Introduction
Measles is an acute viral disease characterised by a prodromal illness of fever and catarrhal symptoms (e.g., cough, coryza and conjunctivitis), followed by the appearance of a generalised maculopapular rash [Reference Knipe1, Reference Moss2]. Although measles is among the most infectious of communicable diseases, no specific antiviral therapy exists. Two doses of measles vaccine are the crucial and best way to prevent and eliminate measles [3]. In other words, measles can be eliminated by sustaining high levels of population immunity in combination with disease surveillance and an integrated laboratory network [3]. Indeed, measles elimination has progressed worldwide. For example, as a result of two doses of measles vaccine and exhaustive surveillance of measles cases, Japan was verified by the World Health Organization (WHO) Western Pacific Regional Office in March 2015 as a country that had achieved measles elimination [4].
Modified measles (M-Me), an atypical and milder version of typical measles (T-Me), should be discovered adequately, especially in measles-eliminated or measles-reduced nations. The illness has been confirmed in persons with incomplete immunity after measles vaccination. The fever is generally low grade. Moreover, the catarrhal symptoms or the rash are usually mild: sometimes so mild as to be nearly imperceptible [Reference Gershon5, Reference Grammens6]. Actually, M-Me is a potential source of infection, as with T-Me [Reference Gershon5], but the risk of transmission by M-Me patients is low [Reference Rota7, Reference Rosen8]. Based on the WHO Global Vaccine Action Plan, which targeted the elimination of measles in at least five of six WHO Regions by 2020 [9], it is expected that cases of T-Me will be decreased drastically worldwide. However, M-Me patients, some of whom show common cold-like symptoms, might increase because of primary or secondary vaccine failure [Reference Hickman10, Reference Holzmann11]. In the near future, early detection of M-Me might become an important challenge that must be tackled in the measles-reduced world.
Early detection of measles virus (MeV) positive specimens using laboratory tests contributes to the prevention of the spread of measles. Currently, an antibody test, the culture of MeV and genetic tests for MeV-RNA are available for detecting measles infection. The antibody test is the most common among methods, but early detection is difficult because MeV antibodies are first detectable when the rash appears [Reference Knipe1]. MeV-culture often needs 3–5 days for plaques to become visible in cell lines [Reference Knipe1]. On the other hand, genetic tests that take only a few hours can detect MeV-RNA, which appears in the early stage of symptoms [Reference Moss2]. Actually, genetic tests are performed routinely in Japan, where specimens from cases suspected of measles are collected in an early stage. Nevertheless, genetic testing in the early stage of measles cases, especially in M-Me cases, has not been investigated adequately.
For this study, we investigated the effectiveness of real-time reverse transcription (rRT) polymerase chain reaction (PCR) in an import-associated measles outbreak dominated by M-Me cases in Yamagata Prefecture, Japan. The aim of this study is the clarification of suitable specimens for detecting MeV-RNA properly through the description of the results of the rRT-PCR and the construction of a strategy for finding measles cases with high infectiousness using the semi-quantitative capability of rRT-PCR. This work was approved by the Ethics Committees of Yamagata Prefectural Institute of Public Health (approval no. YPIPHEC 17-10).
Materials and methods
Outbreak description
During March–April 2017, the largest measles outbreak in Japan since the announcement of measles elimination occurred in Yamagata Prefecture. Among 60 measles cases in the outbreak, we studied 53 cases that had been detected in Yamagata Prefecture, Japan. The detail of the outbreak was described by Komabayashi et al. [Reference Komabayashi12].
Definition of M-Me, T-Me and super-spreader
Based on Japan's Act on the Prevention of Infectious Diseases and Medical Care for Patients with Infectious Diseases, physicians reported a patient as M-Me when the patient showed one or two of three clinical symptoms of T-Me (i.e., fever, catarrhal symptoms and rash) and when a laboratory test of measles produced positive results. When a patient showed the three clinical symptoms, physicians reported the patient as T-Me. Additionally, laboratory testing for measles is necessary for the reported cases in Japan. Genetic testing for MeV was administered for all reported cases in this study. Super-spreader was defined as a patient who transmitted measles to numerous contacts and who caused nine or more measles cases in this study.
Swift and thorough surveillance and specimen collection
During the outbreak, public health centres (PHCs) in Yamagata Prefecture performed swift and thorough surveillance to prevent measles transmission. Public health nurses conducted contact tracing to measles patients, identified their contacts and conducted educational and awareness-raising activities to the contacts including early visits to the hospital for people who had shown measles symptom(s). Fundamentally, PHCs collected four specimens (whole blood, throat swab, urine and serum) from patients suspected of having measles on the day that the patients visited a hospital.
Pretreatment of specimen
For this study, all genetic tests for MeV were conducted at the Yamagata Prefectural Institute of Public Health (YPIPH). The specimens were delivered to YPIPH within 1 day after collection. Pretreatment of specimens was performed based on the manual of the National Institute of Infectious Diseases, Japan (NIID) [13]. Throat swab within 3 mL of transport medium [Reference Mizuta14] was centrifuged at 2000 rpm for 15 min. The supernatant was collected. Approximately 10 ml of urine specimen was centrifuged at 2000 rpm for 15 min. Then the supernatant was removed so that the total volume was 3 ml. The pellet was re-suspended. However, the concentration was not performed for urine specimens containing numerous pellets. Peripheral blood mononuclear cells (PBMC) were extracted from 4 ml of whole blood containing EDTA-2Na using Ficoll-Paque™ Plus (GE Healthcare). Then, the PBMC was suspended with 500 µl of Eagle's MEM Nissui (Nissui Pharmaceutical Co. Ltd., Tokyo, Japan). Serum was used directly. RNA was extracted from 200 µl of each sample using the High Pure Viral RNA Kit (Roche Life Science) according to the manufacturer's instructions. Finally, RNA was collected by 40 µl of an elution buffer.
rRT-PCR
Semi-quantitative rRT-PCR targeted to the N gene of MeV [Reference Hummel15] was conducted based on the NIID manual [13]. In all, 20 µl mixture containing 5 µl of 4 × TaqMan Fast Virus 1-step Master Mix (Thermo Fisher Scientific Inc.), each 400 nM of MVN1139F (5′-TGGCATCTGAACTCGGTATCAC-3′) and MVN1213R (5′-TGTCCTCAGTAGTATGCATTGCAA-3′) primers, 250 nM of MVNP1163P probe (5′-FAM-CCGAGGATGCAAGGCTTGTTTCAGA-TAMRA-3′) and 5 µl of the extracted RNA was analysed using 7500 Real-Time System (Thermo Fisher Scientific Inc.) under the following conditions: 5 min for 50 °C, 20 s for 95 °C, then 50 cycles of 15 s for 95 °C and 1 min for 60 °C. According to the NIID manual [13], 50 and 5 copies of standard MeV-RNA were used as positive control (PC). A sample was judged as positive for MeV when the cycle threshold (C t), which means the number of rRT-PCR cycles that amplification product of MeV genome reached a certain amount, was lower than that of 5 copy PC. When C t was between C t of 5 copy PC and 40.0, the sample was judged as equivocal. If 5 copy PC was not detected from rRT-PCR, then 50 copy PC was used for the borderline of positive and equivocal results [13]. Patients were judged positive for MeV when at least one specimen showed a positive result. Patients were judged as negative for MeV when all specimens showed negative results. The rRT-PCR analysis outcomes were conveyed to PHCs on the same day that the specimens arrived at YPIPH.
Conventional reverse transcription PCR
When at least one sample showed an equivocal result and other specimens were negative rRT-PCR results, conventional RT-PCR of two types, targeted to N and H genes, were executed to the equivocal sample, according to the NIID manual [13]. Patients were regarded as positive for MeV when at least one specimen showed positive results in conventional RT-PCR.
Additional interview of measles patients
After the declaration by Yamagata Prefecture on 17 May 2017 that the measles outbreak had ended, PHCs interviewed the measles patients again to review the overall picture of the outbreak. The reinvestigation included an interview, which revealed that worsening of symptoms was observed after visiting a hospital (i.e., after collecting specimens). Based on the interview results, we identified the patients who were judged as having M-Me at the point of specimen collection, but who finally progressed to T-Me.
Estimation of the number of MeV-RNA copies
We determined a formula to estimate the number of MeV-RNA copies per tube retrospectively from C t in the rRT-PCR. We acquired the formula from the slopes of standard curves drawn three times using the three samples of 5000, 500, 50 and 5 copies of the standard MeV-RNA. Additionally, in throat swab samples, the number of MeV-RNA copies per swab was determined as 24 times of that of per tube, by calculating back from the testing process.
Statistical analysis
To clarify patient characteristics of the three measles groups, we applied one-way analysis of variance or Fisher's exact test. Furthermore, we applied a Tukey–Kramer test when one-way analysis of variance was statistically significant. Since we hypothesised that C t of the M-Me group, mildly ill patients, is higher than those of other groups, we performed a Wilcoxon rank sum test adjusted using the Holm method. The cut-off point of C t in the rRT-PCR was calculated using receiver operating characteristic analysis. We considered P < 0.05 as statistically significant. All statistical analyses were conducted using R version 3.4.3 (R Foundation for Statistical Computing, Vienna, Austria).
Results
During March–May 2017, approximately 3700 people who contacted measles patients in Yamagata Prefecture, Japan represented the scope of surveillance at PHCs [Reference Komabayashi12]. Samples from 137 patients showing measles symptom(s) were genetically tested for MeV. MeV-RNA was detected from samples of 53 patients, of which 44 (83.0%) and nine (17.0%) cases were reported respectively as M-Me and T-Me by physicians. Thereafter, additional interviews of measles patients revealed that nine M-Me patients at the point of sample collection had shown the onset of three T-Me symptoms. We defined the nine patients as being in the prodromal stage of T-Me at the point of sample collection (hereinafter, P-Me). Characteristics of the cases of M-Me, P-Me and T-Me are presented in Table 1. Of the 53 cases, 37 (69.8%) patients were 20–39 years of age. Men were 39 (73.6%) of the 53 patients. Catarrhal symptoms were scarce in the cases of M-Me and P-Me. In the M-Me cases, 27 (77.1%) of the 35 patients received at least one dose of measles vaccine. Measles vaccination was incomplete in P-Me and T-Me cases. In M-Me, 28 (80.0%) of the 35 patients visited hospitals within 2 days after experiencing fever. It is particularly interesting that the days after onset in the P-Me cases were significantly fewer than in the T-Me cases.
Characteristics of three measles groups in a modified measles-dominated outbreak, Yamagata Prefecture, Japan, March–April 2017

a Patients diagnosed with modified measles at the point of sample collection, but who finally progressed to typical measles.
b Result of one-way analysis of variance or Fisher's exact test.
c Significantly longer than the groups of modified measles (P = 0.004) and prodromal stage of typical measles (P = 0.04) by Tukey–Kramer test.
The rRT-PCR results of the 35 M-Me patients revealed that PBMC yielded the highest positive rate, followed by a throat swab, urine and serum (Table 2). Table 3 presents details of genetic tests applied to seven cases for which PBMC samples showed negative or equivocal results by rRT-PCR. In four of the seven cases, the throat swab result by the rRT-PCR recovered the negative or equivocal result of PBMC. Three other cases were judged as positive for MeV by conventional RT-PCR in the samples of PBMC, throat swab, or both. Accordingly, the 35 M-Me cases were detectable by a combination of the results of PBMC and throat swab. Urine and serum did not contribute to find M-Me cases. In addition, the positive rates of the rRT-PCR in PBMC, throat swab, urine and serum in the nine P-Me cases were 100% (nine samples), 71.4% (five of seven samples), 88.9% (eight of nine samples) and 33.3% (three of nine samples), respectively. The respective positive rates in the nine T-Me cases were 100% (nine samples), 88.9% (eight of nine samples), 100% (nine samples) and 50.0% (four of eight samples).
Results of a real-time reverse transcription (rRT) PCR for measles virus among 35 cases of modified measles in Yamagata Prefecture, Japan, March–April 2017

PBMC, peripheral blood mononuclear cells.
Results of genetic tests for measles virus (MeV) in modified measles cases for which the PBMC sample showed a negative or equivocal result by a real-time reverse transcription (RT) PCR in Yamagata Prefecture, Japan, March–April 2017

PBMC, peripheral blood mononuclear cells; ND, not done.
To estimate the number of MeV-RNA copies in the rRT-PCR retrospectively, we determined a formula from the average of three standard curves as shown below:
Additionally, the average efficiency of the rRT-PCR was 96.6%.
Results of the rRT-PCR for respective specimens showed that C t of M-Me cases was significantly higher (i.e., lower amount of MeV-RNA) than those of P-Me and T-Me cases (Fig. 1). It is noteworthy that statistical significance was not confirmed in C t between the cases of P-Me and T-Me for any specimen. The cut-off point of C t between M-Me and others (i.e., P-Me or T-Me) in the rRT-PCR is shown in Table 4. The estimated number of MeV-RNA copies in PBMC was extremely low.
Scatter plot of cycle threshold (C t) in a real-time reverse transcription PCR for measles virus RNA in samples of PBMC (a), throat swab (b) and urine (c), Yamagata Prefecture, Japan, March–April 2017. Measles cases are classified into three groups: modified measles (M-Me), a prodromal stage of typical measles at the point of sample collection (P-Me) and typical measles (T-Me). Black circles represent super-spreaders. Bars represent median C t. Asterisks denote statistical significance (P < 0.05).

Cut-off point of cycle threshold (Ct) between modified measles and other measles in a real-time reverse transcription PCR in a modified measles dominated outbreak, Yamagata, Japan, 2017

AUC, area under the curve; PBMC, peripheral blood mononuclear cells.
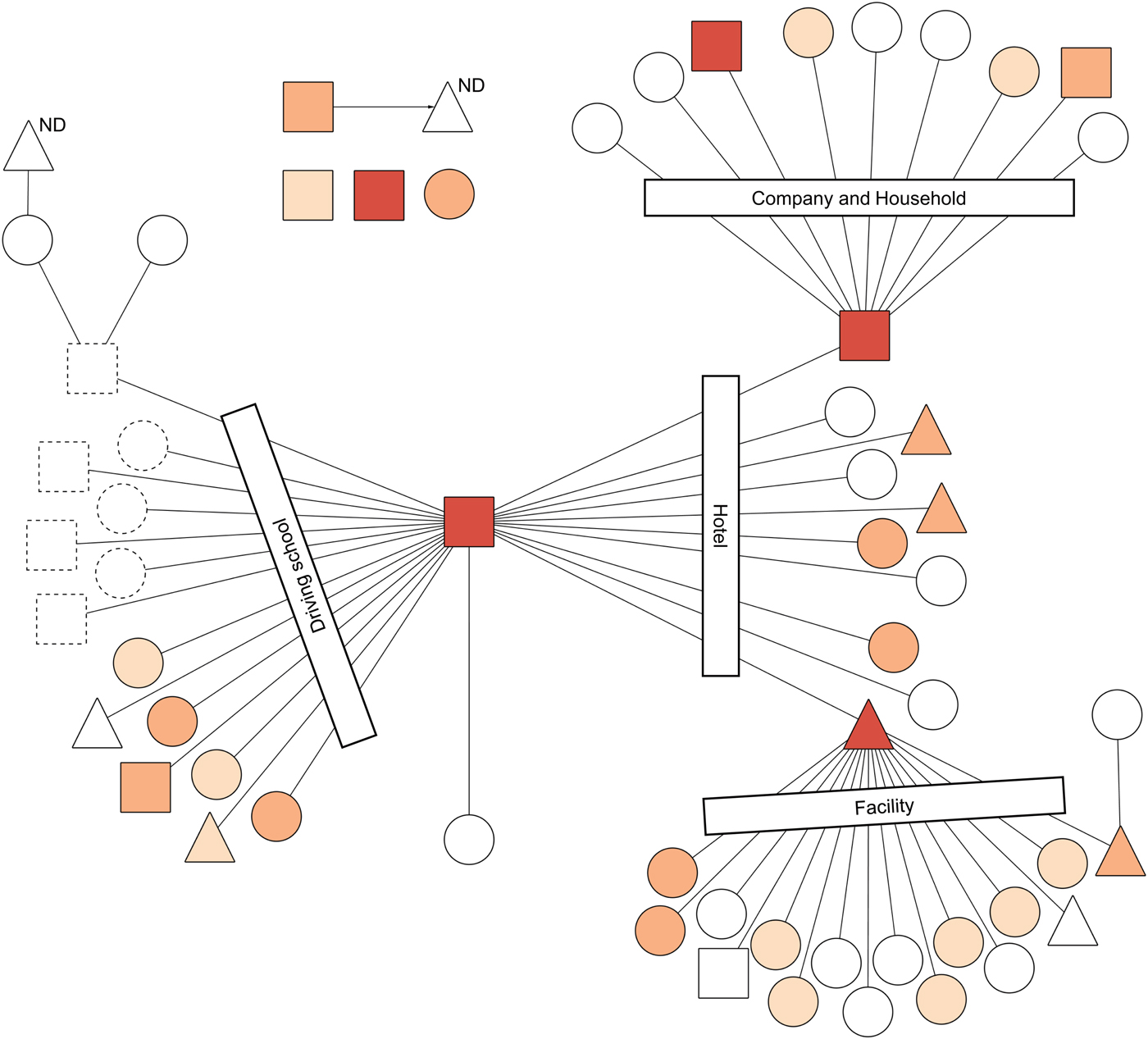
The combination of the transmission network of the outbreak and throat swab's C t in each case indicated that three super-spreaders were estimated as possessing a great amount of MeV in the upper respiratory tract (URT) (Fig. 2). Specifically, the C t of the index case (i.e., measles spread at a driving school, a hotel and a hospital), the spreader of a company and a household and of a facility were 22.61, 24.78 and 16.43, of which the numbers of MeV-RNA copies were calculated, respectively, as 2.9 × 106, 6.7 × 105 and 1.9 × 108 per swab. Importantly, the super-spreader of the facility, a P-Me patient, had the highest amount of MeV in URT. Additionally, the cut-off point of throat swab's C t between super-spreaders and non-super spreaders was found to be 24.78 (sensitivity, 1.00; specificity, 0.93; AUC (95% CI): 0.96 (0.90–1.00)). In two other patients whose C t in the throat swab sample was less than the cut-off point of super-spreaders, a patient was an infant who contacted the super-spreader of a company and a household. Another patient was an adult working at a company. It is noteworthy that only one M-Me patient examined in this study transmitted measles to another person (Fig. 2).
Results of a real-time reverse transcription (rRT) PCR in the throat swab samples of the transmission network of a measles outbreak, Yamagata Prefecture, Japan, March–April 2017. Circles, triangles and squares represent respective cases. Circles denote cases of modified measles (n = 35). Triangles denote cases of prodromal stage of typical measles at the point of sample collection, but which finally progressed to typical measles (n = 9). Squares denote cases of typical measles (n = 9). Lines and an arrow denote transmission routes. Transmission settings for linked cases are shown within rectangles. Symbols with dotted lines were not included in this study because of notification outside Yamagata. Cycle threshold (C t) is represented by differences of colours: C t ⩽ 24.78 is orange; 24.78 < C t ⩽ 32.39 is light orange; and 32.39 < C t ⩽ 40.0 is beige. White symbols represent negative or equivocal results in the rRT-PCR. ND above a triangle means that the rRT-PCR was not done. This figure is modified from a figure by Komabayashi et al. [Reference Komabayashi12].
Discussion
For this study, we investigated the effectiveness of rRT-PCR for MeV in a measles outbreak dominated by M-Me cases in Yamagata Prefecture, Japan. Results indicated that the combination of PBMC and throat swab is useful for detecting MeV-RNA in M-Me patients. Furthermore, we demonstrated the potential of rRT-PCR in throat swab samples for predicting the candidates of super-spreader. The wider use of rRT-PCR with PBMC and throat swab is expected to be effective as a high-sensitivity test, a method for early detection of measles cases and a tool for preventing super-spreading events.
To date, M-Me has not been studied sufficiently. To understand M-Me-related studies among reports of the relevant literature, we searched PubMed on 15 February 2018 with two conditions, finding that English papers including ‘measles’ in the title were 11 030, although those containing ‘modified measles’ in the title were only 21. Moreover, evaluations by genetic testing of samples from M-Me cases are scarce [Reference Nagai16]. Accordingly, we tried to clarify rRT-PCR effectiveness in a measles outbreak dominated by M-Me cases. The WHO Global Vaccine Action Plan will engender a decrease in the number of measles cases worldwide [9]. Alternatively, the proportion of M-Me cases might increase because persons with one dose of measles vaccine, who might have primary or secondary vaccine failure [Reference Hickman10, Reference Holzmann11], account for a certain share of the population worldwide [Reference Trentini17]. Indeed, measles patients who are 20–39 years of age, which includes the generation of one-dose vaccination in Japan [Reference Inaida18, Reference Nishiura19], accounted for the greatest number of measles cases in this study. Our study will bring information for detecting M-Me cases properly, which might become a task for the elimination of measles in the world.
Results strongly suggest that the samples of PBMC and throat swab are fundamentally important for detecting MeV-RNA of M-Me cases in genetic tests. Given that MeV attaching the respiratory tract first becomes incorporated into PBMC (e.g., lymphocytes, monocytes and macrophages) in measles-infected persons [Reference Knipe1], it is reasonable that PBMC derived from M-Me patients, 80% of whom visited hospitals within two days after fever, showed the highest positive rate in the rRT-PCR (Tables 1, 2). Although the estimated number of MeV-RNA copies at the cut-off point of PBMC's C t in M-Me was low (Table 4), the high sensitivity of the rRT-PCR enables us to detect small quantities of MeV-RNA [Reference Hummel15]. Furthermore, throat swab samples showed the second highest positive rate in the rRT-PCR (Table 2) and recovered negative or equivocal results of PBMC samples (Table 3). Although urine was used for genetic tests in T-Me cases in earlier studies [Reference Magurano20, Reference Vaidya21], the specimen might not be suitable for M-Me cases based on our results (Table 2). Taken together, for certain detection of MeV-RNA using rRT-PCR, it is worth considering the specimen collection of whole blood and throat swab from cases suspected of M-Me.
A semi-quantitative capability of rRT-PCR contributes to the estimation of the prognosis of measles patients who immediately received medical attention. In this study, educational and awareness-raising activities by PHCs induced contacts who showed measles symptoms to visit a hospital promptly (Table 1). Consequently, some patients diagnosed as having M-Me eventually showed the three symptoms of T-Me (i.e., fever, catarrhal symptoms and rash). Results show that C t of P-Me was significantly lower than that of M-Me and that it was indistinguishable from that of T-Me (Fig. 1). Those results indicate that measles patients with milder symptoms in the early stage might be classified clinically into M-Me, but some of whom might be classified as P-Me patients based on rRT-PCR results. By sharing the information immediately, health workers can perform adequate treatment for patients who are estimated as having a worse prognosis and can thereby reduce the occurrence of complications, which are expected to improve the patients’ quality of life [Reference Thorrington22]. Furthermore, public health officials can conduct proactive countermeasures for P-Me efficiently as they do for T-Me (e.g., urge patients estimated as having a worse prognosis to remain indoors and giving outbreak response vaccinations to the contacts of patients) and decrease measles transmission further.
Super-spreading events in measles outbreaks might be predicted using rRT-PCR results from throat swab samples. In this study, three super-spreaders were estimated as possessing a great amount of MeV in URT. It is noteworthy that the super-spreader at a facility, diagnosed as M-Me at the point of sample collection, exhibited the highest amount of MeV. Currently, super-spreaders are only categorised retrospectively after epidemiological tracing [Reference Hummel15]. Our result, however, indicated that the results of rRT-PCR using throat swab samples offer some potential to predict super-spreader candidates. Further measles transmission resulting from contacts of the super-spreader candidates might be prevented through aggressive measles control by public health officials.
Evidence-based field epidemiological investigation enables efficient measles control by reducing economic costs of measles outbreaks. In this study, PHCs spent great effort for approximately 3700 subjects. Consequently, 53 (1.4%) cases were found to be positive for MeV. Furthermore, we found that five (0.1%) of the 3700 subjects had great amounts of MeV in URT (Fig. 2). Overall, maximum interventions should be conducted preferentially for measles patients with high infectiousness selected by symptoms, vaccine status, social activities and genetic test results. Conversely, close countermeasures might not be necessary when an M-Me case was judged as presenting low infectiousness. Earlier studies have shown that approximately 40% of economic costs related to measles outbreaks are public health costs [Reference Ghebrehewet23, Reference Suijkerbuijk24]. Field epidemiological investigation using results of genetic tests enables public health officials to administer focused countermeasures, which can ultimately reduce economic costs related to measles outbreaks.
Our study had several potential limitations. First, although we experienced the largest outbreak after measles elimination in Japan, the case numbers are insufficient to generalise the cut-off values of M-Me cases (Table 4) and the super-spreader candidates. Second, the results of rRT-PCR might have a bias because we were unable to ensure the uniform quality of clinical samples. Third, as a result of educational and awareness-raising activities by PHCs, most M-Me patients visited a hospital during the early stage after onset in this study. A positive rate and C t values of rRT-PCR in each specimen might differ greatly by the timing of sample collection. The three limitations that influence the decision of cut-off values are expected to be resolved through further research efforts based on our proposition that rRT-PCR is supportive of elucidating the condition of measles patients. The last limitation is that the classification of M-Me and T-Me by clinical symptoms was difficult when patients had underlying diseases. For instance, a T-Me patient with two doses of vaccine (Table 1), who was negative for rRT-PCR in throat swab sample and who was diagnosed as having catarrhal symptoms actually had hay fever. Conversely, results of rRT-PCR might support clinical diagnosis when patients had subtle symptoms.
In summary, results of this study show that the combination of swift and thorough surveillance and rRT-PCR for MeV enables administration of efficient measles to control through identification of M-Me cases and through estimation of P-Me and super-spreader candidates. Those benefits might engender measles patients’ quality of life improvement, economic cost reduction, measles outbreak elimination during early stages and finally measles eradication worldwide.
Acknowledgements
We thank all parties in public health centers, hospitals and prefectural office for their cooperation in ending the measles outbreak. Particularly, we appreciate the exhaustive work of public health nurses in an epidemiological investigation. This research was supported in part by the Research Program on Emerging and Re-emerging Infectious Diseases from the Japan Agency for Medical Research and Development, AMED (Grant number: JP17fk0108213).
Conflict of interest
All authors declare that they have no conflict of interest related to this study.








