmugam and Cavalleri2022) within the studied cohort (i.e., UKB), and hindering their transferability to new data. In contrast, supervised learning using a reference panel can provide more interpretable results, with outputs directly corresponding to specific reference samples, facilitating the generalizability and transferability of the clustering approach across different datasets, while being less computationally intensive compared to the IBD-based method. This is especially beneficial for the common practice of genetic epidemiology where studies have been conducted in discrete population groups. By leveraging UKB data as a reference dataset for a supervised model, we can apply a consistent clustering approach to different datasets, making the most efficient use of the available data resources to group individuals who are genetically similar to each other and the reference samples. This consequently assists with performing genetic analyses such as genomewide association studies (GWAS) or PRSs in populations of interest, especially underrepresented population groups.
Hence, we propose a clustering approach that applies a supervised classification algorithm and dimensionality reduction technique to the UK Biobank genetic data to cluster individuals with high genetic similarity. It is important to clarify here that we are not introducing a novel method, but rather a practical approach to exploit existing biobank data for finer grouping of genetically similar individuals. Group labels were defined using a combination of self-reported ancestries and countries of birth information. By leveraging an available source of genetic data, we aim to assign more fine-scaled population labels for a wider range of individuals, extending beyond those that fall into the groups represented in 1000 Genomes. This will ultimately help alleviate bias arising from the underrepresentation of populations in the reference panel and improve the accuracy of genetic similarity clustering.
Although the term ‘ancestry’ is widely used and ancestry estimation is a critical step in genetic analyses, ancestry definition is loosely defined and therefore can lead to confusion or misinterpretation. A recent publication discussed the statements of genetic similarity and genetic ancestry groups in research on human genetics (Coop, Reference Coop2023). The article recommended using genetic similarity to provide better descriptions of samples’ genomic data (Coop, Reference Coop2023). The US National Academies of Sciences, Engineering, and Medicine also proposed the use of genetic similarity as preferred population descriptors in all types of genetics and genomics research (National Academies of Sciences, Engineering, and Medicine, 2023). Furthermore, they stated that ‘any description of the genetic ancestry of an individual entails a decision about the relevant time depth at which to describe it’, while the ancestry grouping process in genetic studies was often derived from measuring genetic similarity instead of direct observation (National Academies of Sciences, Engineering, and Medicine, 2023). Therefore, in this study, we did not emphasize inferring the detailed genetic ancestry of each individual. Instead, we focused on identifying sets of individuals who are more genetically similar to each other, bearing in mind that this represents only an approximation to the true continuous nature of genetic variation. However, we considered this approach to be more practical and feasible for the downstream application of clustering study participants for population-based studies, as it allows us to account for genetic differences among populations in the development and validation of genetic testing, facilitating its translation to practice while avoiding the computational complexity of treating genetic variation as a continuum. Accordingly, all of the following ancestry inference results should be understood as ‘most genetically similar to the sample X in the reference panel’. Additionally, it is important to note that all labels used to refer to our results, as well as previous findings we mentioned, should not be taken as ‘genetic ancestry’ but rather as ‘study population/sample’ of respective studies and reference panels, which describe the shared characteristics of participants.
Materials and Methods
UK Biobank Data and Labeling Genetically Similar Groups
We used the UK Biobank dataset with available imputed genetic data (N = 487,180 individuals) as the reference for population clustering based on genetic similarity. The cohort covers individuals from four main self-reported ethnic background groups: White, Asian or Asian British, Black or Black British, and Mixed. Included in each group are different populations: British and Irish in White; Indian, Pakistani, Bangladeshi, and Chinese in Asian; Caribbean and African in Black. Initially, we combined the self-reported ethnic background (Data-Field 21000) and the country of birth (non-UK origin; Data-Field 20115) to create country-based labels. Each group was constructed to contain individuals with self-reported ethnic-label subgroups compatible with countries of birth. For countries located in areas with ethnic backgrounds that are not well defined in the UK Biobank’s ethnic background field, such as the Middle East and South America, we considered that participants’ answers could vary based on personal views or experiences. For example, a Latin American might identify himself as ‘White’, ‘Black’, ‘Mixed’, or ‘Other ethnic group’ in the UK Biobank Ethnic background field. Therefore, in these cases, only individuals who self-identified as one of the incompatible ethnic background groups (e.g., Asian in the case of Latin American) or subgroups such as British, Irish, Indian, Pakistani, Bangladeshi, or Chinese were removed from country-based groups. To ensure analysis power and accuracy, we excluded countries of birth with modern diverse-descent population structures and country-based groups with less than 50 individuals. After exclusion, 56 country-based population groups remained. Phylogenic tree and admixture plots from FRAPPE (FRequentist APProach for Estimating individual ancestry proportion), a software program that estimates founding allele frequencies and individual admixture using maximum likelihood model, separated different regional ancestral groups including Sub-Saharan African, North African-Near East/Middle East, South/Central Asian, European, East Asian, South East Asian-Oceania, and American (natives) (Duda & Jan Zrzavý, Reference Duda and Zrzavý2016; Li et al., Reference Li, Absher, Tang, Southwick, Casto, Ramachandran, Cann, Barsh, Feldman, Cavalli-Sforza and Myers2008). Accordingly, we merged all country-based groups in the Sub-Sahara, North African, and Middle East regions into respective larger groups. For East Asian and South East Asian subclades, we only had one available group each, namely Chinese and Filipino. For the Central/South Asian subclade, we had three country-based groups, which remained in broader classified groups due to a lack of information about geography-based substructure. For the European subclade, a study on European genetic structure identified four main distinct regions, including (1) Finland, (2) the Baltic region, Eastern Russia, and Poland, (3) Central and Western Europe, and (4) Italy. Additionally, correlating genetic distance and geography, they also created four regional barriers separating (1) Finland and the rest, followed by (2) South Italy and the other, then (3) Lithuania, Poland, and Western Russia from Bulgaria, and finally (4) Baltic region and Poland from Sweden and the remainder (Nelis et al., Reference Nelis, Esko, Mägi, Zimprich, Zimprich, Toncheva, Karachanak, Piskácková, Balascák, Peltonen, Jakkula, Rehnström, Lathrop, Heath, Galan, Schreiber, Meitinger, Pfeufer, Wichmann and Metspalu2009). Considering these results and the insular and peninsular geography, we reclassified our European country-based groups into eight main groups, namely, (1) Finnish, (2) Baltic and Eastern European, (3) Balkan (except Greek), (4) Northern and Western European (except British and Irish), (5) Iberian, (6) Italian, (7) Greek, and (8) British and Irish. The remaining country-based groups only included countries in South America and thus were merged into one Latin American group. In general, using research findings of genetic substructures and population differentiation within continents and data availability, we merged the original 56 country-based into 19 groups for further genetic similarity clustering analysis. These 19 population labels were then used as target classes for a multiclass random forest model that was trained on principal components of genetic data to assign individuals’ genetically similar labels.
CARTaGENE Data
To validate our model, we used data from CARTaGENE (https://cartagene.qc.ca/en/; Awadalla et al., Reference Awadalla, Boileau, Payette, Idaghdour, Goulet, Knoppers, Hamet and Laberge2013), a population-based biobank of around 40,000 individuals in Québec, Canada. CARTaGENE contains a diverse population with participants and their parents coming from 258 countries and 12 ethnicities (self-reported) covering all 19 genetically similar population groups we created. First, we used the father’s and mother’s countries of birth as the main criteria to assign individuals to population groups. People who had individuals’ or parents’ self-reported ethnicity that did not match the assumed ethnicity derived from countries of birth were removed. For example, an individual with both parents born in Barbados but had East Asian ethnicity information was excluded. We also removed individuals where their (or their parents’) first languages learned, or languages spoken most often at home were not the official languages of Canada or their countries of birth. Individuals who were not able to be classified as any of our 19 groups were likewise omitted.
1000 Genomes Phase 3 Data
To further validate our model, we used data from the 1000 Genomes Project, a global catalogue of genetic variations present in the human population. Phase 3 of the project provides 26 ethnic/ethnolinguistic populations in five geographic regions (Supplementary Table S1; Sudmant et al., Reference Sudmant, Rausch, Gardner, Handsaker, Abyzov, Huddleston, Zhang, Ye, Jun, Fritz, Konkel, Malhotra, Stütz, Shi, Casale, Chen, Hormozdiari, Dayama, Chen and Korbel2015). Besides using 1000 Genomes as validating data, we also employed it as a reference panel for genetic similarity clustering. More specifically, we utilized 1000 Genomes 26 population labels as target classes for a random forest model trained on genetic principal components to identify genetically similar groups of individuals in testing data, namely CARTaGENE. We then used 1000 Genomes as the reference panel and applied the same approach of combining the random forest algorithm and principal component analysis of genetic data to further evaluate the role of population representation in clustering genetically similar populations.
Statistical Analysis
Merging training and validating dataset
We used UK Biobank and 1000 Genomes imputed data and CARTaGENE directly genotyped data derived from the Global Screening Array (GSA) for the genetic analyses of this study. The hard-called genotypes for imputed data were assigned based on the highest genotype probability. Since the majority of UK Biobank data participants are British and Irish, we avoided the significant imbalance among classified genetically similar groups derived from the UK Biobank reference by randomly selecting 1000 British and 1000 Irish individuals as the training samples for genetic analysis. Before merging, the training data and the validating data were checked for strand, alleles, position, reference/alternative allele assignment, allele frequencies, variant ID, palindromic and duplicated SNPs to ensure consistency using https://www.chg.ox.ac.uk/~wrayner/tools/. Subsequently, the training genetic data (e.g., UK Biobank imputed data) were filtered to keep only variants in the validating dataset (e.g., CARTaGENE GSA-based directly genotyped data). Data then were merged using Plink v1.9 (Purcell et al., Reference Purcell, Neale, Todd-Brown, Thomas, Ferreira, Bender, Maller, Sklar, de Bakker, Daly and Sham2007).
Quality control procedure
After merging the prepared data, variant genotypes were filtered for individual missing call rate (<5%) and minor allele frequency (MAF; >0.05). Subsequently, we removed individuals with low genotype call rate (<90%) before performing linkage disequilibrium (LD) pruning to create an independent SNP set for further analysis. A 200-kilobase window size, a 50-kilobase step size, and an r 2 threshold of .25 were used to reduce the SNP set to avoid potential bias arising from the confounding effects of LD when capturing genetic variation across populations. The quality control and LD pruning process were performed using Plink v1.9.
Estimating genetically similar groups
Using the set of cleaned independent SNPs in the merged data, we ran principal component analysis (PCA) to summarize genetic variation across different population groups with reduced data dimensionality. Subsequently, we inferred genetic similarity information by applying the random forest approach to 80 principal components of cleaned genotype data, a sufficiently large number to capture genetic variation among different groups. PCA was performed using Plink v1.9 and the random forest model was run via the randomForest() R package (Liaw & Wiener, Reference Liaw and Wiener2022) with the default setting of 500 decision trees and bootstrap sampling, which generated satisfactory precision and recall in validation datasets. Individuals’ most genetically similar groups were identified by majority voting. The accuracy of the approach was validated using precision and recall rate.
Measures of fixation index (FST)
To quantify the genetic differentiation between 19 labeled populations in the training UK Biobank data, we calculated pairwise FST between samples using Plink v2.00a5LM. For each sample, we removed variants with missing call rates exceeding 10%, MAF below 1%, and Hardy-Weinberg equilibrium exact test p-value below 10-6. We also excluded from each sample individuals with missing call rates above 10% and possible cryptic relationships (pi-hat > 0.25). We then merged postcleaning data from each sample. The set of variants that were genotyped on all individuals was kept for FST calculation. We measured pair-wise FST between 19 labeled populations using the Weir-Cockerham method. Standard errors were estimated with the block size of 50 adjacent variants.
We applied the same approach to calculate pairwise FST between 19 demographic-based imputed populations in the CARTaGENE data and between 26 available populations in 1000 Genomes data.
Results
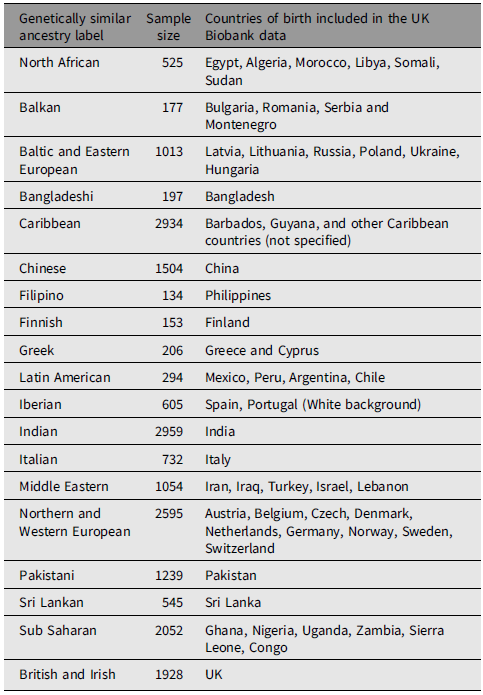
Combining the ethnic background and country of birth information as the new standard for population labeling, we identified 19 genetically similar groups represented in the UK. We applied geography-based names to the 19 groups, including North African, Greek, Balkan (except Greek), Baltic and Eastern European, Bangladeshi, Caribbean, Chinese, Filipino, Finnish, Latin American, Iberian (comprising Spanish and Portuguese), Indian, Italian, Middle Eastern, Northern and Western European (except British and Irish), Pakistani, Sri Lankan, Sub Saharan, British and Irish. The total dataset used as the reference panel comprises 20,846 individuals, with the sample size of each group ranging from 134 to 2959 (details in Table 1).
Training data for random forest model
We applied the random forest model to estimate genetically similar groups based on principal components of genetic data. The training model generated an out-of-bag (OOB) error of 9.87%, which was relatively low considering the setting of the 19-group classification. The confusion matrix demonstrating the detailed classifiers’ predictions for unseen training data is shown in Supplementary Table S2. To measure genetic differentiation between these 19 clusters, we calculated pairwise FST between samples (Supplementary Table S3). Among sub-European clusters, Finnish was the most distinct population, with paired FST values between Finnish and any of the other sub-European groups greater than 0.005. Among intercontinental groups, considerable overlapping values were observed between South Asian groups, Latin American, North African, Middle Eastern, and European populations, as well as between Chinese and Filipino groups, with FST ranging from 0.01 to 0.04.
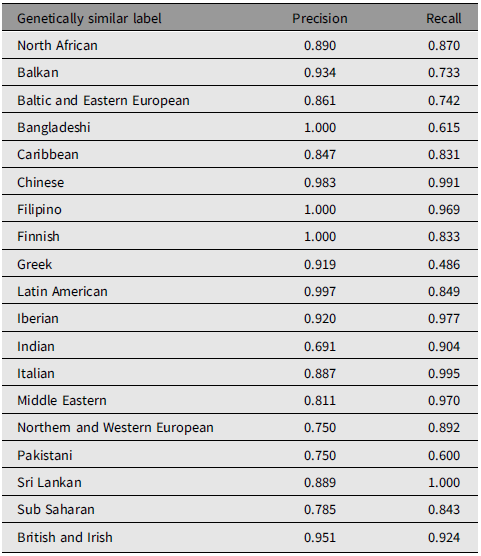
We validated our prediction model on two independent cohorts, CARTaGENE, and 1000 Genomes. Genetically similar clustering performance was evaluated via precision and recall rate. Although CARTaGENE data contains nearly 40,000 individuals, genetic data needed for clustering is only available in 29,248 individuals. We only imputed population labels for this subset of people using the father’s and mother’s countries of birth as the main criteria. Of these 29,248 individuals, 26,350 had parents whose countries of birth either were not covered in 19 UK Biobank-derived population labels (e.g., Canada, USA, Malaysia) or belonged to two different labelled groups and therefore were excluded from validation. The remaining 2898 individuals fell into one of the 19 groups we created. While CARTaGENE participants’ population labels were imputed using the 19-label classification derived from the UK Biobank, the 1000 Genomes project covers 26 populations based on ethnic/ethnolinguistic backgrounds that do not exactly match our training 19 classes. However, the majority of these are associated with countries or regions that we used to create genetically similar labels. Therefore, ESN, GWD, LWK, MSL, YRI were expected to be in the Sub-Saharan group; GBR in British and Irish; CEU in Northern and Western European; FIN in Finnish; IBS in Iberian; TSI in Italian; BEB in Bangladeshi; GIH, ITU in Indian; PJL in Pakistani; STU in Sri Lankan; CDX, CHB, CHS in Chinese; ACB in Caribbean; CLM, MXL, PEL, PUR in Latin American. The three 1000 Genomes populations absent from the training samples, including KHV (Kinh in Vietnam), JPT (Japanese), and ASW (African American), were excluded when calculating precision and recall.
The validation results showed the possibility of identifying people from groups that are underrepresented in commonly used reference panels such as Filipino, Middle Eastern, Baltic and Eastern European, and Balkan people. In CARTaGENE, precision for all genetically similar groups ranged from 69.1% to 100%, while recall exceeded 60%, except for Greek (Table 2). In particular, 11 out of 19 populations demonstrated good performance with both precision and recall of greater than 80%. Results are specifically high for populations with distinct genetic profiles such as Finnish and Filipino. In contrast, Greek had a high precision, at 91.9%, but a low recall of only 48.6% since half of the Greek individuals were assigned to the Italian group (confusion matrix in Supplementary Table S4). Looking further at pairwise FST between demographic-based imputed populations in CARTaGENE (Supplementary Table S5), Greek and Italian clusters showed little differentiation, with an FST value of 0.0004 (SE = 4.66e-05). Similarly, a large proportion of Bangladeshi and Pakistani individuals were predicted to be most genetically similar to the Indian group, resulting in low recall rates in Bangladesh (61.5%) and Pakistan (60%) and low precision in the Indian group (69.1%). The Pakistani cluster also exhibited minimal differentiation from the Indian group in both training data (FST = 0.0005, SE = 5.77e-05) and the validating CARTaGENE data (FST = 0.0010, SE = 0.0002).
Genetic similarity clustering accuracy in CARTaGENE
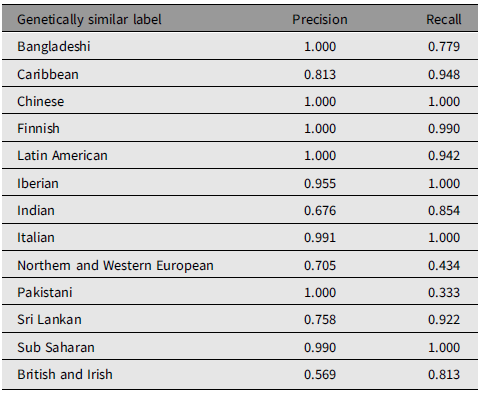
In 1000 Genomes, 23 populations were assigned to one of 13 genetically similar groups as expected. Genetic similarity clustering performance across all populations is illustrated in Table 3. Among the 13 groups analyzed, Chinese, Finnish, Latin American, Iberian, Italian, and Sub-Saharan achieved both precision and recall of greater than 90%. Low precision in the Indian group (67.6%) and low recall in the Pakistani group (33.3%) were observed, which is similar to the validation results in CARTaGENE. In addition, more than half of the CEU population was assigned to the British and Irish group (confusion matrix in Supplementary Table S6), leading to low precision in the British and Irish group (56.9%) and low recall in the Northern and Western Europe group (43.4%). However, the CEU population, which represents people residing in Utah with Northern and Western European origin, may include Irish. Moreover, the observed results may be attributed to factors like location sampling, environmental influences, and migration patterns, all of which could be encapsulated by the genetically similar label. Indeed, the CEU and GBR population in 1000 Genomes showed an extremely low level of genetic differentiation, with an FST value of 0.0003 (SE = 1.76e-05; Supplementary Table S7).
Genetic similarity clustering accuracy in 1000 Genomes
To further evaluate the role of population representation in clustering genetically similar individuals, we applied the same approach of integrating multiclass random forest and PCA of genetic data using 1000 Genomes phase 3 as the reference dataset to cluster genetically similar individuals in CARTaGENE (confusion matrix results in Supplementary Table S8). Among 19 imputed populations in CARTaGENE, over 90% of individuals in Bangladeshi, Finnish, Latin American, Iberian, Pakistani, and Sub-Saharan groups were assigned to 1000 Genomes’ most closely related populations. Expected results were also seen in partially overlapping groups. For example, 50.6% of Caribbean individuals were more similar to ACB, whereas 31.0% were more similar to ASW compared to other reference samples. British and Irish individuals were mainly estimated to be most genetically similar to the GBR (71.4%) and CEU (26.7%) groups. All Indian individuals were predicted to be most genetically similar to South Asian samples, although only 26.9% were assigned to Indian samples, including ITU and GIH. Considering population groups that were not present in the 1000 Genomes reference samples, 93.8% of Filipino individuals were most genetically similar to the KHV sample, while 98.6% of Greek and 99.6% of Middle Eastern individuals were most genetically similar to the TSI sample. Meanwhile, the majority of the three remaining absent groups, North African, Balkan, Baltic and East European, were assigned to different European populations. We increased the prediction threshold to evaluate whether individuals in absent populations were removed from incompatible groups. After replacing majority voting with the prediction probability threshold of 0.8, all North African and 99.3% Middle Eastern individuals were removed from the prediction. However, 25% of Filipinos and 58% of Greeks remained assigned to KHV and TSI samples, respectively. The results suggest that using UK Biobank data enables us to identify missing populations in 1000 Genomes, thus providing a more comprehensive population reference that allows us to match individuals to more genetically similar groups.
Discussion
By referring to the UK Biobank dataset as the reference, we identified reliable categorizations for a wide range of populations that are present in the UK. With 19 group labels, we covered a more diverse set of genetically similar groups from all over the world compared to existing datasets such as 1000 Genomes. It is now possible to identify people from populations that are absent in commonly used reference panels (e.g., the Middle East, the Philippines, the Baltic states and Eastern Europe, and the Balkans). Validating results on CARTaGENE and 1000 Genomes data showed satisfactory precision and recall rates of greater than 70% for most of the labeled groups. Nevertheless, it is still challenging to distinguish closely related groups such as Greeks from Italians, and Bangladeshi and Pakistani from Indians. A study on European population genetic substructure reported the overlap between Italian American and Greek American (FST = 0.000, SD = 0.0011) as well as close relatedness between Greek American and Tuscany (FST = 0.001, SD = 0.0025; Tian et al., Reference Tian, Kosoy, Nassir, Lee, Villoslada, Klareskog, Hammarström, Garchon, Pulver, Ransom, Gregersen and Seldin2009), which is consistent with the historical event of Greek colonization in Southern Italy (Tofanelli et al., Reference Tofanelli, Brisighelli, Anagnostou, Busby, Ferri, Thomas, Taglioli, Rudan, Zemunik, Hayward, Bolnick, Romano, Cali, Luiselli, Shepherd, Tusa, Facella and Capelli2016). Similar fixation indexes were observed in our Italian and Greek clusters in both training data (FST = 0.0010, SE = 2.29e-05) and validating data (FST = 0.0004, SE = 4.66e-05). This can be the explanation for the overlapped prediction results of Greek and Italian individuals that we observed. In the case of India, Bangladesh, and Pakistan, the past and ongoing migration patterns with shared genetic history make it challenging to disentangle genetic differences among the three country-based populations. Genetic distance between Northwest Indian and Pakistani populations was also found to be lower than the measures for Northwest Indian and other North Indian groups (Pathak et al., Reference Pathak, Kadian, Kushniarevich, Montinaro, Mondal, Ongaro, Singh, Kumar, Rai, Parik, Metspalu, Rootsi, Pagani, Kivisild, Metspalu, Chaubey and Villems2018).
Replicating the same approach but using 1000 Genomes as a reference dataset demonstrated the importance of population representation in improving genetic similarity clustering. Typical examples are the results for Filipino and Greek. The prediction model using 1000 Genomes reference, which did not cover Filipino and Greek samples, assigned Filipino and Greek individuals in CARTaGENE to KHV (Vietnam) and TSI (Italy) groups respectively with high probabilities. This indicates that high prediction probability does not necessarily mean an individual is a member of that population. Hence, using more comprehensive reference populations is the key to achieving more accurate genetic similarity clustering and avoiding reference population bias. However, in cases where the reference does not adequately represent targeted populations, the results are still reasonable when interpreting appropriately: for example as, ‘among all population samples in the 1000 Genomes dataset, the individual is most genetically similar to the TSI/KHV sample’. Furthermore, the lack of the Middle Eastern population in 1000 Genomes is a big limitation that we tried to overcome by leveraging the UK Biobank as a reference dataset. Located at the intersection of Europe, Asia, and Africa, the Middle East has a long history of intercontinental migration and trading exchange, all of which are likely to enrich the genetic diversity of the population (Elliott et al., Reference Elliott, Haber, Daggag, Busby, Sarwar, Kennet, Petraglia, Petherbridge, Yavari, Heard-Bey, Shobi, Ghulam, Haj, A., Tikriti, Mohammad, Antony, Alyileili, Alaydaroos, Lau and Barakat2022). From the other perspective, the cultural practice of endogamy, consanguineous marriage, and the prevalence of extended family structures have led to a disproportionately high prevalence of Mendelian recessive disorders (Abou Tayoun et al., Reference Abou Tayoun, Fakhro, Alsheikh-Ali and Alkuraya2021; Elliott et al., Reference Elliott, Haber, Daggag, Busby, Sarwar, Kennet, Petraglia, Petherbridge, Yavari, Heard-Bey, Shobi, Ghulam, Haj, A., Tikriti, Mohammad, Antony, Alyileili, Alaydaroos, Lau and Barakat2022). Due to its unique genetic patterns, the Middle Eastern population holds great potential for studying disease genetics and evaluating the implementations of preventive medicine, early diagnostics, and timely intervention (Abou Tayoun et al., Reference Abou Tayoun, Fakhro, Alsheikh-Ali and Alkuraya2021), making it an important study population for genetic and genomic research.
The number of clusters we created for random forest classification to generate satisfactory precision and recall rates was lower than the cluster number obtained using the Uniform Manifold Approximation and Projection (UMAP) and the Hierarchical Density-Based Spatial 88 Clustering of Applications with Noise (HDBSCAN; Diaz-Papkovich et al., Reference Diaz-Papkovich, Zabad, Ben-Eghan, Anderson-Trocmé, Femerling, Nathan, Patel and Gravel2023; 19 vs. 26) and the IBD-based method (Gilbert et al., Reference Gilbert, Shanmugam and Cavalleri2022); 8 versus 41 sub-European populations). However, while these unsupervised approaches identified more fine-scale population structures within the UK Biobank cohort, distinguishing formed clusters from each other and labeling them proved challenging. For example, clusters 3 and 4 created by the UMAP-HDBSCAN approach exhibited overlapping patterns, with the majority of individuals being born in England and having a white background (including both British and other white backgrounds). Similarly, high proportions of both clusters 17 and 20 were born in England (>80%) and had a British background (>90%). Clusters 9 and 14, on the other hand, contained individuals from various ethnic backgrounds, complicating their characterization. Regarding the IBD-based clustering, among 41 sub-European populations, 15 groups included British and Irish members without further distinguishable characteristics. The lack of clear labels and characterization when using these unsupervised approaches limits the use of the UK Biobank as a reference dataset for genetic similarity clustering in new data and generally hinders the comparison and integration of resulting clusters across different datasets. Although the IBD-based method produced hierarchical results with broader, distinguishable clusters, which were similar to our created clusters, haplotype construction is more computationally intensive, especially for large-scale studies.
Therefore, while these unsupervised clustering approaches reveal more detailed population structures within the UK Biobank, our approach of leveraging UK Biobank data as a reference dataset for genetic similarity clustering offers greater generalizability and transferability in genetic epidemiology studies. It provides a more straightforward means to cluster individuals who are genetically similar to the predefined reference populations of study interest in new data without requiring additional characterization. This allows better comparison and integration across different studies, as well as better matching of genetically similar individuals across different datasets using the same reference. Our approach, hence, holds promising use in not only handling population stratification but also selecting external control for various downstream genetic analyses. Population stratification, which involves differences in allele frequencies, has significant impacts on variant-trait association in GWAS. By providing more detailed genetic similarity information compared to existing resources such as 1000 Genomes, our approach can assist with handling population structure at a smaller scale and correcting its confounding effects. Furthermore, with the inclusion of diverse, predefined reference populations, our approach may also be useful in allocating controls from different datasets to perform GWAS and PRS in underrepresented populations, reducing health disparities and improving overall healthcare outcomes. This is beneficial, especially for diverse nations such as Australia, Canada, the USA, and Fiji. We took Australia as an example. According to the Australian 2021 Census of Population and Housing, Australia was populated by residents of over 300 different ancestries. Noticeably, among the key groups that formed relatively large proportions of the Australian population were underrepresented populations, namely Greek, Italian, Filipino, and Lebanese. Our approach can assist with clustering these populations in genetic studies, facilitating the translation of genetic findings into practice for a wider range of populations.
Our approach has several limitations. First, countries of birth and self-reported ethnic backgrounds at a broad scale are incomplete information to assign ancestry labels. However, the UK Biobank has limited demographic information that we can use to impute individuals’ origins. In addition to that, collecting information on ethnic backgrounds at a subregional scale is difficult. The views of ethnic backgrounds and ancestries may vary among individuals and people may not be aware of their recent detailed ancestral origins, leading to potential subjective or inconsistent responses. Second, we cannot include references for some population groups due to the limited availability of data. Our UK Biobank reference sample only provides one genetically similar ancestry group in East Asia (China) and one in South East Asia (Filipino). Regarding Africa, the most genetically diverse region, we ended up having two genetically similar groups. The rationale behind this division is that genetic studies on population structures and differentiation in Sub-Saharan Africa have often been correlated with ethnolinguistic groups (Gurdasani et al., Reference Gurdasani, Carstensen, Tekola-Ayele, Pagani, Tachmazidou, Hatzikotoulas, Karthikeyan, Iles, Pollard, Choudhury, Ritchie, Xue, Asimit, Nsubuga, Young, Pomilla, Kivinen, Rockett, Kamali and Sandhu2015), making it challenging for us to provide more detailed ancestry groups and validate our approach with available data. Third, the genetic data we used were derived from the UK Biobank participants and might not adequately represent the original population’s genetic variation. Migration flows to the UK may follow specific patterns, such as immigrants coming from particular regions or due to historical events and settling in particular areas. Examples include Caribbean immigrants from the Windrush generation, who were mainly Jamaican. As a result, the principal components we calculated solely reflect the genetic patterns of our sample, but cannot fully capture the genetic diversity of the original populations. Finally, despite the availability of both 1000 Genomes and UK Biobank as references, indigenous populations such as American Indians/Alaska Natives, Australian Aboriginal and Torres Strait Islanders, Māori, and Canadian First Nation people remain uncharacterized in our study. Several barriers, including historical causes for hesitance, different perspectives in communication and decision-making, access concerns related to geographical distance, and cultural and spiritual beliefs, have limited the inclusion of Indigenous people in genetic research and data (Waanders et al., Reference Waanders, Brown, Caron, Plisiewicz, McHugh, Nguyen, Lehmann, Stevens, Storm, Resnick, Davidson, Mueller and Kline2023). Regardless of these challenges, it is essential to include Indigenous people in genetic data to ensure diversity and equity in healthcare research and practice.
Supplementary material
To view supplementary material for this article, please visit https://doi.org/10.1017/thg.2025.15.
Data availability
UK Biobank data is available through the UK Biobank Access Management System https://www.ukbiobank.ac.uk/. The CARTaGENE data is available for access request on the study website https://cartagene.qc.ca/. The 1000 Genomes data is available through the Phase 3 release https://www.internationalgenome.org/category/phase-3/.
Acknowledgments
This research has been conducted using data from the UK Biobank (application number 25331) and the CARTaGENE Biobank (project ID: 488635). We thank the participants and investigators of the CARTaGENE study and the UK Biobank study.
Financial support
Funding was provided by the Australian National Health and Medical Research Council (grant number 1150144).
Competing interests
The authors have no conflicts of interest to declare that are relevant to the content of this article.
Ethics approval
This study utilized de-identified data from UK Biobank, CARTaGENE, and 1000 Genomes. UK Biobank obtained ethics approval from the North West Multi-centre Research Ethics Committee (MREC) (reference number 06/MRE09/65). At the time of enrollment, all participants provided informed consent to take part in the UK Biobank and to be monitored using a signature capture device. CARTaGENE obtained ethics approval from the CHU Sainte-Justine (reference number MP-21-2011-345, 3297). Written consent was acquired from all participants in CARTaGENE. Ethical considerations and informed consent for the 1000 Genomes Project can be found at https://www.internationalgenome.org/sites/1000genomes.org/files/docs/Informed%20Consent%20Background%20Document.pdf.





