Introduction
Probit analysis is often used in seed research to analyse germination data collected in response to a continuous variable that either promotes or prevents – for various reasons including seed death – germination (Hay et al., Reference Hay, Mead and Bloomberg2014). In particular, probit analysis is the basis of the seed viability equation that can be used to predict seed longevity depending on the storage environment (moisture content, temperature; Ellis and Roberts, Reference Ellis and Roberts1980a, Reference Ellis and Roberts1980b; Ellis, Reference Ellis2022). Probit analysis is also used to compare the longevity (shapes of the survival curves showing germination percentage vs. storage period) of different seed lots. Samples of seeds from the different lots are stored under the same conditions and subsamples are removed after different periods of time for a germination test (Hay et al., Reference Hay, Davies, Dickie, Merritt and Wolkis2022). These different ‘seed lots’ may represent different species, genotypes within a species (Lee et al., Reference Lee, Velasco-Punzalan, Pacleb, Valdez, Kretzschmar, McNally, Ismail, Sta. Cruz, Sackville Hamilton and Hay2019) or, for example, seeds that have experienced a different maturation environment or duration (Hay et al., Reference Hay, Timple and van Duijn2015; White et al., Reference White, Hay, Abeli and Mondoni2023), seeds resulting from distinct mating (e.g., self- vs. cross-pollination; Carta et al., Reference Carta, Bedini, Giannotti, Savio and Peruzzi2015), or which have been treated differently in some way after harvest as part of a designed experiment (e.g. Whitehouse et al., Reference Whitehouse, Hay and Ellis2015).
Probit analysis belongs to the generalized linear family of models (McCullagh and Nelder, Reference McCullagh and Nelder1989). In a generalized linear model (GLM), rather than fitting a linear relationship between the dependent variable (in our case, germination percentage) and the explanatory variable (storage period) directly, there is a transformation of the dependent variable and a linear relationship is fitted between this ‘linear predictor’ and the explanatory variable. In probit analysis, the germination data are transformed to the linear predictor probit value using the cumulative normal distribution function, Φ−1. Thus, it is assumed that there is a normal distribution in the response of the seeds to the explanatory variable. Most statistical tools use a maximum-likelihood estimation (MLE) procedure to fit a probit GLM while taking into account the binomial error distribution of germination data, which is determined by the number of seeds tested. The error distribution is binomial since each seed has two possible outcomes: it can either germinate or not germinate. The two parameters that are usually estimated are the intercept, which is the fitted or theoretical initial viability of the seed lot on the probit, linear predictor scale and the slope, the rate of change in the value of the linear predictor. Using the terminology of the viability equations, these parameters are referred to as K i and −1/σ, respectively. Thus, the GLM describing the change in viability during storage can be written:
where y is the proportion of seeds that germinate, v is the linear predictor expressed in probits, K i is the initial viability in probits, p is the storage period and σ is the time for viability to fall by one probit and hence, the standard deviation of the normal distribution of seed deaths over time. After applying probit analysis, the results are usually illustrated by plotting the cumulative normal distribution of germination over time which typically declines and is referred to as the seed ‘survival curve’. By fitting Equation 1 using probit analysis, the time for viability to fall to certain levels may also be estimated. For example, p 50 is the time for viability to fall to 50% which is often used as an index of seed longevity (Hay et al., Reference Hay, Davies, Dickie, Merritt and Wolkis2022).
Despite recent efforts aimed at describing and enhancing the use of free software for germination modelling (Carta et al., Reference Carta, Fernández-Pascual, Gioria, Müller, Rivière, Rosbakh, Saatkamp, Vandelook and Mattana2022; Onofri et al., Reference Onofri, Mesgaran and Ritz2022), seed survival modelling in such an open-source framework is still under-applied (but see Carta et al., Reference Carta, Bottega and Spanò2018). This may limit longevity studies – or the interpretation of such studies – among the seed science community. Here, we show how probit analysis can be applied in the free statistical software environment, R (R Core Team, 2023), to model seed survival data. In addition to showing how to fit the basic GLM to data for one seed lot (Box 1), thereby fitting Equation 1, we also show how to test hypotheses when survival curves for multiple seed lots or treatments are being compared (Supplementary Material 1). Lastly, we present the probit analysis code in which σ (Equation 1) is modelled as a function of the storage environment (Supplementary Material 2).
R script for fitting a simple single survival curve.
This is sample code for fitting a simple single survival curve (data for one seed lot). Original data are from Rezaei et al. (Reference Rezaei, Buitink and Hay2023) and are supplied as Supplementary Material 3.

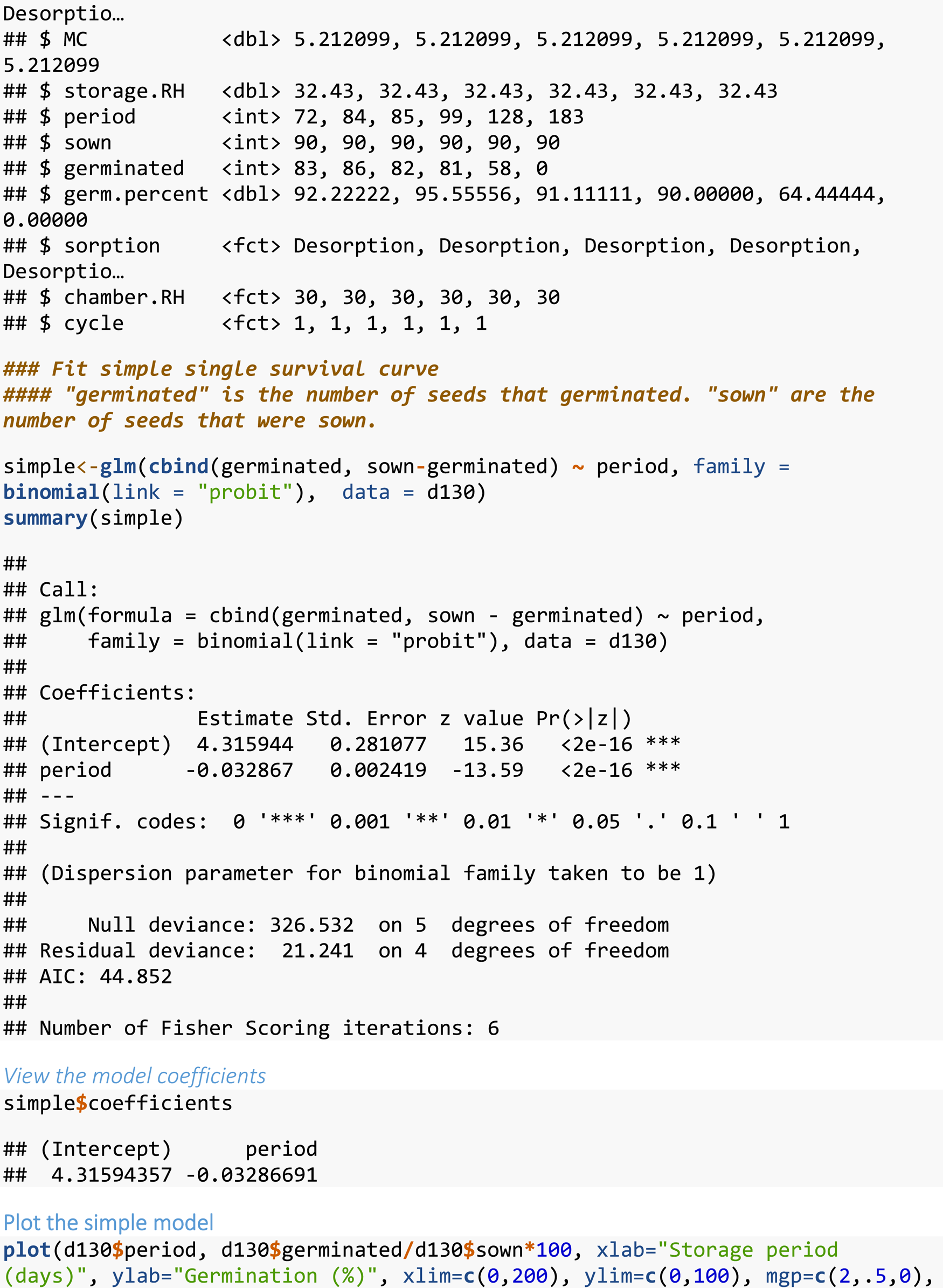
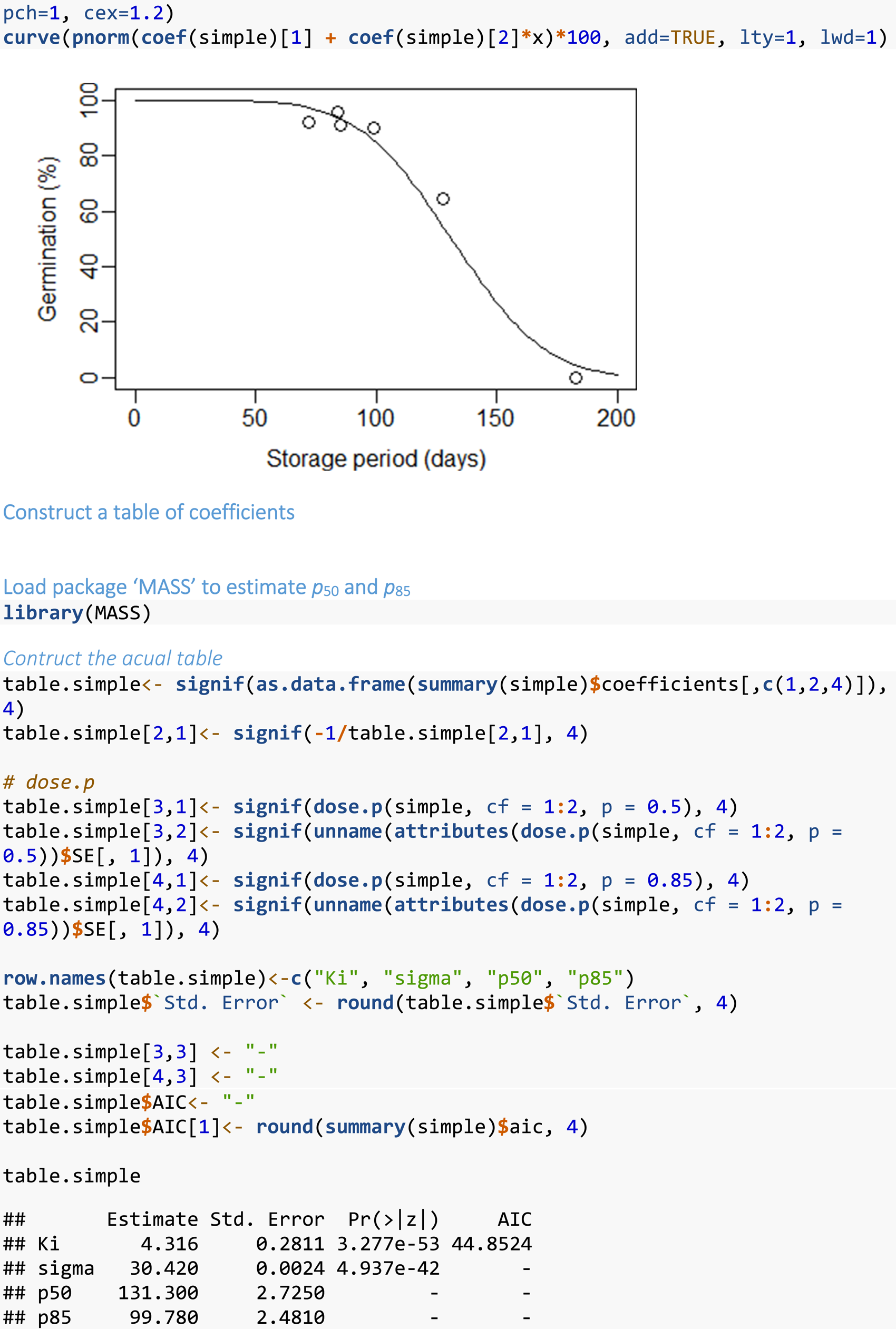
Fitting a simple single survival curve (data for one seed lot)
It may be appropriate to analyse the data for multiple stored seed lots completely independently, i.e., without any hypothesis testing (see below). This might occur for example, in a study where the aim is to screen diverse taxa for seed longevity (e.g., Probert et al., Reference Probert, Daws and Hay2009). In this case, the following code (Box 1) could be run for the data for each seed lot. The data should be arranged as quantity or ‘dose’ of the explanatory variable (in our case, storage period), number of seeds that germinated and number of seeds that were sown. Strings of observations where germination is equal to ~100%, at the start of storage, or to 0% at the end of storage should be excluded from the analysis; this is usually done manually. Although these values have less influence in the MLE fitting process, including a run of 100 or 0% values can be likened to trying to fit a linear relationship to data which has a ‘broken stick’ response, i.e. with a high or low plateau, before or after the slope. This means only including data from the last highest percentage germination value (but see below) to the first zero percentage germination value.
Fitting multiple survival curves (data for multiple seed lots) and hypothesis testing
Often when we study seed longevity, we are interested in the impact of some sort of treatment on subsequent seed longevity. Hence, we conduct storage experiments on multiple seed lots and want to test whether the survival curves are significantly different or not. This hypothesis testing can be done using approximate F-tests. The data from such designed experiments should be arranged as before, but with an additional factor variable which is the treatment code. Different models are then fitted to the data:
(i) the ‘independent model’ with different estimates for both the slope and intercept for each treatment (or species, seed lot, or genotype) (Fig. 1A);
(ii) the number of parameters in the model is reduced by estimating a single, common intercept estimate for all the seed lots (‘common intercept’; Fig. 1B);
(iii) instead of estimating a common intercept for the seed lots, we reduce the number of parameters compared with the independent model, by estimating a common value for the slope parameter for all the seed lots (‘common slope’; Fig. 1C);
(iv) lastly, we fit a model with common estimates for both the slope and intercept among all the seed lots (‘one line’; Fig. 1D).
Examples of fitting multiple survival curves and different models. (A) ‘Independent model’ with different estimates for both the slope and intercept for each treatment (or species, seed lot, or genotype); (B) the number of parameters in the model is reduced by estimating a single, common intercept estimate for all the seed lots (‘common intercept’); (C) instead of estimating a common intercept for the seed lots, we reduce the number of parameters compared with the independent model, by estimating a common value for the slope parameter for all the seed lots (‘common slope’); (D) lastly, we fit a model with common estimates for both the slope and intercept among all the seed lots (‘one line’).

The F-test is used to test whether there is a significant increase in the residual deviance after fitting a model with fewer parameters (the independent model has the most parameters: an intercept, and slope for each seed lot/treatment). Thus, the residual deviance after fitting the common slope or common intercept models is compared against the residual deviance after fitting the independent model; if there has not been a significant increase (F-probability >0.05), the residual deviance after fitting the one-line model is compared with the residual deviance from the common slope and/or common intercept models. If there is a significant increase in residual deviance comparing the one-line model against the common slope or common intercept models (F-probability ≤0.05), then the choice of accepting one of these models can be based on the relative F-probability values for these two models when compared against the independent model (choosing the one with the highest F-probability), unless there is a scientific (i.e., biological) reason why the other is more appropriate. In this way, by the use of F-tests, we determine the model with the fewest number of parameters that adequately describes the data.
In some experiments, there may be a sensible reason why seed lots do not have the same intercept, the most obvious being if seeds from the same bulk seed lot are stored under different storage conditions, in which case the treatment is the different storage environments (e.g., storage moisture content, as described below). Otherwise, this hypothesis is perhaps in general, less relevant to test. In the initial model development that led to the viability equations, it was concluded that the slopes of the survival curves are the same for all seed lots within a species if stored under identical conditions of moisture content and temperature (Ellis and Roberts, Reference Ellis and Roberts1980a, Reference Ellis and Roberts1980b). Although it is now generally accepted that this is not necessarily the case (Whitehouse et al., Reference Whitehouse, Hay and Ellis2018; Lee et al., Reference Lee, Velasco-Punzalan, Pacleb, Valdez, Kretzschmar, McNally, Ismail, Sta. Cruz, Sackville Hamilton and Hay2019; Ellis, Reference Ellis2022), this is still an obvious simplification of the full (independent) model and it is important to test this assumption. If the seed lots (or treatments) being compared are not from the same or closely related species, there is no underlying reason to expect them to respond similarly to a particular storage environment. Having established that it is possible to constrain the survival curves from the different seed lots (or treatments) to either a common intercept or a common slope, the next step is to examine whether it is possible to accept a model where both these parameters are constrained among different seed lots, i.e. whether a single survival curve adequately explains the data obtained for seeds subjected to or derived from different treatments. Testing whether a single curve can be fitted is in fact testing whether there has been a significant effect of the treatment on the subsequent response to seed storage. Often, we summarize this as ‘seed longevity’ although it is not a direct comparison of what we actually normally measure as seed longevity (e.g. p 50).
The R code for fitting this set of models and for the corresponding F-tests is provided in Supplementary Material 1. In the table showing the model comparisons from the F-tests, all the comparisons are significant at the 5% level (F-probability <0.05) except the comparison of the common intercept model vs. the independent model. Hence, in this case, the common intercept model can be accepted.
One-step fitting of the viability equation
The effect of storage environment on seed longevity was originally determined by storing seeds at different moisture contents and temperatures according to a factorial design (Ellis and Roberts, Reference Ellis and Roberts1980b). Then, individual survival curves were fitted to get estimates of the intercept and slope for each combination of storage moisture content and temperature using probit analysis. Lastly, regression analysis was used to model the effect of moisture content or temperature on σ (i.e., −1/slope from the probit analysis). Separately, these two relations can be written as:
at constant temperature, t:
and at constant moisture content, m:
Combined, they become
where m is seed moisture content during storage (f.wt. basis) and t is the temperature of storage; K E, C W, C H and C Q are ‘species constants’ which, it was assumed, do not vary among seed lots within the same species, thereby resulting in the same value for σ when seeds of the same species are stored at the same moisture content and temperature (see above).
However, this is a two-step process which does not take into account the error in the germination data. It also requires a factorial design for the storage experiments, with seeds stored at fixed moisture contents at one or more fixed temperatures, though as a ‘short cut’, temperature can be disregarded since we expect all seed lots to respond similarly to changes in temperature (Dickie et al., Reference Dickie, Ellis, Kraakj, Ryder and Tompsett1990). As an alternative approach, and given the advances in statistical software, it was recommended that a ‘one-step’ approach be used (Hay et al., Reference Hay, Mead, Manger and Wilson2003), to directly fit the full model (i.e., Equation 4 substituted into Equation 1) while taking into account all of the error in the data, including the binomial sampling error. This approach also means that the storage moisture content and temperature are entered as variables and they can vary. So, for example, the moisture content does not need to be exactly the same for samples of seeds stored at different temperatures. The model that is therefore fitted using the one-step approach is:
This can be reduced if seeds are stored at one temperature or moisture content (i.e., fitting Equations 2 or 3, respectively).
Hypothesis testing can also be carried out, to test whether parameters of the equation vary for different seed lots or following different treatments, as for example, done by Hay et al. (Reference Hay, Mead, Manger and Wilson2003) for different ecotypes of Arabidopsis thaliana. The process for this is similar to that for comparing the survival curves for multiple seed lots, above. We provide the code for fitting Equation 2 in Supplementary Material 2, which can easily be modified to fit equations 3 or 5. In this code, we test whether K t and C W are the same depending on whether seeds are losing (desorbing) or gaining (adsorbing) moisture (Rezaei et al., Reference Rezaei, Buitink and Hay2023). In the model comparison table, the F-probabilities are all greater than 0.05, hence in this case, it can be concluded that the one-line relationship adequately explains the data. That is, there are not significant differences in K t (and hence, assuming the effect of temperature on seed longevity is also the same, K E) and C W depending on whether the seeds are de- or adsorbing moisture.
Fitting a modified survival curve
There are two main modifications that might be appropriate to consider, when the data are indicating that either (a) initial viability increases during the first period of storage due to the breaking of dormancy due to ‘after-ripening’ (Baskin and Baskin, Reference Baskin and Baskin2022); or (b) there is a ‘lag’ phase before percentage viability declines, but the viability during this phase is hovering at a level that is significantly below 100% (Mead and Gray, Reference Mead and Gray1999). The ability to model the after-ripening effect is greater when a significant proportion of the seed lot is dormant when the seeds are first put into the storage environment and if there are a number of data points collected during the breaking of dormancy (Whitehouse et al., Reference Whitehouse, Hay and Ellis2015, Reference Whitehouse, Hay and Ellis2018; Ellis, Reference Ellis2022). This may mean that it is helpful to sample more frequently at the start of the storage experiment, if the study of after-ripening is an objective. We have not provided these options here, but are working on the R code, depending on the needs of the seed research community.
Conclusions
Here, we have shown how to apply probit analysis in R to model germination data for stored seeds, either from designed experiments or data that is collected for example, as part of routine viability monitoring of seed/gene banks. This should help advance our understanding of seed longevity: across different taxa, in response to different harvesting and/or post-harvest treatments, and in response to different storage environments. This will contribute to overarching efforts in relation to the conservation of (agro-)biodiversity, as well as improving our understanding of seed physiological quality and the factors that influence the natural capital value of seeds (Mattana et al., Reference Mattana, Ulian and Pritchard2022). As well as generating survival curves from relatively simple experiments, we also provide the code to proceed with one-step fitting of the viability equation describing the effect of change in storage moisture content on the slope of the survival curve. This can be adapted to also (or perhaps separately) model the effect of change in storage temperature on the survival curve or for example, see how non-storage variables are related to seed longevity (White et al., Reference White, Hay, Abeli and Mondoni2023). This is of particularly importance when examining seed longevity variation in natural systems. For instance, if seed longevity was estimated across multiple seed lots and taxa or if storage conditions were not properly known for all seed lots (e.g. seed ‘storage’ in the soil; Moravcová et al., Reference Moravcová, Carta, Pyšek, Skálová and Gioria2022), our codes could be adapted to model seed longevity while accounting for the statistical non-independence of data points due to phylogenetic relatedness of taxa or shared storage conditions among groups of seed lots (see Carta et al. Reference Carta, Fernández-Pascual, Gioria, Müller, Rivière, Rosbakh, Saatkamp, Vandelook and Mattana2022 for a detailed description of this methodological approach). Within seed science, however, it is not just germination data for stored seeds that are modelled using probit analysis. Population-based threshold models similarly assume that there are underlying normal distributions in the germination response of seeds to variables such as time in the germination test, water potential, salinity, etc. (Bradford and Bello, Reference Bradford, Bello, Buitink and Leprince2022). Hence, our codes could also be adapted to analyse data from other experiments where the objective is to understand the seed germination response to a single or multiple continuous variables (Hay et al., Reference Hay, Mead and Bloomberg2014). Additionally, the demonstration we have provided could also be used to model longevity and fit survival curves for pollen as well as fern and bryophyte spores.
Data availability
The data used in this demonstration were originally presented in Rezaei et al. (Reference Rezaei, Buitink and Hay2023). We provided a subset of those data as well as our R script in the supplementary materials.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0960258524000291.
Acknowledgements
The authors wish to thank two anonymous reviewers and the handling editor, Olivier Leprince, whose comments greatly improved the manuscript and the R code. Author institutions provided institutional support.
Financial support
This research received no specific grant from any funding agency, commercial or not-for-profit sectors.
Conflicts of interest
The authors declare none.
Author contributions
FRH and DW conceived the idea, AC, FRH and DW designed it, AC and DW compiled the codes, SR and FRH provided the data and Genstat codes, FRH led the writing with contributions from all authors.



