Introduction
Dryland regions, characterized by erratic rainfall, degraded soils and extreme temperatures, are among the most climate-sensitive agroecosystems (Ahmed et al. Reference Ahmed, Hayat, Ahmad, ul-Hassan, Kheir, ul-Hassan, Ur-rehman, Shaheen, Raza and Ahmad2022). These fragile environments account for a substantial share of global wheat cultivation, yet productivity is persistently constrained by recurrent drought, heatwaves, and soil salinization (Badaeva et al. Reference Badaeva, Konovalov, Knüpffer, Fricano, Ruban, Kehel, Zoshchuk, Surzhikov, Neumann, Graner, Hammer, Filatenko, Bogaard, Jones, Özkan and Kilian2022; Farooq et al. Reference Farooq, Zahra, Ullah, Nadeem, Rehman, Kapoor, Al-Hinani and Siddique2024a, Reference Farooq, Rafique, Zahra, Rehman and Siddique2024b). Given that wheat (Triticum aestivum L.) contributes about 20% of global caloric intake, enhancing its resilience to dryland stresses is an urgent priority for food security (Bohra et al. Reference Bohra, Kilian, Sivasankar, Caccamo, Mba, McCouch and Varshney2022). Modern wheat cultivars possess a relatively narrow genetic base for abiotic stress adaptation due to domestication bottlenecks and intensive selection, resulting in the loss of allelic and structural variants associated with drought tolerance, heat resilience, salinity exclusion, and nutrient-use efficiency (Özkan et al. Reference Özkan, Willcox, Graner, Salamini and Kilian2011; Maccaferri et al. Reference Maccaferri, Harris, Twardziok, Pasam, Gundlach, Spannagl, Ormanbekova, Lux, Prade, Milner, Himmelbach, Mascher, Bagnaresi, Faccioli, Cozzi, Lauria, Lazzari, Stella, Manconi, Gnocchi, Moscatelli, Avni, Deek, Biyiklioglu, Frascaroli, Corneti, Salvi, Sonnante, Desiderio, Marè, Crosatti, Mica, Özkan, Kilian, De Vita, Marone, Joukhadar, Mazzucotelli, Nigro, Gadaleta, Chao, Faris, Melo, Pumphrey, Pecchioni, Milanesi, Wiebe, Ens, MacLachlan, Clarke, Sharpe, Koh, Liang, Taylor, Knox, Budak, Mastrangelo, Xu, Stein, Hale, Distelfeld, Hayden, Tuberosa, Walkowiak, Mayer, Ceriotti, Pozniak and Cattivelli2019; Sharma et al. Reference Sharma, Schulthess, Bassi, Badaeva, Neumann, Graner, Özkan, Werner, Knüpffer and Kilian2021; Badaeva et al. Reference Badaeva, Konovalov, Knüpffer, Fricano, Ruban, Kehel, Zoshchuk, Surzhikov, Neumann, Graner, Hammer, Filatenko, Bogaard, Jones, Özkan and Kilian2022). Consequently, further yield improvement in dryland systems is increasingly constrained by limited genetic diversity for stress-responsive traits.
The wild wheat relatives (WWRs), wild species across the Triticeae tribe, including Triticum, Aegilops and allied genera such as Thinopyrum Secale and Elymus, offer a powerful solution to this bottleneck (Kilian et al. Reference Kilian, Mammen, Millet, Sharma, Graner, Salamini, Hammer and Özkan2011; Brozynska et al. Reference Brozynska, Furtado and Henry2016; Sharma et al. Reference Sharma, Schulthess, Bassi, Badaeva, Neumann, Graner, Özkan, Werner, Knüpffer and Kilian2021). Among these, three taxa emerge as top-tier donors for dryland breeding: Triticum dicoccoides (wild emmer), Aegilops tauschii (D-genome donor) and Ae. crassa. These species combine ecological adaptation, genomic characterization and demonstrated introgression feasibility. These species evolved under strong and heterogeneous selection pressures in the Fertile Crescent and adjacent dry regions, leading to the accumulation of adaptive traits relevant to water-limited and stress-prone environments (Peleg et al. Reference Peleg, Fahima, Krugman, ABBO, Yakir, KOROL and SARANGA2009; Kilian et al. Reference Kilian, Mammen, Millet, Sharma, Graner, Salamini, Hammer and Özkan2011; Munns et al. Reference Munns, James, Xu, Athman, Conn, Jordans, Byrt, Hare, Tyerman, Tester, Plett and Gilliham2012; Brozynska et al. Reference Brozynska, Furtado and Henry2016; Gaurav et al. Reference Gaurav, Arora, Silva, Sánchez-Martín, Horsnell, Gao, Brar, Widrig, John Raupp, Singh, Wu, Kale, Chinoy, Nicholson, Quiroz-Chávez, Simmonds, Hayta, Smedley, Harwood, Pearce, Gilbert, Kangara, Gardener, Forner-Martínez, Liu, Yu, Boden, Pascucci, Ghosh, Hafeez, O’Hara, Waites, Cheema, Steuernagel, Patpour, Justesen, Liu, RuddRudd, Avni, Sharon, Steiner, Kirana, Buerstmayr, Mehrabi, Nasyrova, Chayut, Matny, Steffenson, Sandhu, Chhuneja, Lagudah, Elkot, Tyrrell, Bian, Davey, Simonsen, Schauser, Tiwari, Randy Kutcher, Hucl, Li, Liu, Mao, Xu, Brown-Guedira, Faris, Dvorak, Luo, Krasileva, Lux, Artmeier, Mayer, Uauy, Mascher, Bentley, Keller, Poland and Wulff2021). Ae. tauschii harbours allelic variation for deep rooting, transpiration efficiency and sodium exclusion, including the well-characterized HKT1;5-D allele conferring salinity tolerance – a rare abiotic stress locus with demonstrated deployment relevance (Munns et al. Reference Munns, James, Xu, Athman, Conn, Jordans, Byrt, Hare, Tyerman, Tester, Plett and Gilliham2012; Gaurav et al. Reference Gaurav, Arora, Silva, Sánchez-Martín, Horsnell, Gao, Brar, Widrig, John Raupp, Singh, Wu, Kale, Chinoy, Nicholson, Quiroz-Chávez, Simmonds, Hayta, Smedley, Harwood, Pearce, Gilbert, Kangara, Gardener, Forner-Martínez, Liu, Yu, Boden, Pascucci, Ghosh, Hafeez, O’Hara, Waites, Cheema, Steuernagel, Patpour, Justesen, Liu, RuddRudd, Avni, Sharon, Steiner, Kirana, Buerstmayr, Mehrabi, Nasyrova, Chayut, Matny, Steffenson, Sandhu, Chhuneja, Lagudah, Elkot, Tyrrell, Bian, Davey, Simonsen, Schauser, Tiwari, Randy Kutcher, Hucl, Li, Liu, Mao, Xu, Brown-Guedira, Faris, Dvorak, Luo, Krasileva, Lux, Artmeier, Mayer, Uauy, Mascher, Bentley, Keller, Poland and Wulff2021). T. dicoccoides exhibits substantial intraspecific variation for drought and heat adaptation, including root system plasticity, osmotic adjustment and membrane stability, with several mapped QTL and introgression-derived lines under evaluation (Peleg et al. Reference Peleg, Fahima, Krugman, ABBO, Yakir, KOROL and SARANGA2009; Avni et al. Reference Avni, Nave, Barad, Baruch, Twardziok, Gundlach, Hale, Mascher, Spannagl, Wiebe, Jordan, Golan, Deek, Ben-Zvi, Ben-Zvi, Himmelbach, MacLachlan, Sharpe, Fritz, Ben-David, Budak, Fahima, Korol, Faris, Hernandez, Mikel, Levy, Steffenson, Maccaferri, Tuberosa, Cattivelli, Faccioli, Ceriotti, Kashkush, Pourkheirandish, Komatsuda, Eilam, Sela, Sharon, Ohad, Chamovitz, Mayer, Stein, Ronen, Peleg, Pozniak, Akhunov and Distelfeld2017; Balla et al. Reference Balla, Kamal, Tahir, Gorafi, Abdalla and Tsujimoto2025). Although less widely deployed, Ae. crassa demonstrates coordinated ion homeostasis and stress-responsive gene expression, marking it as a high-potential but still exploratory donor for salinity resilience (Pour-Aboughadareh et al. Reference Pour-Aboughadareh, Bocianowski, Jamshidi and Turkoglu2025). Additional taxa, including Ae. speltoides or T. araraticum, contribute micronutrient enrichment and grain quality traits, though their breeding deployment remains limited and more exploratory (Badaeva et al. Reference Badaeva, Konovalov, Knüpffer, Fricano, Ruban, Kehel, Zoshchuk, Surzhikov, Neumann, Graner, Hammer, Filatenko, Bogaard, Jones, Özkan and Kilian2022; Zeibig et al. Reference Zeibig, Kilian and Frei2022).
Historically, WWR utilization has been uneven. Major progress has been made in single-gene biotic resistance, whereas progress in complex abiotic stress tolerance has been slower due to polygenic inheritance, linkage drag and genotype × environment interactions (Singh et al. Reference Singh, Hodson, Huerta-Espino, Jin, Bhavani, Njau, Herrera-Foessel, Singh, Singh and Govindan2011; Tian et al. Reference Tian, Wang, Liu and Zhao2025). This contrast highlights an important distinction: traits governed by major loci (e.g., disease resistance, HKT1;5-D) are closer to deployment, whereas complex drought, heat, and nutrient-use efficiency traits remain in advanced discovery or pre-breeding stages. Recent genomic advances have strengthened the bridge between discovery and deployment. High-quality reference genomes and wheat pangenomes now capture extensive presence–absence variation (PAV) and structural polymorphisms previously inaccessible (IWGSC 2018; Walkowiak et al. Reference Walkowiak, Gao, Monat, Haberer, Kassa, Brinton, Ramirez-Gonzalez, Kolodziej, Delorean, Thambugala, Klymiuk, Byrns, Gundlach, Bandi, Siri, Nilsen, Aquino, Himmelbach, Copetti, Ban, Venturini, Bevan, Clavijo, Koo, Ens, Wiebe, N’Diaye, Fritz, Gutwin, Fiebig, Fosker, Fu, Accinelli, Gardner, Fradgley, Gutierrez-Gonzalez, Halstead-Nussloch, Hatakeyama, Koh, Deek, Costamagna, Fobert, Heavens, Kanamori, Kawaura, Kobayashi, Krasileva, Kuo, McKenzie, Murata, Nabeka, Paape, Padmarasu, Percival-Alwyn, Kagale, Scholz, Sese, Juliana, Singh, Shimizu-Inatsugi, Swarbreck, Cockram, Budak, Tameshige, Tanaka, Tsuji, Wright, Wu, Steuernagel, Small, Cloutier, Keeble-Gagnère, Muehlbauer, Tibbets, Nasuda, Melonek, Hucl, Sharpe, Clark, Legg, Bharti, Langridge, Hall, Uauy, Mascher, Krattinger, Handa, Shimizu, Distelfeld, Chalmers, Keller, Mayer, Poland, Stein, McCartney, Spannagl, Wicker and Pozniak2020; Tiwari et al. Reference Tiwari, Saripalli, Sharma and Poland2024; Farooq et al. Reference Farooq, Frei, Zeibig, Pantha, Özkan, Kilian and Siddique2025). Population genomics in Ae. tauschii and T. dicoccoides has identified loci associated with root traits, ion homeostasis and stress-responsive pathways directly relevant to dryland agriculture (Avni et al. Reference Avni, Nave, Barad, Baruch, Twardziok, Gundlach, Hale, Mascher, Spannagl, Wiebe, Jordan, Golan, Deek, Ben-Zvi, Ben-Zvi, Himmelbach, MacLachlan, Sharpe, Fritz, Ben-David, Budak, Fahima, Korol, Faris, Hernandez, Mikel, Levy, Steffenson, Maccaferri, Tuberosa, Cattivelli, Faccioli, Ceriotti, Kashkush, Pourkheirandish, Komatsuda, Eilam, Sela, Sharon, Ohad, Chamovitz, Mayer, Stein, Ronen, Peleg, Pozniak, Akhunov and Distelfeld2017; Gaurav et al. Reference Gaurav, Arora, Silva, Sánchez-Martín, Horsnell, Gao, Brar, Widrig, John Raupp, Singh, Wu, Kale, Chinoy, Nicholson, Quiroz-Chávez, Simmonds, Hayta, Smedley, Harwood, Pearce, Gilbert, Kangara, Gardener, Forner-Martínez, Liu, Yu, Boden, Pascucci, Ghosh, Hafeez, O’Hara, Waites, Cheema, Steuernagel, Patpour, Justesen, Liu, RuddRudd, Avni, Sharon, Steiner, Kirana, Buerstmayr, Mehrabi, Nasyrova, Chayut, Matny, Steffenson, Sandhu, Chhuneja, Lagudah, Elkot, Tyrrell, Bian, Davey, Simonsen, Schauser, Tiwari, Randy Kutcher, Hucl, Li, Liu, Mao, Xu, Brown-Guedira, Faris, Dvorak, Luo, Krasileva, Lux, Artmeier, Mayer, Uauy, Mascher, Bentley, Keller, Poland and Wulff2021). Importantly, these discoveries are increasingly supported by practical deployment platforms, including synthetic hexaploid wheat (SHW), structured introgression libraries and advanced backcross populations. These platforms enhance recombination, reduce linkage drag and improve compatibility with speed breeding and genomic selection (GS) pipelines (Trethowan and Mujeeb-Kazi Reference Trethowan and Mujeeb-Kazi2008; Mastrangelo et al. Reference Mastrangelo, Roncallo, Matny, Čegan, Steffenson, Echenique, Šafář, Battaglia, Barabaschi, Cattivelli and Özkan2023).
This review builds recent syntheses of WWR utilization (e.g., Farooq et al. Reference Farooq, Frei, Zeibig, Pantha, Özkan, Kilian and Siddique2025) but provides a targeted focus on dryland agriculture, integrating ecological adaptation, pre-breeding strategies and emerging genomic tools. In addition, it considers how international policy frameworks, including the Convention on Biological Diversity (CBD), the Nagoya Protocol and the International Treaty on Plant Genetic Resources for Food and Agriculture (ITPGRFA), shape access to and utilization of WWR genetic diversity (Engels and Ebert Reference Engels and Ebert2024). By positioning WWR not merely as reservoirs of genetic diversity but as strategic assets for climate-resilient wheat systems, this review highlights pathways to enhance wheat productivity, yield stability and nutritional quality in water-limited environments.
Origins and ecological adaptations of WWRs
WWRs (Triticum and Aegilops spp.) primarily originated in the Fertile Crescent and adjacent regions, encompassing parts of modern-day Türkiye, Palestine, Israel, Jordan, Lebanon, Syria, Iraq and Iran, recognized as a major centre of origin and crop diversity (Vavilov Reference Vavilov1926; Kilian et al. Reference Kilian, Mammen, Millet, Sharma, Graner, Salamini, Hammer and Özkan2011; Avni et al. Reference Avni, Nave, Barad, Baruch, Twardziok, Gundlach, Hale, Mascher, Spannagl, Wiebe, Jordan, Golan, Deek, Ben-Zvi, Ben-Zvi, Himmelbach, MacLachlan, Sharpe, Fritz, Ben-David, Budak, Fahima, Korol, Faris, Hernandez, Mikel, Levy, Steffenson, Maccaferri, Tuberosa, Cattivelli, Faccioli, Ceriotti, Kashkush, Pourkheirandish, Komatsuda, Eilam, Sela, Sharon, Ohad, Chamovitz, Mayer, Stein, Ronen, Peleg, Pozniak, Akhunov and Distelfeld2017; Eastwood et al. Reference Eastwood, Tambam, Aboagye, Akparov, Aladele, Allen, Amri, Anglin, Araya, Arrieta-Espinoza, Asgerov, Awang, Awas, Barata, Boateng, Magos Brehm, Breidy, Breman, Brenes Angulo, Burle, Castañeda-Álvarez, Casimiro, Chaves, Clemente, Cockel, Davey, De la Rosa, Debouck, Dempewolf, Dokmak, Ellis, Faruk, Freitas, Galstyan, García, Ghimire, Guarino, Harker, Hope, Humphries, Jamora, Jatoi, Khutsishvili, Kikodze, Kyratzis, León-Lobos, Liu, Mainali, Mammadov, Manrique-Carpintero, Manzella, Mat Ali, Medeiros, Mérida Guzmán, Mikatadze-Pantsulaia, Mohamed, Monteros-Altamirano, Morales, Müller, Mulumba, Nersesyan, Nóbrega, Nyamongo, Obreza, Okere, Orsenigo, Ortega-Klose, Papikyan, Pearce, Pinheiro de Carvalho, Prohens, Rossi, Salas, Singh Shrestha, Siddiqui, Smith, Sotomayor, Tacán, Tapia, Toledo, Toll, Vu, Vu, Way, Yazbek, Zorrilla and Kilian2022). The harsh and fluctuating environments of the Fertile Crescent, characterized by recurrent drought, extreme temperatures and nutrient-poor soils, exerted strong selective pressures that shaped the adaptive evolution of wild populations (Fig. 1; Peleg et al. Reference Peleg, Fahima, Korol, Abbo and Saranga2011; Brozynska et al. Reference Brozynska, Furtado and Henry2016). Domestication and subsequent breeding narrowed the genetic base of cultivated wheat, eliminating allelic and structural variants still present in wild taxa such as T. dicoccoides and Ae. tauschii (Özkan et al. Reference Özkan, Willcox, Graner, Salamini and Kilian2011 Avni et al. Reference Avni, Nave, Barad, Baruch, Twardziok, Gundlach, Hale, Mascher, Spannagl, Wiebe, Jordan, Golan, Deek, Ben-Zvi, Ben-Zvi, Himmelbach, MacLachlan, Sharpe, Fritz, Ben-David, Budak, Fahima, Korol, Faris, Hernandez, Mikel, Levy, Steffenson, Maccaferri, Tuberosa, Cattivelli, Faccioli, Ceriotti, Kashkush, Pourkheirandish, Komatsuda, Eilam, Sela, Sharon, Ohad, Chamovitz, Mayer, Stein, Ronen, Peleg, Pozniak, Akhunov and Distelfeld2017; Gaurav et al. Reference Gaurav, Arora, Silva, Sánchez-Martín, Horsnell, Gao, Brar, Widrig, John Raupp, Singh, Wu, Kale, Chinoy, Nicholson, Quiroz-Chávez, Simmonds, Hayta, Smedley, Harwood, Pearce, Gilbert, Kangara, Gardener, Forner-Martínez, Liu, Yu, Boden, Pascucci, Ghosh, Hafeez, O’Hara, Waites, Cheema, Steuernagel, Patpour, Justesen, Liu, RuddRudd, Avni, Sharon, Steiner, Kirana, Buerstmayr, Mehrabi, Nasyrova, Chayut, Matny, Steffenson, Sandhu, Chhuneja, Lagudah, Elkot, Tyrrell, Bian, Davey, Simonsen, Schauser, Tiwari, Randy Kutcher, Hucl, Li, Liu, Mao, Xu, Brown-Guedira, Faris, Dvorak, Luo, Krasileva, Lux, Artmeier, Mayer, Uauy, Mascher, Bentley, Keller, Poland and Wulff2021). Diversity losses were particularly pronounced in low-recombination genomic regions enriched with adaptive haplotypes (Maccaferri et al. Reference Maccaferri, Harris, Twardziok, Pasam, Gundlach, Spannagl, Ormanbekova, Lux, Prade, Milner, Himmelbach, Mascher, Bagnaresi, Faccioli, Cozzi, Lauria, Lazzari, Stella, Manconi, Gnocchi, Moscatelli, Avni, Deek, Biyiklioglu, Frascaroli, Corneti, Salvi, Sonnante, Desiderio, Marè, Crosatti, Mica, Özkan, Kilian, De Vita, Marone, Joukhadar, Mazzucotelli, Nigro, Gadaleta, Chao, Faris, Melo, Pumphrey, Pecchioni, Milanesi, Wiebe, Ens, MacLachlan, Clarke, Sharpe, Koh, Liang, Taylor, Knox, Budak, Mastrangelo, Xu, Stein, Hale, Distelfeld, Hayden, Tuberosa, Walkowiak, Mayer, Ceriotti, Pozniak and Cattivelli2019; Badaeva et al. Reference Badaeva, Konovalov, Knüpffer, Fricano, Ruban, Kehel, Zoshchuk, Surzhikov, Neumann, Graner, Hammer, Filatenko, Bogaard, Jones, Özkan and Kilian2022). These patterns underscore the significance of conserving wild populations as both evolutionary archives and reservoirs of alleles relevant for modern agriculture.
Genetic diversity and trait introgression from wild wheat relatives.

Drought adaptation
Drought represents one of the most pervasive selective pressures across the native landscapes of WWRs (Peleg et al. Reference Peleg, Fahima, Krugman, ABBO, Yakir, KOROL and SARANGA2009; Nevo and Chen Reference Nevo and Chen2010). T. dicoccoides, the progenitor of tetraploid wheat, has evolved robust drought-adaptive traits, including deep, highly branched root systems, efficient osmotic adjustment and superior water-use efficiency (Peleg et al. Reference Peleg, Fahima, Krugman, ABBO, Yakir, KOROL and SARANGA2009; Avni et al. Reference Avni, Nave, Barad, Baruch, Twardziok, Gundlach, Hale, Mascher, Spannagl, Wiebe, Jordan, Golan, Deek, Ben-Zvi, Ben-Zvi, Himmelbach, MacLachlan, Sharpe, Fritz, Ben-David, Budak, Fahima, Korol, Faris, Hernandez, Mikel, Levy, Steffenson, Maccaferri, Tuberosa, Cattivelli, Faccioli, Ceriotti, Kashkush, Pourkheirandish, Komatsuda, Eilam, Sela, Sharon, Ohad, Chamovitz, Mayer, Stein, Ronen, Peleg, Pozniak, Akhunov and Distelfeld2017). Experimental evaluations confirm that WWR accessions exhibit greater biomass stability and root plasticity under drought stress than modern cultivars (Pour-Aboughadareh et al. Reference Pour-Aboughadareh, Kianersi, Poczai and Moradkhani2021). In a pot study, screening of diverse Aegilops–Triticum accessions under water stress (90, 50, 25% FC) indicates that accessions with alien genomic constitutions (e.g., Ae. umbellulata, Ae. crassa, Ae. markgrafii, Ae. cylindrica, T. boeoticum) displayed strong induction of antioxidant enzymes and maintained higher pigment concentrations, suggesting biochemical buffering as a key adaptive mechanism (Ahmadi et al. Reference Ahmadi, Pour-Aboughadareh, Ourang, Mehrabi and Siddique2018). Recent integrative analyses combining physiological, biochemical, and transcriptomic data further demonstrate that drought responses in wheat are strongly genotype dependent, with tolerant cultivars exhibiting enhanced antioxidant activity and activation of ABA-mediated stress signalling pathways (Cevher-Keskin et al. Reference Cevher-Keskin, Yıldızhan, Sekmen, Fayetorbay, Sezerman, Özer, Onarıcı, Türkan and Tör2025). Complementing this, a two-year field study across irrigated and rainfed conditions revealed significant genotypic variation in drought-response traits across 110 Triticum accessions; water limitation reduced grain yield by 21% and stomatal conductance by 45%, with wild relatives maintaining superior performance via physiological regulation. Notably, T. dicoccoides, T. urartu, T. boeoticum and T. araraticum expressed isohydric strategies (water-conservative regulation), whereas T. monococcum subsp. sinskajae and T. durum cv. Sambadur showed anisohydric behaviour (maintaining gas exchange under stress), revealing multiple contrasting drought-adaptation strategies within the Triticum gene pool (Pantha et al. Reference Pantha, Kilian, Özkan, Zeibig and Frei2025). Together, pot and field evidence demonstrate that drought tolerance in WWR arises from integrated architectural, biochemical and stomatal regulation mechanisms, confirming their value as functional donors for synthetic wheat development and targeted trait introgression for low-water environments.
Heat and thermal stress adaptation
Triticum dicoccoides is a viable genetic reservoir for heat-stress resilience due to its direct lineage to domesticated durum wheat (T. durum) and contribution to the A and B subgenomes of bread wheat (T. aestivum), enabling the transfer of agronomic, physiological, and yield-related traits relevant to thermal tolerance (Peng et al. Reference Peng, Sun, Peng and Nevo2013). The feasibility of using this diversity has been demonstrated across both northern and southern evolutionary lineages of T. dicoccoides, corresponding to populations from the northern Fertile Crescent (e.g., Türkiye and the northern Levant) and southern regions (e.g., Palestine and adjacent areas), providing multiple allelic routes for improvement. Multi-environment, field-based evaluations across Japan and Sudan indicate that introgression from T. dicoccoides captures intraspecific variation for heat resilience in durum wheat × T. dicoccoides backcross families. Several families maintained high harvest index, elevated chlorophyll content, and reduced canopy temperature under severe heat stress, while others exhibited increased biomass and seed number, despite overall stress pressure (Balla et al. Reference Balla, Kamal, Tahir, Gorafi, Abdalla and Tsujimoto2025). These superior-performing lines contained introgressed chromosomal segments on chromosomes 1A, 2B, 5B, 6B and 7B, providing candidate regions for heat tolerance absent from elite durum backgrounds (Balla et al. Reference Balla, Gorafi, Kamal, Abdalla, Tahir and Tsujimoto2022). Moreover, WWR and wild-derived genotypes often outperformed elite cultivars for photosynthetic performance, with studies reporting a 2.3-fold range in CO₂ and light-saturated photosynthesis (Amax) and higher capacities for carboxylation, electron transport, and Rubisco activity, alongside faster dynamic responses to light via stomatal and non-photochemical quenching mechanisms (McAusland et al. Reference McAusland, Vialet‐Chabrand, Jauregui, Burridge, Hubbart‐Edwards, Fryer, King, King, Pyke, Edwards, Carmo‐Silva, Lawson and Murchie2020). Notably, the negative impact of heat stress on global wheat production is intensified by the genetic uniformity of modern cultivars, a consequence of breeding within narrow germplasm pools, particularly in developed countries (Fu et al. Reference Fu2015), emphasizing the urgency of reintegrating wild diversity. Harnessing T. dicoccoides enables the recovery of genetic variation lost during domestication and provides actionable alleles for high-temperature resilience, positioning WWR as essential assets for future-proofing wheat improvement.
Salinity and ion toxicity tolerance
Salinity is a major constraint to wheat productivity in dryland and irrigated systems, where ion toxicity and osmotic imbalance disrupt growth, photosynthesis and yield. The adaptive traits of WWRs are particularly relevant to resilience to abiotic stress (Table 1). WWRs, particularly those harbouring the D genome or alien genomic constitutions, provide mechanistically distinct pathways of salinity tolerance. Salinity tolerance in Ae. tauschii is conferred by alleles of the high-affinity potassium transporter HKT1;5, which maintain Na+/K+ balance and improve ionic homeostasis in wheat introgression lines under saline-soil conditions (Table 1; Munns et al. Reference Munns, James, Xu, Athman, Conn, Jordans, Byrt, Hare, Tyerman, Tester, Plett and Gilliham2012). In a pot study, Ae. crassa (MMDD genome) exhibited a broader, system-level salinity response (150–200 mM NaCl), maintaining higher biomass, chlorophyll content, and PSII efficiency (Fv/Fm, Fv/Fo) than other tested materials. This response was accompanied by elevated expression of salt overly sensitive (SOS1/2/3), Na+/H+ exchanger (NHX) and high-affinity potassium transporter (HKT) genes, indicating a coordinated, genotype-specific response involving ion exclusion, vacuolar sequestration, and membrane transport signalling rather than a species-wide trait (Pour-Aboughadareh et al. Reference Pour-Aboughadareh, Bocianowski, Jamshidi and Turkoglu2025). This response exceeded that of Ae. cylindrica and Ae. tauschii, identifying specific Ae. crassa genotypes as promising donors for salinity tolerance introgression. T. dicoccoides similarly exhibits adaptive variation relevant to saline environments. In a comparative assessment of 30 T. dicoccoides and 14 durum wheat genotypes, the salt tolerance index (0.21–0.97) exceeded that of durum (0.46–0.78) and was strongly associated with shoot K+/Na+ ratios and Na+ exclusion, but not shoot K+ concentration (Feng et al. Reference Feng, Cui, Lv, Bian, Wang, Song and Nie2018), confirming ion exclusion and homeostatic regulation as principal mechanisms. Root system traits, particularly length, surface area, and volume, were strongly correlated with tolerance, underscoring below-ground ion avoidance as a complementary adaptation.
Genetic contributions of wild wheat relatives to stress adaptation and resistance

Nutrient acquisition and efficiency
Improving nitrogen (N) and phosphorus (P) use efficiency is essential for wheat grown in low-input dryland systems, where nutrient limitations interact with heat and drought stress. WWRs, particularly D-genome donors, provide alleles that enhance nutrient capture and metabolic efficiency under stress. In a multi-environment field study in Sudan, Ae. tauschii-derived multiple-synthetic-derivative wheat lines showed wide variation in performance under heat × N stress; despite reductions in grain yield (−14%) and grain N uptake (−28%) under zero N, several lines maintained yield stability. Genome-wide association analysis (GWAS) identified 34 marker-trait associations (mostly in the D subgenome), including a locus on chromosome 5A influencing grain growth and a major locus on 3D explaining 88% of variation in grain N uptake, linked to stress signalling and nitrogen metabolism genes (Emam et al. Reference Emam, Tahir, Kamal, Gorafi, Tsujimoto and Ishii2025). These results demonstrate the deployable breeding value of Ae. tauschii introgressions for nitrogen use efficiency (NUE).
Pot experiments under reduced N–P–K fertilization further show that certain wild taxa sustain productivity by increasing NUE. Diploids were generally inefficient (except Ae. tauschii), while tetraploids (T. carthlicum, T. dicoccoides, T. timopheevii) and hexaploid T. sphaerococcum combined high uptake efficiency with nutrient-stress tolerance, marking them as priority donors (Gorny and Garczyński Reference Gorny and Garczyński2008). Similar patterns appear in alien chromosome addition lines from Leymus, Agropyron, Hordeum, Secale and others, where N deficiency reduced biomass and N uptake, while P deficiency constrained plant height and nutrient content (Liu et al. Reference Liu, Guo, Du and Hu2018), confirming that NUE/PUE (Phosphorus use efficiency) expression is genotype- and nutrient-dependent. Since NUE is multigenic, involving uptake, transport, assimilation, and remobilization (Islam et al. Reference Islam, Zhang, Zhao, She and Ma2021; Haider et al. Reference Haider, Farooq, Shah, Malghani, Awan, Habib-ur-rahman, Khalid and Ghaffar2023), WWRs provide valuable entry points for improving these pathways. Micronutrient acquisition further expands the value of WWR. T. dicoccoides is recognized as a strategic germplasm resource for increasing grain iron (Fe) and zinc (Zn) concentrations in cultivated wheat, with diploid and tetraploid wild taxa consistently outperforming modern T. aestivum and T. durum cultivars for grain micronutrient content (Liu et al. Reference Liu, Huang, Li, Liu, Yan, Tang, Zheng, Liu and Wu2021; Zeibig et al. Reference Zeibig, Kilian and Frei2022, Reference Zeibig, Kilian, Özkan, Pantha and Frei2024a). GWAS-based dissection in emmer-derived lines has identified genomic regions associated with higher Fe and Zn accumulation, reinforcing the biofortification potential of these taxa. Some wild relatives also demonstrate preferential Zn partitioning within tissues and grains (Fan et al. Reference Fan, Wang, Hu, Li, Liu, Liu, Ma and Liu2023), adding mechanistic targets for trait stacking and mineral-use-efficiency improvement.
Nutritional and grain quality traits
Beyond stress resilience, WWRs are valuable reservoirs of nutritional and grain-quality traits. Multiple studies highlight their superior micronutrient composition, particularly Fe, Zn and grain protein content (GPC). Diploid and tetraploid wild wheats consistently outperformed modern T. aestivum and T. durum for grain Fe and Zn under field conditions (Zeibig et al. 2024). Notably, diploid einkorn wheats (T. monococcum and T. urartu) and the tetraploid T. dicoccoides exhibited substantial genetic diversity for these minerals (Fu et al. Reference Fu, Bohuslavskyi and Atramentova2024; Zeibig et al. 2024), complementing findings that diploid taxa and T. timopheevii-lineage species (e.g., T. araraticum) possess higher micronutrient content and distinct gluten protein profiles relative to modern cultivars (Zeibig et al. Reference Zeibig, Kilian and Frei2022). These traits position WWRs as key contributors to global biofortification initiatives aimed at combating micronutrient malnutrition (Tadesse et al. Reference Tadesse, Gataa, Rachdad, Baouchi, Kehel and Alemu2023). Micronutrient enrichment is further supported by enhanced uptake and preferential grain partitioning, with several wild relatives showing superior Zn allocation to grain tissues compared to common wheat (Fan et al. Reference Fan, Wang, Hu, Li, Liu, Liu, Ma and Liu2023). GPC, which is a key trait for flour functionality and nutritional value, also benefits from WWR introgression. For example, chromosome 1Ns from Leymus mollis has been transferred into elite wheat, elevating GPC via improved storage protein subunit composition (Li et al. Reference Li, Liang, Huo, Yuan, Tang, Yang, Jia, Wang, Zhao and Li2025). Dietary fibres, such as β-glucan and arabinoxylan, which reduce glycemic index and support cardiovascular health, remain limited in the gene pool of modern wheat but show greater variation and enhancement potential in wild relatives (Kapoor et al. Reference Kapoor, Panigrahi, Singh, Kumar, Singh, Jan, Mir and Kumar2025). Genetic mapping efforts reinforce this potential: GWAS in T. timopheevii-derived lines have identified loci associated with gluten content and technologically relevant flour properties (Leonova et al. Reference Leonova, Kiseleva, Berezhnaya, Orlovskaya and Salina2023). Moreover, volatile compound profiles from T. timopheevii may offer opportunities for improving wheat flour aroma and flavour, adding a sensory dimension to quality improvement (Agarwal et al. Reference Agarwal, MacNaughtan, Liu, King and Foster2024).
Linking abiotic and biotic stress relevance in drylands
Climate-driven drought and heat can intensify pathogen risks by altering crop phenology, canopy microclimate (temperature and humidity gradients), and host susceptibility windows, thereby influencing epidemic development and host–pathogen interactions. WWRs have coevolved with diverse disease-causing pathogens, providing a wide range of resistance genes. For instance, T. dicoccoides, Ae. speltoides and Ae. geniculate contribute stem rust (Sr2), leaf rust (Lr51) and stripe rust (Yr40) resistance genes, respectively (Singh et al. Reference Singh, Hodson, Huerta-Espino, Jin, Bhavani, Njau, Herrera-Foessel, Singh, Singh and Govindan2011; Yaniv et al. Reference Yaniv, Raats, Ronin, Korol, Grama, Bariana, Dubcovsky, Schulman and Fahima2015; Steadham et al. Reference Steadham, Schulden, Kalia, Koo, Gill, Bowden, Yadav, Chhuneja, Erwin, Tiwari and Rawat2021). Recently, Mastrangelo et al. (Reference Mastrangelo, Roncallo, Matny, Čegan, Steffenson, Echenique, Šafář, Battaglia, Barabaschi, Cattivelli and Özkan2025) mapped resistance loci in T. dicoccoides populations and identified novel genomic regions associated with resistance to emerging stem rust pathotypes, expanding the genetic toolbox for breeders.
Several WWR-derived resistance loci can confer broad-spectrum protection; however, durability varies across environments, genetic backgrounds, and pathogen populations, highlighting the importance of pyramiding and multi-environment validation (Farooq et al. Reference Farooq, Frei, Zeibig, Pantha, Özkan, Kilian and Siddique2025). While abiotic stress tolerance is often polygenic, biotic resistance spans a continuum of genetic architectures: major-effect genes enable relatively straightforward introgression, whereas adult plant and quantitative rust resistance are polygenic and require more complex breeding strategies (McGee et al. Reference McGee, Zaleski‐Cox, Jayawardana, Tillman, Wally, Esquivel‐Garcia, Fernando, Raman, Bariana, Copley, Carter and Hoyos‐Villegas2025). Recognizing this spectrum improves alignment between trait discovery and deployment in dryland breeding programmes.
Barriers and bottlenecks to trait deployment
Despite the encouraging progress in identifying stress-resilient traits, the deployment of adaptive alleles from WWRs into modern breeding remains constrained by several biological and technical bottlenecks.
Many key targets – including drought, salinity, and heat tolerance – are polygenic, involving numerous small-effect loci that complicate introgression and stable phenotypic expression in elite backgrounds (Sharma et al. Reference Sharma, Schulthess, Bassi, Badaeva, Neumann, Graner, Özkan, Werner, Knüpffer and Kilian2021; Kashyap et al. Reference Kashyap, Garg, Tanwar, Sharma, Gupta, Ha, Bhattacharya, Mason and Rao2022; Tian et al. Reference Tian, Wang, Liu and Zhao2025). Complex genetic architectures require the simultaneous transfer and stabilization of multiple alleles across several breeding cycles, making selection less predictable than for single-gene traits. These challenges are further intensified by genotype × environment interactions, particularly in dryland agroecosystems, where trait expression fluctuates across locations and seasons, reducing breeding predictability and slowing selection for broad adaptation (Chairi et al. Reference Chairi, Aparicio, Serret and Araus2020; Nyine et al. Reference Nyine, Davidson, Adhikari, Clinesmith, Wang, Akhunova, Fritz and Akhunov2025). Addressing these biological complexities requires large multi-environment trials and GS strategies that capture polygenic signals across variable environments.
Genomic constraints (linkage drag, recombination suppression)
Introgression from WWR is often hindered by linkage drag, especially when beneficial alleles reside in low-recombination or pericentromeric regions. Recombination suppression in these regions makes separating beneficial alleles from adjacent deleterious chromatin extremely difficult, frequently necessitating extensive backcrossing and large breeding populations (Summers and Brown Reference Summers and Brown2013; Murray Reference Murray2024). Effective utilization of WWR alleles may therefore require advanced strategies, including targeted introgression, large mapping populations or the identification of recombination-friendly genomic intervals to facilitate allele transfer while minimizing unwanted genomic baggage.
Translational constraints: phenotyping and validation gaps
Beyond biological and genomic barriers, translational bottlenecks arise from limited phenotyping capacity and field validation. Many promising physiological traits – such as root architecture, osmotic adjustment or water-use efficiency – are often characterized under controlled conditions or in limited field trials. However, the expression of these traits can vary substantially across diverse dryland production systems, where multiple stresses frequently occur simultaneously. Limited multi-environment validation, therefore, restricts the translation of these discoveries into breeding pipelines, as the agronomic stability of candidate traits remains uncertain (Zeibig et al. Reference Zeibig, Kilian, Özkan, Pantha and Frei2024b; Nyine et al. Reference Nyine, Davidson, Adhikari, Clinesmith, Wang, Akhunova, Fritz and Akhunov2025).
Recent genomic advances provide a pathway to overcome these bottlenecks. High-quality pangenomes and genome assemblies reveal extensive PAV and structural rearrangements between wild and cultivated wheats, enabling targeted allele mining (Avni et al. Reference Avni, Nave, Barad, Baruch, Twardziok, Gundlach, Hale, Mascher, Spannagl, Wiebe, Jordan, Golan, Deek, Ben-Zvi, Ben-Zvi, Himmelbach, MacLachlan, Sharpe, Fritz, Ben-David, Budak, Fahima, Korol, Faris, Hernandez, Mikel, Levy, Steffenson, Maccaferri, Tuberosa, Cattivelli, Faccioli, Ceriotti, Kashkush, Pourkheirandish, Komatsuda, Eilam, Sela, Sharon, Ohad, Chamovitz, Mayer, Stein, Ronen, Peleg, Pozniak, Akhunov and Distelfeld2017; Walkowiak et al. Reference Walkowiak, Gao, Monat, Haberer, Kassa, Brinton, Ramirez-Gonzalez, Kolodziej, Delorean, Thambugala, Klymiuk, Byrns, Gundlach, Bandi, Siri, Nilsen, Aquino, Himmelbach, Copetti, Ban, Venturini, Bevan, Clavijo, Koo, Ens, Wiebe, N’Diaye, Fritz, Gutwin, Fiebig, Fosker, Fu, Accinelli, Gardner, Fradgley, Gutierrez-Gonzalez, Halstead-Nussloch, Hatakeyama, Koh, Deek, Costamagna, Fobert, Heavens, Kanamori, Kawaura, Kobayashi, Krasileva, Kuo, McKenzie, Murata, Nabeka, Paape, Padmarasu, Percival-Alwyn, Kagale, Scholz, Sese, Juliana, Singh, Shimizu-Inatsugi, Swarbreck, Cockram, Budak, Tameshige, Tanaka, Tsuji, Wright, Wu, Steuernagel, Small, Cloutier, Keeble-Gagnère, Muehlbauer, Tibbets, Nasuda, Melonek, Hucl, Sharpe, Clark, Legg, Bharti, Langridge, Hall, Uauy, Mascher, Krattinger, Handa, Shimizu, Distelfeld, Chalmers, Keller, Mayer, Poland, Stein, McCartney, Spannagl, Wicker and Pozniak2020; Kilian et al. Reference Kilian, Dempewolf, Guarino, Werner, Coyne and Warburton2021). Pre-breeding platforms – including synthetic hexaploids, introgression libraries and advanced backcross populations – help bridge wild and elite germplasm (Trethowan and Mujeeb-Kazi Reference Trethowan and Mujeeb-Kazi2008; Mastrangelo et al. Reference Mastrangelo, Roncallo, Matny, Čegan, Steffenson, Echenique, Šafář, Battaglia, Barabaschi, Cattivelli and Özkan2023; Li et al. Reference Li, Zhu, Fan, Li, Liu, Shaheen, Nie, Li, Liu, Li, Liu, Yang, Guo, Zhu, Bu, Li, Liang, Bai, Ma, Guo, Zhang, Huang, Zhou and Song2024a, Reference Li, Zeng, Zhao, Li, Chen and Wang2024b; Farooq et al. Reference Farooq, Frei, Zeibig, Pantha, Özkan, Kilian and Siddique2025). Precision approaches, such as gene editing, speed breeding and de novo domestication, can accelerate the deployment of alleles once causal variants are identified and functionally validated (Özkan et al. Reference Özkan, Brandolini, Schäfer-Pregl and Salamini2002; Kilian et al. Reference Kilian, Dempewolf, Guarino, Werner, Coyne and Warburton2021; Farooq et al. Reference Farooq, Frei, Zeibig, Pantha, Özkan, Kilian and Siddique2025; Mastrangelo et al. 2025). Collectively, integrating conservation, pre-breeding and genomics-informed validation pipelines is essential to translate WWR-derived diversity into climate-resilient wheat for dryland agriculture.
WWR as a resource for dryland agriculture
WWRs represent highly valuable genetic resources for improving wheat’s resilience in dryland agroecosystems. Originating in the Fertile Crescent and nearby areas, these plants have adapted to unpredictable rainfall, temperature extremes, soil salinity and nutrient deficiencies, resulting in adaptive traits that remain underexploited in breeding pipelines, particularly root system architecture (RSA), ion homeostasis, micronutrient-use efficiency and heat-responsive photosynthetic regulation (Fig. 1; Özkan et al. Reference Özkan, Willcox, Graner, Salamini and Kilian2011; Kilian et al. Reference Kilian, Mammen, Millet, Sharma, Graner, Salamini, Hammer and Özkan2011; Farooq et al. Reference Farooq, Frei, Zeibig, Pantha, Özkan, Kilian and Siddique2025). While domestication and intensive selection improved agronomic performance, they also narrowed wheat’s genetic base; modern cultivars generally lack adaptive alleles retained in wild progenitors and close relatives (Farooq et al. Reference Farooq, Frei, Zeibig, Pantha, Özkan, Kilian and Siddique2025). This section advances prior syntheses by distinguishing between demonstrated breeding gains and traits still under evaluation, identifying mechanisms underlying stress adaptation, and clarifying the biological and technical bottlenecks limiting deployment.
Demonstrated successes: synthetic wheat and introgressions
SHW, developed by crossing Ae. tauschii with durum wheat (T. durum or T. dicoccon), has become a major strategy for expanding D-genome diversity in bread wheat breeding programmes. Major initiatives led by the International Maize and Wheat Improvement Center (CIMMYT) and partner institutions have generated large SHW libraries that primarily serve as pre-breeding resources rather than direct cultivar replacements (Warburton et al. Reference Warburton, Crossa, Franco, Kazi, Trethowan, Rajaram, Pfeiffer, Zhang, Dreisigacker and Ginkel2006; Mujeeb-Kazi et al. Reference Mujeeb-Kazi, Gul, Farooq, Rizwan and Ahmad2008). These synthetics have introduced alleles conferring improved tolerance to abiotic stresses and foliar diseases, although performance remains uneven and most lines are still embedded in pre-breeding pipelines (Tian et al. Reference Tian, Wang, Liu and Zhao2025). The lines derived from synthetic wheat exhibit excellent RSA traits (e.g., root length, volume, and surface area), which directly support water capture in low-rainfall systems and improve performance under conservation agriculture, thereby benefiting dryland production systems (Gaikwad et al. Reference Gaikwad, Yadav, Kumar, Kumar, Babu, Singh, Ansari and Pandey2023). Introgressions from WWRs have further shaped modern wheat adaptation (Fig. 1). Both intentional breeder-mediated introgressions and historical gene flow events have contributed to the genomic and phenotypic diversity of elite cultivars. For example, an Ae. Tauschii-derived introgression increased yield by >50% under heat stress and reduced canopy temperature by 2 °C in field trials, without a yield penalty under favourable conditions (Molero et al. Reference Molero, Coombes, Joynson, Pinto, Piñera-Chávez, Rivera-Amado, Hall and Reynolds2023).
Comparative pangenome analyses reveal extensive PAV and structural polymorphisms in elite wheat germplasm, reflecting past introgression events from wild relatives (He et al. Reference He, Pasam, Shi, Kant, Keeble-Gagnere, Kay, Forrest, Fritz, Hucl, Wiebe, Knox, Cuthbert, Pozniak, Akhunova, Morrell, Davies, Webb, Spangenberg, Hayes, Daetwyler, Tibbits, Hayden and Akhunov2019; Walkowiak et al. Reference Walkowiak, Gao, Monat, Haberer, Kassa, Brinton, Ramirez-Gonzalez, Kolodziej, Delorean, Thambugala, Klymiuk, Byrns, Gundlach, Bandi, Siri, Nilsen, Aquino, Himmelbach, Copetti, Ban, Venturini, Bevan, Clavijo, Koo, Ens, Wiebe, N’Diaye, Fritz, Gutwin, Fiebig, Fosker, Fu, Accinelli, Gardner, Fradgley, Gutierrez-Gonzalez, Halstead-Nussloch, Hatakeyama, Koh, Deek, Costamagna, Fobert, Heavens, Kanamori, Kawaura, Kobayashi, Krasileva, Kuo, McKenzie, Murata, Nabeka, Paape, Padmarasu, Percival-Alwyn, Kagale, Scholz, Sese, Juliana, Singh, Shimizu-Inatsugi, Swarbreck, Cockram, Budak, Tameshige, Tanaka, Tsuji, Wright, Wu, Steuernagel, Small, Cloutier, Keeble-Gagnère, Muehlbauer, Tibbets, Nasuda, Melonek, Hucl, Sharpe, Clark, Legg, Bharti, Langridge, Hall, Uauy, Mascher, Krattinger, Handa, Shimizu, Distelfeld, Chalmers, Keller, Mayer, Poland, Stein, McCartney, Spannagl, Wicker and Pozniak2020). However, the functional significance of many PAVs remains uncertain, requiring further validation to confirm their phenotypic contributions. For example, the cultivar Pamukale carries an ∼14 Mb segment on chromosome 7D introgressed from a wild donor, conferring powdery mildew resistance (PmPam), yet suppressed recombination restricts fine-mapping and limits segment optimization for breeding purposes (Heuberger et al. Reference Heuberger, Bernasconi, Said, Jung, Herren, Widrig, Šimková, Keller, Sánchez-Martín and Wicker2024). This study exemplifies both the value and the challenges of WWR introgression.
Mechanisms underpinning adaptation to dryland stresses
Ion homeostasis and salinity
Abiotic stress resilience derived from WWR remains underutilized compared with disease resistance, though recent studies underscore its potential. The Ae. tauschii-derived HKT1;5 allele, a canonical example, reduces sodium accumulation and enhances salinity tolerance (Munns et al. Reference Munns, James, Xu, Athman, Conn, Jordans, Byrt, Hare, Tyerman, Tester, Plett and Gilliham2012). Synthetic vs. bread wheat comparisons under salt stress confirm that SHW lines retain higher chlorophyll levels, photosynthetic activity, and antioxidant enzyme activity, along with favourable Na+/K+ ratios (Alghabari and Shah Reference Alghabari and Shah2024). T. dicoccoides displays variation in salt tolerance index correlated with Na+ exclusion and root architectural traits (Feng et al. Reference Feng, Cui, Lv, Bian, Wang, Song and Nie2018). These mechanistic pathways offer targets for controlled introgression but require careful selection to avoid co-transfer of undesirable linked loci.
Drought adaptation and root system architecture
Root traits from WWRs, including deeper rooting angles, enhanced branching, and improved root hydraulic conductivity, facilitate access to subsoil moisture and enhance transpiration efficiency (Peleg et al. Reference Peleg, Fahima, Krugman, ABBO, Yakir, KOROL and SARANGA2009). Thinopyrum elongatum 7DL translocations enhance root biomass and drought tolerance, though some backgrounds carry growth penalties under optimal conditions (Placido et al. Reference Placido, Campbell, Folsom, Cui, Kruger, Baenziger and Walia2013). These outcomes underscore a persistent trade-off between stress resilience and productivity, challenging the assumption that deeper or larger root systems are universally beneficial. In Aegilops accessions, drought tolerance is often attributed to coordinated osmotic adjustment, antioxidant defence, and stomatal regulation (Sultan et al. Reference Sultan, Hui Liu, Yang and XIAN2012; Jadidi et al. Reference Jadidi, Pour-Aboughadareh, Bocianowski, Jamshidi and Akbari2025). Nevertheless, these responses are typically controlled by complex, multi-layered regulatory networks, limiting their predictable transfer into elite germplasm. Consequently, trait stacking remains a major bottleneck in drought breeding, as simultaneous optimization of root architecture, physiological buffering, and yield stability has proven difficult to achieve without unintended trade-offs.
Heat tolerance and photosynthetic stability
Heat resilience in wild relatives is associated with canopy temperature regulation, maintenance of chlorophyll content, and stable light-saturated photosynthesis (Amax). Emmer wheat (T. dicoccon)-derived hexaploid progenies with 1–43% emmer genomic contribution outperform recurrent parents under reproductive-stage heat stress, though responses vary across environments (Ullah et al. Reference Ullah, Bramley, Mahmood and Trethowan2021). These findings emphasize the role of polygenic buffering mechanisms, including Rubisco efficiency and dynamic stomatal responses (McAusland et al. Reference McAusland, Vialet‐Chabrand, Jauregui, Burridge, Hubbart‐Edwards, Fryer, King, King, Pyke, Edwards, Carmo‐Silva, Lawson and Murchie2020). The WWRs exhibit a set of adaptive mechanisms, including antioxidant defences, osmotic adjustment, stomatal regulation and improved RSA, which translate into improved survival and yield stability under suboptimal growth conditions (Sultan et al. Reference Sultan, Hui Liu, Yang and XIAN2012; Akman et al. Reference Akman, Akgun and Tamkoc2017; Jadidi et al. Reference Jadidi, Pour-Aboughadareh, Bocianowski, Jamshidi and Akbari2025). Nevertheless, their use in elite backgrounds is constrained by polygenic complexity, frequent yield trade-offs, and genotype × environment interactions. The plant’s adaptive traits under stressful environments are often negatively associated with productivity, creating a breeding dilemma that has limited widespread adoption.
Biotic stress resistance
Biotic stress resistance demonstrates clearer breeding tractability because major-effect genes govern many resistance traits. Well-documented examples include Sr2 and Yr15 from T. dicoccoides, SrTA10171 and SrTA10187 from Ae. tauschii, and diverse rust resistance loci from Aegilops taxa (Singh et al. Reference Singh, Hodson, Huerta-Espino, Jin, Bhavani, Njau, Herrera-Foessel, Singh, Singh and Govindan2011; Olson et al. Reference Olson, Rouse, Pumphrey, Bowden, Gill and Poland2013). These successes contrast with the slower progress in abiotic stress improvement and underscore why disease resistance has historically dominated WWR contributions. However, durability and pleiotropic effects, such as the yield-quality trade-offs associated with the 1RS rye (Secale cereale) translocation, highlight the need for judicious deployment and trait stacking strategies (Kaur et al. Reference Kaur, Jindal, Kaur, Chhuneja, Gosal and Wani2018).
Collectively, WWR-derived alleles have been most successfully deployed for single-gene biotic resistance and a limited set of abiotic mechanisms such as ion homeostasis and root traits. In contrast, complex drought, heat and nutrient-related traits remain difficult to translate into stable yield gains. These contrasting outcomes highlight the need for mechanism-based introgression, haplotype-level selection and robust multi-environment validation to overcome polygenic control, linkage drag and genotype × environment interactions in dryland wheat breeding.
Challenges, trade-offs and barriers
Despite the success reported in earlier sections, the utilization of WWR faces both biological and breeding-system constraints. From a biological perspective, many adaptive traits relevant to dryland agriculture (e.g., drought, salinity, heat and nutrient-use efficiency) are polygenic and strongly influenced by genotype × environment interactions, which reduces predictability of trait expression across heterogeneous dryland gradients (Chairi et al. Reference Chairi, Aparicio, Serret and Araus2020; Nyine et al. Reference Nyine, Davidson, Adhikari, Clinesmith, Wang, Akhunova, Fritz and Akhunov2025). Yield penalties associated with WWR-derived alleles are therefore genotype- and environment-dependent rather than universal, often becoming evident under favourable rather than stress-prone conditions (Murray Reference Murray2024; Tian et al. Reference Tian, Wang, Liu and Zhao2025). Cytogenetic constraints further limit deployment, including linkage drag, suppressed recombination in pericentromeric regions and large alien introgressions. Such constraints are exacerbated in tertiary gene pool species, including Thinopyrum and Elymus, due to structural rearrangements, homoeologous interference, and low crossover frequency (Wijerathna-Yapa et al. Reference Wijerathna-Yapa, Ramtekey, Ranawaka and Basnet2022).
Beyond intrinsic biological barriers, the effective deployment of WWRs is constrained by several breeding-system limitations. Modern wheat cultivars possess a narrow genetic base due to prolonged selection for yield and uniformity, reducing adaptive diversity and increasing vulnerability to emerging stresses (Przewieslik-Allen et al. Reference Przewieslik-Allen, Wilkinson, Burridge, Winfield, Dai, Beaumont, King, Yang, Griffiths, Wingen, Horsnell, Bentley, Shewry, Barker and Edwards2021; Wani et al. Reference Wani, Khan, Riaz, Joshi, Hussain, Rana, Kumar, Athiyannan, Singh, Ali, Kang, Rajaram, Mujeeb-Kazi, Bishnoi and Aziz2022). Although WWRs harbour extensive genetic variation, their integration into elite germplasm is impeded by reproductive isolation barriers, including low crossability, hybrid inviability and sterility, often requiring prolonged backcrossing to mitigate linkage drag and restore agronomic performance (Mondal et al. Reference Mondal, Rutkoski, Velu, Singh, Crespo-Herrera, Guzmán, Bhavani, Lan, He and Singh2016). These challenges are compounded by the difficulty of detecting and tracking introgressed segments within the large (≈17 Gb), highly repetitive hexaploid wheat genome, where even modern single-nucleotide polymorphism (SNP) based genotyping platforms, including Kompetitive Allele Specific PCR (KASP) assays, may struggle to resolve small but beneficial introgressions (Ma et al. Reference Ma, Zhang, Zhang, Zhou, Han, Liu, Yang, Li and Li2019; Keilwagen et al. Reference Keilwagen, Lehnert, Berner, Badaeva, Himmelbach, Börner and Kilian2022). In addition, slow breeding cycles, limited high-throughput phenotyping for complex stress-adaptive traits, and uncertainty in identifying donor species for unknown introgressions further delay the translation of WWR-derived variation into elite cultivars (Mondal et al. Reference Mondal, Rutkoski, Velu, Singh, Crespo-Herrera, Guzmán, Bhavani, Lan, He and Singh2016; Keilwagen et al. Reference Keilwagen, Lehnert, Berner, Badaeva, Himmelbach, Börner and Kilian2022). Collectively, these constraints highlight the need for integrated genomics, phenotyping, and pre-breeding pipelines to more efficiently harness WWR diversity (Wani et al. Reference Wani, Khan, Riaz, Joshi, Hussain, Rana, Kumar, Athiyannan, Singh, Ali, Kang, Rajaram, Mujeeb-Kazi, Bishnoi and Aziz2022).
Genomics and molecular tools for harnessing WWR diversity
Recent genomic advances are reshaping the utilization of WWR, enabling more strategic and predictive breeding approaches to address genetic erosion, climate change, and food security challenges (Özkan et al. Reference Özkan, Brandolini, Schäfer-Pregl and Salamini2002; Kilian et al. Reference Kilian, Dempewolf, Guarino, Werner, Coyne and Warburton2021; Sharma et al. Reference Sharma, Schulthess, Bassi, Badaeva, Neumann, Graner, Özkan, Werner, Knüpffer and Kilian2021; Farooq et al. Reference Farooq, Frei, Zeibig, Pantha, Özkan, Kilian and Siddique2025; Mastrangelo et al. 2025). These tools can be organized hierarchically according to their roles in tracking introgressions, trait discovery, deployment acceleration and precision modification.
Tracking introgressions and diversity
Marker-assisted selection (MAS) remains a cornerstone for tracing desirable alleles from WWR into elite wheat backgrounds, particularly in marker-assisted backcrossing schemes aimed at minimizing linkage drag (Sanjay and Prakash Reference Sanjay and Prakash2024). High-throughput SNP platforms and chromosome-specific KASP markers have enabled precise detection of Aegilops and T. urartu introgressions, facilitating rapid screening of large breeding populations (Przewieslik-Allen et al. Reference Przewieslik-Allen, Burridge, Wilkinson, Winfield, Shaw, McAusland, King, King, Edwards and Barker2019; Grewal et al. Reference Grewal, Guwela, Newell, Yang, Ashling, Scholefield, Hubbart-Edwards, Burridge, Stride, King and King2021). Beyond marker systems, sequence-based detection approaches such as coverage analysis allow genome-wide identification of introgressions and inference of donor species, even in complex or unknown material (Keilwagen et al. Reference Keilwagen, Lehnert, Berner, Badaeva, Himmelbach, Börner and Kilian2022). Application of these methods has revealed that introgressions from wild relatives can increase bread wheat genetic diversity by 4–32%, underscoring their substantial but often cryptic contribution (Zhou et al. Reference Zhou, Zhao, Li, Xu, Bi, Kang, Xu, Chen, Wang, Wang, Liu, Jiao, Lu, Wang, Yin, Jiao and Lu2020).
Discovery: Pangenomes and GWAS
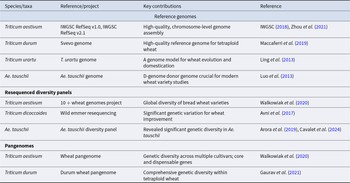
Pangenomic frameworks and GWAS have substantially expanded the discovery of adaptive variation in WWR (Walkowiak et al. Reference Walkowiak, Gao, Monat, Haberer, Kassa, Brinton, Ramirez-Gonzalez, Kolodziej, Delorean, Thambugala, Klymiuk, Byrns, Gundlach, Bandi, Siri, Nilsen, Aquino, Himmelbach, Copetti, Ban, Venturini, Bevan, Clavijo, Koo, Ens, Wiebe, N’Diaye, Fritz, Gutwin, Fiebig, Fosker, Fu, Accinelli, Gardner, Fradgley, Gutierrez-Gonzalez, Halstead-Nussloch, Hatakeyama, Koh, Deek, Costamagna, Fobert, Heavens, Kanamori, Kawaura, Kobayashi, Krasileva, Kuo, McKenzie, Murata, Nabeka, Paape, Padmarasu, Percival-Alwyn, Kagale, Scholz, Sese, Juliana, Singh, Shimizu-Inatsugi, Swarbreck, Cockram, Budak, Tameshige, Tanaka, Tsuji, Wright, Wu, Steuernagel, Small, Cloutier, Keeble-Gagnère, Muehlbauer, Tibbets, Nasuda, Melonek, Hucl, Sharpe, Clark, Legg, Bharti, Langridge, Hall, Uauy, Mascher, Krattinger, Handa, Shimizu, Distelfeld, Chalmers, Keller, Mayer, Poland, Stein, McCartney, Spannagl, Wicker and Pozniak2020; Leonova et al. Reference Leonova, Kiseleva, Berezhnaya, Orlovskaya and Salina2023). Sequencing of key wild progenitors, such as Ae. tauschii (D-genome donor), T. dicoccoides and Secale cereale, has revealed PAV, copy number variation and introgressed haplotypes linked to stress adaptation (Table 2; Avni et al. Reference Avni, Nave, Barad, Baruch, Twardziok, Gundlach, Hale, Mascher, Spannagl, Wiebe, Jordan, Golan, Deek, Ben-Zvi, Ben-Zvi, Himmelbach, MacLachlan, Sharpe, Fritz, Ben-David, Budak, Fahima, Korol, Faris, Hernandez, Mikel, Levy, Steffenson, Maccaferri, Tuberosa, Cattivelli, Faccioli, Ceriotti, Kashkush, Pourkheirandish, Komatsuda, Eilam, Sela, Sharon, Ohad, Chamovitz, Mayer, Stein, Ronen, Peleg, Pozniak, Akhunov and Distelfeld2017; Keilwagen et al. Reference Keilwagen, Lehnert, Berner, Badaeva, Himmelbach, Börner and Kilian2022). Pangenome analyses show that many adaptive loci lost during domestication remain conserved in WWR populations, providing a reservoir of alleles for crop improvement (Table 1; Walkowiak et al. Reference Walkowiak, Gao, Monat, Haberer, Kassa, Brinton, Ramirez-Gonzalez, Kolodziej, Delorean, Thambugala, Klymiuk, Byrns, Gundlach, Bandi, Siri, Nilsen, Aquino, Himmelbach, Copetti, Ban, Venturini, Bevan, Clavijo, Koo, Ens, Wiebe, N’Diaye, Fritz, Gutwin, Fiebig, Fosker, Fu, Accinelli, Gardner, Fradgley, Gutierrez-Gonzalez, Halstead-Nussloch, Hatakeyama, Koh, Deek, Costamagna, Fobert, Heavens, Kanamori, Kawaura, Kobayashi, Krasileva, Kuo, McKenzie, Murata, Nabeka, Paape, Padmarasu, Percival-Alwyn, Kagale, Scholz, Sese, Juliana, Singh, Shimizu-Inatsugi, Swarbreck, Cockram, Budak, Tameshige, Tanaka, Tsuji, Wright, Wu, Steuernagel, Small, Cloutier, Keeble-Gagnère, Muehlbauer, Tibbets, Nasuda, Melonek, Hucl, Sharpe, Clark, Legg, Bharti, Langridge, Hall, Uauy, Mascher, Krattinger, Handa, Shimizu, Distelfeld, Chalmers, Keller, Mayer, Poland, Stein, McCartney, Spannagl, Wicker and Pozniak2020; Fernandez et al. Reference Fernandez, Nestor, Danilevicz, Gill, Petereit, Bayer, Finnegan, Batley and Edwards2022; Awan et al. Reference Awan, Pervaiz, Rasheed, Amin, Saeed, Dhugga and Mansoor2022). For example, alien introgressions that still segregate in elite cultivars confer resistance to rusts and mildew while also affecting RSA and osmotic regulation.
Reference genomes, resequenced diversity panels, and pangenomes of domesticated and wild wheat types

Despite these advances, several methodological limitations persist. The large and polyploid wheat genome reduces GWAS mapping resolution due to extensive linkage disequilibrium and homoeologous gene copies, complicating the identification of causal variants for complex traits (Maccaferri et al. Reference Maccaferri, Harris, Twardziok, Pasam, Gundlach, Spannagl, Ormanbekova, Lux, Prade, Milner, Himmelbach, Mascher, Bagnaresi, Faccioli, Cozzi, Lauria, Lazzari, Stella, Manconi, Gnocchi, Moscatelli, Avni, Deek, Biyiklioglu, Frascaroli, Corneti, Salvi, Sonnante, Desiderio, Marè, Crosatti, Mica, Özkan, Kilian, De Vita, Marone, Joukhadar, Mazzucotelli, Nigro, Gadaleta, Chao, Faris, Melo, Pumphrey, Pecchioni, Milanesi, Wiebe, Ens, MacLachlan, Clarke, Sharpe, Koh, Liang, Taylor, Knox, Budak, Mastrangelo, Xu, Stein, Hale, Distelfeld, Hayden, Tuberosa, Walkowiak, Mayer, Ceriotti, Pozniak and Cattivelli2019). Rare alleles, often enriched in wild relatives, may be missed due to limited statistical power (Alqudah et al. Reference Alqudah, Sallam, Baenziger and Börner2019). Environmental variability further challenges stress-related GWAS, as genotype × environment interactions strongly influence drought, salinity, and heat tolerance traits, potentially obscuring genetic signals and reducing reproducibility unless multi-environment phenotyping is conducted (Reynolds et al. Reference Reynolds, Lewis, Ammar, Basnet, Crespo-Herrera, Crossa, Dhugga, Dreisigacker, Juliana, Karwat, Kishii, Krause, Langridge, Lashkari, Mondal, Payne, Pequeno, Pinto, Sansaloni, Schulthess, Singh, Sonder, Sukumaran, Xiong and Braun2021; Tene et al. Reference Tene, Adhikari, Cobo, Jordan, Matny, Del Blanco, Roter, Ezrati, Govta, Manisterski, Yehuda, Chen, Steffenson, Akhunov and Sela2022). Additionally, many WWR species, such as Ae. searsii and Elymus spp., remain poorly represented in genomic databases, leaving critical gaps in the catalogue of stress-adaptive alleles (Table 1).
Complementing pangenomic resources, GWAS in T. dicoccoides and related taxa has identified loci underlying resistance to biotic stresses and tolerance to drought and salinity, highlighting the value of combining population-level diversity with high-resolution phenotyping (Tene et al. Reference Tene, Adhikari, Cobo, Jordan, Matny, Del Blanco, Roter, Ezrati, Govta, Manisterski, Yehuda, Chen, Steffenson, Akhunov and Sela2022). Collectively, pangenomics and GWAS provide complementary tools for dissecting the complex genetic architecture of adaptive traits, while also revealing priorities for future resource development (Fernandez et al. Reference Fernandez, Nestor, Danilevicz, Gill, Petereit, Bayer, Finnegan, Batley and Edwards2022; Tene et al. Reference Tene, Adhikari, Cobo, Jordan, Matny, Del Blanco, Roter, Ezrati, Govta, Manisterski, Yehuda, Chen, Steffenson, Akhunov and Sela2022).
Deployment accelerators: Speed breeding and introgression platforms
The deployment of WWR-derived variation can be accelerated through speed breeding, which manipulates photoperiod and temperature to shorten generation time, enabling up to four to five wheat generations per year under controlled conditions (Li et al. Reference Li, Zhu, Fan, Li, Liu, Shaheen, Nie, Li, Liu, Li, Liu, Yang, Guo, Zhu, Bu, Li, Liang, Bai, Ma, Guo, Zhang, Huang, Zhou and Song2024a). Importantly, speed breeding primarily accelerates generation turnover and does not increase recombination frequency or resolve recombination suppression in low-recombination genomic regions. When combined with marker-assisted introgression and GS, speed breeding facilitates rapid fixation and evaluation of wild-derived alleles for stress adaptation and yield stability (Gudi et al. Reference Gudi, Kumar, Singh, Tanin and Sharma2022). However, accelerated generation cycling alone does not guarantee genetic gain unless coupled with accurate phenotyping and predictive selection. Complex traits such as drought, salinity, and heat tolerance still require multi-environment testing to capture genotype × environment interactions, which cannot be fully replicated under controlled conditions (Watson et al. Reference Watson, Ghosh, Williams, Cuddy, Simmonds, Rey, Asyraf Md Hatta, Hinchliffe, Steed, Reynolds, Adamski, Breakspear, Korolev, Rayner, Dixon, Riaz, Martin, Ryan, Edwards, Batley, Raman, Carter, Rogers, Domoney, Moore, Harwood, Nicholson, Dieters, DeLacy, Zhou, Uauy, Boden, Park, Wulff and Hickey2018; Potts et al. Reference Potts, Jangra, Michael and Wu2023).
Complementary introgression platforms, such as structured populations derived from Ae. tauschii, enable systematic transfer of D-genome diversity into bread wheat, supporting targeted exploration of adaptive alleles for stress resilience and agronomic performance (Li et al. Reference Li, Zhu, Fan, Li, Liu, Shaheen, Nie, Li, Liu, Li, Liu, Yang, Guo, Zhu, Bu, Li, Liang, Bai, Ma, Guo, Zhang, Huang, Zhou and Song2024a). Integrating these platforms with pangenomic resources, MAS, and GS provides a translational bridge from allele discovery in wild relatives to practical breeding deployment. Together, pipelines combining genomic discovery, structured introgression, speed breeding and predictive selection offer a feasible path to accelerate the utilization of WWR diversity in climate-resilient wheat improvement.
Precision modification: Opportunities and limitations
CRISPR/Cas genome editing offers precise allele modification, including gene knockouts, base editing, and targeted deletions relevant to domestication and stress-response traits (Kelliher et al. Reference Kelliher, Starr, Su, Tang, Chen, Carter, Wittich, Dong, Green, Burch and McCuiston2019; Borah et al. Reference Borah, Singh, Chattopadhyay, Kaur and Bari2024). In wheat, the hexaploid genome complicates editing because most genes exist in three homoeologous copies across the A, B, and D subgenomes (Peng et al. Reference Peng, Sun, Peng and Nevo2013). Achieving a complete phenotypic effect often requires simultaneous editing of all homoeologs, increasing the technical complexity of multiplex genome editing. Practical limitations also include low transformation efficiency, regulatory hurdles, and challenges in achieving consistent edits across homoeologous in the large, repetitive genome (Zhou et al. Reference Zhou, Bai, Li, Sun, Zhang, Ma, Zhao, Nie, Li, Chen, Lv, Zhu, Fan, Ge, Shaheen, Guo, Zhang, Ma, Liang, Qiu, Hu, Sun, Hou, Xu, Xue, Jiang, Huang, Li, Zou and Song2023; Awan et al. Reference Awan, Pervaiz, Rasheed, Amin, Saeed, Dhugga and Mansoor2022). Multiplex editing can partially overcome these challenges, but often requires extensive screening to identify fully edited lines.
Limitations also arise from sgRNA design tools trained on non-plant datasets, variable delivery systems, and cell-type-specific DNA repair pathways (Tan et al. Reference Tan, Wang and Liu2024), all of which reduce predictability and increase off-target risk. Consequently, genome editing is currently better suited for validating and fine-tuning already identified loci rather than for discovering new adaptive variation (Kelliher et al. 2019; Borah et al. Reference Borah, Singh, Chattopadhyay, Kaur and Bari2024). In wheat improvement pipelines, CRISPR is most effective for confirming candidate gene function, modifying regulatory elements, or optimizing favourable alleles identified through genomics or pre-breeding (Awan et al. Reference Awan, Amin, Rasheed, Saeed and Mansoor2024; Borah et al. Reference Borah, Singh, Chattopadhyay, Kaur and Bari2024; Tan et al. Reference Tan, Wang and Liu2024). Advances in delivery technologies, tissue culture optimization, and DNA-free editing (e.g., ribonucleoprotein-mediated systems) offer promising avenues to improve efficiency while mitigating regulatory constraints.
Pre-breeding pipelines: Bridging wild and elite germplasm
Pre-breeding provides the critical interface between WWR and elite breeding programmes by converting exotic diversity into usable genetic materials. Global initiatives at CIMMYT, International Center for Agricultural Research in the Dry Areas (ICARDA), and partner institutions have established structured pipelines that emphasize population development, controlled introgression, and early-stage evaluation of wild-derived alleles. A cornerstone of these efforts is SHW, which enables efficient transfer of D-genome diversity while partially overcoming crossability barriers (Warburton et al. Reference Warburton, Crossa, Franco, Kazi, Trethowan, Rajaram, Pfeiffer, Zhang, Dreisigacker and Ginkel2006; Mujeeb-Kazi et al. Reference Mujeeb-Kazi, Gul, Farooq, Rizwan and Ahmad2008; Aberkane et al. Reference Aberkane, Payne, Kishi, Smale, Amri and Jamora2020).
In addition to SHW, introgression libraries, advanced backcross populations and nested association mapping panels are increasingly used to expose wild alleles in agronomically relevant backgrounds. These resources allow breeders to evaluate stress-adaptive variation under managed and target environments, accelerating the identification of promising donor segments. Nevertheless, translating wild-derived variation into cultivars remains a long-term process. From the initial development of SHW or wide-cross derivatives to the release of improved cultivars, breeding pipelines typically require 10–15 years, reflecting multiple generations of selection, backcrossing, and multi-environment field validation. Furthermore, only a small proportion of early-generation introgression lines ultimately progress to advanced breeding programmes due to linkage drag, poor agronomic performance, or instability of stress-adaptive traits (Mondal et al. Reference Mondal, Rutkoski, Velu, Singh, Crespo-Herrera, Guzmán, Bhavani, Lan, He and Singh2016; Gudi et al. Reference Gudi, Kumar, Singh, Tanin and Sharma2022).
Accurate phenotyping represents an additional bottleneck in effectively utilizing wild germplasm. While genomic tools facilitate the detection of introgressed regions, validating adaptive traits requires robust phenotyping under realistic stress conditions. Recent advances in high-throughput phenotyping – including UAV-based remote sensing, canopy temperature and spectral reflectance measurements, root phenotyping platforms, and managed drought or salinity screening systems – are increasingly enabling large-scale evaluation of physiological traits relevant to dryland adaptation, such as water-use efficiency, canopy temperature depression, and root architecture (Gill et al. Reference Gill, Gill, Saini, Chopra, de Koff and Sandhu2022). Emerging molecular and genomic tools are beginning to strengthen pre-breeding efficiency. Functionally informed markers and validated regulatory modules, such as salinity-related Na+ exclusion pathways, enable more targeted introgression and reduce reliance on large, undirected wild segments (Wang et al. Reference Wang, Cheng, Wu, Chen, Liu, Wang, Ma, Guo, Li, Di, Zhang, Han, Kronzucker, Xia and Shi2024). Genome editing approaches offer additional opportunities to refine pre-breeding outputs, particularly for removing undesirable genomic regions during wide-cross derivatives, although their routine use in wheat remains technically and institutionally constrained (Schulthess et al. Reference Schulthess, Kale, Liu, Zhao, Philipp, Rembe, Jiang, Beukert, Serfling, Himmelbach, Fuchs, Oppermann, Weise, Boeven, Schacht, Longin, Kollers, Pfeiffer, Korzun, Lange, Scholz, Stein, Mascher and Reif2022; Borah et al. Reference Borah, Singh, Chattopadhyay, Kaur and Bari2024). Collectively, integrating genomic prediction, targeted introgression and high-throughput phenotyping platforms is expected to accelerate the conversion of wild wheat diversity into deployable breeding materials.
Functional impacts and gaps in utilization
Functional validation of WWR-derived alleles demonstrates substantial potential for improving wheat resilience under abiotic and biotic stress. Well-characterized examples include the HKT1;5-D allele from Ae. tauschii, validated in elite wheat backgrounds and multi-environment field trials for enhanced salinity tolerance (Munns et al. Reference Munns, James, Xu, Athman, Conn, Jordans, Byrt, Hare, Tyerman, Tester, Plett and Gilliham2012). Similarly, several root-related QTLs from T. dicoccoides have been identified in mapping populations and pre-breeding materials, associated with improved root architecture and water-use efficiency under drought. In addition, introgression of a 7DL chromosome segment from Agropyron elongatum (Thinopyrum elongatum) enhances drought adaptation by increasing root biomass and improving water uptake under stress (Placido et al. Reference Placido, Campbell, Folsom, Cui, Kruger, Baenziger and Walia2013). Chromosomal translocations from Secale cereale, particularly the widely deployed 1RS arm, have been extensively incorporated into commercial cultivars worldwide, providing durable disease resistance and improved stress robustness (Liu et al. Reference Liu, Li, Bernardo, Amand, Zhang, Sehgal and Bai2023).
Despite these successes, the practical use of WWR in breeding programmes remains concentrated in a small subset of species and traits. Historically, most successful introgressions have targeted major biotic stress resistance genes, such as rust and pest resistance from species including Ae. tauschii, S. cereale and Thinopyrum spp., whereas complex adaptive traits like drought tolerance, heat resilience and RSA remain comparatively underutilized in modern cultivars (Dempewolf et al. Reference Dempewolf, Baute, Anderson, Kilian, Smith and Guarino2017; Reynolds et al. Reference Reynolds, Lewis, Ammar, Basnet, Crespo-Herrera, Crossa, Dhugga, Dreisigacker, Juliana, Karwat, Kishii, Krause, Langridge, Lashkari, Mondal, Payne, Pequeno, Pinto, Sansaloni, Schulthess, Singh, Sonder, Sukumaran, Xiong and Braun2021).
Several taxa with documented adaptive traits remain underrepresented in breeding pipelines. For example, Ae. searsii harbours powdery mildew resistance and quality-related alleles but remains largely confined to pre-breeding research due to limited genomic resources and marker development (Gong et al. Reference Gong, Gong, Han, Li, Sehgal, Li, Liu, Song, Song, Liu and Liu2016). Likewise, species within the genus Elymus exhibit exceptional drought tolerance and perennial growth habits, yet their distant phylogenetic relationship to wheat imposes severe crossing and ploidy barriers (Khan et al. Reference Khan, Ali, Ullah, Ali, Kaushik, Alyemeni, Rasheed and Sher2022). These challenges illustrate that biological barriers and limited genomic resources still constrain the broader deployment of many wild relatives.
A critical bottleneck in translating wild diversity into breeding gains is not the discovery of genetic variation, but the phenotypic validation of adaptive traits. While genotyping and genome sequencing of wild germplasm collections have expanded rapidly over the past decade, large-scale phenotypic characterization under realistic field conditions has progressed more slowly. Traits such as drought tolerance, root architecture, and water-use efficiency require multi-environment testing and precise physiological measurements, making them considerably more difficult to evaluate than simple Mendelian resistance traits (Sansaloni et al. Reference Sansaloni, Franco, Santos, Percival-Alwyn, Singh, Petroli, Campos, Dreher, Payne, Marshall, Kilian, Milne, Raubach, Shaw, Stephen, Carling, Pierre, Burgueño, Crosa, Li, Guzman, Kehel, Amri, Kilian, Wenzl, Uauy, Banziger, Caccamo and Pixley2020; Reynolds et al. Reference Reynolds, Lewis, Ammar, Basnet, Crespo-Herrera, Crossa, Dhugga, Dreisigacker, Juliana, Karwat, Kishii, Krause, Langridge, Lashkari, Mondal, Payne, Pequeno, Pinto, Sansaloni, Schulthess, Singh, Sonder, Sukumaran, Xiong and Braun2021; Farooq et al. Reference Farooq, Frei, Zeibig, Pantha, Özkan, Kilian and Siddique2025). Consequently, limited phenotyping capacity across diverse environments remains a principal barrier to fully exploiting wild wheat diversity in modern breeding programmes. Addressing this imbalance will require coordinated efforts to expand genomic resources for underutilized species, integrate high-throughput phenotyping platforms with genomic analyses, and prioritize traits with clear agronomic relevance for dryland and stress-prone environments.
Integrated pathways for climate-resilient wheat
Technological pipelines for harnessing WWR diversity
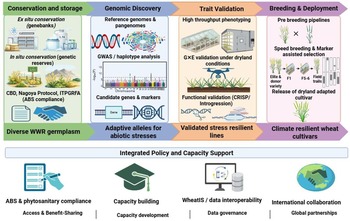
Maximizing WWR’s contribution to climate-resilient wheat requires clearly defined technological pipelines that link allele discovery, validation and deployment rather than treating these processes independently (Fig. 2). Genomic resources, including reference genomes, pangenomes and GWAS, enable the discovery of adaptive loci associated with drought tolerance, ion homeostasis, RSA and heat-responsive photosynthetic regulation (Avni et al. Reference Avni, Nave, Barad, Baruch, Twardziok, Gundlach, Hale, Mascher, Spannagl, Wiebe, Jordan, Golan, Deek, Ben-Zvi, Ben-Zvi, Himmelbach, MacLachlan, Sharpe, Fritz, Ben-David, Budak, Fahima, Korol, Faris, Hernandez, Mikel, Levy, Steffenson, Maccaferri, Tuberosa, Cattivelli, Faccioli, Ceriotti, Kashkush, Pourkheirandish, Komatsuda, Eilam, Sela, Sharon, Ohad, Chamovitz, Mayer, Stein, Ronen, Peleg, Pozniak, Akhunov and Distelfeld2017; IWGSC 2018; Walkowiak et al. Reference Walkowiak, Gao, Monat, Haberer, Kassa, Brinton, Ramirez-Gonzalez, Kolodziej, Delorean, Thambugala, Klymiuk, Byrns, Gundlach, Bandi, Siri, Nilsen, Aquino, Himmelbach, Copetti, Ban, Venturini, Bevan, Clavijo, Koo, Ens, Wiebe, N’Diaye, Fritz, Gutwin, Fiebig, Fosker, Fu, Accinelli, Gardner, Fradgley, Gutierrez-Gonzalez, Halstead-Nussloch, Hatakeyama, Koh, Deek, Costamagna, Fobert, Heavens, Kanamori, Kawaura, Kobayashi, Krasileva, Kuo, McKenzie, Murata, Nabeka, Paape, Padmarasu, Percival-Alwyn, Kagale, Scholz, Sese, Juliana, Singh, Shimizu-Inatsugi, Swarbreck, Cockram, Budak, Tameshige, Tanaka, Tsuji, Wright, Wu, Steuernagel, Small, Cloutier, Keeble-Gagnère, Muehlbauer, Tibbets, Nasuda, Melonek, Hucl, Sharpe, Clark, Legg, Bharti, Langridge, Hall, Uauy, Mascher, Krattinger, Handa, Shimizu, Distelfeld, Chalmers, Keller, Mayer, Poland, Stein, McCartney, Spannagl, Wicker and Pozniak2020). The MAS and GS translate these discoveries into breeding populations, while speed breeding accelerates the fixation of favourable alleles and shortens breeding cycles (Mwadzingeni et al. Reference Mwadzingeni, Figlan, Shimelis, Mondal and Tsilo2017; Watson et al. Reference Watson, Ghosh, Williams, Cuddy, Simmonds, Rey, Asyraf Md Hatta, Hinchliffe, Steed, Reynolds, Adamski, Breakspear, Korolev, Rayner, Dixon, Riaz, Martin, Ryan, Edwards, Batley, Raman, Carter, Rogers, Domoney, Moore, Harwood, Nicholson, Dieters, DeLacy, Zhou, Uauy, Boden, Park, Wulff and Hickey2018). GS is particularly valuable for WWR-derived traits because many adaptive responses to drought, heat and salinity are polygenic and strongly influenced by genotype × environment interactions. By leveraging genome-wide marker information to predict breeding values, GS enables the selection of complex stress-adaptive traits before extensive field evaluation, thereby improving selection efficiency during pre-breeding and introgression programmes (Budhlakoti et al. Reference Budhlakoti, Kushwaha, Rai, Chaturvedi, Kumar, Pradhan, Kumar, Kumar, Juliana, Mishra and Kumar2022). However, prediction accuracy can decline under strong environmental variation, underscoring the need to train populations across representative stress environments.
Stepwise conservation-to-deployment pipeline for harnessing wild wheat relatives in dryland wheat improvement.

Emerging tools such as speed introgression and genome editing provide opportunities to fine-tune wild alleles, for example, by reducing the size of salinity-tolerance introgressions, such as HKT1;5, to minimize yield penalties under non-stress conditions (Munns et al. Reference Munns, James, Xu, Athman, Conn, Jordans, Byrt, Hare, Tyerman, Tester, Plett and Gilliham2012; Li et al. Reference Li, Zhu, Fan, Li, Liu, Shaheen, Nie, Li, Liu, Li, Liu, Yang, Guo, Zhu, Bu, Li, Liang, Bai, Ma, Guo, Zhang, Huang, Zhou and Song2024a). Complementary case studies illustrate the deployment spectrum: CRISPR/Cas9-mediated knockout of the yield-related negative regulator TaD27 in an elite wheat cultivar increased tiller number and grain yield without morphological penalties, demonstrating how targeted editing can optimize major-effect genes directly in breeding-ready backgrounds while avoiding linkage drag (Awan et al. Reference Awan, Amin, Rasheed, Saeed and Mansoor2024). In contrast, high-throughput whole-genome speed-introgression platforms have enabled rapid transfer of the Ae. tauschii genome into elite wheat within approximately two years, providing scalable access to complex, polygenic stress-adaptive variation that remains difficult to capture through conventional introgression (Li et al. Reference Li, Zhu, Fan, Li, Liu, Shaheen, Nie, Li, Liu, Li, Liu, Yang, Guo, Zhu, Bu, Li, Liang, Bai, Ma, Guo, Zhang, Huang, Zhou and Song2024a). Together, these approaches indicate that genome editing is most effective for precise trait optimization, whereas accelerated introgression pipelines remain essential for mobilizing polygenic resilience traits from WWR. Nevertheless, their practical deployment is constrained by regulatory uncertainty surrounding genome editing, limited high-throughput phenotyping capacity, and high operational costs, particularly in dryland breeding programmes of low- and middle-income countries, highlighting the need for coordinated investment and capacity building to translate technological advances into field-level impact (Louafi and Welch Reference Louafi and Welch2021; Zhu et al. Reference Zhu, Jhingan, Chen and Stein2026).
Conservation modalities: from static preservation to dynamic use
Ex situ and in situ conservation approaches are complementary rather than interchangeable, and both are required to sustain WWR’s adaptive potential under climate change (Fig. 2; Maxted et al. Reference Maxted, Amri, Castañeda-Álvarez, Dias, Dulloo, Fielder, Ford-Lloyd, Iriondo, Brehm, Nilsen and Thormann2016; Dempewolf et al. Reference Dempewolf, Baute, Anderson, Kilian, Smith and Guarino2017). Ex situ genebank collections provide immediate and well-documented access to germplasm for genomic analysis, pre-breeding, and trait discovery. However, evidence shows that such collections often capture only a portion of the diversity present in natural populations, frequently exhibiting reduced heterozygosity and allelic richness when sampling is limited (Fu Reference Fu2017; Wei and Jiang Reference Wei and Jiang2021). Sampling strategies that include 30–50 individuals distributed across a species’ geographic range substantially improve the genetic representativeness of ex situ collections (Wei and Jiang Reference Wei and Jiang2021). Similarly, Özkan et al. (Reference Özkan, Kafkas, Sertac Ozer and Brandolini2005) argue that for Hordeum spontaneum, priority should shift toward sampling a greater number of populations rather than increasing the number of individuals within a single population, in order to better capture and safeguard genetic diversity in the wild barley gene pool. We consider this population-focused approach to be equally applicable to WWR. In situ conservation complements these limitations by maintaining WWR populations within their native habitats, where they continue to evolve under environmental pressures such as drought, salinity and pathogen stress (Maxted et al. Reference Maxted, Amri, Castañeda-Álvarez, Dias, Dulloo, Fielder, Ford-Lloyd, Iriondo, Brehm, Nilsen and Thormann2016; Bellon et al. Reference Bellon, Dulloo, Sardos, Thormann and Burdon2017). Global gap analyses indicate that nearly 30% of crop wild relatives are underrepresented in genebanks, and more than 70% require additional collecting, underscoring significant gaps in current conservation coverage (Castañeda-Álvarez et al. Reference Castañeda-Álvarez, Khoury, Achicanoy, Bernau, Dempewolf, Eastwood, Guarino, Harker, Jarvis, Maxted, Müller, Ramirez-Villegas, Sosa, Struik, Vincent and Toll2016). Targeted international initiatives have begun addressing these gaps; for example, coordinated collecting missions between 2013 and 2018 secured over 4,500 new crop wild relative accessions representing 355 taxa across 25 countries, many of which are now incorporated into pre-breeding programmes (Eastwood et al. Reference Eastwood, Tambam, Aboagye, Akparov, Aladele, Allen, Amri, Anglin, Araya, Arrieta-Espinoza, Asgerov, Awang, Awas, Barata, Boateng, Magos Brehm, Breidy, Breman, Brenes Angulo, Burle, Castañeda-Álvarez, Casimiro, Chaves, Clemente, Cockel, Davey, De la Rosa, Debouck, Dempewolf, Dokmak, Ellis, Faruk, Freitas, Galstyan, García, Ghimire, Guarino, Harker, Hope, Humphries, Jamora, Jatoi, Khutsishvili, Kikodze, Kyratzis, León-Lobos, Liu, Mainali, Mammadov, Manrique-Carpintero, Manzella, Mat Ali, Medeiros, Mérida Guzmán, Mikatadze-Pantsulaia, Mohamed, Monteros-Altamirano, Morales, Müller, Mulumba, Nersesyan, Nóbrega, Nyamongo, Obreza, Okere, Orsenigo, Ortega-Klose, Papikyan, Pearce, Pinheiro de Carvalho, Prohens, Rossi, Salas, Singh Shrestha, Siddiqui, Smith, Sotomayor, Tacán, Tapia, Toledo, Toll, Vu, Vu, Way, Yazbek, Zorrilla and Kilian2022). Nevertheless, regional analyses still reveal significant underrepresentation. In the Fertile Crescent, assessments of 441 priority crop wild relatives found that 30% are absent from genebanks and 57% are represented by fewer than 10 accessions, highlighting the vulnerability of adaptive diversity and the need for targeted conservation strategies (Vincent et al. Reference Vincent, Wiersema, Kell, Fielder, Dobbie, Castañeda-Álvarez, Guarino, Eastwood, Leόn and Maxted2013; Zair et al. Reference Zair, Maxted, Brehm and Amri2021). Importantly, ongoing evolution in natural populations – particularly under the pressures of climate change – generates new adaptive alleles that may be critical for future crop improvement. The loss of wild populations through habitat degradation therefore risks eliminating valuable genetic variation before it can be characterized or incorporated into breeding programmes.
An integrated conservation framework that links static preservation with dynamic evolutionary processes is therefore essential. Genebanks should increasingly function as genomics-informed resources supported by broad geographic sampling and improved characterization, while in situ genetic reserves should be strategically established in diversity hotspots relevant to dryland agriculture (Vincent et al. Reference Vincent, Hole and Maxted2022; Wambugu et al. Reference Wambugu and Henry2022). Aligning conservation priorities with breeding needs can ensure that WWRs are preserved not only as biodiversity assets but also as practical reservoirs of climate-adaptive variation for future wheat improvement.
Integrated conservation-to-deployment pathways
Translating WWR diversity into climate-resilient cultivars requires coordinated conservation-to-deployment pipelines that integrate genomic discovery, phenotyping, and breeding. A practical framework includes: (i) targeted gap-filling of underrepresented WWR gene pools through coordinated ex situ and in situ conservation; (ii) genomic and haplotype-based discovery of adaptive alleles using pangenomes and GWAS; (iii) multi-environment phenotyping to evaluate yield–stress tolerance trade-offs and (iv) deployment through pre-breeding platforms such as synthetic wheats, introgression libraries and genomic prediction (Fig. 2; Mujeeb-Kazi et al. Reference Mujeeb-Kazi, Gul, Farooq, Rizwan and Ahmad2008; Warburton et al. Reference Warburton, Crossa, Franco, Kazi, Trethowan, Rajaram, Pfeiffer, Zhang, Dreisigacker and Ginkel2006; Kilian et al. Reference Kilian, Dempewolf, Guarino, Werner, Coyne and Warburton2021; Farooq et al. Reference Farooq, Frei, Zeibig, Pantha, Özkan, Kilian and Siddique2025).
The efficiency of this pipeline depends not only on scientific advances but also on the governance frameworks that regulate access to WWR germplasm. Delays or uncertainty in access procedures can prolong the establishment of pre-breeding populations, thereby slowing the introgression and fixation of adaptive alleles and delaying the assessment of yield stability across target environments.
Future innovations, including haplotype-informed pangenome selection and de novo domestication, hold high potential but differ markedly in feasibility and time horizon, largely due to differences in regulatory frameworks, infrastructure requirements, and phenotyping capacity (Kilian et al. Reference Kilian, Dempewolf, Guarino, Werner, Coyne and Warburton2021; Zhu et al. Reference Zhu, Jhingan, Chen and Stein2026). In particular, access conditions may vary depending on the taxonomic status of the material: while many cultivated Triticum resources and some closely associated Aegilops materials are exchanged through the ITPGRFA multilateral system, numerous wild relatives fall outside Annex 1 and are therefore subject to bilateral access and benefit-sharing (ABS) procedures under the CBD and Nagoya Protocol (Sherman and Henry Reference Sherman and Henry2021; Ebert et al. Reference Ebert, Engels, Schafleitner, Hintum and Mwila2023; Engels and Ebert Reference Engels and Ebert2024). These distinctions influence how rapidly WWR diversity can move from conservation collections into pre-breeding and applied breeding pipelines. Ensuring equitable participation in this pipeline is therefore critical. Without improved access to conserved germplasm, genomic tools, and phenotyping platforms in dryland countries, the benefits of WWR-derived resilience are unlikely to reach the regions most vulnerable to climate stress (Louafi and Welch Reference Louafi and Welch2021). Aligning genebanks, international breeding programmes, and enabling policy frameworks into a unified pipeline can reposition WWR from passive genetic reservoirs into active drivers of climate-resilient wheat improvement, while maintaining realistic expectations about yield stability and adoption timelines (Özkan et al. Reference Özkan, Brandolini, Schäfer-Pregl and Salamini2002; Kilian et al. Reference Kilian, Dempewolf, Guarino, Werner, Coyne and Warburton2021; Mastrangelo et al. 2025).
Policy and international cooperation
Scientific advances in genomics, phenotyping, and pre-breeding have substantially increased WWR’s potential to support climate-resilient wheat improvement. However, translation into breeding impact ultimately depends on policy environments and international cooperation (Fig. 2). Access to WWR, the movement of germplasm and data, and the equitable sharing of benefits are governed by complex legal and governance frameworks that can either enable or constrain innovation. Integrating these governance considerations with breeding pipelines is therefore essential to ensure that WWR-derived diversity is conserved, accessed and deployed efficiently, particularly in dryland regions where climate adaptation needs are most urgent.
Harmonizing access, benefit-sharing, and digital sequence information
Effective conservation and utilization of WWR for climate-resilient wheat improvement depend on coherent implementation of international ABS frameworks, particularly the CBD, the Nagoya Protocol and ITPGRFA. As discussed in the conservation-to-deployment pathway above, differences between the ITPGRFA multilateral system and bilateral ABS procedures can influence the speed at which WWR germplasm enters pre-breeding and applied breeding programmes. Greater harmonization across these frameworks is required to reduce legal uncertainty for pre-breeding and genomics-assisted research.
The rapid expansion of genomic sequencing has intensified debates around Digital Sequence Information (DSI), as breeders increasingly access adaptive alleles through sequence data rather than physical germplasm. Current ABS mechanisms inadequately address DSI, creating risks of bypassing benefit-sharing obligations and eroding trust with provider countries (Sherman and Henry Reference Sherman and Henry2021; Lawson et al. Reference Lawson, Humphries and Rourke2024). Internationally agreed guidance that links DSI use to transparent benefit-sharing mechanisms is therefore essential to sustain genomic research on WWR while ensuring equity.
Enabling equitable utilization: Capacity, infrastructure and data
Policy effectiveness is ultimately determined by national and regional capacity to conserve, characterize and deploy WWR. Many diversity-rich, dryland regions lack well-resourced genebanks, molecular characterization facilities and high-throughput phenotyping platforms, limiting their ability to participate meaningfully in WWR-based breeding pipelines (Ebert et al. Reference Ebert, Engels, Schafleitner, Hintum and Mwila2023). Recent multi-country assessments applying standardized monitoring and evaluation frameworks across 20 Global South countries demonstrate that systematic genebank performance reviews can identify operational gaps, guide targeted capacity development and improve the efficiency and effectiveness of ex situ conservation systems. Thus, strengthening genebank infrastructure and investing in genomic and phenotypic characterization are critical prerequisites for effective WWR utilization.
Beyond physical infrastructure, interoperable data systems are essential to translate conserved diversity into usable breeding knowledge. The value of ex situ collections depends on the systematic integration of passport, phenotypic, environmental and omics data within federated digital infrastructures that enable discovery, reuse and traceability (Shaw et al. ). The Wheat Information System (WheatIS) exemplifies this approach by linking genomic and phenotypic datasets to accession-level metadata across distributed repositories, allowing climate-adaptive traits identified in WWR to be traced through pre-breeding materials and into applied breeding pipelines (Langridge et al. Reference Langridge, Alaux, Almeida, Ammar, Baum, Bekkaoui, Bentley, Beres, Berger, Braun, Brown-Guedira, Burt, Caccamo, Cattivelli, Charmet, Civáň, Cloutier, Cohan, Devaux, Doohan, Dreccer, Ferrahi, Germán, Goodwin, Griffiths, Guzmán, Handa, Hawkesford, He, Huttner, Ikeda, Kilian, King, King, Kirkegaard, Lage, Le Gouis, Mondal, Mullins, Ordon, Ortiz-Monasterio, Özkan, Öztürk, Pereyra, Pozniak, Quesneville, Quincke, Rebetzke, Christoph Reif, Saavedra-Bravo, Schurr, Sharma, Singh, Singh, Snape, Tadesse, Tsujimoto, Tuberosa, Willis and Zhang2022; Tian et al. Reference Tian, Wang, Liu and Zhao2025). Consistent with the Global Information System vision, WheatIS operates as part of a federated network rather than a centralized database, relying on shared standards, application programming interfaces, and persistent identifiers such as Digital Object Identifiers to address long-standing challenges of data fragmentation and germplasm traceability (Shaw et al. ). However, the effectiveness of such platforms depends on equitable global participation. Many diversity-rich, climate-vulnerable regions remain underrepresented in global datasets, reinforcing data asymmetries that limit trait discovery and bias downstream utilization. Expanding WheatIS-like infrastructures through sustained engagement with provider countries, alongside support for data curation, standardized phenotyping, and identifier assignment, is therefore critical to ensure that WWR-derived knowledge contributes meaningfully to climate-resilient wheat improvement where it is most urgently needed.
Cooperative models, regional hubs and long-term sustainability
International cooperation provides practical pathways to translate policy frameworks into operational impact (Fig. 2). The Consultative Group for International Agricultural Research (CGIAR) genebanks demonstrate how large-scale conservation, standardized germplasm exchange, and coordinated benefit-sharing can support global crop improvement (Engels and Ebert Reference Engels and Ebert2024). Complementing these operational models, coordination platforms such as the Wheat Initiative (https://www.wheatinitiative.org/) play a critical enabling role by aligning research priorities, data standards and pre-breeding efforts across public institutions, genebanks and breeding programmes worldwide. Regional initiatives such as the European Genebank Integrated System further highlight the value of coordinated conservation tailored to specific ecological contexts.
Building on these models, the establishment of regional conservation–breeding hubs in WWR diversity hotspots offers a promising pathway forward. However, many of these diversity hotspots, including parts of the Fertile Crescent and West Asia, are affected by geopolitical instability, which can threaten both in situ conservation and long-term access to genetic resources, underscoring the importance of distributed conservation strategies. Embedding WWR explicitly within national climate adaptation strategies can further align conservation priorities with breeding demand (Reynolds et al. Reference Reynolds, Lewis, Ammar, Basnet, Crespo-Herrera, Crossa, Dhugga, Dreisigacker, Juliana, Karwat, Kishii, Krause, Langridge, Lashkari, Mondal, Payne, Pequeno, Pinto, Sansaloni, Schulthess, Singh, Sonder, Sukumaran, Xiong and Braun2021; Bohra et al. Reference Bohra, Choudhary, Bennett, Joshi, Mir and Varshney2024). Long-term funding commitments for genebanks and genetic reserves, coupled with transparent tracking of germplasm utilization, are essential to sustain trust and ensure fair benefit-sharing. Initiatives such as the Crop Trust’s BOLD Project (Biodiversity for Opportunities, Livelihoods and Development) – a decade-long, Norway-funded programme designed to strengthen conservation, documentation, and use of crop diversity in collaboration with national genebanks and breeding partners – illustrate how stable resourcing and coordinated capacity support can enhance both conservation and use of diversity by breeders and farmers alike (https://bold.croptrust.org/). Critically, co-development of pre-breeding pipelines with provider countries rather than one-directional germplasm extraction will be key to transforming WWR from conserved assets into active drivers of climate-resilient wheat improvement.
Conclusion
The WWRs constitute an indispensable genetic foundation for enhancing wheat resilience to drought and salinity in dryland agroecosystems. This review advances previous syntheses by integrating adaptive trait biology, modern genomic deployment tools, and policy and access constraints into a unified framework that links WWR discovery to practical breeding outcomes. The review highlights that while pangenomics, GS, speed introgression and genome editing have markedly improved the precision and pace of WWR utilization, their impact remains uneven due to persistent challenges, including polygenic stress tolerance, genotype × environment interactions, linkage drag, regulatory uncertainty around genome editing, and limited pre-breeding and phenotyping capacity in many dryland regions.
A realistic pathway forward requires prioritization rather than breadth. Key near-term opportunities lie in well-characterized gene pools such as Ae. tauschii and T. dicoccoides, targeted deployment in representative dryland agroecologies, and focused mitigation of bottlenecks in allele validation and introgression. Strengthening conservation–breeding linkages through genomics-enabled ex situ and in situ strategies, alongside equitable access frameworks under the CBD, Nagoya Protocol, and ITPGRFA, will be essential to sustain the flow of adaptive diversity. Ultimately, WWR should not be viewed as auxiliary resources but as core components of future dryland wheat improvement, without which durable adaptation to increasing climatic variability is unlikely to be achieved. Progress in WWR utilization can be monitored through practical indicators, including the number of WWR-derived introgressions released in commercial cultivars, the expansion of introgression libraries in breeding programmes, and the frequency of WWR alleles deployed in multi-environment trials.
Declaration of AI Assistance in Language Editing
The authors used ChatGPT to improve language clarity and to correct minor grammatical and stylistic errors. All suggestions were carefully reviewed, and the authors retain full responsibility for the content of this publication.