INTRODUCTION
Staphylococcus aureus is a common community and hospital pathogen in humans. It is virulent, easily transmissible, carried for long periods, and rapidly acquires resistance to antibiotics. Reports showed that the number of S. aureus community-onset infections rose significantly in the late 1990s and early 2000s [Reference Munckhof1, Reference Nimmo2], particularly in the USA where very high attack rates are described for emergent epidemic strains [Reference Miller and Kaplan3]. While there is some evidence that the rates have been decreasing [Reference Landrum4], clinicians are still having to manage patients with S. aureus infections arising in the community that are both more virulent and more resistant to antibiotics than previously encountered.
In Australia most of the individual patient data concerning resistance and disease burden are derived from hospital series of S. aureus bacteraemia. While this represents only a small fraction of all S. aureus infections, there are an estimated 6900 episodes of S. aureus bacteraemia annually (35/100 000), associated with 25–30% mortality; at least 12% of which are community-onset methicillin-resistant S. aureus (MRSA) infections [Reference Collignon5]. In a subsequent 2009 Australasian survey of invasive S. aureus disease in hospital patients, 6% of all cases were caused by isolates with a resistance profile consistent with non-multiresistant MRSA (nmMRSA) (resistant to <3 non-β-lactam antibiotics) [Reference Turnidge6] and there have been worrying reports of rapidly lethal infections in otherwise healthy young people [Reference Peleg and Munckhof7]. However, not all community-onset MRSA strains are equally virulent and there is marked regional variation in the distribution of both virulence and resistance in Australia [Reference Tong8].
Although S. aureus possesses a suite of virulence factors, most interest has been invested in one major determinant, Panton–Valentine leukocidin (PVL), because emergent virulent community-acquired MRSA isolates in the USA, Europe and Australia are frequently PVL positive [Reference Lina9–Reference Nimmo and Coombs11]. However, the exact role that PVL plays in pathogenesis remains a topic of intense controversy [Reference Lo12–Reference Wehrhahn17].
In this study we investigated prevalence and resistance of S. aureus isolates identified in a cohort of patients with community-onset S. aureus infections who were not admitted to major hospitals. In the first study of its kind, a stratified sampling strategy was used that was based on private laboratory results for infections that presented to community doctors and small community hospitals purposefully enriched for community MRSA infections.
METHODS
Study population
Clinical specimens submitted between April and September 2006 to a large community-based pathology service in Melbourne, Australia (Dorevitch Pathology) were used to identify cases of S. aureus infection. All S. aureus isolates were from routine clinical specimens processed according to standard laboratory protocols. Screening and duplicate specimens were excluded, together with specimens collected from residents of long-term care facilities, such as nursing homes, and hospital patients where the specimen was collected more than 48 h after admission or within 2 weeks post-discharge. Patient, specimen collection and isolate details were extracted from laboratory records (demographic information, specimen source, isolate resistance patterns) for all cases. All eligible MRSA cases were included. Because methicillin-sensitive S. aureus (MSSA) cases were far more numerous, a frequency-matched computer-generated random subsample of MSSA cases, representing about 5% of all MSSA, were selected for comparison. Confirmation that the infection was community-onset, and collection of further clinical details (infection site, severity as judged by the treating doctor, initial and subsequent antibiotic treatment, use of surgical drainage, history of recent antibiotic use, number of return visits) were obtained from the doctor who referred the specimen by telephone interview. Study patients were then allocated to one of three groups for analysis [MSSA, multi-resistant (m)MRSA, nmMRSA] according to the resistance pattern of their initial S. aureus isolate.
Microbiology
The following definitions were used to categorize S. aureus infections:
-
MSSA: Methicillin sensitive.
nmMRSA: Non-multiresistant. Resistant to methicillin but resistant to fewer than three antibiotics on a panel of eight antibiotics: (tetracycline, erythromycin, trimethoprim, ciprofloxacin, gentamicin, rifampin, fusidic acid and mupirocin).
mMRSA: Multiresistant. Resistant to methicillin and three or more other antibiotics on the same panel.
Dorevitch Pathology, Melbourne, performed all initial laboratory work on clinical isolates. S. aureus were identified by conventional methods; catalase, DNAse and latex tests. Any discordant results between the DNAse and latex tests were resolved by the tube coagulase test. Antibiotic susceptibilities were determined by disc diffusion on Sensititre agar according to the CDS disc method of antibiotic sensitivity testing [Reference Bell18]: tetracycline (10 μg), erythromycin (5 μg), ciprofloxacin (2·5 μg), gentamicin (10 μg), rifampin (1 μg), trimethoprim (5 μg), fusidic acid (2·5 μg), mupirocin (5 μg).
Of the 106 isolates corresponding to patients included in the final analysis, 102 were subjected to molecular characterization. Pulsed-field gel electrophoresis (PFGE) was performed on all isolates as described previously [Reference O'Brien, Udo and Grubb19], using a contour-clamped homogeneous electric field (CHEF) DR III system (Bio-Rad Laboratories Pty Ltd, USA). Chromosomal patterns were examined visually, scanned with a Quantity One device (Bio-Rad), and digitally analysed using FPQuest (Bio-Rad). S. aureus strain NCTC 8325 was used as a reference strain.
PCR for the detection of PVL determinants was performed on all isolates as described previously [Reference Fey20]. Putative MRSA multilocus sequence types were attributed on the basis of previously characterized MRSA PFGE types [Reference Nimmo21]. Multilocus sequence typing (MLST) was performed according to the strategy previously described [Reference Enright22] on all novel MRSA PFGE types and all PVL positive MSSA. Chromosomal DNA for (MLST) was prepared using a DNeasy tissue kit (Qiagen Pty Ltd, Germany). The sequences were submitted to http://www.mlst.net/ where an allelic profile was generated and a sequence type (ST) assigned. Clonal complex (CC) was determined using the eBURST V3 algorithm at the same website. Clones that diverged in only one of the seven MLST loci were considered to belong to the same CC. Double locus variants (DLVs) were included if the linking single locus variant (SLV) was present in the MLST database. Staphylococcal cassette chromosome mec (SCCmec) typing was performed on novel MRSA PFGE types as described previously [Reference Coombs23].
Statistical analysis
χ 2 and t tests were performed and exact 95% confidence intervals (CIs) computed to compare patient and infection characteristics by resistance group and PVL status. Logistic regression was used to estimate odds ratios (ORs) for PVL and clinically initiated surgical drainage treatment adjusted for age and infection type and site. Infection site categories were collapsed into ‘axilla’, ‘head and neck’, and ‘other’ for multivariate analysis. Analysis of PVL positivity and associations with patient and infection characteristics was restricted to 86 nmMRSA and MSSA cases where PVL results were available. The mMRSA infections were excluded from this comparative analysis as PVL genes had not been reported in these strains.
Estimating the overall prevalence of MRSA among community-onset infections had to take into account that all MRSA were included in the study, while only a random sample of MSSA cases was included. We first estimated how many of the total MSSA isolates from the collection period were likely to be associated with community-onset infections by applying the MSSA eligibility fractions for the study sample to the total MSSA isolates. This was used to reconstruct the probable profile of S. aureus isolates by resistance type associated with community-onset infections through this period. The 95% CIs for the prevalence estimates were also computed and weighted according to the original sampling fractions. All computations were performed with Stata version 11 (StataCorp., USA) and SPSS version 20 (SPSS Inc., USA).
Ethical approval
The project was approved by the University of Melbourne Human Research Ethics Committee (HREC no. 060125).
RESULTS
Recruitment outcomes
In total, 2094 S. aureus were isolated by Dorevtich Pathology in the 6-month study period, 11·7% were MRSA, 6·4% mMRSA and 5·3% nmMRSA (Fig. 1). The majority of referring doctors were general practitioners. We achieved a high response rate with 87% of the doctors approached agreeing to participate in the study. A number of doctors elected to contact their patients for permission before agreeing to participate, although this was not a requirement. The recruitment rate was slightly higher for those doctors who had referred a MRSA infection, but still approached 80% in the MSSA group. The median time lag from specimen collection to interview was 33·5 days, ranging from 9 to 84 days. There was no discernible difference in the demographic profile of patients of those doctors who decided to participate and those who did not.
Recruitment. mMRSA, Multiresistant methicillin-resistant S. aureus; nmMRSA, non-multiresistant MRSA; MSSA, methicillin-sensitive S. aureus.

The majority of mMRSA infections were reported by the doctors to be associated with a hospital admission (70%) and therefore did not meet eligibility criteria. Although the majority of nmMRSA infections were initially thought to be community onset, 45% of patients reported recent hospital exposure. All but 9% of MSSA cases were confirmed as community-onset. The final number of S. aureus isolates included that were confirmed to be community-onset infections was 106; 15 mMRSA, 34 nmMRSA and 57 MSSA (Table 1).
Clinical features by antibiotic resistance group
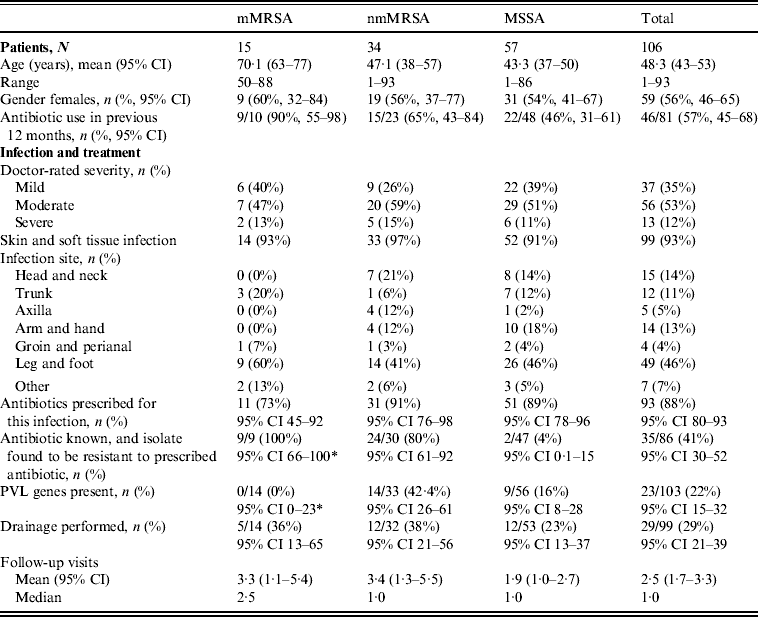
* One-sided 97·5% CI.
mMRSA, Multiresistant methicillin-resistant S. aureus; nmMRSA, non-multiresistant MRSA; MSSA, methicillin-sensitive S. aureus; CI, confidence interval; PVL, Panton–Valentine leukocidin.
Patient and infection characteristics
The majority of patients presented with skin and soft tissue infections (93%). Table 1 presents the patient profile and infections characteristics stratified by the different antibiotic resistance groups. Patients with mMRSA infections were on average older and, consistent with older age, were more likely to be female. They were also more likely to have been prescribed antibiotics in the previous 12 months.
Doctors were asked to self-rate the severity of their patient's infection as ‘mild’, ‘moderate’ or ‘severe’. The proportion of infections that were rated as ‘severe’ did not differ across resistance groups. Infections were less often reported as ‘mild’ for nmMRSA, but again this did not approach statistical significance.
The distribution of infection site did differ between the three resistance groups, with mMRSA infections largely confined to the trunk, leg and foot, while nmMRSA and MSSA infections were more widely distributed and included the head and neck, although leg and foot remained the most common infection sites.
Initial patient management did not differ by resistance group, despite the differences noted in patient profile and infection site. In this sample, any MRSA infection was more likely to have led to multiple follow-up visits than MSSA, but the differences in average visits in the isolate groups were not statistically significant.
Drainage was performed in less than a third of all cases, even though incision and drainage alone may be the optimal treatment for skin and soft tissue infections. On the other hand doctors reported that they had prescribed antibiotics for nearly all infections (88%, 95% CI 80–93). Most cases were treated with flucloxacillin, which was the common practice in Australia at the time (71/94, 95% CI 66–84). Figure 2 shows that the overall pattern of antibiotic prescribing was the same across all cases, regardless of the antibiogram of the isolate causing the infection, indicating that these infections did not present any differently to the treating doctor to signal the likelihood of flucloxacillin resistance.
Antibiotic resistance and antibiotic treatment by S. aureus resistance type. Shaded areas denote antibiotic resistance; ●, antibiotic initially prescribed for the infection; ○, change of antibiotic. mMRSA, Multiresistant methicillin-resistant S. aureus; nmMRSA, non-multiresistant MRSA; MSSA, methicillin-sensitive S. aureus.
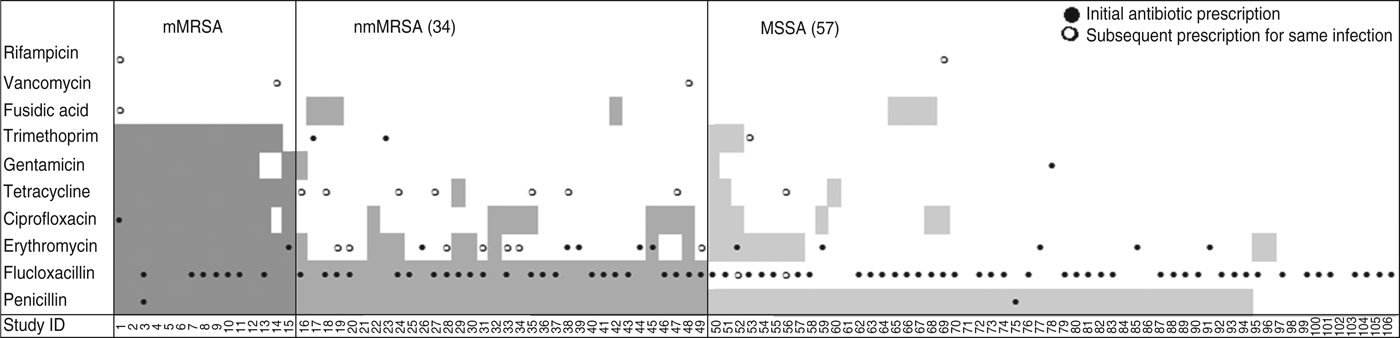
The isolate responsible for the infection was resistant to the prescribed antibiotic 100% of the time for mMRSA infections, 73% of the time for nmMRSA, and 3% of the time for MSSA. Given that we have under-sampled MSSA, the majority of doctors still prescribed microbiologically active antibiotics (estimated at 92·5% adjusting for sampling fractions). Subsequent prescription choices, and/or a decision to incise and drain, appeared to follow patient failure to respond to initial treatment and/or were influenced by the antibiotic resistance details provided subsequently on the laboratory report following the wound swab.
Presence of PVL encoding genes
The overall prevalence of PVL encoding genes (lukS-PV/lukF-PV) was 22% (23/103); 42·4% of nmMRSA (95% CI 26–61), 16% of MSSA (95% CI 8–28) and none of the MRSA (one-sided 97·5% CI 0–23). Patient and infection characteristics were further analysed by PVL status for pooled nmMRSA and MSSA strains (Table 2). Patients with PVL-positive infections were more likely to be younger with a 29-year difference in average age (P < 0·001). The distribution of infection site also varied according to PVL (P = 0·005), with PVL-positive infections seen more frequently in the head, neck, trunk and axilla, and less frequently in the leg and foot than for PVL-negative infections.
Clinical features by PVL status for nmMRSA and MSSA infections (n = 89)
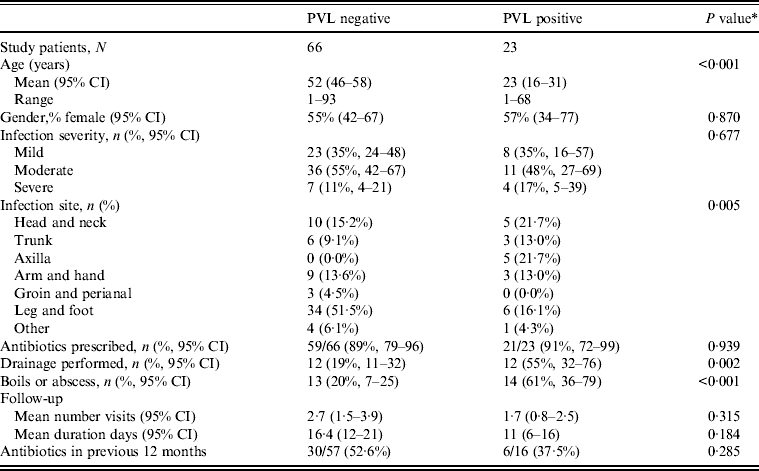
PVL, Panton–Valentine leukocidin; nmMRSA, non-multiresistant methicillin-resistant S. aureus; MSSA, methicillin-sensitive S. aureus; CI, confidence interval.
Antibiotic prescription was the initial treatment choice for 90% of cases in both groups, but interestingly drainage was performed in 55% of infections that were subsequently identified as PVL positive compared to only 19% of PVL negative (P = 0·005). Hence initial management decisions appeared to be influenced by isolates carrying PVL genes because in this study PVL isolates often presented as boils or abscesses.
Multivariate analysis was used to model PVL as a predictor of drainage being performed, adjusted for infection type, site and patient's age (Table 3). Most factors that were significantly associated with a decision to incise and drain in univariate analyses became non-significant in the multivariate model (PVL, infection site, type, patient's age). However, the presence of PVL-encoding genes remained strongly associated with a decision to lance and drain, with patients 4·8 times more likely to have an infection caused by a PVL-positive isolate (95% CI 1·5–15·2) if the doctor performed drainage, independent of infection type, site or patient's age.
Logistic regression model for drainage treatment (n = 76)
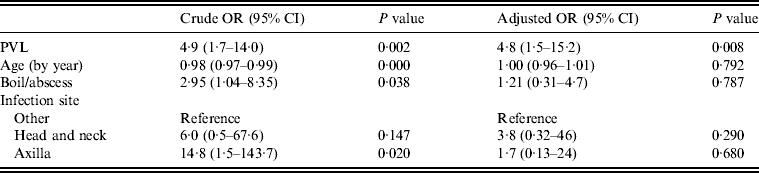
OR, Odds ratio; CI, confidence interval; PVL, Panton–Valentine leukocidin.
MRSA prevalence in community-onset infections
In order to estimate the proportion of community-onset infections caused by MRSA, the eligibility fractions for community onset that were determined in this study (Fig. 1) were applied to the total number of isolates believed to be associated with individual infection episodes. We estimated that over this 5-month period 3·5% (95% CI 2·2–4·9) of a total of 2094 infections referred to this community-based laboratory were due to MRSA.
Clonal diversity
The PFGE pattern for all 14 mMRSA characterized was equivalent to ST239-MRSA-III, the dominant healthcare-associated MRSA clone isolated in Melbourne hospitals [Reference Nimmo21]. All were PVL negative (see Table 4).
Molecular characterization of S. aureus isolates

PFGE, Pulsed-field gel electrophoresis; MLST, multilocus sequence type; SCCmec, staphylococcal cassette chromosome mec; CC, clonal complex; PVL, Panton–Valentine leukocidin; mMRSA, multiresistant methicillin-resistant S. aureus; nmMRSA, non-multiresistant MRSA; MSSA, methicillin-sensitive S. aureus; ST, sequence type; DLV, double locus variant.
* MLST inferred from PFGE pattern.
The 33 nmMRSA consisted of 13 clones: two healthcare-associated MRSA (HA-MRSA) and 11 community-associated MRSA (CA-MRSA). The two HA-MRSA clones, EMRSA-15 (three isolates) and EMRSA-16 (one isolate), were PVL negative. The CA-MRSA clones included three international PVL-positive clones; USA300 (one isolate), Taiwan CA-MRSA (one isolate) and South West Pacific (SWP CA-MRSA) (two isolates). The remaining eight CA-MRSA clones included the five dominant CA-MRSA clones isolated in Australia [Reference Nimmo and Coombs11]; PVL-positive Queensland (Qld) CA-MRSA (ST93-MRSA-IV), WA1 CA-MRSA (ST1-MRSA-IV), of which two isolates were PVL positive, WA23 CA-MRSA (ST45-MRSA-IV), WA2 CA-MRSA (ST78-MRSA-IV), and WA3 CA-MRSA (ST5-MRSA-IV). Single isolates of WA5 CA-MRSA (ST8-MRSA-IV), PVL-positive WA15 (ST59-MRSA-IV) and a novel CA-MRSA, PVL-positive ST5-MRSA-IV were also characterized. Overall, the nmMRSA population consisted of eight CCs and one singleton.
The MSSA population consisted of 25 different PFGE pulsotypes. The PVL determinants were found in eight different lineages, consisting of at least four clonal clusters and one singleton (ST1, ST DLV, ST20, ST30, ST93, ST121, ST152, ST772). Three of the PVL-positive lineages (ST1, ST30, ST93) were also identified in the nmMRSA population (WA1 CA-MRSA, SWP CA-MRSA, Qld CA-MRSA).
DISCUSSION
The aim of the study was to obtain an epidemiological snapshot of S. aureus infections in the community without making a priori assumptions about disease severity, resistance or virulence. This approach has enabled us to examine infections that are not captured through hospital admissions alone, and to understand the way these infections present and are managed in the community. In this study of community-based staphylococcal infections, the treating clinicians, mainly general practitioners, were asked to rate severity, report their initial and subsequent treatment decisions and describe the type and location of the infections they were managing. By including all patients with MRSA infection and a random sample of MSSA infections, we were able to create and compare three comparative groups with community-onset mMRSA, nmMRSA or MSSA infections.
We compared clinical presentation and management of nmMRSA and MSSA case-patients by the PVL status of their isolate. At the time of presentation, clinicians have to make treatment decisions without knowledge of the resistance profile, although this is generally available within 48–72 h. PVL status was not available to any of the clinicians during the study as it was only determined later for research purposes. The strength of this study therefore is that any differences in clinical presentation and initial management were likely to be linked to intrinsic resistance and/or virulence of the responsible strains, and not influenced by practitioner preconceptions about the significance of MRSA or PVL.
The majority of our patients with S. aureus infections in all groups presented with skin and soft tissue infections, and in this regard our results are complementary to hospital-based studies which often focus on bacteraemia [Reference Collignon5, Reference Turnidge6, Reference Blaine24–Reference Wang27]. Our principal finding was that CA-MRSA infections were still relatively uncommon in suburban Melbourne in late 2006, contributing an estimated 3·5% (95% CI 2·2–9·5) of all community-onset S. aureus infections, and were caused by multiple different clones. Moreover, when they did occur, the treating clinicians did not perceive increased severity. However, when nmMRSA and MSSA cases were combined and analysed by PVL status, it was clear that clinicians recognized something about these infections that led them to initiate incision and drainage procedures more frequently than for PVL-negative infections. This was also recently reported in hospital-based studies in New Zealand and Spain [Reference Daskalaki28, Reference Muttaiyah29]. In our study this association persisted even when we controlled for the presence of boils, age and infection site.
Some reports have suggested that PVL is a stable marker of the presence of methicillin resistance in community-onset infections [Reference Vandenesch10], but in contrast we found that 58% of nmMRSA isolates were PVL negative and that 15% of our MSSA isolates were PVL positive. Furthermore, it was the presence of PVL that correlated with a perceived need to undertake incision and drainage, not the detection of MRSA per se, as also reported in studies from Spain, New Zealand, Sweden and France [Reference Wang27, Reference Muttaiyah29–Reference Gillet32].
There is an emerging view that the biological role of PVL may be to allow direct invasion of human skin so that CA-MRSA may disseminate via clinically active skin infection rather than relying solely on asymptomatic colonization of mucus membranes [Reference Cohen33–Reference Munckhof37]. In support of this view are reports that PVL is less likely to be present in blood culture isolates than isolates obtained from aggressive skin infections or necrotizing pneumonia [Reference Ellington26, Reference Muttaiyah29]. Our data would fit this model but suggest that the presence of PVL is responsible for this independently of the methicillin resistance-encoding mecA gene. Whether PVL itself is directly responsible or is a marker of other virulence determinants with similar properties is still not completely determined [Reference Villaruz15], although the majority view now is that PVL is a principal virulence determinant [Reference Vandenesch14].
We also observed that PVL positive infections were more likely to occur in young people than non-PVL infections and more likely to involve the upper body, particularly the axilla or head and neck. The association between PVL and young age has also been noted by other investigators [Reference Muttaiyah29, Reference Gillet31, Reference Munckhof37]. Possible explanations include age-related acquisition of PVL-specific immunity. However, a recent study has reported that children acquire neutralizing antibody to PVL in an age-specific fashion reaching adult levels by the age of 3–5 years, but anti-PVL did not appear to protect against skin and soft tissue infections with PVL-expressing S. aureus [Reference Hermos, Yoong and Pier38]. Other explanations such as specific behaviours that increase contact between individuals carrying virulent clones may offer an alternative explanation for the age-specific incidence and site of infection location [Reference Cohen33].
The limitations of our study include the relatively small numbers of case-patients, the reliance on detection of PVL genes rather than gene expression in our isolates, the possible presence of other virulence factors linked to PVL that we have not yet assessed, and our inability to follow-up all patients; however, our response rate was high (87%). The fact that the patterns of initial antibiotic treatment are consistent across the three antibiotic resistance groups in these community-onset infections suggests that doctors are not seeing these patients differently, and therefore there are unlikely to be patient or infection characteristics apparent to a treating physician that could reliably predict whether a community-onset infection is caused by a MRSA strain.
The cross-sectional nature of the data also limited our ability to investigate the timing of treatment decisions in relation to the laboratory test results being made available to the doctor. While initial treatment decisions were not influenced by knowledge of the resistance phenotype, subsequent management decisions would have been influenced by knowledge of the resistance profile. However, this would not have applied to PVL status where the results were not determined until some time later and were not reported to the treating doctor.
As we identified patients with community-onset infections via laboratory test requests, it is also possible we over-selected for doctors who swab skin infections and/or prescribe antibiotics. Caution must therefore be exercised in extrapolating the prevalence of antibiotic prescription and drainage to the wider medical community managing community-onset infections. To address the issue of sample size and the limitations of cross-sectional analyses, a larger more comprehensive longitudinal study is currently underway based on the same community-based recruitment strategy.
Our study has shown that observational studies that avoid a priori assumptions about the S. aureus isolates on which they are based can provide a more complete picture of the relative incidence and importance of resistance and virulence in this ubiquitous human pathogen. Despite the undeniable importance of emerging resistance and virulence, most staphylococcal infections are milder than is apparent if only reports from hospital-based studies are considered. We estimate that 80% of S. aureus causing clinically significant infections in this Melbourne-based study were both methicillin sensitive and PVL negative, and that appropriately active antibiotics were prescribed in 92·5% of cases using clinical judgement and existing protocols.
CONCLUSION
We report high levels of clonal diversity and PVL positivity in nmMRSA and MSSA isolates responsible for community-onset infections. Infections with PVL-positive S. aureus were more common in younger patients and presented more often as axillary, head or neck infections. Treating doctors seem to detect something different in the presentation of PVL-positive infections and surgically lanced and drained these infections more frequently. The association between PVL and drainage is independent of the age of patient and type and site of the skin infection. While we did not find doctor-rated infection severity to be associated with PVL positivity, the observed increase in frequency of drainage procedures performed in PVL-positive infections is suggestive of a relationship between PVL positivity and the ability to form a drainable collection.
ACKNOWLEDGEMENTS
We thank the laboratory staff at Dorevitch Pathology and the Australian Collaborating Centre for Enterococcus and Staphylococcus Typing and Research, and the participating doctors who made this study possible. This project was funded by The University of Melbourne Research Grant Scheme (grant no. 5038860).







