INTRODUCTION
During infection of the mammalian host, trypanosomatids are exposed to a variety of oxidizing environments. The reduced form of the thiol trypanothione, which is unique to trypanosomatids, is thought to play a central role in the redox defence systems of these parasites to such stresses (Flohé, Reference Flohé2012; Peloso et al. Reference Peloso, Gonçalves, Silva, Ribeiro, Piñeyro, Robello and Gadelha2012; Paiva and Bozza, Reference Paiva and Bozza2014). This reduced form is generated by the action of trypanothione reductase (TRed), an enzyme belonging to the flavoprotein disulphide oxidoreductase family. The substrate of this enzyme is trypanothione disulphide. TRed is a specific NADPH-dependant homodimeric protein that requires flavin-adenine dinucleotide (FAD) as a coenzyme and a redox-active cysteine disulphide in the active site (Fairlamb and Cerami, Reference Fairlamb and Cerami1992). TRed and peroxidases play a role in protecting these parasites against oxidative stress that may arise internally as a result of their aerobic metabolism and externally by the action of the host immune response (Fairlamb et al. Reference Fairlamb, Blackburn, Ulrich, Chait and Cerami1985; Shames et al. Reference Shames, Fairlamb, Cerami and Walsh1986).
Among the trypanosomatids that are trypanothione-dependent organisms, Trypanosoma rangeli is a non-virulent protozoan parasite of mammals that is genomically related to Trypanosoma cruzi, the aetiologic agent of Chagas disease (Guhl and Vallejo, Reference Guhl and Vallejo2003; Stoco et al. Reference Stoco, Wagner, Talavera-Lopez, Gerber, Zaha, Thompson, Bartholomeu, Lückemeyer, Bahia, Loreto, Prestes, Lima, Rodrigues-Luiz, Vallejo, Filho, Schenkman, Monteiro, Tyler, de Almeida, Ortiz, Chiurillo, de Moraes, de Cunha, Mendonça-Neto, Silva, Teixeira, Murta, Sincero, Mendes, Urmenyi, Silva, DaRocha, Andersson, Romanha, Steindel, de Vasconcelos and Grisard2014). Compared with T. cruzi, T. rangeli possesses striking new genomic features such as (1) considerably less amplification of the gene copy number within multigene virulence factor families, such as mucin-associated proteins (MASPs), trans-sialidases and mucins; (2) the presence of vestigial orthologues of the interference RNA (RNAi) machinery, which are insufficient to constitute a functional pathway; and (3) a reduced repertoire of genes encoding antioxidant defence enzymes (Stoco et al. Reference Stoco, Wagner, Talavera-Lopez, Gerber, Zaha, Thompson, Bartholomeu, Lückemeyer, Bahia, Loreto, Prestes, Lima, Rodrigues-Luiz, Vallejo, Filho, Schenkman, Monteiro, Tyler, de Almeida, Ortiz, Chiurillo, de Moraes, de Cunha, Mendonça-Neto, Silva, Teixeira, Murta, Sincero, Mendes, Urmenyi, Silva, DaRocha, Andersson, Romanha, Steindel, de Vasconcelos and Grisard2014). These parasites also occur in sympatry in a wide overlapping area in South and Central America, infecting triatomine bugs, humans and a variety of sylvatic and domestic mammals (Coura et al. Reference Coura, Fernandes, Arboleda, Barrett, Carrara, Degrave and Campbell1996; Grisard et al. Reference Grisard, Steindel, Guarneri, Eger-Mangrich, Campbell and Romanha1999; Guhl and Vallejo, Reference Guhl and Vallejo2003). Moreover, T. rangeli and T. cruzi can share at least 60% of their soluble antigenic constitution, which can lead to a false-positive diagnosis of chagasic infection in people carrying T. rangeli, with consequent socio-economic effects (Azambuja et al. Reference Azambuja, Ratcliffe and Garcia2005; Ferreira et al. Reference Ferreira, Lorenzo, Elliot and Guarneri2010).
Trypanosoma cruzi and Trypanosoma brucei express high levels of TRed (Piacenza et al. Reference Piacenza, Zago, Peluffo, Alvarez, Basombrio and Radi2009b ), supporting its essential role in parasite viability. Trypanosoma brucei parasites became more susceptible to H2O2-induced oxidative stress and were avirulent only when TRed levels were reduced to <10% (Krieger et al. Reference Krieger, Schwarz, Ariyanayagam, Fairlamb, Krauth-Siegel and Clayton2000). Interestingly, TRed levels were not increased in T. cruzi during metacyclogenesis (Piacenza et al. Reference Piacenza, Alvarez, Peluffo and Radi2009a , Reference Piacenza, Zago, Peluffo, Alvarez, Basombrio and Radi b ).
Little is known regarding the behaviour of T. rangeli in response to oxidative stress or the relevance of TRed to survival and adaptation in response to the new environments encountered by the parasite during its life cycle. Thus, in the present study, we characterized the T. rangeli TRed gene and its expression levels during the parasite life cycle and under oxidative stress conditions. The relevance of this enzyme for the parasite response to reactive oxygen species (ROS) is highlighted.
MATERIALS AND METHODS
Parasites
Trypanosoma rangeli (Choachí strain) and T. cruzi (Y strain) epimastigotes were maintained at 27·5 °C in liver infusion tryptose medium (LIT) supplemented with 10% heat-inactivated fetal bovine serum (FBS), 100 units mL−1 penicillin and 100 µg mL−1 streptomycin by weekly passages after cyclic mouse-triatomine-mouse passages. Polymerase chain reaction (PCR) showed that all samples tested negative for the presence of Mycoplasma sp. For DNA or protein extraction and for in vitro oxidative stress experiments, epimastigotes in the exponential growth phase were harvested during the late log phase and washed twice with sterile PBS (pH 7·2). Trypanosoma rangeli trypomastigotes were obtained in vitro using previously described conditions (Koerich et al. Reference Koerich, Emmanuelle-Machado, Santos, Grisard and Steindel2002). Trypanosoma cruzi culture-derived trypomastigotes (TCT) were obtained by infection of Vero cells as previously described (Eger-Mangrich et al. Reference Eger-Mangrich, De Oliveira, Grisard, De Souza and Steindel2001).
Isolation of DNA and RNA
Total RNA was extracted from T. rangeli epimastigotes using the TRIzol method (Invitrogen) and then treated with DNase I, according to the manufacturer's instructions. Trypanosoma rangeli and T. cruzi genomic DNA (gDNA) was isolated using a standard phenol–chloroform method (Sambrook et al. Reference Sambrook, Fritch and Maniatis2001). The purity and quality of the obtained DNA and RNA were assessed spectrophotometrically and via agarose gel electrophoresis.
Protein extraction
Epimastigote or trypomastigote forms were washed once with PBS (pH 7·2), lysed by repeated pipetting in ice-cold lysis buffer (0·25 m sucrose, 0·25% Triton X-100, 10 mm EDTA) containing a protease inhibitor cocktail (2 mm 4-(2-aminoethyl)benzenesulfonyl fluoride hydrochloride, 0·3 µ m aprotinin, 116 µ m bestatin, 14 µ m E-64, 1 µ m leupeptin and 1 mm EDTA) (Sigma-Aldrich). Cellular debris were removed by centrifugation at 12 000 g for 20 min at 4 °C (Romero et al. Reference Romero, Téllez, Yamanaka, Steindel, Romanha and Grisard2014). The protein concentration was determined by the Bradford method (Bio-Rad) using BSA as a standard (Bradford, Reference Bradford1976). Parasite extracts were stored at −20 °C until use.
cDNA Synthesis and qPCR
For first-strand cDNA synthesis, 1 µg of total RNA was used in a 20-μL final reaction volume containing 200 U Moloney Murine Leukaemia Virus Reverse Transcriptase (M-MLV RT), 1X First Strand Buffer, 200 µ m dNTP, 10 pmoles oligodT-Anchor, 10 mm dithiothreitol (DTT), 40 U RNaseOUT™ (Invitrogen). The mixture was incubated at 37 °C for 50 min, heat-inactivated at 70 °C for 15 min and then diluted 5-fold in nuclease-free water. The diluted cDNA was used for qPCR amplification on ABI Prism® 7900HT Sequence Detection System (Applied Biosystems) equipment using specific primers (qTRedTr-F: 5′-CGA AAG ACT GAT CAC ACC CG-3′ and qTRedTr-R: 5′- CGA GTG CCG TCT TCT CTA TTC CTC-3′). GAPDH (GenBank AUPL01004827) (GAPDH-F: 5′-GCG ACA CCA GCA TCA AAG AG-3′/GAPDH-R: 5′- CTG TGC TCA CAA GTT CCT CG-3′) and RNA60S (GenBank AUPL01005494) (RNA60S-F: 5′-CGA TGA AGC TCA AGT GGA CC-3′/RNA60S-R: 5′-CGG TTG TAC TTG ACG GGA AC-3′) genes were used as references, and a negative control containing no cDNA was included in each reaction set. Reactions consisted of 0·3 µ m of each qTrTRed primer, Maxima® SYBR Green/ROX qPCR Master Mix reagent (Thermo Scientific), 2 µL template cDNA and nuclease-free water up to 10 µL. The temperature profile for all reactions was 95 °C for 10 min, followed by 40 cycles of denaturation (95 °C for 15 s) and annealing/extension (61 °C for 1 min). A final step was included to obtain the dissociation curve (95 °C, 60 °C and 95 °C for 15 s each). The reaction efficiency for each primer set was calculated after a 5-point serial dilution (1:2) of cDNA pools and gDNA. All qPCR assays were performed using biological and technical triplicates. Raw quantification cycle (C q) results were obtained using SDS 2·4 data analysis software (Applied Biosystems) and normalized to the geometrical average value of two reference genes for each sample. The amplification efficiency was obtained with the E = 10(−1 slope−1) formula (Pfaffl, Reference Pfaffl2001). The relative quantification method (Livak and Schmittgen, Reference Livak and Schmittgen2001) was used to assess variations in transcript levels. Comparison of transcript levels between two samples was conducted using the unpaired Student's t-test with Prism 5·0 (GraphPad) and P < 0·05.
Identification and Cloning of T. rangeli TRed (TrTRed)
Using the T. cruzi TRed gene as the query (GenBank Accession Number CAA78360), the complete sequence of the TrTRed gene was retrieved via Blast from T. rangeli genomic and transcriptomic sequences available in GenBank and TriTrypDB (www.tritrypdb.org) (Aslett et al. Reference Aslett, Aurrecoechea, Berriman, Brestelli, Brunk, Carrington and Wang2010), as well as from sequences recently generated using the Illumina platform at the Science for Life Laboratory, Karolinska Institutet, Sweden. The entire TrTRed open reading frame (ORF) was PCR-amplified using primers TrTRedBglII (5′-AGA TCT ATG AAA GCC TTT GAT TTG GTT G-3′) and TrTRedKpnI (5′- GGT ACC AGC CTC TAG AGC CGT TTC CG-3′), containing the respective restriction enzyme sites to allow downstream cloning. PCR assays were performed on a Mastercycler® Gradient (Eppendorf) thermocycler using 35 cycles of the following thermal profile for denaturation (95 °C, 1 min), annealing (64 °C, 1 min) and extension (72 °C, 1 min), including a final extension step at 72 °C for 5 min. The PCR product was cloned into pGEM-T-Easy vector (Promega), and the resulting constructs were verified by sequencing on an Abi3500 automated DNA sequencer (Life Technologies) using the BigDYE 3·1 kit (Life Technologies) according to the manufacturer's instructions. Both DNA strands were sequenced for each obtained clone, and those presenting a high quality (Phred ≥ 20), as indicated by the Phred/Phrap/Consed package (Ewing et al. Reference Ewing, Hillier, Wendl and Green1998), were clustered and compared with the public databases using the BLAST algorithm (Altschul et al. Reference Altschul, Gish, Miller, Myers and Lipman1990).
Comparative analysis of TRed expression by T. rangeli and T. cruzi
Soluble protein extracts (50 µg) of different life cycle stages of T. rangeli and T. cruzi were resolved by sodium dodecyl sulphate- 12% polyacrylamide gel electrophoresis (SDS- 12% PAGE) and electroblotted onto nitrocellulose membranes (GE Healthcare) using an appropriate buffer (25 mm Tris; 192 mm glycine; 20% v/v methanol, pH 8·3). After blocking with 5% non-fat milk in blotting buffer (25 mm Tris–HCl pH 7·4, 150 mm NaCl, and 0·1% Tween-20) overnight at 4 °C (Gallagher et al. Reference Gallagher, Winston, Fuller and Hurrell2008), the membranes were incubated for 1 h with an anti-rTcTRed mouse polyclonal antiserum (1:500) (kindly provided by Dr. Silvane Murta – CPqRR/Fiocruz) or anti α-tubulin monoclonal antibody (Cell Signaling Technology) (1:1000) as a loading control. Detection of expressed TRed was conducted using anti-mouse IgG conjugated to horseradish peroxidase (HRP) (1:10 000), followed by the enhanced chemiluminescence (ECL) kit (Pierce) according to the manufacturer's recommendations. The Western blots were digitally recorded and analysed using the ImageJ 1·463r software package. The relative protein expression was determined by the ratio of the TRed and α-tubulin band intensities.
Assessment of the enzymatic activity of T. rangeli TRed
The TrTRed activity was assessed spectrophotometrically by measuring nicotinamide adenine dinucleotide phosphate (NADPH) oxidation at 340 nm as formerly described (Jockers-Scherübl et al. Reference Jockers-Scherübl, Schirmer and Krauth-Siegel1989). Kinetic assays were conducted in 96-well microplates at 27 °C in an incubation medium containing 20 mm HEPES pH 7·4, 30 mm NaCl, 0·1 mm EDTA, 150 µ m NADPH, 50 µ m T(S)2 and 1·5 µg µL−1 of soluble T. rangeli epimastigote protein extract in a final volume of 200 µL. The reaction was started by the addition of trypanothione disulphide (T(S)2) and followed for 3 min. A negative (no substrate) and positive control (0·1 µg µL−1 of rTcTred) were included in each enzymatic assay. One unit of TRed activity was defined as the amount of enzyme required to oxidize 1 µ m of NADPH to NADP+ per minute at 27 °C. The results were expressed as μ m min−1 mg of protein−1. The reference values used were an NADPH molar extinction coefficient εM = 6220 m −1 cm−1 and assuming a 1-cm path length.
Oxidative stress induced by H2O2 in T. rangeli
Exponential growth phase epimastigotes (5 × 107 cells) were harvested by centrifugation (3000 g for 10 min) and washed twice in sterile PBS pH 7·2. The parasites were then resuspended in LIT medium supplemented with 10% FBS and incubated in the presence of 67 µ m H2O2 for 30, 60 or 90 min at 27 °C. The IC50 was determined as previously described (Romero et al. Reference Romero, Téllez, Yamanaka, Steindel, Romanha and Grisard2014). Considering that such a concentration of H2O2 in a single dose could induce bias in an in vitro system, the parasites were exposed to increasing concentrations of H2O2 every 20 min up to 1 h while another sample received no H2O2 (negative control). After incubation, the parasites were centrifuged (3000 g for 10 min) and washed twice with PBS (pH 7·4), and the pellet was stored at −20 °C for protein extraction. TRed expression and activity were measured as described above.
Detection of intracellular ROS in T. rangeli epimastigotes
The presence of intracellular ROS in T. rangeli and T. cruzi epimastigotes was assessed using the 2′−7′-dichlorodihydrofluorescene diacetate (DCFH-DA) probe (Sigma-Aldrich) as previously described (Wang and Joseph, Reference Wang and Joseph1999). Briefly, 5 × 105 parasites well−1 (200 µL) were incubated in LIT medium supplemented with 10% heat-inactivated FBS in the presence of different H2O2 concentrations (0–1 mm) in black 96-well plates. After 10 min, 10 µ m of DCFH-DA well−1 was added, and the fluorescence intensity was determined in a TECAN Infinite M200 spectrofluorometer (Switzerland) for 1 h using an excitation wavelength of 485 nm and an emission wavelength of 535 nm. Untreated parasites (no H2O2) and 10 µ m of DCFH-DA probe in lysis buffer were used as negative controls. Two independent experiments were performed in quadruplicate. The results are expressed as relative fluorescence units that are directly proportional to the increase in the ROS concentration over the time course.
NADPH production through the measurement of glucose-6-phosphate dehydrogenase (G6PD) and 6-phosphogluconate dehydrogenase (6PGD) activities
NADPH production was determined as previously described (Mielniczki-Pereira et al. Reference Mielniczki-Pereira, Chiavegatto, López, Colli, Alves and Gadelha2007). Briefly, cells were collected by centrifugation and resuspended in PBS pH 7·2 in the presence of a protease inhibitor cocktail (Cocktail Set III, Calbiochem, Catalogue #: 539134). The combined activities of G6PD and 6PGD were measured by NADP+ reduction at 340 nm after the addition of parasites (5 × 107 cells mL−1) to the reaction mixture (50 mm Tris–HCl, pH 7·6, 50 mm KCl, 0·1% Triton X-100, 250 µ m NADP+, 2 mm MgCl2, 1 mm 6-phosphogluconate and 1 mm glucose-6-phosphate). 6PGD activity was assessed in the same medium but in the absence of glucose-6-phosphate.
Determination of hydrogen peroxide release
H2O2 release by T. rangeli epimastigotes was determined as previously described (Peloso et al. Reference Peloso, Vitor, Ribeiro, Piñeyro, Robello and Gadelha2011). Briefly, 108 cells mL−1 were incubated in PBS pH 7·2 1 mm −1 MgCl2 in the presence of 5 mm succinate, 60 µ m digitonin, 1 U mL−1 of HRP and 25 µ m Amplex Red (Molecular Probes). The fluorescence was monitored at excitation and emission wavelengths of 563 nm and 587 nm, respectively, using a Hitachi F2500 fluorescence spectrophotometer with continuous stirring. The quantitative correlation between the fluorescence and H2O2 release by the cells was determined as previously described (Barros et al. Reference Barros, Bandy, Tahara and Kowaltowski2004).
Effect of antioxidants on the growth of T. rangeli and TRed expression
Epimastigotes (2 × 106 mL−1) were grown in LIT medium (10% FBS) for 15 days at 27 °C in the presence of reduced glutathione (GSH) or N-acetylcysteine (NAC) (Sigma-Aldrich) at two different concentrations (1 and 2·5 mm). The experiments were performed in triplicate, and the parasites were counted every day in a Neubauer chamber. Samples were collected on days 1, 3, 5, 7, 9 and 11 to analyse TRed expression by Western blotting using anti-rTcTRed and anti α-tubulin antibodies.
Overexpression of TRed in T. rangeli
The TrTRed gene was excised from the pGEM-T-Easy vector using BglII/KpnI restriction enzymes and then subcloned into the pLEXSY-NEO2 vector (Jena). All constructs were checked for their restriction profiles and sequenced prior to transfection into T. rangeli epimastigotes by electroporation using e Nucleofector® device (Lonza). Briefly, 5 × 106 epimastigotes in the log growth phase were washed twice in PBS (pH 7·2), mixed with 100 µL of human T cell Nucleofector buffer (Lonza) and 10 µg of pLEXSY-NEO2 –TrTRed construct DNA and transfected using program U-033. After transfection, the parasites were transferred to Neal, Novy, Nicolle-LIT medium and grown at 27·5 °C for 48 h. The transfectants were then selected via incremental exposure to geneticin (Gibco), from 25 µg mL−1 up to a final concentration of 300 µg mL−1. Parasites transfected with empty vector were generated and compared with wild-type parasites in terms of growth and TRed expression levels. Overexpression of TrTRed (TrTRed+) by transfected parasites was assessed by Western blotting using an anti-His6-tagged antibody (1:200) (Santa Cruz Biotechnology). After confirmation, the TrTRed+ parasites were harvested during the exponential and late log phases for protein extraction and for the in vitro oxidative susceptibility assay as described above, using non-transfected wild-type (WT) parasites as a control. Additionally, WT and TrTRed+-overexpressing parasites were comparatively evaluated according to their in vitro growth curve and differentiation into trypomastigote forms.
Cytolocalization of the rTrTRed expression sites
The expression sites of TrTRed+ by T. rangeli were assessed by indirect immunofluorescence assays (IFA). Exponential growth phase WT and TrTRed+ epimastigotes were harvested from LIT by centrifugation (3000 g , 10 min), washed twice with PBS pH 7·2 and adhered onto 13-mm circular glass coverslips for 20 min. The parasites were fixed with 4% paraformaldehyde (w/v) in PBS and then permeabilized with 0·1% NP-40 or 0·1%Triton X-100. The cells were then blocked with 2% BSA diluted in PBS and probed with anti-His6-tagged antibody diluted 1:100 in blocking solution. After 90 min, the slides were washed extensively with PBS and then incubated for 1 h with Alexa Fluor 488-conjugated goat-anti-mouse IgG diluted 1:10 000 in blocking solution. After washing 3× with PBS, the parasites were stained for 5 min with 1 µg mL−1 of DAPI (4,6-diamidino-2-phenylindole), and the coverslips were mounted over microscopy slides using Hydromount (National Diagnostics). Images were obtained using an Olympus Bx40–FL fluorescence microscope (Olympus).
Susceptibility of TRed-overexpressing T. rangeli epimastigotes to oxidative stress
The susceptibility of T. rangeli-overexpressing TrTRed to oxidative stress was compared with that of WT cells using Alamar Blue (AB) assays as previously described (Räz et al. Reference Räz, Iten, Grether-Bühler, Kaminsky and Brun1997; Decuypere et al. Reference Decuypere, Vanaerschot, Brunker, Imamura, Müller, Khanal, Rijal, Dujardin and Coombs2012). Briefly, 5 × 105 T. rangeli and T. cruzi epimastigotes were exposed to hydrogen peroxide (Sigma-Aldrich) prepared at different dilutions (0–1 mm) and incubated for 48 h at 26 °C in quadruplicate in 96-well plates. After incubation for 24 h, 20 µL of AB reagent (Invitrogen) was added to each well to assess the parasite viability via fluorescence emission at 600 nm. Data from treated and non-treated cultures were used to calculate the IC50 by a sigmoidal regression analysis (with a variable slope) using GraphPad Prism v.5.0. Untreated control parasites and reagent blanks were included in each test plate.
In vitro interaction of THP-1 cells and T. rangeli overexpressing TrTRed
Cells of the human acute monocytic leukaemia cell line THP-1 (ATCC#TIB-202) were cultured at 37 °C in a 5% CO2 atmosphere using RPMI 1640 medium (HiMedia Laboratories) supplemented with 1% glutamine, 10% heat-inactivated FBS, 12·5 mm HEPES (Gibco), 100 U mL−1 penicillin, 100 µg mL−1 streptomycin (Gibco), 2 mm Glutamax® (Gibco) and 1 mm sodium pyruvate (Gibco). Monocytes were harvested during the logarithmic growth phase and transferred to medium containing 50 µg mL−1 phorbol myristate acetate (Sigma-Aldrich) to induce adherence and differentiation of the macrophages (Schwende et al. Reference Schwende, Fitzke, Ambs and Dieter1996). Prior to the in vitro infection assays with T. rangeli, a total of 1 × 105 macrophages were seeded onto 13-mm circular glass coverslips placed in 6-well plates and then incubated for 72 h at 37 °C in a 5% CO2 atmosphere (Romero et al. Reference Romero, Saraiva and Walker2005). After removal of non-adherent cells by washing the plate wells twice with sterile PBS (pH 7·2), THP-1-derived macrophages were incubated with T. rangeli trypomastigotes overexpressing TrTRed for 1 h at 37 °C in a 5% CO2 atmosphere using a parasite-to-cell ratio of 25:1. Non-adherent parasites were removed by washing with PBS, which was considered the initial time point of infection (T0), and the progression of the parasite–cell interaction was observed at 2 (T2), 4 (T4), 6 (T6) and 24 hours (T24) after infection. The cells were then stained with Giemsa, mounted on microscope slides with Entellan® (Merck) and observed by light microscopy. The number of infected cells was determined by counting 200 randomly chosen cells per coverslip for each of the time points, strains and species. The assays were performed using biological triplicate samples and non-infected cells were used as controls.
Statistical analyses
All experiments were performed in triplicate, and the results are presented as the mean and standard deviation (s.d.m.) or standard error of the mean (s.e.m.). Significant differences were assessed by one-way analysis of variance (ANOVA) followed by the Bonferroni post-hoc test or Student's t-test using GraphPad Prism v.5.0 software as indicated in the figure legends.
RESULTS
In silico analysis revealed that the T. rangeli TRed gene (TrTRed) has an ORF of 1473 nt encoding a predicted protein of 491 aa (~53·5 kDa) with a predicted isoelectric point of 6·33. TrTRed is present as a single copy in the parasite genome and displays conserved catalytic domains, particularly the disulphide-binding site, FAD and NADH binding domains among different kinetoplastid species (Fig. 1).
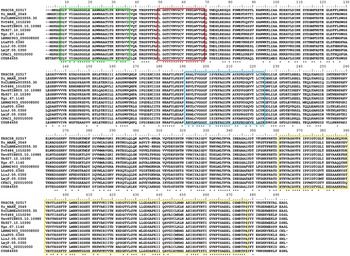
Fig. 1. Trypanosoma rangeli trypanothione reductase (TRed) has conserved catalytic domains. Alignment of the amino acid sequences of TRed from T. rangeli and other kinetoplastid species. Red rectangle- catalytic site; Green rectangle- FAD binding domain; Blue rectangle- NAD(P) binding domains; Yellow rectangle- dimerization domain. Accession numbers for TRed from T. rangeli – TRSC58_02517; Trypanosoma cruzi marinkellei – Tc_MARK_3549; T. cruzi CLBrener – TcCLBNEs503555·30; Trypanosoma vivax- TvY486_1010290; Trypanosoma evansi – TevSTIB805.10.10980; Trypanosoma brucei -Tb927.10.10390; Trypanosoma grayi – Tgr.67·1140; Leishmania braziliensis – LBRM2903_050008500; Leishmania tarentolae – LtaP05·0360; Leishmania infantum -LinJ.05·0350; Leishmania major – LmjF.05·0350; Crithidia fasciculata – CFAC1_020010000; Bodo saltans – CUG84655.
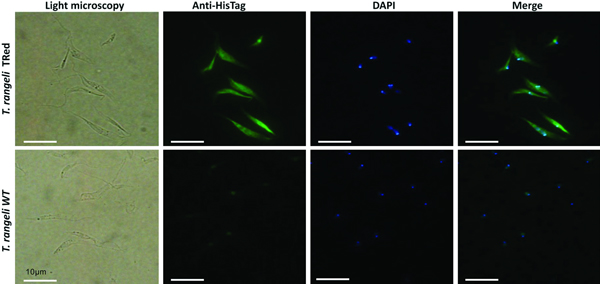
Assessment of the intracellular localization of TrTRed in T. rangeli epimastigotes overexpressing the enzyme by IFA revealed dispersed fluorescence throughout the parasite cytoplasm, suggesting a cytosolic localization (Fig. 2), which is consistent with the in silico prediction for the native protein.
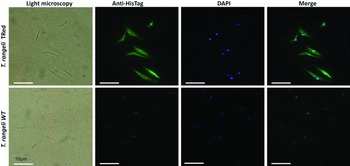
Fig. 2. Homologous trypanothione reductase overexpressed by Trypanosoma rangeli has a cytosolic localization. Immunolocalization of the expression sites of rTrTRed in T. rangeli epimastigotes (T. rangeli TRed) by IFA assays using anti-His6-tag antibody. T. rangeli wild type (WT) parasites were used as controls. (1) Light microscopy, (2) Detection of rTrTRed using anti-His6-tagged antibodies, (3) DAPI staining and (4) merged images. Bars indicate 10 µm.
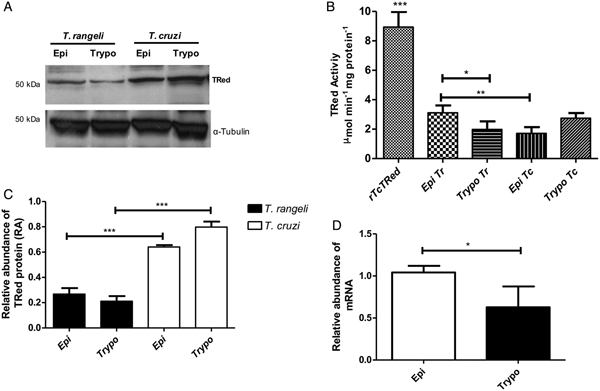
The relative abundance of the TrTRed protein assessed by Western blotting in T. rangeli epimastigotes and trypomastigotes revealed no significant differences (Fig. 3A). The absence of TRed stage-specific expression was also observed for the homologous protein in T. cruzi (TcTRed).
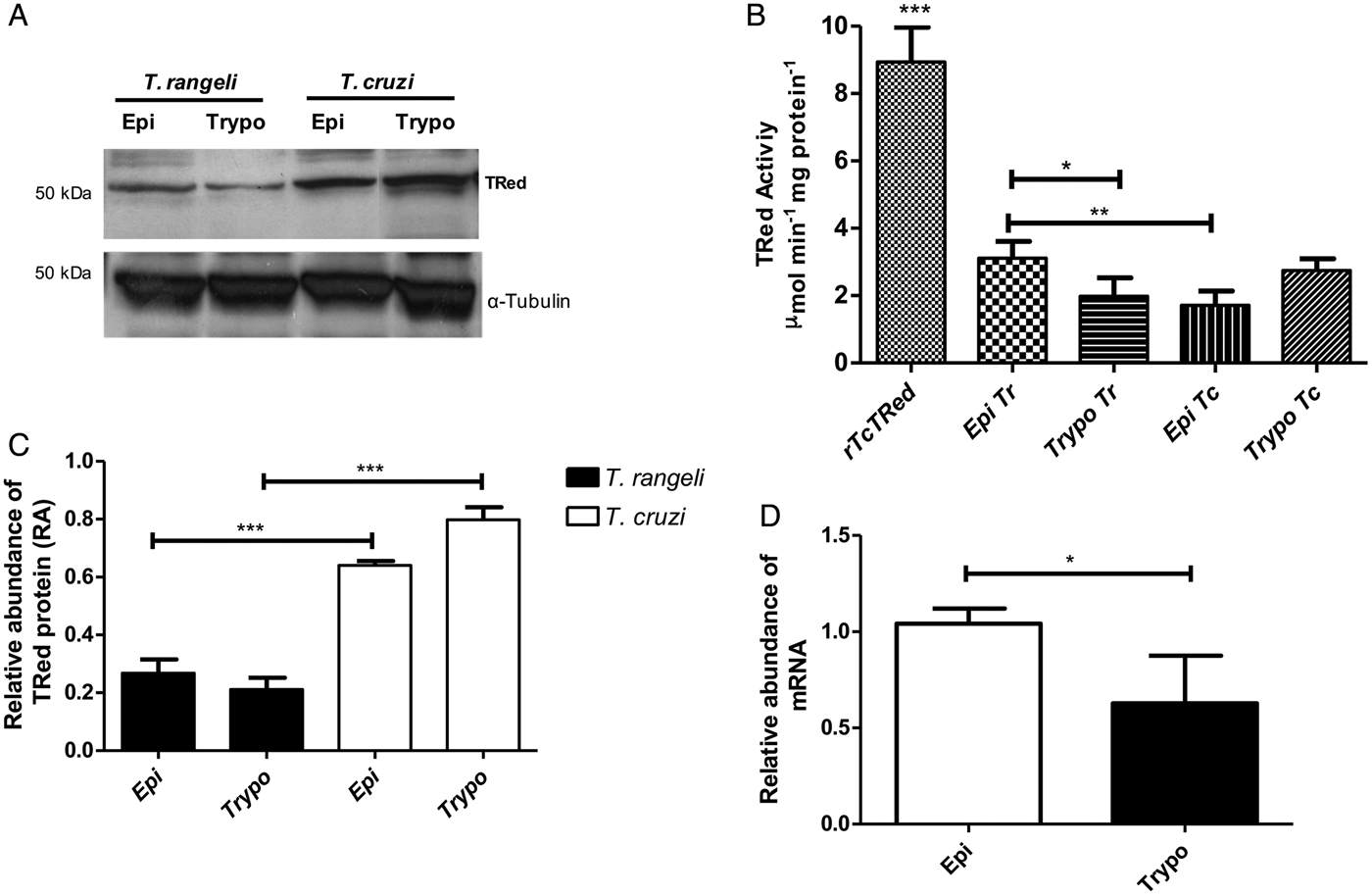
Fig. 3. TRed expression and activity are mainly observed in Trypanosoma rangeli epimastigotes. (A) Western blot analysis of TRed expression in soluble T. rangeli and T. cruzi protein extracts obtained from epimastigote (Epi) and trypomastigote (Trypo) forms and comparative densitometric analysis of the blotting results using ImageJ software. Significant differences were determined by one-way ANOVA followed by the Bonferroni test (***P < 0·001). Detection of α-tubulin served as a loading control and for normalization of the densitometry analysis. (B) Enzymatic activity of TRed in soluble protein extracts of T. rangeli and T. cruzi epimastigotes and trypomastigotes. The recombinant T. cruzi TRed (rTcTRed) was used as a positive control, and the results represent the mean ± s.d.m. (standard deviation of the mean) from 3 independent experiments performed in triplicate. Significant differences were assessed by one-way ANOVA followed by the Bonferroni test (*P < 0·05, **P < 0·01, ***P < 0·001). (C) Detection of TrTRed mRNA levels in T. rangeli epimastigotes and trypomastigotes by qPCR using GAPDH and RNA60S for normalization. Data represent the mean ± s.d.m. from two independent experiments conducted with technical quadruplicates. Significant differences were assessed by Student's t-test (*P < 0·05).
Trypanosoma rangeli TRed was active in both epimastigotes and trypomastigotes (3·11 and 1·82 µ m min−1 mg of protein−1, respectively), suggesting that its activity is significantly increased in the replicative parasite forms (Fig. 3B). Interestingly, T. cruzi TRed was more active in trypomastigotes than in epimastigotes (2·74 and 1·71 µ m min−1 mg of protein−1, respectively), but no significant differences were observed between the infective and replicative forms of the parasite. Considering this difference, we also comparatively assessed the TrTREd mRNA levels between replicating epimastigotes and infective trypomastigotes. The results are shown in Fig. 3C and indicate a significant decrease in the mRNA levels of TrTRed in trypomastigotes.
Having determined the basal TrTRed transcriptional and expression levels, we also evaluated whether TrTRed expression by T. rangeli epimastigotes was influenced by H2O2-induced oxidative stress. The Western blotting shown in Fig. 4A indicates that there was no significant increase in TrTRed expression up to 1 h after exposure to 67 µ m H2O2. This concentration of H2O2 was based on the IC50 determined in previous studies (Stoco et al. Reference Stoco, Wagner, Talavera-Lopez, Gerber, Zaha, Thompson, Bartholomeu, Lückemeyer, Bahia, Loreto, Prestes, Lima, Rodrigues-Luiz, Vallejo, Filho, Schenkman, Monteiro, Tyler, de Almeida, Ortiz, Chiurillo, de Moraes, de Cunha, Mendonça-Neto, Silva, Teixeira, Murta, Sincero, Mendes, Urmenyi, Silva, DaRocha, Andersson, Romanha, Steindel, de Vasconcelos and Grisard2014; Romero et al. Reference Romero, Téllez, Yamanaka, Steindel, Romanha and Grisard2014). For all times and conditions tested, no significant differences were observed in the densitometric analysis, which indicated that TrTRed expression by parasites remained constant regardless of exposure to oxidative stress.

Fig. 4. Trypanosoma rangeli and Trypanosoma cruzi epimastigotes exhibit similar responses to in vitro oxidative stress induced by H2O2. (A) Western blot analysis of TrTRed expression in soluble protein extracts obtained at different time points from T. rangeli epimastigotes under oxidative stress induced by 67 µ m H2O2. Comparative densitometric analysis of the blotting results using ImageJ software. Data represent the mean ± s.d.m. from three independent experiments. Significant differences were assessed by one-way ANOVA followed by the Bonferroni post-hoc test (*P < 0·05). Detection of α-tubulin served as loading control and for normalization in the densitometry analysis. (B) Detection of fluorescence emitted by the DCFH-DA probe in parasites exposed to increasing concentrations of hydrogen peroxide. The results are expressed as the mean ± s.d.m. from two independent experiments performed using technical quadruplicates. Significant differences were detected by one-way ANOVA following by the Bonferroni post-hoc test (***P < 0·001).
Modulation of enzyme expression was evaluated during oxidative stress induced by hydrogen peroxide in T. rangeli, also considering the endogenous production of H2O2 by the parasite. Using the DCFH-DA probe, we observed that oxidative stress induced by concentrations of H2O2 up to 125 µ m did not induce changes in fluorescence emission among treated and non-treated parasites. However, at concentrations of 250 µ m and higher, an effective and significant effect of the stressor was observed for both T. rangeli and T. cruzi in comparison with non-H2O2-treated parasites (Fig. 4B).
The amount of NADPH produced by T. rangeli epimastigotes, as determined by the combined measurement of G6PDH and 6PGD activities, was 1·1 ± 0·05 nm min−1 107 cells−1, whereas T. cruzi epimastigotes produced 2·13 ± 0·09 nm NADPH min−1 107 cells−1. This difference was significantly different (P < 0·05) and indicated that the pentose pathway is more effective in T. cruzi than in the other parasite. Based on this difference and considering the distinct virulence of these species to their mammalian hosts, we determined the amount of mitochondrial hydrogen peroxide released by these parasites. Using digitonin-permeabilized cells and succinate as a substrate for the mitochondrial respiratory chain, we were able to determine that T. rangeli epimastigotes released 14·2 ± 1·6 pm of H2O2 min−1 107 cells−1 (data not shown), which is higher than the levels observed for T. cruzi under the same experimental conditions (Silva et al. Reference Silva, Peloso, Vitor, Ribeiro and Gadelha2011).
The presence of antioxidants improved the in vitro growth of T. rangeli epimastigotes (Fig. 5A). Both treatments with NAC (1 and 2·5 mm) induced dramatic growth of the parasites from day 3–7 (P < 0·001). Parasites grown in the presence of 1 mm NAC reached the largest number of parasites (5·8 × 107 parasites mL−1) on the 5th day of culture. Treatments with GSH (1 and 2·5 mm) also improved parasite growth from day 5–12 (P < 0·01). Although important differences in parasite growth were observed in the presence of antioxidants, the reducing environment did not affect the expression of TRed (Fig. 5B). The protein levels remained constant in parasites grown under different conditions on the days evaluated.
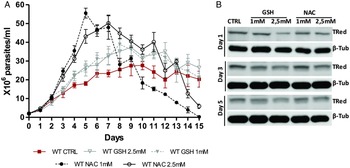
Fig. 5. N-acetylcysteine treatment improves the growth of Trypanosoma rangeli epimastigotes. (A) Comparative in vitro growth curve of T. rangeli submitted to treatments with GSH and NAC at 1 and 2·5 mm. The results are expressed as the mean ± s.d.m. of triplicates. Significant differences were detected using two-way ANOVA followed by the Bonferroni post-hoc test. (B) Representative Western blot analysis of TrTRed expression in soluble protein extracts from T. rangeli with different treatments at days 1, 3 and 5 revealed by anti-TcRed antibody. Detection of α-tubulin served as a loading control and for normalization in the densitometry analysis.
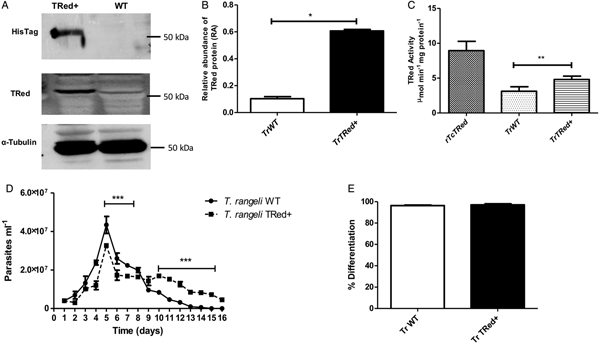
Overexpression of homologous TrTRed by T. rangeli epimastigotes revealed a 5-fold increase in the protein expression level compared with wild type parasites (Fig. 6A and B). As expected, the specific TRed activity in parasites overexpressing this gene was 1·3-fold higher than the TRed activity in control WT parasites (Fig. 6C). Overexpression of TrTRed by T. rangeli revealed no major differences in in vitro growth when compared with WT parasites (Fig. 6D). The maximum number of parasites for both WT (4·3 × 107 parasites mL−1) and overexpressing parasites (3·3 × 107 parasites mL−1) was observed on day 5 of culture. While the growth curve of the wild type strain exhibited a typical decrease in the number of live parasites from day 6 forward, overexpressing parasites appeared to be sustained in culture up to day 14 and then started to decrease. Significant differences were observed in comparisons of the number of parasites between days 4–7 and 10–15. We may conclude that TrTRed-overexpressing parasites have slightly higher growth and survival rates in vitro compared with WT parasites. It is noteworthy that overexpression of TrTRed had no effect on the ability of parasites to undergo in vitro differentiation into infective trypomastigotes compared with the parental WT strain (Fig. 6E).
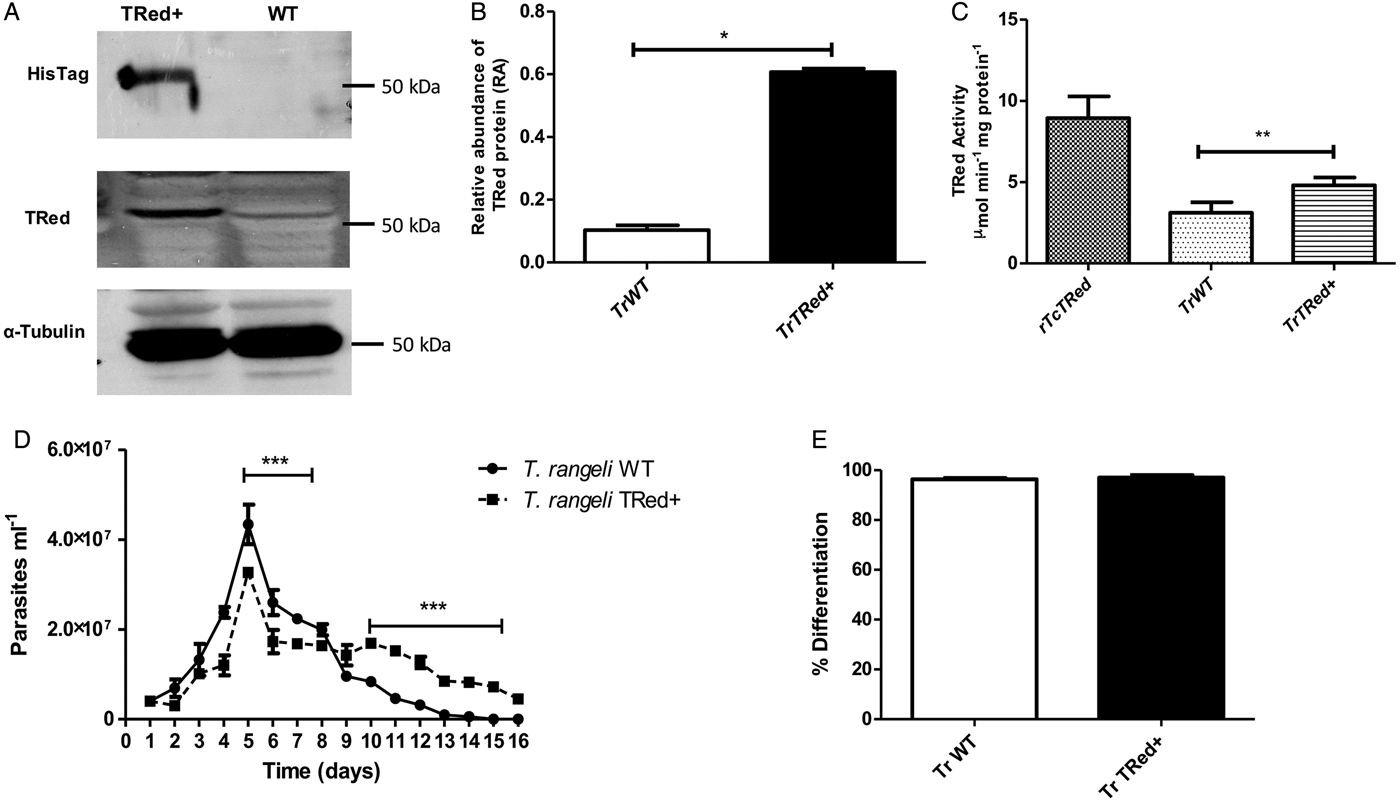
Fig. 6. Overexpression of TRed by Trypanosoma rangeli epimastigotes does not affect parasite biology in vitro. (A) Western blot analysis of TrTRed expression in soluble protein extracts from T. rangeli overexpressing TrTRed+ and WT revealed by anti-His6-tag and anti-TcRed antibodies. Detection of α-tubulin served as a loading control and for normalization in the densitometry analysis. (B) Comparative densitometric analysis of the blotting results using ImageJ software. (C) TRed enzymatic activity in protein extracts from T. rangeli WT and TrTRed+-overexpressing epimastigotes. rTcTRed was used as a control. (D) Comparative in vitro growth curve of TrTRed+ and WT T. rangeli strains starting from 4 × 106 parasites mL−1. (E) Comparative percentages of the in vitro differentiation of T. rangeli WT and TRed+ mutants. The results shown in panels B and C represent the mean ± s.d.m. obtained from three independent experiments using biological triplicates. For panels B and C, significant differences were assessed using Student's t-test (*P < 0·05, **P < 0·01). The results in panel D represent the mean ± s.d.m. obtained from three independent experiments performed in duplicate. Significant differences were assessed by two-way ANOVA (***P < 0·001).
Parasites overexpressing TrTRed protein were also more resistant (2·3 times) to H2O2 than wild-type parasites. The dose-response analysis revealed a marked difference in resistance to H2O2 between T. rangeli WT and mutant strains at the 125 µ m H2O2 concentration, in which the viability of the mutant parasites was 13·4-fold higher compared with the WT parasites (Fig. 7A).

Fig. 7. Overexpression of homologous TrTRed by Trypanosoma rangeli epimastigotes increases parasite survival under oxidative stress in vitro. (A) Viability of T. rangeli WT and TrTRed+ epimastigotes in response to increasing concentrations of H2O2 assessed by Alamar Blue staining. The bars represent the survival ratio between TrTRed+ and WT at 62,5 and 125 µ m. (B) The percentages of THP-1 cells infected by T. rangeli WT or TrTRed+ mutants during the time course of the in vitro cell-parasite interaction. (C) Light micrographs of distinct time points (T0-T24 hours) of THP-1-derived human macrophages infected by T. rangeli WT or TrTred+ mutants. Trypanosoma cruzi-infected THP-1 cells were used as a control. Cells were stained with Giemsa, black arrows indicate parasites and bars represent 10 µ m. The results shown in panels A and B represent the mean ± s.d.m. from three independent experiments performed in triplicate.
Due to this increased resistance to H2O2 stress, we also assessed whether the increased expression of TrTRed by T. rangeli would modulate its ability to infect and survive within a phagocytic cell. Although TrTRed-overexpressing parasites seemed to be more infective on T0, T4 and T24, the kinetic study of the interaction of WT and mutant parasites with THP1-derived macrophages in vitro revealed no significant differences between the parasite strains (P < 0·05) in terms of the number of infected cells (Fig. 7B and C). No signs of T. rangeli multiplication were observed for WT or mutant parasites. In conclusion, increased expression of homologous TRed by T. rangeli enhanced its resistance to oxidative stress but did not induce detectable phenotypic changes in the in vitro parasite-THP-1 cell interactions.
DISCUSSION
Members of the Trypanosomatidae family are digenetic parasites with a life cycle that presents multiple differentiation forms alternating between invertebrate and vertebrate hosts (Teixeira and DaRocha, Reference Teixeira and Darocha2003). During each stage of differentiation, a stage-specific protein expression pattern participates in the invasion and survival processes of the parasite (Macleod et al. Reference Macleod, Maudlin, Darby and Welburn2007; Díaz et al. Reference Díaz, Solari and González2011).
Redox homoeostasis appears to be efficiently regulated in kinetoplastids because the parasites can successfully survive the oxidative burst generated at the onset of infection and adapt to the different metabolic and environmental conditions (Peloso et al. Reference Peloso, Gonçalves, Silva, Ribeiro, Piñeyro, Robello and Gadelha2012). At present, antioxidant defence is directly related to the success of the infection process because the proteins making up this system are among those that are most expressed during differentiation from replicative epimastigotes to infective trypomastigotes (Atwood et al. Reference Atwood, Weatherly, Minning, Bundy, Cavola, Opperdoes, Orlando and Tarleton2005; Piñeyro et al. Reference Piñeyro, Parodi-Talice, Arcari and Robello2008; Piacenza et al. Reference Piacenza, Alvarez, Peluffo and Radi2009a , Reference Piacenza, Zago, Peluffo, Alvarez, Basombrio and Radi b ; Gretes et al. Reference Gretes, Poole and Karplus2012). In T. cruzi, increased expression of antioxidant defence-related proteins enables the parasite to survive in the complex and highly oxidative environment within macrophages (Atwood et al. Reference Atwood, Weatherly, Minning, Bundy, Cavola, Opperdoes, Orlando and Tarleton2005; Araujo and Teixeira, Reference Araujo and Teixeira2011). According to Irigóin et al. (Reference Irigoín, Cibils, Comini, Wilkinson, Flohé and Radi2008), this increase is a preadaptation of the parasite prior to invasion of the vertebrate host in preparation for environments in which it may be exposed to ROS and RNS (reactive nitrogen species) such as those generated by immune cells. However, an evaluation of antioxidant enzymes in T. cruzi virulent strains compared with attenuated ones revealed that the protein contents observed in TcTRed remained constant across the different parasites stages (Piacenza et al. Reference Piacenza, Alvarez, Peluffo and Radi2009a , Reference Piacenza, Zago, Peluffo, Alvarez, Basombrio and Radi b ).
The susceptibility of T. rangeli to oxidative stress was evaluated by subjecting epimastigotes to H2O2-induced stress conditions in vitro according to Stoco et al. (Reference Stoco, Wagner, Talavera-Lopez, Gerber, Zaha, Thompson, Bartholomeu, Lückemeyer, Bahia, Loreto, Prestes, Lima, Rodrigues-Luiz, Vallejo, Filho, Schenkman, Monteiro, Tyler, de Almeida, Ortiz, Chiurillo, de Moraes, de Cunha, Mendonça-Neto, Silva, Teixeira, Murta, Sincero, Mendes, Urmenyi, Silva, DaRocha, Andersson, Romanha, Steindel, de Vasconcelos and Grisard2014) and Romero et al. (Reference Romero, Téllez, Yamanaka, Steindel, Romanha and Grisard2014). These authors demonstrated that T. rangeli was more sensitive than T. cruzi to oxidative stress, showing an IC50 of 53 µ m, which is significantly less than the IC50 obtained for T. cruzi epimastigotes (188·3 µ m). Previous studies using distinct T. cruzi strains have shown a wide range of IC50 for the H2O2 stress, varying from 98 to 190 µ m (Finzi et al. Reference Finzi, Chiavegatto, Corat, Lopez, Cabrera, Mielniczki-Pereira, Colli, Alves and Gadelha2004; Mielniczki-Pereira et al. Reference Mielniczki-Pereira, Chiavegatto, López, Colli, Alves and Gadelha2007).
Taylor et al. (Reference Taylor, Kelly, Champman, Fairlamb and Miles1994) described an absence of Leishmania donovani TRed expression modulation during the induction of oxidative stress by H2O2. These authors state that in contrast to peroxidases, which are upregulated due to direct involvement in the detoxification of H2O2-induced stress, TRed is not directly involved in such detoxification. For T. rangeli, TRed appears to be constitutively expressed during the parasite life cycle, as described in this study and in accordance with Taylor et al. (Reference Taylor, Kelly, Champman, Fairlamb and Miles1994). It also appears to participate in the mechanisms that protect the parasite from ROS. Although the results obtained herein were obtained from only one strain of each species (T. rangeli Choachí and T. cruzi Y), they yielded some interesting insights regarding comparisons of the antioxidant defence of these parasites.
Although expressed during the entire life cycle of T. rangeli, elevated activity of TrTRed was observed during the replicative epimastigote stage. Unlike T. cruzi, which faces oxidative stresses in the mammalian host and within the digestive tract of the triatomine vector, T. rangeli epimastigotes are found within the intestinal tract and haemolymph of triatomines, where the parasite is exposed to successive and distinct oxidative and/or nitrosative stresses. In this study, reduced TrTRed expression was observed for the in vitro-derived trypomastigote forms. Because these parasites are biochemically distinct from the T. rangeli metacyclics observed within the triatomine salivary glands and bloodstream trypomastigotes found in infected mammals, the reduced TrTRed expression observed in vitro-derived trypomastigotes remains to be examined in in vivo parasites.
The present observations can be related to a reduction in NADPH production in T. rangeli compared with T. cruzi epimastigotes. Unlike T. cruzi, T. rangeli is able to evade the intestinal tract of the triatomine by penetrating through the gut epithelium towards the haemocoel, where the parasites undergo successive division cycles prior to invading the salivary glands (Stoco et al. Reference Stoco, Wagner, Talavera-Lopez, Gerber, Zaha, Thompson, Bartholomeu, Lückemeyer, Bahia, Loreto, Prestes, Lima, Rodrigues-Luiz, Vallejo, Filho, Schenkman, Monteiro, Tyler, de Almeida, Ortiz, Chiurillo, de Moraes, de Cunha, Mendonça-Neto, Silva, Teixeira, Murta, Sincero, Mendes, Urmenyi, Silva, DaRocha, Andersson, Romanha, Steindel, de Vasconcelos and Grisard2014). These highly oxidative environments (Garcia et al. Reference Garcia, Castro, Figueiredo, Genta and Azambuja2009) demand increased synthesis of NADPH by the parasite as a primary source of reducing equivalents for biosynthetic reactions and the oxidation-reduction involved in protecting against the toxicity of ROS (Igoillo-Esteve et al. Reference Igoillo-Esteve, Maugeri, Stern, Beluardi and Cazzulo2007; Leroux et al. Reference Leroux, Maugeri, Opperdoes, Cazzulo and Nowicki2010). In this sense, several enzymes are also co-responsible for the maintenance of this reducing environment, such as the NADP-linked glutamate dehydrogenase (Barderi et al. Reference Barderi, Campetella, Frasch, Santomé, Hellman, Pettersson and Cazzulo1998), the NADP-linked malic enzyme (Cannata et al. Reference Cannata, Frasch, Cataldi De Flombaum, Segura and Cazzulo1979) and the two dehydrogenases in the pentose phosphate pathway (PPP) (Leroux et al. Reference Leroux, Maugeri, Opperdoes, Cazzulo and Nowicki2010), which is the major source of reduced TRed (Krauth-Siegel and Comini, Reference Krauth-Siegel and Comini2008; Leroux et al. Reference Leroux, Maugeri, Opperdoes, Cazzulo and Nowicki2010; Peloso et al. Reference Peloso, Vitor, Ribeiro, Piñeyro, Robello and Gadelha2011).
By analysing the PPP-related enzymes in different T. cruzi life stages, Maugeri and Cazzulo (Reference Maugeri and Cazzulo2004) demonstrated increased expression in trypomastigotes that probably resulted from the need to respond to the oxidative stress created by the mammalian host.
Trypanosoma cruzi and T. brucei depend on their NADP-linked dehydrogenases, such as malic enzymes, for NADPH production in low or no glucose environments, in the gut of their insect vectors or within cells, because under such conditions, the PPP cannot act as a source of essential redox mechanisms (Hellemond et al. Reference Hellemond, Opperdoes and Tielens2005; Leroux et al. Reference Leroux, Maugeri, Opperdoes, Cazzulo and Nowicki2010; Allman et al. Reference Allman, Morand, Ebikeme, Gales, Biran, Hubert, Brennand, Mazet, Franconi, Michels, Portais, Boshart and Bringaud2013). The relative roles of the PPP and alternative reactions to provide NADPH have not yet been investigated in T. rangeli.
The major and continuous source of cellular ROS, including H2O2, is the mitochondrial electron transport chain. H2O2 can easily diffuse through the plasma membrane and interact with membrane-bound components (Cosentino-Gomes et al. Reference Cosentino-Gomes, Russo-Abrahão, Fonseca-de-Souza, Ferreira, Galina and Meyer-Fernandes2009). As determined in this study, endogenous production of H2O2 was higher in T. rangeli than in T. cruzi (Peloso et al. Reference Peloso, Vitor, Ribeiro, Piñeyro, Robello and Gadelha2011). This distinct behaviour could be due to differences in the mitochondrial antioxidant defences of the two species. In addition to the reduced TRed expression compared with T. cruzi, the reduced NAPDH production and higher susceptibility to H2O2-induced stress by T. rangeli allowed us to hypothesize that these two species have distinct superoxide dismutase expression profiles. Thus, it is noteworthy that overexpression of superoxide dismutase by T. cruzi induces a reduced sensitivity to trypanocidal agents (Temperton et al. Reference Temperton, Wilkinson, Meyer and Kelly1998).
N-acetylcysteine (NAC) has been widely used as an antioxidant in vivo and in vitro. It is a powerful scavenger of hypochlorous acid (H--OCl) and reacts with hydroxyl radical and H2O2 (Aruoma et al. Reference Aruoma, Halliwell, Hoey and Butler1989; Rossato et al. Reference Rossato, Zecchin, La Guardia, Ortega, Alberici, Costa, Catharino, Graner, Castilho and Vercesi2014). GSH is the major intracellular antioxidant thiol that protects mammalian cells against oxidative stress induced by oxygen- and nitrogen-derived reactive species (Romão et al. Reference Romão, Fonseca, Hothersall, Noronha-Dutra, Ferreira and Cunha1999). Our results demonstrated that the presence of these antioxidants (NAC and GSH) promoted improved growth of epimastigotes cultivated in LIT medium. Conversely, the presence of the same antioxidants dramatically impaired T. cruzi epimastigote proliferation (Nogueira et al. Reference Nogueira, Saraiva, Sultano, Cunha, Laranja, Justo, Sabino, Coelho, Rossini, Atella and Paes2015), and the presence of the pro-antioxidant heme induced epimastigote proliferation. Thus, T. rangeli appeared to require a reduced environment for epimastigote proliferation, whereas T. cruzi required an oxidative environment. Both parasites develop inside triatomines; however, differences related to the biological cycle of these parasites in this host could explain the differences in proliferation. Epimastigotes from T. cruzi multiply and colonize the entire gut (oxidative environment), whereas T. rangeli epimastigotes rapidly cross the gut epithelium and invade the haemolymph. The haemolymph of Rhodnius prolixus has been described to have ten times more urate than human plasma (Souza et al. Reference Souza, Petretski, Demasi, Bechara and Oliveira1997; Graça-Souza et al. Reference Graça-Souza, Maya-Monteiro, Paiva-Silva, Braz, Paes, Sorgine, Oliveira and Oliveira2006). Urate is considered as one of the most important antioxidants in human plasma. Thus, the effect of the antioxidants NAC and GSH in T. rangeli cultures would improve their ability to grow by simulating the role of urate in the haemolymph. Despite the improved growth and reducing environment, we did not observe any modifications in the expression levels of TrTRed.
The role of TRed in T. rangeli, like in T. cruzi and L. donovani (Kelly et al. Reference Kelly, Taylor, Smith, Hunter and Fairlamb1993; Tovar et al. Reference Tovar, Cunningham, Smith, Croft and Fairlamb1998), has been investigated by overexpressing the homologous enzyme. Kelly et al. (Reference Kelly, Taylor, Smith, Hunter and Fairlamb1993) transfected L. donovani and T. cruzi with an expression vector containing the LdTRed gene and achieved 10–14-fold enzyme activity in transformed cells. Trypanosoma cruzi cells overexpressing the sense construct displayed a 4-fold increase in TRed protein levels and enzyme activity (Tovar and Fairlamb, Reference Tovar and Fairlamb1996). The difference between the species is probably due to the distinct basal expression levels of TRed.
In addition to numerous tools used to unravel the mechanisms of T. cruzi infection, such as cellular tropism, infectivity and intracellular replication, in vitro infection of cell cultures has been utilized (Duran-Rehbein et al. Reference Duran-Rehbein, Vargas-Zambrano, Cue´llar, Puerta and Gonzalez2014). Non-epithelial cells such as monocytes/macrophages are highly parasitized after very short incubation times, providing insight into the initial stages of T. cruzi infection and the subsequent immune response. Furthermore, survival within active macrophages seems to be highly dependent on the efficiency of ROS reduction (Leirião et al. Reference Leirião, Rodrigues, Albuquerque and Mota2004; Piacenza et al. Reference Piacenza, Peluffo, Alvarez, Kelly, Wilkinson and Radi2008, Reference Piacenza, Peluffo, Alvarez, Martínez and Radi2013).
By overexpressing the homologous TrTRed, we have demonstrated an apparent increase in the ability of T. rangeli to address the intracellular reducing environment within human macrophages, where there is a high demand for trypanothione (Dumas et al. Reference Dumas, Ouellette, Tovar, Cunningham, Fairlamb, Tamar, Olivier and Papadopoulou1997). This study also indicates that TRed does not appear to be a pivotal molecule involved in the response of T. rangeli to antioxidant environments in vertebrate host cells; however, it plays a role in other parts of the parasite life cycle, such as in the insect gut or haemocytes where ROS production limits the infection and colonization of R. prolixus by T. rangeli (Cosentino-Gomes et al. Reference Cosentino-Gomes, Rocco-Machado and Meyer-Fernandes2014).
ROS have been traditionally viewed as a ‘necessary evil’ in the battle against pathogens, and their production is coupled to antioxidant responses that are important for mitigating oxidative damage in cells and tissues (Andrews, Reference Andrews2012). However, Paiva et al. (Reference Paiva, Feijó, Dutra, Carneiro, Freitas, Alves, Mesquita, Fortes, Figueiredo, Souza, Fantappié, Lannes-Vieira and Bozza2012) and Goes et al. (Reference Goes, Rocha, Diniz, Aguiar, Machado and Vieira2016) have demonstrated that oxidative stress actually enhances infection by the protozoan T. cruzi. In others words, T. cruzi seems to thrive in an oxidative environment, and antioxidant responses suppress infections by this protozoan parasite. Likewise, Leishmania donovani promastigotes treated with sub-lethal doses of H2O2 have been found to be more virulent and more potent for infecting macrophages ex vivo via a metabolic reconfiguration designed to replenish the NADPH pool for protection against oxidative challenges.
Therefore, future investigations will be designed to elucidate the role of ROS during T. rangeli infection. In this work, the role of H2O2 appeared to be a determinant in the host protection against parasite infection, and the inability of the parasite to respond to oxidative stress is directly related to its virulence.
ACKNOWLEDGEMENTS
To Dr. Silvane Murta (CPqRR/Fiocruz) for the anti-TcTRed antibody. To Dr. Ibeth Romero, Dr. Milene Höehr de Moraes, Ana Paula Nascimento, MSc., Carime Lessa Mansur Pontes, MSc and Carine Lais Moreira for technical assistance and discussions.
FINANCIAL SUPPORT
To Conselho Nacional de Desenvolvimento Científico e Tecnológico – CNPq (E.C.G.), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior – CAPES (E.C.G.) and Fundação de Amparo à Pesquisa do Estado de São Paulo – FAPESP (F.R.G.) – Brazilian Government Agencies. To the STINT (Sweden)/CAPES (Brazil) joint program (B.A./E.C.G.).
CONFLICT OF INTEREST
The authors declare no conflicts of interest.
ETHICAL STANDARDS
All procedures involving animals were previously approved by the Ethical Committee on Animal Use of Federal University of Santa Catarina (Reference number 23080·025618/2009-81).