INTRODUCTION
Streptococcus pneumoniae is the leading cause of bacterial infection in infants and young children. In 2005, the World Health Organization (WHO) estimated that 1·6 million people die of pneumococcal disease annually, including 0·7–1 million children (aged <5 years) mostly from developing countries [1]. This situation has been worsened by the rapid spread of antimicrobial-resistant S. pneumoniae and is now a global problem.
To date, more than 93 serotypes have been identified in S. pneumoniae. Before the first pneumococcal conjugate vaccine was introduced, the antibiotic-resistant serotype 23F was one of the most common types worldwide [Reference Pilishvili2, Reference Barnes3]. Spain23F-1 clone or sequence type (ST) 81 first emerged in 1970 and became the first antibiotic-resistant S. pneumoniae clone whose standard nomenclature was given by the Pneumococcal Molecular Epidemiology Network (PMEN) (http://www.sph.emory.edu/PMEN/index.html). This clone was one of the first pandemic penicillin (PEN)-resistant isolates from Spain in the 1980s [Reference Muñoz4] and in the late 1990s, it constituted about 40% of PEN-resistant isolates in the USA [Reference Corso5]. Previous studies have reported this clone to be resistant to tetracycline (TCY) and chloramphenicol (CHL), and to be associated with the emergence of fluoroquinolone and macrolide resistance [Reference Pletz6, Reference Reinert7]. Although widespread in Europe, Asia, Africa, and the USA [Reference Croucher8], ST81 has recently been identified in 7·7% of S. pneumoniae isolates from Chongqing, southern China [Reference Zhang9]. Since the 1990s, serotype 23F has been one of the most resistant types recovered in Beijing [Reference Wang, Chen and Huebener10, Reference Yao and Yang11] and almost all isolates collected in 2010 were ST81 [Reference Zhou12]. However, whether ST81 was predominant before 2010 is unknown. In the present study, we analysed serotype 23F S. pneumoniae isolates collected in Beijing from 1997 to 2006 and 2010 and report our findings on their antibiotic susceptibility, genotype distribution, and temporal trends.
METHODS
Clinical isolates
The study protocol was in accordance with the ethical standards of the responsible regional committee on human experimentation and the Helsinki Declaration of 1975 (revised 1983); it was approved by the ethics committee of the Beijing Children's Hospital. Nasopharyngeal swabs were taken with written informed consent from the parents or legal guardians of the children. A total of 1132 S. pneumoniae isolates were collected from outpatients aged <5 years diagnosed with acute upper respiratory infection in Beijing Children's Hospital. A total of 101 isolates were identified as serotype 23F based on the Quellung reaction using a set of factor antisera (Statens Serum Institute, Copenhagen, Denmark) and of these 99 were investigated further owing to the loss of two strains collected in 1999.
Anticrobial susceptibility testing
The minimum inhibitory concentrations (MICs) for PEN, amoxicillin–clavulanic acid (AMC), ceftriaxone (CRO), cefuroxime (CXM), erythromycin (ERY), and vancomycin (VAN) were determined using E-test strips (AB Biodisk, Sweden) [Reference Kelly13], and for TCY, sulfamethoxazole–trimethoprim (SXT), and CHL by the disk diffusion method. Breakpoints were determined in accordance with the Clinical and Laboratory Standards Institute 2010 criteria [14] using S. pneumoniae ATCC49619 as the quality control. Isolates were considered multidrug resistant (MDR) if they were not susceptible to three or more classes of antimicrobials.
Detection of macrolide resistance genes
The ermB and mefA/E resistance genes were amplified by polymerase chain reaction (PCR) for all ERY non-susceptible strains using the primers and PCR conditions as previously described [Reference Sutcliffe, Tait-Kamradt and Wondrack15]. Each PCR reaction contained 500 ng template DNA, 50 mm KCl, 10 mm Tris–HCl (pH 8·3), 200 μm of each deoxynucleotide triphosphate, 2·5 U Taq DNA polymerase (Dalian, Takara Bio, China), 1·5 mm MgCl2, and 1·5 μm of each primer. PCR products were visualized by 1·5% agarose gel electrophoresis and gold-view staining.
Multilocus sequence typing (MLST)
Internal fragments ∼450 bp long from the aroE, gdh, gki, recP, spi, xpt, and ddl genes were amplified by PCR as previously described [Reference Enright and Spratt16]. Alleles that were not present in the pneumococcal MLST database were verified by resequencing the gene fragments on both strands. These alleles were submitted to the MLST S. pneumoniae database for designation. The eBURST algorithm (http://eburst.mlst.net) was used to estimate the relationships among isolates. Various STs were subdivided into one group as a clonal complex (CC) only if they shared six identical alleles of the seven MLST loci with another ST in the group.
Pulsed-field gel electrophoresis (PFGE)
Pneumococci from overnight cultures were suspended in low-melting-point agarose at a concentration of 1·2 in a Dade turbidity meter to form plugs. The bacteria were incubated in 500 mm EDTA/1% N-lauroylsarcosine overnight at 55 °C and DNA was digested with SmaI (New England Biolabs, USA) at 30 °C overnight according to the protocol described by Lefevre et al. [Reference Lefevre17]. PFGE was performed in 1·1% agarose (SeaKem GTG agarose, Switzerland) and 0·5× Tris-borate-EDTA buffer using an electrophoresis system (CHEF-DR III, Bio-Rad Laboratories, USA) at 6 V/cm (3–40 s switch time) for 23 h. Gels were stained with ethidium bromide for 30 min and visualized under UV light. The image was captured using the Gel Doc system (Cell-Bio Sciences, USA), and saved dendrograms of unweighted-pair group method with arithmetic means were constructed using Bionumerics software (Applied Maths, Belgium). PFGE patterns were clustered for isolates with ⩾80% genetic relatedness on the dendrogram. An optimization value of 1% and a position tolerance of 2% were used for the analysis.
Statistical analysis
Drug susceptibility data were analysed using WHONET v. 5.6 software, recommended by the WHO. The χ2 test, calculated using SPSS software v. 10.0 (SPSS Inc., USA), was used for statistical comparisons. A two-tailed cut-off of P < 0·05 indicated statistical significance.
RESULTS
Frequency of serotype 23F
In the entire study period, serotype 23F was identified in 8·9% (101/1132) S. pneumoniae isolates and their frequencies for different years were 10·7% (n = 25), 6·7% (n = 12), 10·4% (n = 21), 6·6% (n = 16), 10·2% (n = 14), and 9·3% (n = 13), respectively. No temporal trend was found (χ2 = 30·00, P > 0·05).
Antimicrobial susceptibility testing
All 99 strains were susceptible to VAN and AMC. The number of intermediate and resistant strains, as well as their frequency against seven other antimicrobials, is shown in Table 1. Only one isolate was resistant to PEN (MIC 16 μg/ml). Non-susceptibility to CXM significantly increased from 4% (1997) to 28·5% (2006) to 92·4% (2010). A total of 97 (98·0%) isolates were resistant to ERY with high MICs (94 isolates had a MIC ⩾256 μg/ml); 95 carried only the ermB gene and two carried both ermB and mefA/E genes. No strain was positive for the mefA/E gene alone. Moreover, 90 isolates had multidrug resistance to ERY, SXT, TCY, and 25 isolates in addition were resistant to β-lactam antibiotics.
Susceptibility against seven antimicrobials of the 99 serotype 23F strains isolated in Beijing, n (%)
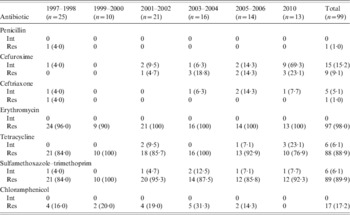
Int, Intermediate; Res, resistant.
MLST and PFGE
MLST analysis revealed 16 STs, six of which were novel (ST6825, ST7909–7913). The most common STs were ST342 (45·5%), ST81 (17·2%), ST802 (8·1%), ST2624 (6·1%), ST242 (5·1%), and ST6321 (5·1%). The predominant STs varied during the study period. The first ST81 was isolated in 2001, and its frequency increased thereafter, reaching 84·6% in 2010. In 1997–2006, ST342 was predominant and fluctuated over the study with a decreasing trend; none was identified in 2010 (Fig. 1).
[colour online]. Distribution of different sequence types (STs) during the study period.

eBURST analysis found four CCs and five singletons. CC342 comprising five STs was the most common group and accounted for 59·6% of all the isolates and CC81 for 18·2%; only one of this complex was not identified as ST81. CC802 (nine isolates), CC242 (seven isolates), and singletons are shown in the population snapshot in Figure 2. Seven different PFGE patterns were found in all isolates and the four major patterns (A–D) corresponded respectively to CC242, CC81, CC802, and CC342 (Supplementary Fig. S1, available online).
[colour online]. Population snapshot of 99 S. pneumoniae strains as revealed by eBURST analysis. Lines indicate the presence of single locus variant links in particular sequence types indicated by circles. The size of the circle corresponds to the number of isolates belonging to a sequence type.

Except for one isolate, all isolates identified as CC81 and CC242 were non-susceptible to CXM and showed higher PEN MICs than the other CC/STs (Table 2, Supplementary Fig. S1). Most of the strains (96%, 24/25) in these two CCs were consistently resistant to β-lactam, ERY, SXT, and TCY. By contrast, all CC342 strains were susceptible to the four tested β-lactam antimicrobials, except for one isolate that was classified as intermediate to CRO (MIC 2 μg/ml). The CC342 strains were frequently resistant to ERY, SXT, and TCY.
Susceptibility of the 97 serotype 23F strains to seven antimicrobial in different clonal complexes (CCs) and sequence types (STs)
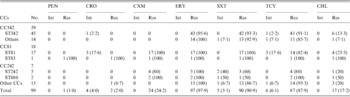
PEN, Penicillin; AMC, amoxicillin–clavulanic acid; CXM, cefuroxime; CHL, chloramphenicol; SXT, sulfamethoxazole–trimethoprim; CRO, ceftriaxone; ERY, erythromycin; VAN, vancomycin; TCY, tetracycline; Int, Intermediate; Res, resistant.
DISCUSSION
The present study showed that the frequency of serotype 23F in all the S. pneumoniae isolates was 8·9%, which is similar to previously reported data from Beijing [Reference Yao and Yang11, Reference Xue18, Reference Liu19]. No temporal trend was found. A significant decrease in serotype 23F was reported in Western countries after the global introduction of the 7-valent conjugate pneumococcal vaccine (PCV7) [Reference Pilishvili2, Reference Whitney20, Reference Messina21]. Since September 2008, PCV7 has become available in the private sector in China but to date, only a few parents have sought immunization for their children because of the very high cost (860 RMB or US$136 per dose). Thus, PCV7 immunization is not yet ‘universally administered’ in China and hence, no decrease in the frequency of serotype 23F was evident in our study.
ST81, in the present study, was found to be the predominant ST in 2010. Zhang et al. [Reference Zhang9] also collected ST81 isolates from 2009 to 2010 in southern Chin but this clone proved to be rather uncommon during the period from 1997 to 2006. ST81 was first identified in 1978 by Spanish researchers [Reference McDougalL22] and since then has been found to have spread globally. This ST was very common in the USA and Europe before the introduction of PCV [Reference Croucher8, Reference Ardanuy23]. Some earlier surveys indicated that ST81 had emerged and remained stable in some Asian countries or regions from the 1990s [Reference Ip24–Reference Byung28] and thus the clone may have spread into the hinterland of China from other countries in the region, and may have the potential to spread further. This epidemic spread may be a result of the increasing movements of populations, which contributes to the emergence of antimicrobial resistance worldwide. This situation emphasizes the need for vaccination to interrupt the transmission of resistant clones.
In the present study, ST342 was found to have been the predominant strain before 2006 but its frequency decreased slightly from 1997 to 2006 and unexpectedly was found in 2010, but was largely displaced by ST81. Considering the high β-lactam resistance of ST81 (or CC81), the low resistance of ST342 (or CC342), and the common use of PEN and cephalosporin in China [Reference Zhang29], the genotype replacement in serotype 23F may be driven by antibiotic use. In our previous reports [Reference Gao30–Reference Yao32], we emphasized that antibiotic abuse increased serogroup 19 (especially 19A) and subtype 6B-II before the introduction of PCV7 in China. The present study reconfirms this phenomenon because the genotype replacement described here cannot be associated with vaccine administration. A similar genotype replacement was discovered in other serotypes and in other countries where serotype replacement was identified before the introduction of PCV [Reference Choi33].
In Asia, most ERY-resistant strains that carry both ermB and mefA/E are serotypes 19F and 14 [Reference Song34]. The prevalence of macrolide-resistant genes varies in different countries and regions. For example, the dominant gene in the hinterland of China and Taiwan is ermB, whereas in Hong Kong it is mefA/E [Reference Song34–Reference Ko and Song37]. Therefore, almost all ERY-resistant strains expressing 23F (97·9%) carried only the ermB gene. However, the very low carriage of mefA/E in serotype 23F (2%) should be considered. Ip et al. [Reference Ip24] found that the mefA/E gene was expressed in serotype 23F isolates with ST81 in Hong Kong in 1994. Recently, Byung et al. [Reference Byung28] reported for the first time in Korea three isolates belonging to CC81 (with serotypes 23F, 15B/C, and 6A) that were positive for both ermB and mefA/E genes. In the present study, two ST81 isolates (CC81) harbouring both genes were identified in 2004 and 2010. These findings suggest that the mefA/E gene may have been transferred from another clone to ST81 when it spread into Asia.
A limitation of the present study is the disruption of survey continuity from 2007 to 2009 and further that all strains were isolated from a single hospital. Considering the high number of patient visits to the largest paediatric hospital in China (5000–6000 visits per day on average) and the total number of pneumococcal isolates, we believe that the present results objectively demonstrate the epidemiological changes of serotype 23F S. pneumoniae in Beijing.
In summary, serotype 23F remains the most common serotype in children with respiratory infections in Beijing. However, the genotype component has changed because of antibiotic misuse. The MDR PMEN1 clone (ST81) replaced another MDR ST342 to become the predominant clone in Beijing. Further long-term surveys of S. pneumoniae serotype 23F are required to monitor the ST prevalence and antimicrobial resistance of this important human pathogen.
SUPPLEMENTARY MATERIAL
For supplementary material accompanying this paper visit http://dx.doi.org/10.1017/S0950268812002269.
ACKNOWLEDGEMENTS
This study was supported by the Special fund for high-level personnel development in the Beijing Health System (Y. H. Yang, 2011-3-052) and the Beijing Nova Program (K. H. Yao, 2008B64). The authors thank all the participants from the Beijing Children's Hospital.
DECLARATION OF INTEREST
None.





