Hand, foot, and mouth disease (HFMD) is a common childhood illness characterized by fever and vesicular lesions on hands, feet and in the mouth. It is caused by members of the genus Enterovirus, belonging to the Picornaviridae family. Two serotypes from human enterovirus species A (HEV-A), coxsackievirus A16 (CVA16) and enterovirus 71 (EV71), are the main enteroviruses associated with HFMD [Reference Wong1, Reference Hosoya2], although cases involving other types have been reported [Reference Scott and Stone3–Reference Osterback5]. Infections are usually sporadic, but epidemics can also occur. Indeed, this disease is endemic in Asia where large outbreaks of EV71-associated HFMD with severe neurological complications have occurred in recent years [Reference Chang6, Reference Ding7]. Onychomadesis is complete nail shedding at the proximal end; it follows nail matrix arrest and is a rare disorder in children. It has been related to a variety of drug exposures and systemic illnesses, including infections. It is a recognized complication of HFMD, although up to now, there are few studies in the literature associating the two conditions [Reference Clementz and Mancini8, Reference Bernier9]. However, in Spain, two onychomadesis outbreaks with a clinical history of HFMD were reported in 2008 [Reference Salazar10, Reference Redondo, Torres and Izquierdo11].
In April 2009, onychomadesis cases occurred 2 months after clinical HFMD in a nursery school in Arzúa, a small town in La Coruña region (Spain). The aim of this study was to relate both illnesses and to attempt to identify the causative agent of the outbreak.
Clinical samples (stools) were collected from 34 children at the school and from eight staff members. Specimens were sent for analysis to the Enterovirus Laboratory of the National Centre for Microbiology in Madrid. Twelve children developed nail shedding, 11 (92%) of whom exhibited symptoms of HFMD between 36 and 69 days earlier. In addition, six other children had clinical HFMD but no signs of onychomadesis. Samples were collected 5–20 days after nail shedding. The rest of the individuals (16 children, eight adults) had no clinical symptoms. The symptomatic patients' mean age was 1·8 years (range 8 months to 3 years).
Viral RNA was extracted from specimens using a QIAamp Viral RNA kit, according to the manufacturer's instructions (Qiagen, Germany) and 5 μl were analysed for enterovirus detection by a RT-nested PCR in the 5′-non-coding region (5′-NCR) of the viral genome [Reference Casas12]. For enterovirus typing, a species HEV A-, B- and C-specific RT-nested PCR in the 3′-VP1 region, as previously described [Reference Cabrerizo13], was performed.
Enterovirus was detected in 14 clinical samples, from eight (47%) of the 17 HFMD cases (four with onychomadesis) and six (25%) of the 24 asymtomatic individuals (two children, four adults). However, none tested positive by the typing assay, probably because the sensitivity of the RT–PCR in VP1 region is from 10- to 100-fold less than that the 5′-NCR RT–PCR [Reference Cabrerizo13]. To increase the amount of virus, isolation from cell culture was performed with the 14 positive samples. For this, an aliquot of each stool was suspended in 4 ml of Eagle's minimum essential medium with 2% fetal bovine serum, and centrifuged at 15 000 rpm/4°C for 25 min. Then, 0·2 ml of each clarified sample was inoculated in two different cell lines, rhabdomyosarcoma (RD) and human embryo fibroblast (HEF). The culture supernatants were collected on the tenth day, although no cytopathic effect was observed. However, positive results with the RT–PCR assay in VP1 region were obtained in six of the isolates (two in RD, four in HEF), corresponding to three cases (two with HFMD and onychomadesis symptoms and one with HFMD only) and three asymptomatic individuals (two children, one adult). The virus type in all samples was successfully identified by sequencing and BLAST (www.ncbi.nlm.gov/BLAST) analysis as coxsackievirus B1 (CVB1).
In order to determine the relationships between the six CVB1 strains obtained, a phylogenetic analysis was performed, including reference strain, sequences deposited in the GenBank from different countries, and also several Spanish CVB1 sequences characterized by our laboratory in 2008 and 2009. These latter strains were from patients with aseptic meningitis or fever, and were isolated from May to July in Spanish regions other than La Coruña. Multiple sequence alignments were performed by the ClustalW program. Genetic distances were calculated using the Kimura two-parameter model of nucleotide substitution, and statistical significance of phylogenies estimated by bootstrap analysis with 1000 pseudoreplicate datasets. A phylogenetic tree was constructed using the neighbour-joining method in MEGA 3.0 software.
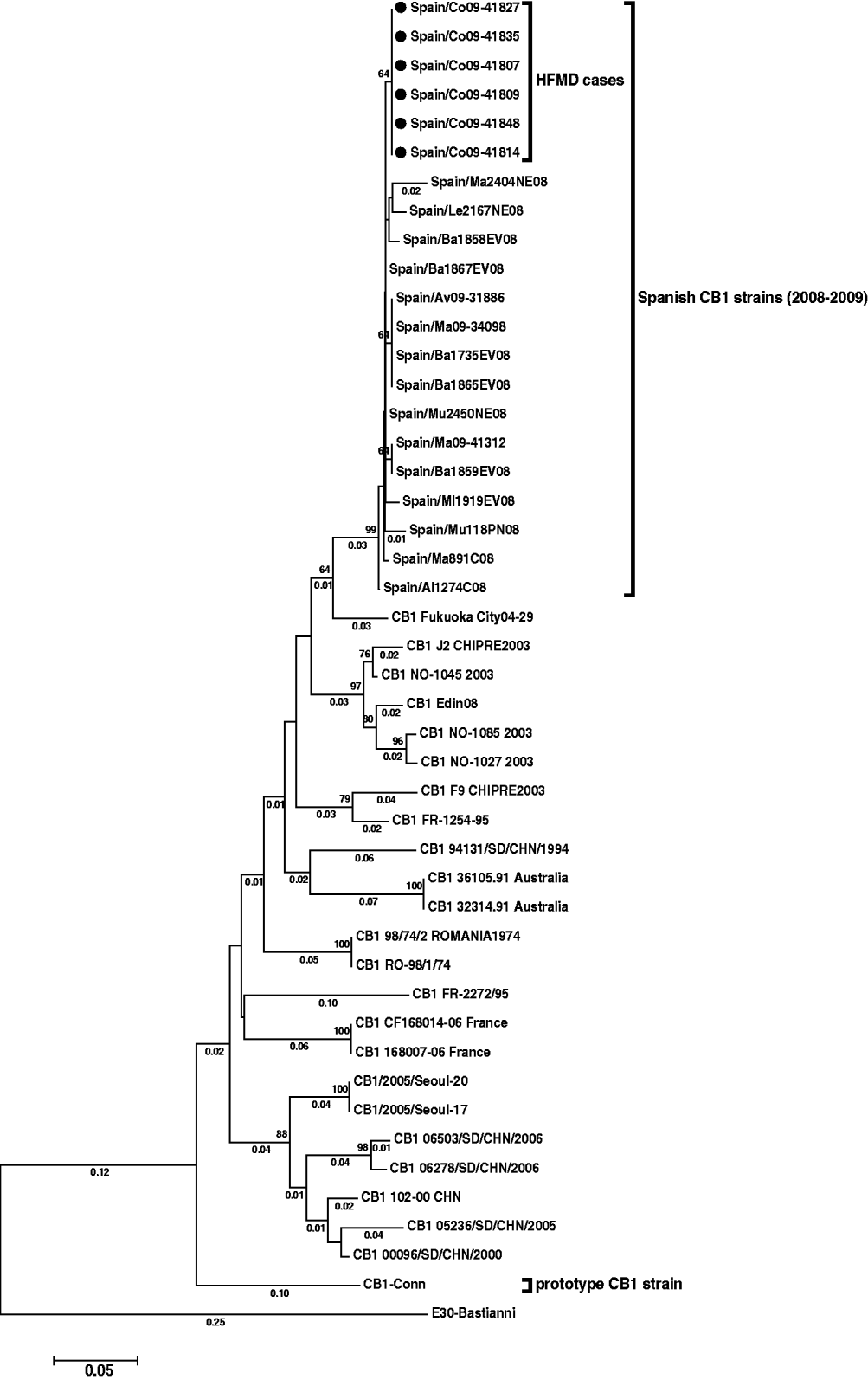
The reconstructed tree (Fig. 1) showed the six CVB1 strains from the outbreak to be closely related (98·3–99·5% identity), and form a subgroup (nucleotide distance <0·01) within a major cluster represented by all Spanish CVB1 strains from 2008 and 2009 (bootstrap value of 99%).
Phylogenetic tree of CVB1 3′-VP1 sequences (390 nucleotides) showing the relationship between the Spanish strains [six detected in this study (black spots) and 15 from other cases], several CVB1 sequences from other countries available in Genbank, and the prototype Conn strain. The tree is rooted with E30 Bastianni strain. Dendograms were constructed by the neighbour-joining method, with 1000 bootstrap pseudoreplicates. Only bootstrap values >64% are shown at nodes. Genetic distances were calculated with a Kimura two-parameter model of evolution (values <0·01 are not shown), and horizontal branch lengths are drawn to scale. The Spanish sequences have been deposited in the GenBank database, under accession numbers HM584455–HM584475.
Onychomadesis, as a complication in the course of enteroviral HFMD, has rarely been reported [Reference Osterback5, Reference Clementz and Mancini8, Reference Bernier9]. However, in 2008 two large Spanish outbreaks were described [Reference Salazar10, Reference Redondo, Torres and Izquierdo11] one of which appeared to be produced by a CA10 strain [Reference Salazar14]; no typing analysis was performed in the other outbreak.
The results of this study suggest that the onychomadesis cases which occurred in a nursery school from a town in the northwest of Spain in April 2009 were a consequence of clinical HFMD, as 92% of the cases with nail shedding had suffered from this viral infection 2 months previously. The clinical and epidemiological characteristics of the patients from this study were similar to those described in the other Spanish outbreaks which occurred in 2008 [Reference Salazar10, Reference Redondo, Torres and Izquierdo11].
Enterovirus was detected in 47% of patients with HFMD. Stool specimens collected during onychomadesis illness are not the most appropriate clinical samples for viral diagnosis of HFMD but, unfortunately, no vesicular samples at the acute phase of the primary illness were available. Moreover, only six of the 14 detected enteroviruses (two from patients with HFMD and onychomadesis and one from a patient with HFMD only) could be further typed as CVB1. Thus, there was insufficient, and only speculative, information to confirm the tentative hypothesis regarding this serotype as the causative agent of the HFMD onychomadesis outbreak described in this study. However, some data might help to support this. First, enterovirus detection 2 months after the initial infection is in accord with a recent report in which the authors detected an enterovirus in a fragment of nail shed 2 months after a primary HFMD infection [Reference Osterback5]. Second, HFMD cases associated with different echovirus or coxsackievirus B serotypes, including CVB1, have been previously reported, although infrequently [Reference Scott and Stone3]. Finally, phylogenetic results showed that the six sequences detected had very close identity and formed a cluster separate from other Spanish CVB1 strains (Fig. 1).
In conclusion, this report strongly suggests that onychomadesis can be a complication during the course of viral infections presenting clinically as HFMD, but further case-control studies with suitable samples (mainly exanthematic specimens as vesicular fluids or shed nail fragments) during the onychomadesis symptoms and/or primary HFMD infection are needed to know what enterovirus serotypes are implicated in this type of infection.
ACKNOWLEDGEMENTS
We thank Isidoro Bustillo, Almudena Otero and Hortensia del Pozo for their technical assistance and Ingrid M. Outschoorn for text editing. This study was funded in part by grant DGEG-1304/08 from the Spanish Ministry of Health.
DECLARATION OF INTEREST
None.


