Introduction
Depressive disorders emerged as a major contributor to the global disability burden across all populations. Ranked second among non-communicable diseases (NCDs), depressive disorders caused 56.3 million years lived with disability (YLDs) in 2021 (GBD 2021 Diseases and Injuries Collaborators, 2024). Major depressive disorder (MDD) is one of the most disabling mental disorders, affecting over 35 million people worldwide. MDD is characterized by significant and persistent low moods and is driven by a combination of genetic and environmental factors (Liu et al., Reference Liu, Wang, Chen, Zhang, Zhang and Xie2023). Nonmalignant gastrointestinal diseases, characterized by chronic inflammation and tissue damage, affect over 5% of the population cumulatively, with annual incidence rates reaching 0.3%. These conditions cause debilitating symptoms like chronic pain and malnutrition, severely impairing quality of life while generating substantial healthcare costs through both direct medical expenditures and indirect productivity losses. Their persistent burden underscores an urgent need for improved management strategies (Bai et al., Reference Bai, Wang, Shen, An, Yang and Mo2025).
Previous research has mainly viewed psychological disorders, such as depression, as secondary complications of chronic diseases. However, few studies have evaluated them as independent risk factors that contribute to the onset or progression of somatic diseases. This conceptual gap highlights the need to investigate the potential causal role of depression in physical health outcomes. Most observational studies have investigated the role of gastrointestinal diseases in the development of depression (Haag et al., Reference Haag, Senf, Häuser, Tagay, Grandt, Heuft and Holtmann2008; Panara et al., Reference Panara, Yarur, Rieders, Proksell, Deshpande, Abreu and Sussman2014), but have been limited to the reverse impact. While Mendelian randomization (MR) studies have advanced causal insights into this bidirectional relationship across 24 gastrointestinal conditions (Ruan et al., Reference Ruan, Chen, Sun, Zhang, Zhao, Wang and Larsson2023), critical gaps persist for specific endpoints. Several gastrointestinal diseases, such as esophagitis, dyspepsia, gastritis, and duodenitis, remain understudied. Furthermore, the application of machine learning (ML)-based predictive models to untangle complex psychosomatic risk interactions remains scarce.
MR strengthens causal inference by leveraging genetic instruments to mitigate confounding and reverse causality (Davey Smith & Hemani, Reference Davey Smith and Hemani2014; Davies, Holmes, & Davey Smith, Reference Davies, Holmes and Davey Smith2018; Palmer et al., Reference Palmer, Sterne, Harbord, Lawlor, Sheehan, Meng and Didelez2011; Verduijn et al., Reference Verduijn, Siegerink, Jager, Zoccali and Dekker2010). And predictive modeling synthesizes findings with multi-dimensional data to develop personalized risk stratification tools that identify high-risk subgroups and evaluate the potential clinical impact of mental health interventions.
Our three-stage study, including the Cox proportional hazards model, MR, and ML-based predictive modeling, offers a powerful and synergistic approach to unraveling the complex bidirectional relationship between psychological disorders and somatic diseases. This multi-method approach overcomes the limitations of individual methods to provide comprehensive evidence for both causality and clinical utility in psychosomatic medicine. Based on a large-scale prospective cohort study at UK Biobank, the Psychiatric Genomics Consortium and the FinnGen Biobank, our study takes depression as the exposure and six gastrointestinal diseases as the outcome, with the aim of (1) evaluating the relationship between depression and the risk of gastrointestinal diseases; (2) investigating the causal relationship between depression and gastrointestinal diseases; (3) developing predictive models for incident gastrointestinal diseases in clinically depressed populations.
Methods
Study design and population
The UK Biobank is a large population-based cohort study that enrolled >0.5 million volunteers aged 37–73 at 22 assessment centers in England, Wales, and Scotland between 2006 and 2010 (Sudlow et al., Reference Sudlow, Gallacher, Allen, Beral, Burton, Danesh and Collins2015). Participants were asked to provide various health-related information through a touch-screen questionnaire, a brief face-to-face interview, physical and functional measurements, and biological samples at baseline or follow-up assessments (Collins, Reference Collins2012). Among 502,129 total participants, 44,189 were excluded in the present cohort study, including 27,543 with prevalent depression after baseline, 7713 with missing data, 7636 who preferred not to answer, and 1297 who were lost to follow-up (Supplementary Figure S1). The identification of depression occurring after baseline was performed sequentially, first using self-reported follow-up data and then validated against hospital-record ICD-10 diagnoses (F32–F33), to ensure the temporal accuracy of exposure classification. Additionally, participants diagnosed with six predefined gastrointestinal diseases at baseline were separately excluded to ensure disease-free cohorts for each outcome analysis.
Ascertainment of depression
In the UK Biobank, follow-up for all participants involved linkage with hospital admissions data and the national death registry records to identify the date of the first known diagnosis of gastrointestinal diseases. The International Classification of Diseases ICD-10 codes (F32 and F33) were used to identify participants with depression if one or more of these codes were recorded as a primary or secondary diagnosis in the health records (Prigge, Wild, & Jackson, Reference Prigge, Wild and Jackson2022). Individuals whose depression diagnosis occurred after the baseline assessment were excluded to avoid temporality issues in causality interpretation. This exclusion was consistent with the criteria described in Section “Study design and population”.
Identification of gastrointestinal diseases
Gastrointestinal diseases were identified according to International Classification of Diseases, Tenth Revision (ICD-10) codes obtained from hospital inpatient and death registry records, ensuring consistent case ascertainment across all outcomes. After assessing both data sufficiency and quality metrics, six representative gastrointestinal disorders were inclusively identified: esophagitis, gastroesophageal reflux disease (GERD), gastric ulcer, duodenal ulcer, gastritis and duodenitis, and dyspepsia. The International Classification of Diseases ICD-10 codes (K20, K21, K25, K26, K29, and K30) were used to identify participants with gastrointestinal diseases. Esophagitis was defined as code K20. GERD was defined as code K21. A gastric ulcer was defined as code K25. Duodenal ulcer was defined as code K26. Gastritis and duodenitis were defined as code K29. Dyspepsia was defined as code K30.
Assessment of covariates
We used the baseline touchscreen questionnaire and nurse-led interview to define the following potential confounders. Covariates were selected a priori based on previous literature and causal reasoning using a directed acyclic graph (DAG) framework to identify potential confounders of the association between depression and gastrointestinal diseases. The included variables – age, sex, education level, Townsend Deprivation Index (TDI), smoking status, alcohol intake frequency, physical activity, and body mass index (BMI) – were considered established risk factors for both depression and gastrointestinal disorders, as supported by prior evidence (Firth et al., Reference Firth, Solmi, Wootton, Vancampfort, Schuch, Hoare and Stubbs2020).
Data resources for Mendelian randomization analysis
Summary statistics on depression were obtained from the Psychiatric Genomics Consortium. To avoid the bias that sample overlap caused in the causal relationship investigation, the gastrointestinal diseases summary statistics were obtained from the FinnGen Biobank. All genome-wide association study (GWAS) data involved are de-identified, publicly available, and ethical approval and consent to participate were obtained by each GWAS, which was described in the original publications. The summary of the GWAS data used in our study is presented in Supplementary Table S1.
Instrumental variable selection and data harmonization
Genetic instruments for depression were selected as the 197 independent genome-wide significant single nucleotide polymorphisms (SNPs) (P < 5 × 10−8), after excluding variants in linkage disequilibrium (r2 < 0.001 or distance >10,000 kb) or shared with genetic instruments for the six gastrointestinal diseases (esophagitis, GERD, gastric ulcers, duodenal ulcer, dyspepsia, and gastritis-duodenitis). We calculated F-statistics for each exposure in univariable MR, with F-statistics >10 considered suggestive of adequate instrument strength (Palmer et al., Reference Palmer, Lawlor, Harbord, Sheehan, Tobias, Timpson and Sterne2012).
K is the number of SNPs, and N is the sample size.
R is the correlation coefficient, and it quantifies the strength and direction of the relationship between the instrument and the exposure. EAF is the frequency of the effect allele in the population for a given genetic variant, and β is the regression coefficient that quantifies the relationship between the genetic variant and the exposure.
Machine learning classifiers
We implemented seven ML algorithms – LightGBM (LGBM), Gradient Boosting Machine (GBM), Random Forest (RF), Logistic Regression (LR), K-Nearest Neighbors (KNN), Multilayer Perceptron (MLP), and XGBoost – to classify participants as either class 0 (predicted to remain gastrointestinal disease-free) or class 1 (predicted to develop an incident gastrointestinal disease). ML models were established among participants without gastrointestinal diseases at baseline to predict incident cases during follow-up. ML models were developed in the total population, with depression status incorporated as a key predictor to evaluate its utility as an early warning indicator for incident gastrointestinal diseases. All sociodemographic, lifestyle, and metabolic covariates used in the Cox models were also included as predictors to improve model performance. Postprocessing isotonic regressions were adopted to calibrate the raw predicted probabilities into actual incident gastrointestinal disease risks. We used the best-performing method, LGBM, as the final algorithm.
Statistical analysis
Associations between depression and gastrointestinal diseases
Multivariable Cox proportional hazards regression models were used to estimate hazard ratios (HRs) and 95% confidence intervals (CIs) for depression associated with six gastrointestinal diseases. Follow-up time was considered as the timescale, which was calculated from the recruitment date to the first diagnosis of six gastrointestinal diseases, death, loss to follow-up, or end of the current follow-up (November 30, 2022), whichever came first.
Two-sample Mendelian randomization
Two-sample MR was performed to estimate the effect of depression on six gastrointestinal diseases. All MR analyses fulfilled three critical assumptions: (1) the genetic variants are vigorously associated with depression; (2) the genetic variants are independent of any potential confounders; (3) the effect of genetic variants on gastrointestinal diseases only passes through each exposure (Hemani, Bowden, & Davey Smith, Reference Hemani, Bowden and Davey Smith2018). We used the inverse variance weighted (IVW) as the main method. Meanwhile, to ensure the robustness of the statistical results, we used both the weighted median (WM) and the MR-Egger based on Egger regression to perform sensitivity analyses of the statistical results.
Sensitivity analysis
We performed the following sensitivity analyses to assess the stability of the primary results. First, we excluded participants who had gastrointestinal disease events during the first 3 years of follow-up to minimize the influence of potential reverse causality. Second, we define the presence of any of the six gastrointestinal diseases as a single category, simplifying analysis to examine the overall risk of digestive diseases associated with depression. Third, to investigate the extent to which any attrition bias from the missing data affected our results, we repeated the main analyses in a full cohort with imputed data. Finally, possible pleiotropic effects on the selection of SNPs as IVs were analyzed using MR-Egger regression, which may provide a valuable assessment of whether horizontal pleiotropy is affecting the analysis (Burgess & Thompson, Reference Burgess and Thompson2017). To analyze the degree of heterogeneity present among the various forms of inheritance, the Cochran Q statistic was computed, and heterogeneity was considered significant when P < 0.05. We performed a leave-one-out sensitivity assessment to find potentially significant SNPs, where each SNP was excluded one by one from the MR. The forest plot, scatter plot, and funnel plot were plotted for sensitivity analysis.
Machine learning for incident gastrointestinal diseases
Models were trained and optimized through a nested leave-one-region-out cross-validation strategy. Specifically, the data was partitioned into 10 folds based on the geographical locations of participants’ recruitment centers. The geographical locations included East Midlands, London, North-East, North-West, Scotland, South-East, South-West, Wales, West Midlands, and Yorkshire and Humber. Each time, nine folds of data (training set) were used to develop a model, and the rest was used as a testing set. After 10 iterations, all folds of data were used as testing sets, and they were aggregated for evaluation.
The model’s performance was assessed using discrimination and calibration. Discrimination was evaluated by the area under the receiver operating characteristic curve (AUC). Calibration was visually depicted using calibration plots of decile subgroups, and Brier scores were calculated. Metrics of accuracy, sensitivity, specificity, precision, and F1-score were reported as well. In addition, we leveraged Shapley Additive explanations (SHAP) plots to interpret the relationship between predictor values and model output.
Results
Baseline characteristics
The baseline characteristics of participants are illustrated in Table 1. Out of 457,940 eligible participants, 9563 incident esophagitis events, 36,420 incident gastroesophageal reflux disease, 5469 incident gastric ulcer, 3096 incident duodenal ulcer, 37,225 incident gastritis and duodenitis, and 9153 incident dyspepsia were diagnosed, respectively, during a median follow-up of 13.7 years.
Baseline characteristics of study participants by incident depression

Associations between depression and gastrointestinal disease risk
Table 2 shows the association between depression and the risk of six gastrointestinal diseases. Compared to those with no depression, individuals with depression were associated with an increased risk of esophagitis (HR = 3.069, 95% CI: 2.888, 3.263), gastroesophageal reflux disease (HR = 2.907, 95% CI: 2.760, 3.063), gastric ulcer (HR = 3.184, 95% CI: 2.991, 3.389), duodenal ulcer (HR = 3.168, 95% CI: 2.972, 3.376), gastritis and duodenitis (HR = 2.925, 95% CI: 2.779, 3.080), dyspepsia (HR = 2.980, 95% CI: 2.803, 3.167), respectively, in age- and sex-adjusted model. When socio-demographic characteristics and lifestyle were further adjusted for in model 2, however, the HRs for six gastrointestinal diseases were slightly attenuated. In model 2, people with depression were associated with an increased risk of esophagitis (HR = 2.246, 95% CI: 2.111, 2.388), gastroesophageal reflux disease (HR = 2.220, 95% CI: 2.107, 2.340), gastric ulcer, respectively, (HR = 2.269, 95% CI: 2.130, 2.416), duodenal ulcer (HR = 2.264, 95% CI: 2.123, 2.415), gastritis and duodenitis (HR = 2.222, 95% CI: 2.110, 2.341), dyspepsia (HR = 2.209, 95% CI: 2.077, 2.349), respectively.
Multivariable HRs and 95% CIs of associations of depression on gastrointestinal diseases

Note: Model 1 was adjusted for age and sex. Model 2 was further adjusted for Townsend deprivation index, educational level, body mass index (BMI), smoking status, alcohol intake frequency, and physical activity. CI, confidence interval; HR, hazard ratio.
Effects of depression on gastrointestinal diseases
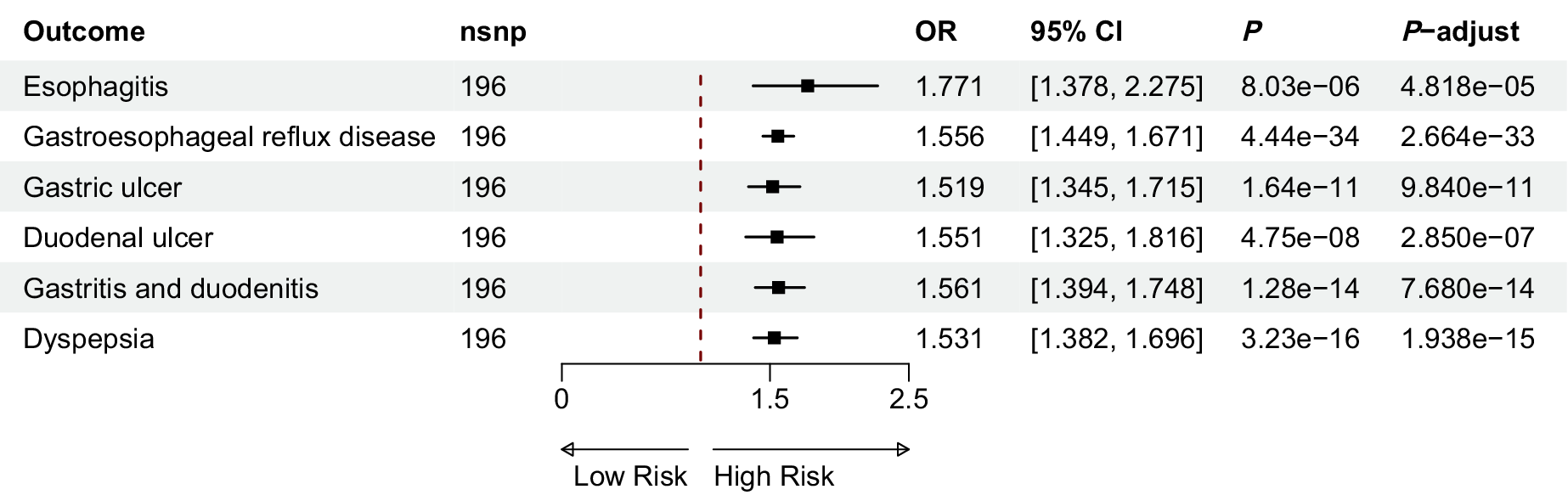
The results of the IVW analysis showed that depression was associated with a higher risk of esophagitis (OR = 1.771; 95% CI: 1.378, 2.275), gastroesophageal reflux disease (OR = 1.556; 95% CI: 1.449, 1.671), gastric ulcer (OR = 1.519; 95% CI: 1.345, 1.715), duodenal ulcer (OR = 1.551; 95% CI: 1.325, 1.816), gastritis and duodenitis (OR = 1.561; 95% CI: 1.394, 1.748) and dyspepsia (OR = 1.531; 95% CI: 1.382, 1.696), respectively (Figure 1).
Effect of depression on outcome in two sample MR.

Sensitivity analysis
In our sensitivity analyses, the effect of depression to gastrointestinal diseases did not change significantly when those who experienced gastrointestinal diseases within the first 3 years of the follow-up were not included (Supplementary Table S2), when we define the presence of any of the six gastrointestinal diseases as a single category (Supplementary Table S3), and when we repeated the main analyses in a full cohort with imputed data (Supplementary Table S4). Similarly, the associations remained largely unchanged after excluding participants using antidepressant medications and after additionally adjusting for participants using antidepressant medications as a covariate (Supplementary Tables S5–S6).
Scatter and funnel plots (Supplementary Figures S2–S3) indicated balanced sample selection. Forest and leave-one-out analyses (Supplementary Figures S4–S5) demonstrated stable results, with no single SNP exerting a significant influence on the estimates. No obvious bias was observed, and Cochran’s Q test detected heterogeneity only for GERD. In addition, the MR Egger method showed that there was no pleiotropy (Supplementary Table S7).
Performance of gastrointestinal diseases risk prediction
The ML models demonstrated differential predictive accuracy across the six gastrointestinal diseases investigated. Specifically, duodenal ulcers achieved the highest discriminative performance with an AUC of 0.76, followed closely by gastric ulcers (AUC = 0.75). Moderately robust predictions were observed for esophagitis (AUC = 0.74) and dyspepsia (AUC = 0.73), though these estimates did not reach the pre-specified clinical relevance threshold of AUC ≥0.75. However, the models showed reduced precision in classifying gastroesophageal reflux disease (AUC = 0.70) and gastritis and duodenitis (AUC = 0.70) (Figure 2).
Performance of gastrointestinal diseases risk prediction: (a) esophagitis, (b) gastroesophageal reflux disease (GERD), (c) gastric ulcer, (d) duodenal ulcer, (e) gastritis and duodenitis, and (f) dyspepsia.

Model interpretation
We leveraged SHAP values to assess the effect visually. Figure 3 aims to interpret the specific impact of each predictor by its value magnitude (coded by a gradient of colours) and tendency direction on the horizontal axis (the likelihood of developing gastrointestinal diseases). Higher depression levels were associated with an increased risk of each condition, with the strongest impact observed in esophagitis and dyspepsia, where SHAP values approached +1.5 and +2.0, respectively. Depression also had a notable but slightly weaker effect in GERD and gastric ulcer, with SHAP values around +1.0 to +1.2, and continued to contribute positively in duodenal ulcer, as well as gastritis and duodenitis. In addition, calibration curves (Supplementary Figure S6) showed good calibration, and performance metrics (Supplementary Table S8) suggested moderate discrimination with balanced precision and recall across outcomes.
Dierctional SHAP of risk predictors for gastrointestinal diseases: (a) esophagitis, (b) gastroesophageal reflux disease (GERD), (c) gastric ulcer, (d) duodenal ulcer, (e) gastritis and duodenitis, and (f) dyspepsia.

Discussion
Our three-stage study provides compelling evidence for the causal role of depression in elevating the risk of gastrointestinal diseases. Cox models establish the temporal relationship by demonstrating depression’s association with subsequent digestive disorders, while MR analysis strengthens causal inference by showing that genetically predicted depression leads to higher odds of developing gastrointestinal conditions. ML techniques further validate these findings by developing predictive models that quantify depression’s differential impact across various digestive diseases, revealing particularly strong associations with certain ulcerative conditions. Taken together, these complementary approaches provide robust evidence that depression is an independent and clinically meaningful risk factor for gastrointestinal pathology, with the magnitude of association differing across specific digestive disorders. The findings underscore the importance of considering depression in both the prediction and prevention of gastrointestinal diseases.
The multivariable-adjusted Cox proportional hazards model demonstrates that depression is robustly associated with heightened risks of six gastrointestinal diseases. Our findings align with several recent studies, which similarly found that individuals with depression tend to have a higher risk of developing GERD, peptic ulcers (including gastric and duodenal ulcers), or dyspepsia (He et al., Reference He, Wang, Yao, Li and Bai2022; Kao et al., Reference Kao, Sung, Huang, Lin, Chen, Lin and Stewart2021; Kim, Min, Oh, & Choi, Reference Kim, Min, Oh and Choi2020). However, the associations between depression and the other two gastrointestinal diseases (esophagitis, gastritis, and duodenitis) are novel findings. The robust association between depression and gastrointestinal diseases persisted even after comprehensive adjustment for sociodemographic (age, sex, TDI, education level), lifestyle (smoking, alcohol intake frequency, physical activity), and metabolic (BMI) factors. This pattern suggests depression may influence gastrointestinal pathophysiology through both behavioral and biological pathways. The attenuation of hazard ratios after covariate adjustment highlights the contribution of these factors to the observed associations. Lower educational attainment is potentially partly due to diminishing access to economic, cultural, and social resources (X. Zhang et al., Reference Zhang, Yang, Zhang, Yin, Man and Lu2023), whose reduction is relevant to risk factors for smoking, alcohol intake, and BMI (Marmot et al., Reference Marmot, Friel, Bell, Houweling and Taylor2008; Phelan, Link, & Tehranifar, Reference Phelan, Link and Tehranifar2010). High levels of physical activity have been associated with lower insulin levels (Helmerhorst et al., Reference Helmerhorst, Wijndaele, Brage, Wareham and Ekelund2009), which have been indicated to play a role in the development of many gastrointestinal diseases (Chen et al., Reference Chen, Yuan, Fu, Ruan, Qiao, Wang and Larsson2023b; Rubenstein et al., Reference Rubenstein, Morgenstern, McConell, Scheiman, Schoenfeld, Appelman and Inadomi2013; Watanabe, Hojo, & Nagahara, Reference Watanabe, Hojo and Nagahara2007). High BMI contributes to gastrointestinal diseases by inducing chronic inflammation via elevated pro-inflammatory markers (Park, Park, & Yu, Reference Park, Park and Yu2005), dysregulating insulin/IGF-1 signaling to promote tumorigenesis (Samani, Yakar, LeRoith, & Brodt, Reference Samani, Yakar, LeRoith and Brodt2007), and altering gut microbiota composition through high-fat diets, which amplifies inflammation and metabolic dysfunction (Cani et al., Reference Cani, Amar, Iglesias, Poggi, Knauf, Bastelica and Burcelin2007; Ley, Turnbaugh, Klein, & Gordon, Reference Ley, Turnbaugh, Klein and Gordon2006). These findings highlight that patients with depression, particularly those with lower educational attainment or adverse lifestyle factors, may benefit from targeted gastrointestinal screening. Interventions addressing both psychological distress and modifiable risk factors (e.g. smoking cessation, dietary counseling) could be considered in high-risk groups.
Our MR study has newly identified causal associations between depression and esophagitis, dyspepsia, as well as gastritis and duodenitis, and we corroborated previous MR studies’ findings that depression was associated with an increased risk of gastroesophageal reflux disease, gastric ulcer, and duodenal ulcers (Chen et al., Reference Chen, Zhang, Huang and Jia2023a; Ruan et al., Reference Ruan, Chen, Sun, Zhang, Zhao, Wang and Larsson2023). Several interconnected biological mechanisms may underlie these causal relationships. First, depression-induced autonomic dysfunction disrupts gastric acid regulation, creating a pathological microenvironment conducive to mucosal injury across gastrointestinal segments (S. Zhang et al., Reference Zhang, Xu, Gao, Wu, Li, Liu and Zhang2012). Second, the chronic neuroendocrine activation characteristic of depression leads to sustained cortisol elevation, which alters gut microbial composition and promotes inflammatory cascades (Agirman, Yu, & Hsiao, Reference Agirman, Yu and Hsiao2021). Third, depression-associated gut dysbiosis and metabolite disturbances compromise intestinal barrier integrity, establishing a pro-inflammatory milieu that may manifest differently across anatomical regions, from esophageal inflammation to gastroduodenal ulceration (Ghosh, Whitley, Haribabu, & Jala, Reference Ghosh, Whitley, Haribabu and Jala2021; Kronsten, Tranah, Pariante, & Shawcross, Reference Kronsten, Tranah, Pariante and Shawcross2022).
The ML model developed in this study can serve as a tool for predicting gastrointestinal diseases to evaluate those suspected individuals who may benefit from effective preventive measures. Our study developed and compared multiple ML models – including LightGBM (LGBM), Gradient Boosting Machine (GBM), Random Forest (RF), Logistic Regression (LR), K-Nearest Neighbors (KNN), Multilayer Perceptron (MLP), and XGBoost – for predicting gastrointestinal disease risk in depression. While LR offered interpretability but limited nonlinear modeling, KNN struggled with scalability, and MLP required extensive tuning. Tree-based ensembles (RF, GBM, XGBoost) showed robust performance but were outperformed by LGBM, which combined computational efficiency (via histogram-based learning), automatic missing-data handling, and superior capacity to model complex interactions – advantages that traditional Cox regression lacks. The LGBM-based framework provides a clinically actionable tool for identifying high-risk individuals who may benefit from targeted screening and preventive intervention.
This work has several strengths. First, our study leveraged a population-based prospective design with a large sample size and long-term follow-up. Second, we used independent GWAS sources for depression and gastrointestinal diseases, including the Psychiatric Genomics Consortium and the FinnGen biobank. The robustness of the IVW estimates in this study is supported by multiple MR sensitivity analyses, each of which accommodates different assumptions about genetic pleiotropy (Burgess & Thompson, Reference Burgess and Thompson2017). Finally, the LGBM algorithm outperformed traditional Cox regression in handling large datasets, missing values, and nonlinear interactions. Developed using comprehensive, contemporary European population data (N > 500,000 with long-term follow-up), the model demonstrates enhanced accuracy, adaptability, and clinical validity.
There are some limitations to this study. First, in the section on the Cox proportional hazards model, residual confounding from unknown or unmeasured factors is possible. Second, the constant presence of SNP heterogeneities could potentially bias and affect the robustness of our MR results. Third, although the ML-based predictive model was well-calibrated across different recruitment centers, its effectiveness in clinical practice should be validated in independent prospective cohorts to confirm that its use actually improves patient outcomes. Finally, the majority of the data used in the analysis were collected in European populations from high-income countries. Therefore, generalizations of our findings to other ethnic groups or low- and middle-income countries should be further investigated.
In conclusion, depression acts as a modifiable risk factor for gastrointestinal diseases, underscoring the imperative of upstream psychological prevention in chronic disease control. Embedding universal mental health interventions in primary care systems, especially for socioeconomically disadvantaged populations, may mitigate gastrointestinal morbidity burdens and alleviate long-term healthcare disparities. Such integration serves as a cost-effective blueprint for optimizing population-level health outcomes through holistic prevention frameworks.
Supplementary material
The supplementary material for this article can be found at http://doi.org/10.1017/S003329172610316X.
Acknowledgments
The authors thank all participants and the UK Biobank team for their contribution. We also acknowledge the collaborative efforts of the Psychiatric Genomics Consortium and FinnGen Consortium, and other authors, as well as participants, for providing publicly available GWAS data. This research has been conducted using the UK Biobank Resource under project number 170304.
Author contribution
Conceptualization: B.Z., Y.W., X.Z.; Data Curation: D.L., X.Z.; Funding Acquisition: H.Y., Y.W., X.Z.; Formal Analysis: B.Z., D.Z.; Investigation: D.L.; Methodology: B.Z., D.L., D.Z., Y.Y., H.Y., Y.G., S.L.; Project Administration: Y.W., X.Z.; Resources: D.L.; Software: D.L., Y.Y., H.Y., Y.G., S.L.; Supervision: Y.Y., H.Y., Y.G., Y.W., X.Z.; Validation: D.Z.; Visualization: B.Z.; Writing – Original Draft: B.Z.; Writing – Review and Editing: B.Z., D.L., D.Z., Y.Y., H.Y., Y.G., S.L., Y.W., X.Z.
Funding statement
This work was supported by the National Natural Science Foundation of China (No.72342017, No. 72472155, and No. 71904142).
Competing interests
All authors declare: no other relationships or activities that could appear to have influenced the submitted work.