Introduction
Economic evaluation models are key tools for health technology assessment (HTA) (Reference Briggs, Claxton and Sculpher1), as they support policy-makers and HTA practitioners in comparing the costs and health outcomes of alternative interventions. For methodological rigor and relevance, such models require a coherent integration of clinical, epidemiological, and economic knowledge. Traditionally, building these models has been resource-intensive, often requiring substantial time and specialized expertise, due to extensive data gathering and expert interpretation.
Ontologies, defined as formal representations of domain knowledge, have been proposed as a solution (Reference Prieto-González, Castilla-Rodríguez, González-González and de la Luz Couce-Pico2), providing a semantic framework to define entities such as diseases, interventions, costs or outcomes, and their relationships (Reference Kuziemsky and Lau3;Reference Zhang, Gou and shu Zhou4). This structure enhances consistency, interoperability, and reusability, allowing HTA practitioners to standardize model conceptualization across evaluation contexts.
In parallel, generative artificial intelligence (GenAI), particularly large language models (LLMs), has transformed knowledge creation and management (Reference Alavi, Leidner and Mousavi5). LLMs efficiently summarize, extract, and generate domain-specific content, thereby reducing manual effort in evidence synthesis, automating processes that previously required substantial expertise (Reference Lee, Bubeck and Petro6).
This article explores how combining ontologies with GenAI enables scalable and automated ontology population. Ontologies are conceived not as static repositories but as dynamic guides for LLMs, whose outputs are constrained by semantic rules to generate relevant, context-aware individuals. This addresses a critical bottleneck in HTA modeling: the costly and error-prone manual curation of domain-specific evidence.
Accelerating the synthesis of accurate clinical and economic data requires expert review of literature and guidelines, creating a gap that limits scalability. GenAI’s natural language capabilities can mitigate this by automatically extracting and synthesizing knowledge. Yet controlling LLM behavior remains difficult due to their stochastic nature and the risk of hallucinations, defined here as the generation of content that is syntactically plausible but factually incorrect, unsupported by evidence, or inconsistent with the target knowledge base.
Prompt engineering has improved control over LLM outputs through structured prompts and iterative refinement (Reference Liu, Yuan and Fu7). However, its use for ontology-guided individual generation is still underexplored. Most GenAI healthcare research focuses on text generation and decision support (Reference Bhuyan, Solanki and Malik8;Reference Biswas9), whereas robust methodological frameworks for systematic model population remain scarce (Reference Ouédraogo, Tapsoba, Sabane, Koné, Sere and Kouamé10;Reference Brank, Grobelnik and Mladenic11), limiting reproducibility and reliability in regulated HTA domains.
This study proposes combining the expressive power of ontologies with the generative capacity of LLMs, steered by prompt engineering, to streamline the development of evaluation models. We hypothesize that ontologies can structure and constrain LLM output, producing valid, semantically rich individuals aligned with the target ontology, Ontology for Simulation of Diseases (OSDi) (Reference Castilla Rodríguez and González12). An experimental framework was designed to generate individuals via ontology-driven prompts, testing strategies for syntactic validity, semantic completeness, and practical utility in accelerating health economic evidence synthesis.
The remainder of the paper is organized as follows: Section “State of the art” reviews prior work on ontology-based modeling, GenAI, and prompt engineering; Section “The OSDi ontology” introduces the OSDi ontology; Section “Methodology” details the methodology and evaluation criteria; Section “Results and discussion” presents and discusses results; and Section “Conclusions and further work” concludes with implications and future directions.
State of the art
GenAI systems (exemplified by ChatGPT-4, Claude, Perplexity, and Mixtral) are transforming healthcare by automating data integration and analysis (Reference Moulaei, Yadegari and Baharestani13). This enables advances in evidence synthesis, personalized medicine, and population health management (Reference Xu and Wang14;Reference Rouzrokh, Alkhaldi and Mohammadi15). Current LLMs can efficiently extract terminologies, uncover relationships, and generate context-sensitive knowledge (Reference Aggarwal, Salatino, Osborne and Motta16–Reference Taboada, Rivas and Martinez18). In this sense, LLMs are increasingly being used in evidence screening and synthesis, allowing to automate health economic modeling (Reference Reason, Rawlinson and Langham19). Indeed, ISPOR has established a dedicated group reflecting the growing importance of this topic (https://www.ispor.org/member-groups/task-forces/genai-for-heor-slrs-task-force).
Recent research on GenAI in healthcare focuses mainly on automated report generation, question answering, decision support, and evidence synthesis. Reviews highlight both the benefits (efficiency, automation, triage) and risks (hallucinations, bias, privacy breaches) of these tools. Chustecki (Reference Chustecki20) summarizes opportunities and regulatory challenges, whereas Panteli et al. (Reference Panteli, Adib and Buttigieg21) emphasize GenAI’s role in public health surveillance and communication, noting concerns about equity, privacy, and governance.
In HTA, Fleurence et al. (Reference Fleurence, Bian and Wang22) and Reason et al. (Reference Reason, Klijn and Rawlinson23) offer insights on how GenAI supports literature reviews and evidence synthesis. Studies such as Qureshi et al. (Reference Qureshi, Shaughnessy and Gill24), Reason et al. (Reference Reason, Benbow and Langham25), and Li et al. (Reference Li, Deng and Sun26) show that LLMs can summarize and extract information efficiently but still require human validation. Gartlehner et al. (Reference Gartlehner, Kahwati and Hilscher27) and Schopow et al. (Reference Schopow, Osterhoff and Baur28) report similar findings: high accuracy but a need for semi-automated workflows. Szabó et al. (Reference Szabó, Pinsent and Slim29) explore data extraction from cost-effectiveness models in HTA reports, finding limited performance in complex modeling assumptions.
Prompt engineering emerges as a key skill for the reliable use of GenAI in medicine (Reference Meskó30–Reference Wang, Jiang and Zeng32). Well-designed, iterative prompts can enhance domain specificity and compliance with medical standards. Recent frameworks, including programmatic and feedback-driven approaches, improve model consistency while balancing computational efficiency (Reference Taboada, Rivas and Martinez18;Reference Ng31;Reference Wang, Jiang and Zeng32).
However, none of these studies address how LLMs can interact with formal semantic technologies. Ontologies structure knowledge through defined entities (diseases, interventions, costs, outcomes) and relationships, facilitating integration across domains. Frameworks like SNOMED CT (https://www.snomed.org/) demonstrate how ontologies enhance decision support, predictive analytics, and population health systems. Ambalavanan et al. (Reference Ambalavanan, Snead and Marczika33) highlight challenges in scalability, governance, and privacy, suggesting AI-assisted ontology maintenance as a future direction. In health economics, automatically generating ontology-aligned individuals could streamline decision trees and simulations, promoting standardized evaluations among HTA agencies. Persistent challenges include ensuring semantic accuracy, reducing bias, and maintaining ethical oversight (Reference Howell34).
Emerging studies combine LLMs with ontologies for enrichment, text-to-OWL transformation, and alignment, showing potential but lacking methodological consistency. Ouédraogo et al. (Reference Ouédraogo, Tapsoba, Sabane, Koné, Sere and Kouamé10) propose an automated pipeline where ChatGPT-3 converts treatment guidelines for multidrug-resistant tuberculosis into OWL axioms, producing a richer ontology than a semi-automated baseline. Complementarily, prompt engineering research provides structured methodologies such as the Goal-Prompt-Evaluation-Iteration (GPEI) framework by Velásquez Henao et al. (Reference Velásquez Henao, Franco Cardona and Cadavid35), which guides iterative prompt refinement through defined objectives, evaluation, and correction.
The OSDi ontology
The ontology used in this study is OSDi (Reference Castilla Rodríguez and González12), an extension of the earlier RaDIOS ontology (Reference Prieto-González, Castilla-Rodríguez, González-González and de la Luz Couce-Pico2). OSDi provides a flexible semantic framework to represent knowledge about diverse diseases and healthcare interventions, organizing clinical and economic concepts into classes and relationships that capture disease characteristics, progression, and intervention effects. For its development from the earliest versions of the research, the authors adopted selected steps from the NeOn methodological framework (Reference Suárez-Figueroa36), because it offers a set of flexible alternatives that can be tailored to the specific needs of the ontology developer. Among its proposed scenarios, “Scenario 1” which focuses on developing an ontology from scratch without relying on pre-existing knowledge resources aligns with the objectives of our work. The process in this case comprised the following main steps: (i) domain and scope definition, (ii) identification of core concepts and relationships, (iii) ontology formalization in OWL, (iv) validation and refinement with domain experts, and (v) ontology population with individuals. Section “Methodology” focuses in more detail on step (vi), namely the semi-automated creation of ontology individuals using LLMs, which constitutes a key contribution of this work.
At its core, OSDi models disease dynamics through classes such as Development, Stage, and Manifestation, allowing for the representation of disease presentation, evolution, and severity (e.g., distinguishing mild from severe developments or defining symptom probabilities at different stages). Disease progression is further detailed through Pathways, which describe alternative routes between stages or manifestations, conditioned by previous events, interventions, or probabilistic parameters.
The ontology also represents Populations, Interventions, and Health resources involved in HTA, enabling the formal specification of clinical follow-up and treatment processes.
Moreover, OSDi provides mechanisms to represent Parameters used in health economic modeling, including resources, distributions, rates, and risks. It incorporates utilities and disutilities, that is, quantitative measures of health-related quality of life, typically ranging from 0 (death) to 1 (perfect health), or the corresponding loss due to disease, adverse events, or side effects.
Compared with RaDIOS, OSDi adds greater detail to parameter characterization, distinguishing between FirstOrder, SecondOrder, and DeterministicParameters, and allowing CalculatedParameters defined by expressions. These enhancements enable not only the storage of numerical values but also the encoding of statistical properties and inferential rules, thereby supporting richer and more realistic simulation models.
Overall, OSDi standardizes the description of disease modeling components, from clinical presentation and progression to uncertainty parameterization and intervention effects. It is particularly suited to economic evaluation models, such as decision trees and discrete-event simulations, providing a reusable structure for health economics.
Currently, OSDi includes individuals for two diseases: profound biotinidase deficiency (PBD) and type 1 diabetes mellitus (T1DM). PBD is an autosomal recessive disorder of biotin metabolism manifesting in childhood, diagnosed via biochemical, enzymatic, and genetic tests, and treated with biotin supplementation (Reference Wolf37). The corresponding individuals (https://w3id.org/ontologies-ULL/OSDi/individuals/PBD.ttl) replicate the PBD submodel from Vallejo-Torres et al. (Reference Vallejo-Torres, Castilla and Couce38). T1DM, in contrast, is a chronic autoimmune disease marked by beta-cell destruction and absolute insulin deficiency (Reference Eisenbarth39), managed through continuous insulin therapy and glucose monitoring. Its individuals (https://w3id.org/ontologies-ULL/OSDi/individuals/T1DM.ttl) mirror the model by Castilla-Rodríguez et al. (Reference Castilla-Rodríguez, Arnay, González-Cava, Bruzzone, Frascio, Longo and Novak40).
Methodology
The methodology combines factorial experimental design with prompt engineering techniques (Reference Velásquez Henao, Franco Cardona and Cadavid35). A full-factorial setup was used to generate ontology individuals representing disease progression (pathogenesis, stages, manifestations) and their associated utilities and disutilities for PBD, forming the basis for estimating health indicators such as life expectancy and quality-adjusted life-years.
LLM interactions were conducted through the Perplexity AI platform (https://www.perplexity.ai/) under a Pro subscription, which provides controlled experimental “spaces” for uploading and querying documents. Both the Perplexity deep research model and ChatGPT-4.1 (https://chatgpt.com/) (Reference Shvets, Murtazin, Piho and Meeter41) were used. The platform’s ability to cite sources proved valuable, especially when no external references were supplied. Each experiment included contextual information and explicit task instructions for the model.
AI-generated individuals were incorporated into separate ontology versions while maintaining the shared OSDi structure. Ontology management and editing were performed using Protégé (Reference Musen42) and Visual Studio Code (https://code.visualstudio.com/). Integration involved adding AI-generated individuals and relationships in OWL format, which were subsequently visualized and validated in Protégé.
Validation followed an expert-driven approach. A domain specialist (who was the developer of both the reference model and the original ontology individuals, with over 15 years of HTA experience) evaluated the GenAI outputs. Previously defined individuals were used as a reference and benchmark for assessing the completeness and semantic adequacy of the generated content.
Factorial design
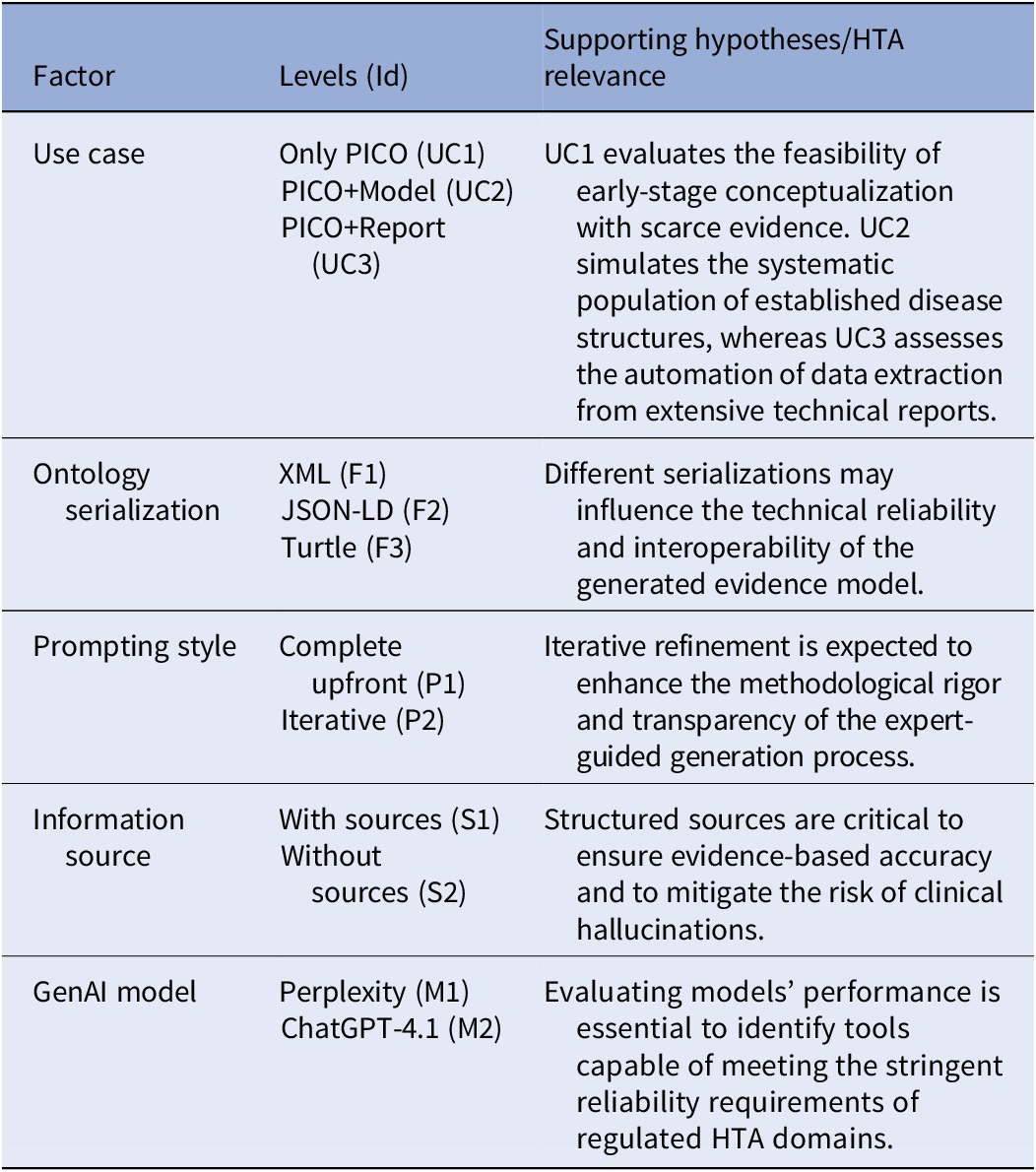
A factorial experimental design was implemented to evaluate how multiple variables influence the ability of GenAI models to generate valid ontology individuals. Based on an analysis of the problem involving both ontologies and GenAI systems, a set of relevant factors was identified, along with their levels for factorial design and the corresponding initial hypotheses, as summarized in Table 1. This design provides a structured framework to analyze how variables interact and jointly affect output quality.
Factorial matrix of factors and levels aligned with HTA modeling challenges

Three use cases were established to represent realistic scenarios in health economic modeling, each simulating different stages of the evidence synthesis process. All were formulated using the PICO framework (43), which is commonly used in evidence-based medicine to define the problem (P), intervention (I), comparator (C), and outcomes (O).
In the first case (UC1), designed to address exploratory or early-stage modeling where specific evidence is scarce, GenAI received only the PICO-formulated research question. This case tests the model’s ability to act as a preliminary conceptualization tool with minimal context.
In the second (UC2), the PICO question was accompanied by previously generated conceptual model individuals for PBD, representing a scenario where an initial disease structure is already established but requires systematic population of parameters.
In the third (UC3), GenAI was given a complete published technical report (Reference Vallejo Torres, Castilla-Rodríguez and Cuéllar Pompa44), reflecting the real-world task of extracting structured data from extensive HTA documentation to automate the transition from already published evidence to computational implementation.
Other experimental factors included the prompting style (iterative vs. single prompt), the inclusion or exclusion of specific information sources, and the choice of GenAI model (Perplexity deep research ChatGPT-4.1). Using explicit information sources, such as references on biotinidase deficiency (Reference Salbert, Astruc and Wolf45–Reference Grünewald, Champion, Leonard, Schaper and Morris53), allowed for comparison with free-text generation based solely on model knowledge. However, it was determined that the application of this factor made sense only in UC1: UC2 represented a scenario where it should be expected to have already collected proper references, and UC3 made use of a report with validated data and references.
A full-factorial combination of these factors would yield 72 experimental runs. However, serialization was added as an additional factor in a subsequent phase of the experimentation, when the version of Perplexity used in the previous experiments was not available anymore. Consequently, to ensure comparability, JSON-LD and Turtle were only tested with ChatGPT-4.1, thus reducing the number of experiments to 48. These figures were reduced in turn to 32 (16 runs for UC1, 8 for UC2, and 8 for UC3) due to the dependency among the information source factor and the use case.
Prompt design and interaction with GenAI
The interaction with GenAI models followed prompt engineering principles (Reference Velásquez Henao, Franco Cardona and Cadavid35), aiming to maximize the quality, reliability, and reproducibility of outputs. Prompt design adopted a modular structure, where each module controlled a specific aspect of the interaction (context, tasks, or information) enhancing the generation of valid ontology individuals. For each experiment, the total GenAI response time (excluding human interaction) and the number of corrections performed were recorded.
The main modules were: (i) a “role module,” assigning the model the role of a health economics expert specialized in HTA, ensuring consistent and domain-appropriate responses; (ii) a “context module,” describing the ontology’s structure and concepts, that is, clinical manifestations, interventions, parameters, utilities, and costs; (iii) a “PICO question module,” guiding the model through the PICO framework; (iv) an “information sources module,” used only in UC1, specifying whether external references or internal model knowledge should be applied; (v) a “task module,” defining objectives and expected ontology individuals; and (vi) an “iteration module,” determining whether the process followed a sequential or “one-shot” interaction. Iterative prompting, as noted in Velásquez Henao et al. (Reference Velásquez Henao, Franco Cardona and Cadavid35), improves accuracy and reduces hallucinations by enabling stepwise corrections. The number of individuals generated per interaction depended on the prompting strategy. In the single-prompt (non-iterative) setting, the model generated multiple individuals in a single response. In contrast, under the iterative prompting approach, individuals were generated progressively across successive interactions, with each iteration producing a subset of the required individuals.
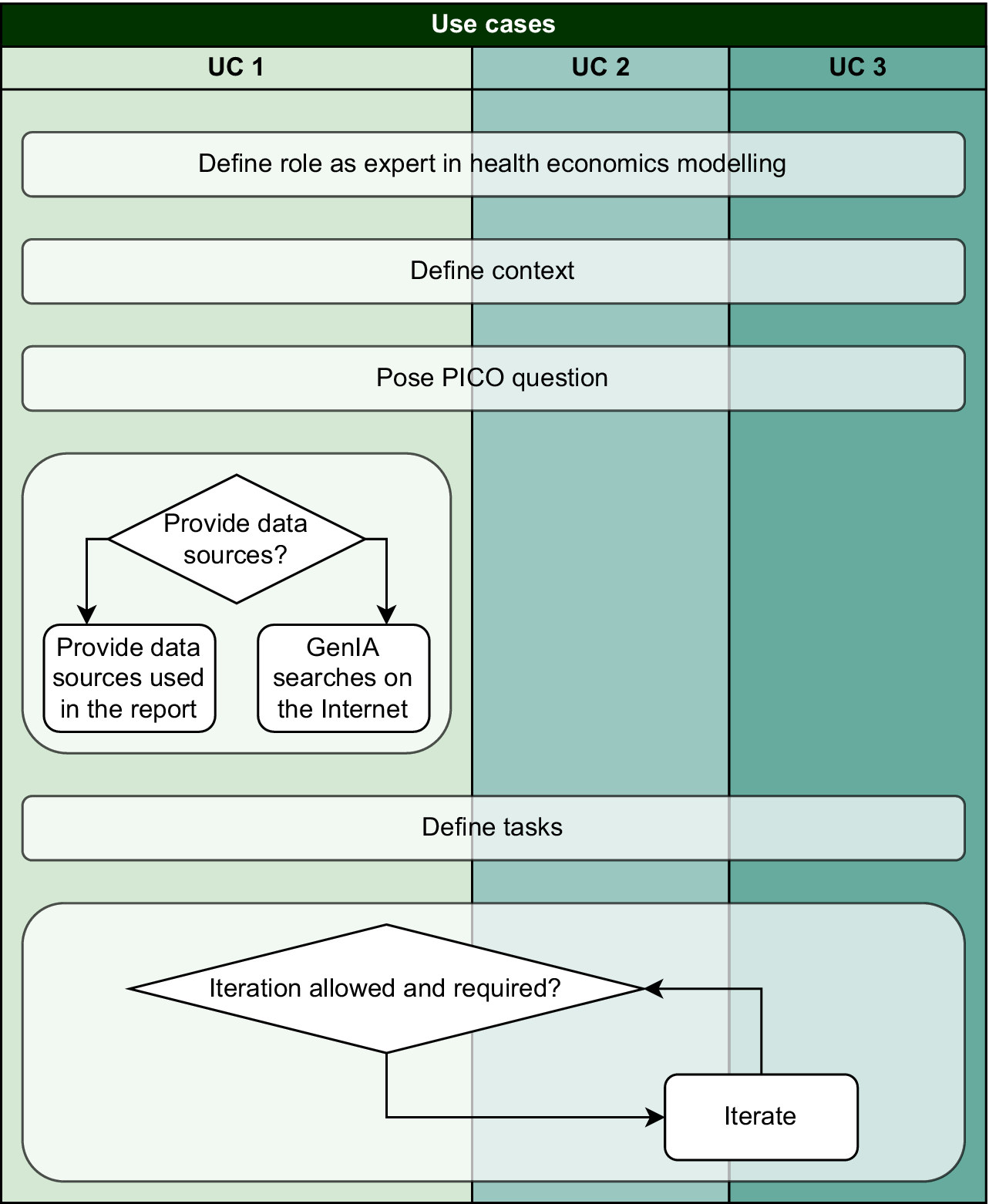
This modular architecture ensured flexibility and scalability across experimental conditions while maintaining methodological rigor. Figure 1 illustrates the general design, and full prompt details for each use case are provided in the supplementary material (Supplementary Material Appendix B).
Modular structure of prompt design used in the experiments.

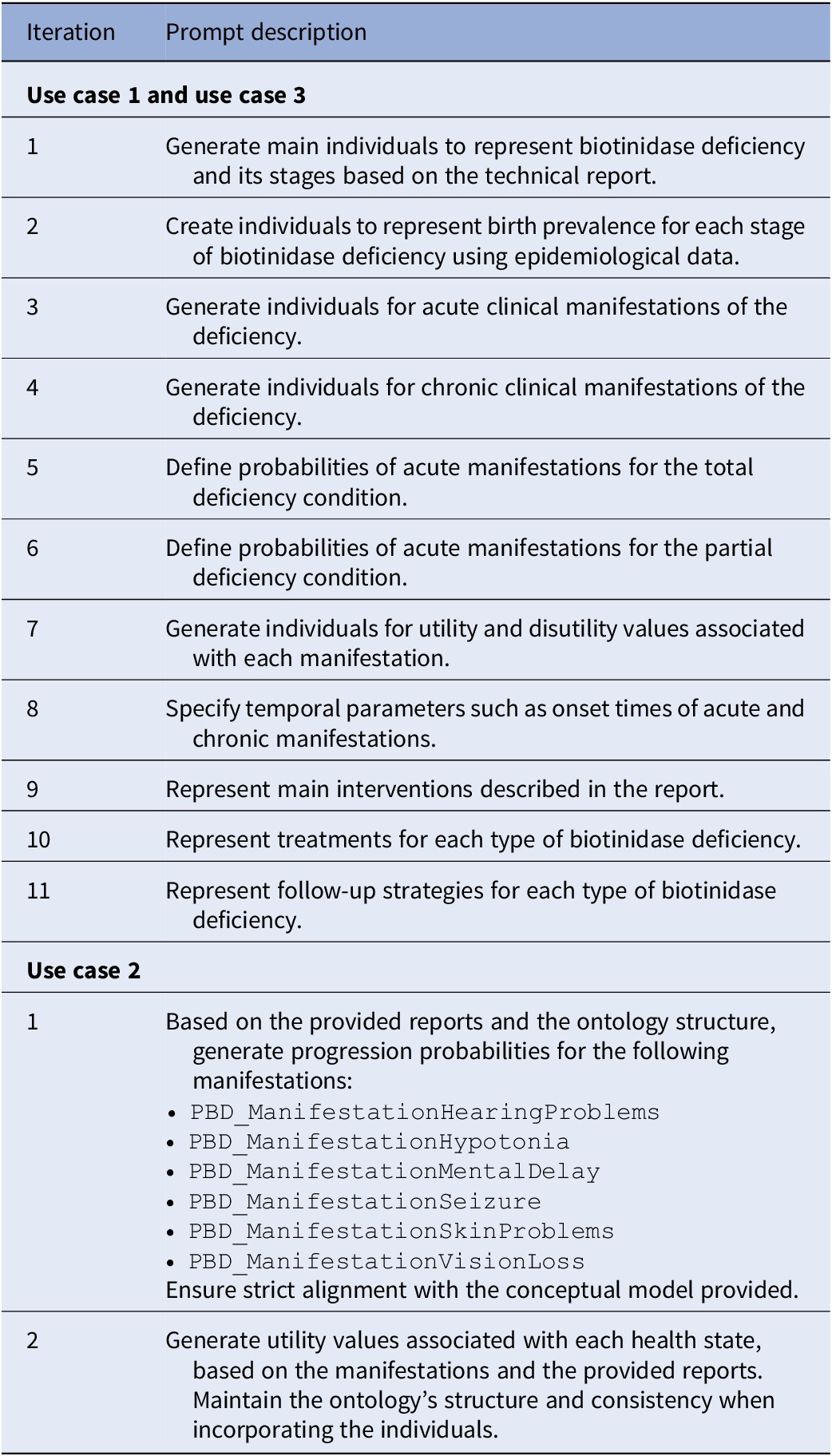
An iterative approach was used in Use Cases 1 and 3 (UC1, UC3), where GenAI progressively generated all individuals needed to represent biotinidase deficiency. In Use Case 2 (UC2), which already included a conceptual model, a simplified iterative procedure was applied to complete missing parameters. Table 2 summarizes the iterations for each case.
Prompts per iteration and use case

During early tests, models struggled to interpret OWL syntax, which led us to develop a corrective prompt. By providing explicit examples based on existing type 1 diabetes mellitus individuals, the corrective version helped GenAI better understand the ontology structure and relationships. This enhancement reduced errors and enabled the semi-automated generation of consistent and semantically valid individuals.
Statistical analysis of results
The analysis evaluated each experimental scenario to assess the quality of individuals generated by the models through qualitative and quantitative approaches. Most variables (both those defining the problem and those describing outcomes) were categorical, taking values from limited discrete sets that facilitated interpretation and statistical processing.
Validation involved two stages. First, an expert reviewed each of the above 1300 generated individuals, classifying them into categories (risks, manifestations, and interventions) and identifying hallucinations. Attending to the observed problems in the generated individuals, and based on existing literature on hallucinations in large language models (Reference Ji, Lee and Frieske54–Reference Alkaissi and McFarlane56), the expert classified the identified hallucinations into four categories:
-
i. Structural hallucinations: generating entities or properties that do not exist in the reference ontology, violating schema constraints.
-
ii. Semantic hallucinations: combining valid ontology terms in logically inconsistent ways, for example, violating domain, range, or class disjunction constraints.
-
iii. Contextual hallucinations: producing outputs that are structurally and logically valid but fail to meet input specifications, such as interventions for populations or conditions not requested.
-
iv. Evidence-based (factuality) hallucinations: producing literals, numerical values, or citations that appear plausible but are unsupported or factually incorrect according to clinical evidence.
After the individual validation, a holistic experiment assessment was performed. The qualitative assessment of each experiment involved four variables rated on a five-point Likert scale (1 = strongly disagree, 5 = strongly agree): “Consistency,” that is, internal coherence across individuals; “Relevance,” that is, alignment with the research question; “Completeness,” that is, coverage of all required concepts; and “Adequacy,” that is, practical usability without major revisions.
Cramer’s V was employed to assess associations between categorical variables, as it effectively captures nonlinear dependencies and accommodates features with multiple levels. This analysis was supplemented with contingency tables and heatmaps to further explore the direction of the identified relationships.
We assessed the internal consistency of the four Likert-rated quality dimensions using Cronbach’s alpha. Given the low internal reliability, we did not aggregate them into a single composite score. Instead, we conducted a multivariate analysis of variance (MANOVA) to test for global effects of the experimental factors on the joint quality profile, using Pillai’s trace as the primary statistic due to the small sample size and potential deviations from multivariate normality. To identify dimension-specific effects, we then estimated separate linear models for each outcome variable. All models were fitted using heteroscedasticity-robust (HC3) standard errors. Finally, we derived reduced, more parsimonious specifications by removing clearly non-informative predictors, balancing model fit (AIC/BIC), statistical stability, and consistency with the experimental design.
We finally explored the relationship between the experimental factors and the proportion of hallucinations. Because the dependent variable was a proportion, we fitted binomial logistic regression models with a logit link function, using the number of generated individuals as binomial weights. Given the evidence of overdispersion and consistently with previous analyses, we estimated heteroscedasticity-robust (HC3) standard errors.
A full list of variables used in the evaluation and statistical analyses is available in the supplementary material (Supplementary Material Appendix A).
Results and discussion
Two out of the 32 planned experiment runs had to be discarded because the output of the GenAI was impossible to parse and convert into a usable ontology format.
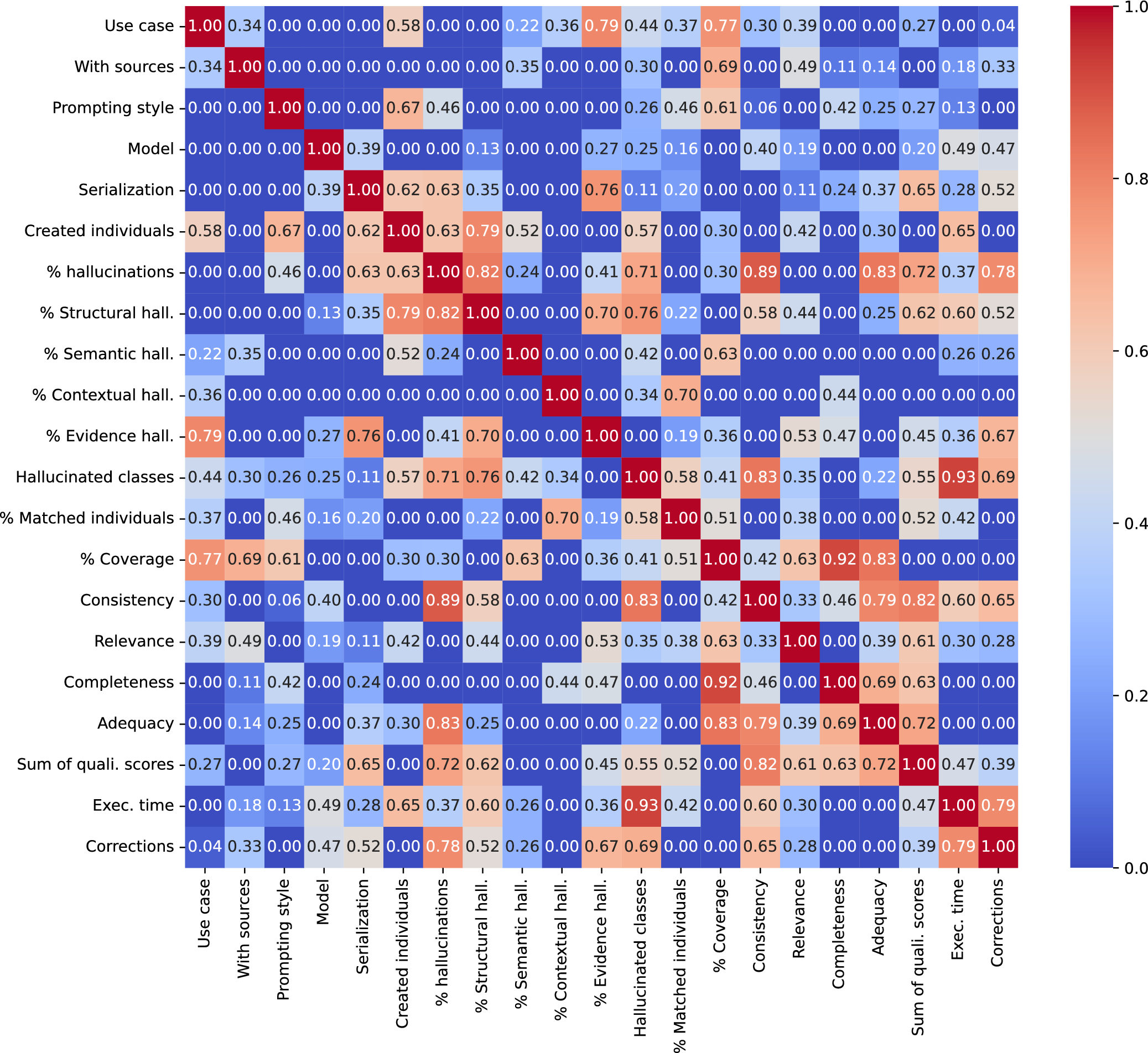
The preliminary analysis with Cramer’s V (Figure 2) helped identifying a series of key findings:
-
i. The use case presents a strong association with the percentage of evidence hallucinations, number and coverage of the created individuals, but seems independent from other types of hallucinations, completeness, or adequacy.
-
ii. Providing sources is related to the coverage of concepts and, moderately, to relevance, but barely related to any other of the qualitative assessments.
-
iii. The prompting style shows a strong association with the number of created individuals and the coverage.
-
iv. The model used shows a moderate association with execution time and consistency.
-
v. Serialization is related to hallucinations (especially structural and evidence ones), the total qualitative score, and also to the number of corrections.
Cramer’s V heatmap for the evaluated variables.

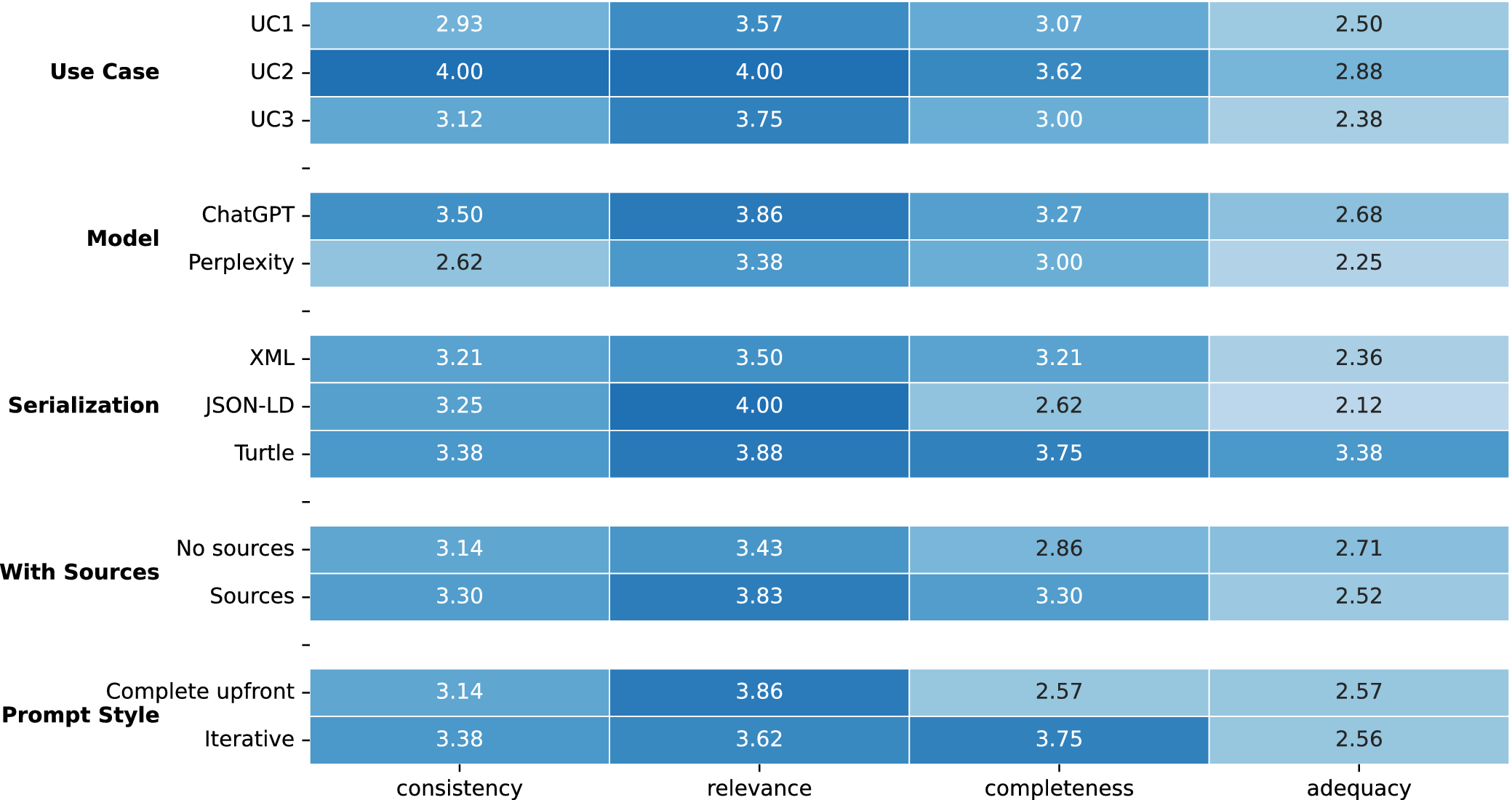
The analysis of the average qualitative assessment per experimental factor (Figure 3) was not conclusive due to the relatively small sample size but remarks a number of trends. It is observed that UC2 consistently outperforms the other cases in all categories. Regarding the model, ChatGPT shows higher average scores than Perplexity across all dimensions. Although serialization using JSON-LD scores higher in relevance, Turtle is the best performing serialization format in general. Providing sources scores better in every dimension but adequacy. The iterative prompting approach achieves a much higher score in completeness but performs similarly to the complete upfront approach in the rest of dimensions.
Heatmap of average qualitative assessment per experimental factor.

The MANOVA reveals a significant multivariate effect of prompt style on the overall quality profile, whereas other factors show weaker or dimension-specific patterns. Subsequent univariate models clarify these effects: iterative prompting produces a substantial increase in completeness, model choice primarily affects consistency, and Turtle serialization is positively associated with adequacy (and, to a lesser extent, completeness). In contrast, relevance remains comparatively stable across experimental conditions. Overall, the results suggest that different experimental factors selectively influence distinct aspects of quality rather than producing a uniform improvement across all dimensions (see Supplementary Material Appendix A for details on the fitted models).
Regarding the proportion of hallucinations, the reduced model retains variables that exhibited substantial effect magnitudes in the full model, despite their initial lack of statistical significance. This selection criterion accounts for the constraints of the sample size and ensures that practically relevant factors are not overlooked. Once the model was refined, Turtle serialization was associated with a substantial decrease in the likelihood of hallucinations (
![]() $ OR $
= 0.15,
$ OR $
= 0.15,
![]() $ p $
-value
$ p $
-value
![]() $ < $
.001), corresponding to an approximate 85 percent reduction in the odds relative to XML. No other experimental factor showed statistically significant effects, although a marginal positive tendency was observed for UC3.
$ < $
.001), corresponding to an approximate 85 percent reduction in the odds relative to XML. No other experimental factor showed statistically significant effects, although a marginal positive tendency was observed for UC3.
An emergent finding was that hallucinations were more frequent in structurally complex ontology components, whereas well-defined classes were generated more reliably. For instance, GenAI typically struggles with defining parameters, such as costs, that involve second-order uncertainty. Such instances require classification as subclasses of both Cost and SecondOrderParameter. Furthermore, characterizing the uncertainty itself necessitates additional parameters to accurately model the underlying probability distribution. This pattern suggests that GenAI performance may reflect differences in ontological complexity, highlighting a potential complementary role in ontology evaluation.
The practical implementation of the experiments faced several limitations, mainly technical. The Perplexity API lacked features for controlled experimental setups, could not process files, and required manual insertion of ontology and source content due to token limits. Additional credit constraints further limited testing.
During early experiments, both models showed limitations in handling ontology files provided directly in OWL or RDF formats. This was identified through follow-up queries asking whether the ontology content had been successfully processed, to which the models indicated that they could not access or interpret the uploaded ontology files in their native format. As a result, ontology content was converted into plain text representations. Although this approach partially improved performance, the issue did not fully disappear, suggesting that additional factors such as input size, token limitations, or structural complexity may also have influenced the models’ ability to process the information. Providing sample individuals and simplified representations ultimately proved to be the most effective workaround in our experimental setup.
Additionally, previous studies (Reference Soares, Saraiva and Pires57) have reported a decrease in the reliability of LLM-generated OWL outputs when multiple individuals are modeled simultaneously within a single prompt. In our experimental design, we did not explicitly evaluate this factor, as our focus was on other dimensions such as prompting strategy, serialization format, and information sources. Therefore, we did not observe clear evidence of this limitation under our specific setup, although the number of individuals generated per interaction was indirectly controlled through the iterative prompting strategy. This aspect represents an interesting direction for future work, particularly in relation to prompt scalability and output consistency.
We also observed occasional incomplete responses in which the model summarized groups of individuals and deferred the generation of remaining elements (e.g., “…include the remaining items here”), suggesting possible limitations related to response length or generation constraints. Although we did not explicitly analyze the effect of input or output size, these observations may be consistent with previous findings on performance degradation in more verbose formats such as XML. This aspect warrants further investigation in future work.
Manual expert validation of above 1,300 individuals was labor-intensive and potentially biased. Although alternative approaches, such as automatic ontology comparison (e.g., OntoSim (https://gitlab.inria.fr/moex/ontosim)) and GenAI-based validation, were explored, they proved impractical or unreliable for the requirements of this study.
Conclusions and further work
This study evaluated the feasibility, reliability, and methodological implications of GenAI models for ontology population within HTA workflows. The results demonstrate that GenAI models can effectively generate structured ontology individuals, but their performance varies across distinct quality dimensions and is strongly influenced by experimental design choices. Rather than uniformly improving overall quality, factors such as prompting strategy and serialization format selectively affected completeness, adequacy, consistency, and the likelihood of hallucinations. In particular, iterative prompting significantly improved completeness, supporting its use as a controlled refinement mechanism during ontology generation. Additionally, Turtle serialization was associated with a substantially lower likelihood of hallucinations, suggesting that formal representation choices play a critical role in ensuring structural reliability and semantic validity (Reference Cao, Wang and Zhang58;Reference Bashah, Salem and Al-waqeerah59;Reference Niel, Dookhun and Caliment60).
For the HTA community, this approach offers a scalable solution for early-stage or exploratory modeling, where evidence is often dispersed and resource-intensive to synthesize manually. By providing a semi-automated pipeline to extract and structure clinical and economic knowledge, agencies and industry can reduce the time-to-insight without compromising methodological rigor.
However, the results also highlight important limitations. Both evaluated GenAI systems showed reduced reliability when generating individuals associated with more complex or abstract ontology structures, indicating that human oversight remains essential, especially for high-stakes or structurally complex modeling tasks. Furthermore, technical constraints related to model interfaces, input handling, and validation workflows currently limit full automation. These findings reinforce the importance of positioning GenAI as an assistive tool within expert-driven HTA processes rather than a fully autonomous solution.
Future research aims to evolve this prototype into a production tool by broadening the validation scope. This evolution seeks to replace individual review with a panel of HTA experts to standardize the evaluative criteria and explore the use of AI-generated vignettes to support expert elicitation. We also propose mitigating the manual workload through automated tools like OntoSim and LLM-based alignment (e.g., Agent-om (Reference Qiang, Wang and Taylor61)), which would streamline the workflow and improve interoperability. Integrating comprehensive metrics as per Encord (Reference Yu, Alégroth, Chatzipetrou and Gorschek62) will ensure the generative performance aligns with high-level quality standards.
Ultimately, integrating GenAI with formal semantic structures like OSDi provides HTA practitioners with a robust framework to manage the complexity of modern healthcare evidence, ensuring that automated outputs remain transparent, reproducible, and aligned with domain-specific standards. Lastly, evaluating the computational models automatically generated from these ontologies (a project already in progress (https://github.com/JaDES-ULL/JaDES-HTA)) will provide a complete evidence-to-model lifecycle of the proposed approach.
Supplementary materials
The supplementary material for this article can be found at http://doi.org/10.1017/S0266462326103754.
Data availability
All the data produced during the experiments described in this document, together with the details on the validation process, are publicly available at https://github.com/ontologies-ULL/OSDi-GenAI/.
Acknowledgements
The authors thank the researchers from the Evaluation and Planning Service of the Canary Islands Health Service (www.sescs.es) for their contribution to the definition of the PICO question and their comments during the validation process.
Author contribution
-
• E.G.G. Conceptualization, Methodology, Writing – Original Draft.
-
• I.C.R. Conceptualization, Methodology, Validation, Formal Analysis, Writing – Original Draft.
-
• J.A.D.H. Software, Validation, Investigation, Data Curation, Writing – Original Draft.
Funding statement
No funding was received for conducting this study.
Competing interests
The authors have no relevant financial or nonfinancial interests to disclose.