Scientists from many disciplines around the world have shelved their research plans so they can contribute to efforts to halt the spread of COVID-19 or assist in the medical crises it has created. Materials scientists are no exception—for there are many ways they can contribute. These range from the manufacture of protective clothing and the invention of antiviral surface treatments to developing platforms for COVID-19 diagnostic tests and for the delivery of antiviral drugs. Materials research could potentially supply some of the key enabling technologies that might allow the world to take off the pause button and resume something like economic and social normality.
“There is an increase in R&D on materials and processes related to COVID-19,” said George Stylios, a specialist in smart textiles at Heriot-Watt University in Edinburgh, Scotland. “If these studies are not just ad hoc but can be pulled together in an organized way, we will get good and fast benefits.”
Protective materials
The pandemic caused by the spread of the SARS-CoV-2 coronavirus that causes COVID-19 has left no part of life untouched. Already the virus has claimed more than 570,000 lives worldwide. By early July there were around 13 million confirmed cases of infection, and the real numbers, which depend on the extent of testing and of asymptomatic spreading, are probably considerably higher. The outbreak began after the respiratory syndrome now called COVID-19—caused by a coronavirus that jumped from (almost certainly) bats to humans—was identified in a hospital in Wuhan, China, at the end of December 2019. Most countries followed China's example by implementing lockdown policies in the spring that closed businesses and schools, advised people to stay at home as much as possible, and brought travel and economic activity almost to a standstill. It seems inevitable that the immediate consequences, including further waves of infection and more lockdowns, will persist until the end of 2020; the longer-term impacts will be felt for decades.
The crisis has brought science to the fore. Politicians have depended on the predictions of epidemiological models; virologists are studying the characteristics of the virus, and immunologists are looking at how it attacks the human body; and biochemists and pharmaceutical scientists are scrambling to develop drugs that confer some protection, whether antiviral agents that inhibit the proliferation and harmful effects of the virus or—a longer-term but essential goal—vaccines that will confer immunity in populations.
The pandemic has made it more clear than perhaps any previous infectious outbreak in modern history how much an effective response relies not just on medicines but on protection and testing—and in particular, on the capacity to provide both of these things on a massive scale.
Some of the challenges have been painfully basic: in many countries, hospital staff have lacked the personal protective equipment (PPE), such as face masks and clothing, that can prevent them from being exposed to the virus. With insufficient standing stocks, PPE has had to be manufactured from scratch. Three-dimensional (3D) printing has come into its own, with tales of individuals, even schoolchildren, using home printers to boost supplies of PPE.
To confer reliable protection from virus particles transmitted on exhaled or coughed droplets, masks should ideally conform to the US National Institute for Occupational Safety and Health N95 specification, meaning that they filter out 95% of airborne particles. They also need to make a good seal around the face.
“Unfortunately the low supply against unprecedented demands has lowered the compliance characteristics of PPE,” said Stylios—with the common attitude that “something is better than nothing.” So “we see masks and gowns made out of anything, and goggles and gloves of any type as long as they can cover eyes and hands,” he said.
Most masks simply filter the virus while preserving breathability, said Jared DeCoste of the US Army Combat Capabilities Development Command Chemical Biological Center in Maryland. “This is adequate to protect the wearer from infection via respiratory means, so long as the equipment is properly fit to the individual,” he said. But “every virus is unique in its size, shape, dose needed for infection, and viability on surfaces,” he added. In particular, it is important with such an infectious pathogen to think about the danger posed by viable virus remaining on clothing and equipment when they are put on and taken off. In a case like this, said DeCoste, “we must reevaluate between government, industry, and academia whether we need to pursue new technologies that can be mass-produced.”
The efforts to create new protective materials solutions for the pandemic, said Stylios, include means to detect the virus, filters that last longer and are designed specifically for the SARS-CoV-2 coronavirus or that even actively neutralize it, face masks that are more comfortable and that fit better, and masks and gowns that are better sealed along their seams. To that end, Stylios has developed a reusable, washable face mask using automated, seamless “whole garment” knitting technology (Figure 1). It fits closely around the face and accommodates a filter that can be removed and replaced daily. He is now collaborating with companies in Scotland that produce knitwear and non-woven garments to manufacture the masks at large scale. He is also exploring the use of collagen nanoparticle coatings in filters, which seem able to absorb virus particles efficiently.
A face mask produced at the Centre of Excellence, Scottish Borders by seamless knitting technology. Credit: A. Young and G.K. Stylios.

The coronavirus can survive for a long time on surfaces in laboratory conditions— for more than 72 hours, and perhaps as long as a week, on metals and plastics. This creates a persistent hazard of infection in public spaces such as trains and buses, and on door handles. Antiviral surface treatments could reduce these risks, and might also be useful for making more effective face masks and other PPE.
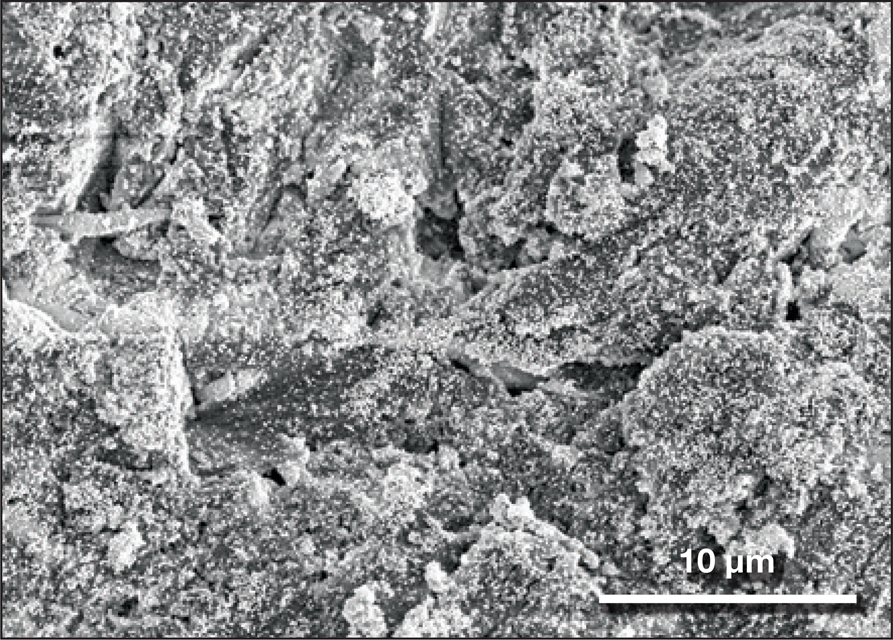
Thomas J. Webster of Northeastern University in Boston is exploring the use of selenium nanoparticles to deactivate viruses if they attach to surfaces. “We have seen that selenium can replace sulfur in the chemical structure of viruses,” he said, “changing their structure so that they are deactivated even if transferred to human skin.” Webster's group has shown previously that surfaces sprayed with such nanoparticles can render influenza type B virus inactive on metal and synthetic surfaces ranging from counters, doorknobs, and bed railings to money and iPhone screens (Figure 2). The selenium treatment “is very versatile, and can be coated on any surface we have tried so far,” said Webster. “We don't see any signs of toxicity or rubbing off.” But his group has not yet tested the approach for SARS-CoV-2.
Aluminum coated with antiviral selenium nanoparticles, a possible solution to deactivate viruses on surfaces. Credit: Thomas J. Webster, Northeastern University.

In an emergency situation like this, said DeCoste, there are inevitably tradeoffs to be faced. While it is good to look for useful new materials, it is not just about efficacy but also cost and availability. “When in a rapid-response crisis, you obviously evaluate that tradeoff a little differently than when in standard operation,” he said. “When a demand surge as large as this pandemic hits, there will be large strains put on every aspect of the supply chains.”
Even though many countries have now exited from lockdown after the first wave of infection, restarting economies may depend crucially on the ability to test people: to see if they have the virus, or if they have immunity, so as to establish quickly who can safely return to or stay at work.
Tests for determining if an individual has been infected by the coronavirus were developed with extraordinary speed once it has been identified and its genome sequenced. Being able to determine quickly if a person was infected was crucial to the effective response to the outbreak in South Korea and Taiwan—that way, the individual's recent contacts could also be traced and tested.
Testing and detection of COVID-19 symptoms will remain essential until a vaccine is available, and could be highly disruptive to daily life—but materials science might help to make it more convenient, enabling continuous rather than sporadic monitoring. Stylios is working on wearable electronics that will monitor the body for symptoms such as changes in respiration, heartbeat, and temperature. “The idea is that if the wearer goes out of the physiological ranges of normal breathing or temperature,” he said, “it triggers a warning” so the wearer can seek more definitive testing. One of his prototype products is an elastic chest band bearing a 9-gram box that contains three electrodes for measuring electrocardiograms (heartbeat), along with a temperature sensor. It can communicate these data via Bluetooth. Stylios has also developed a sports vest with unobtrusive textile-based sensors on the front and back. It is possible that smart garments like this might become standard items of clothing for continuous monitoring of our health in the wake of the pandemic.
Nanoscience for new therapies
Can materials help with medicines against COVID-19? One of the most popular strategies for therapies, said nanomedicine specialist Catherine Fromen of the University of Delaware, is to find agents that bind to the virus and disrupt its life cycle, for example by blocking its ability to enter and infect cells. At present, some of the antivirals being tested are pharmaceutical agents developed for treating other viruses. Remdesivir, for example, a drug created to treat Ebola (without success), has already shown promise in clinical tests; dexamethasone, a known anti-inflammatory agent, seems to reduce deaths among people seriously ill from the disease and needing ventilation. Such “repurposed” drugs have often cleared some stages of clinical trials (such as safety studies), giving them a head start for rapid deployment.
A relatively new approach to antiviral therapies uses nanoparticles as carriers for the active agent, which might be modified to target infected cells or the virus itself. “We are exploring the use of a wide range of nanoparticles,” said Webster, “such as gold, silver, iron oxide, and selenium, as well as self-assembled peptide nanomaterials, that can attach to the virus and stop it attaching to cell membranes, entering the cell, and replicating.” Simply getting a nanoparticle to attach to a virus can block its entry to a cell and so prevent replication, he said. Once nanoparticles have stuck to a virus and deactivated it, they would remain attached until the virus is excreted. Webster is now developing his nanoparticle approach with Audax Medical Inc. of Concord, Mass. They currently have an injectable product, but are working on one that can be inhaled so that it can reach the lungs directly.
Among the advantages of nanomaterials, he said, are that they are of the same size as the coronavirus, and can be functionalized with one or more regions to attach to specific parts of the virus. Multiple binding sites could be important for preventing the virus from mutating into more resistant forms. (Most existing strategies for blocking the virus target just one region, typically the “spike” protein that docks onto the cell membrane; Figure 3.) “We also believe we would have a faster approval process,” said Webster, “since some of the nanomaterials have already been approved for other medical applications.”
The structure of the coronavirus. The spike protein protruding from the lipid coat, which provides the point of attachment and entry to cells, is the target for many antiviral therapies. Credit: Protein structures adapted from D. Wrapp et al., Science 367 (2020), p.1260.

Viruses might also be destroyed with nanoparticles of materials such as gold and iron oxide that can absorb radiation (such as tissue-penetrating near-infrared) and heat up. The particles can act as both diagnostic—iron, for example, can be sensed by magnetic resonance imaging—and therapeutic agents. This “photothermal” approach depends on ensuring that the particles hit the right target: Webster said that such particles have already been developed for treating cancers, where they target and destroy only the tumor cells. His team has developed such systems for treating the influenza B virus, and although they do not yet have results for the coronavirus, he said “we are confident that a photothermal approach would work.”
Fromen is also developing functional nanoparticles as therapeutic agents, in collaboration with biomedical engineer Jason Gleghorn at the University of Delaware. Drawing on her past experience with nanoparticle carriers for vaccines and injectable nanoparticles for treating acute lung injury, Fromen is aiming to produce biodegradable, inhalable particles that can sequester the virus in the nose or upper airways, preventing the infection from moving into the deeper airspaces (Figure 4). “In our observations, most current vaccine and therapeutic approaches are not leveraging any inhalation therapies, despite this being the main site of viral infection,” she said.
Biodegradable poly(ethylene glycol) diacrylate-based hydrogel nanoparticles, which are potential vehicles for drug delivery and immune engineering against COVID-19. Credit: Catherine Fromen, Fromen Research Group, University of Delaware.

One potential approach would take advantage of the fact that the coronavirus enters cells in the airways and lungs by binding to the membrane protein called ACE2, a blood-pressure modulator. Nanoparticles studded with ACE2—or perhaps just with small peptide sequences from it—might thus be expected to mop up the virus. Such virus-attached nanoparticles would then need to be cleared from the airspace, which might happen much as it does for other inhaled particulates: by entrapment in the layer of mucus lining the membranes, which is swept along into the digestive tract. The lungs also contain specialized immune cells—aveolar macrophages—that will break down foreign particles (Figure 5).
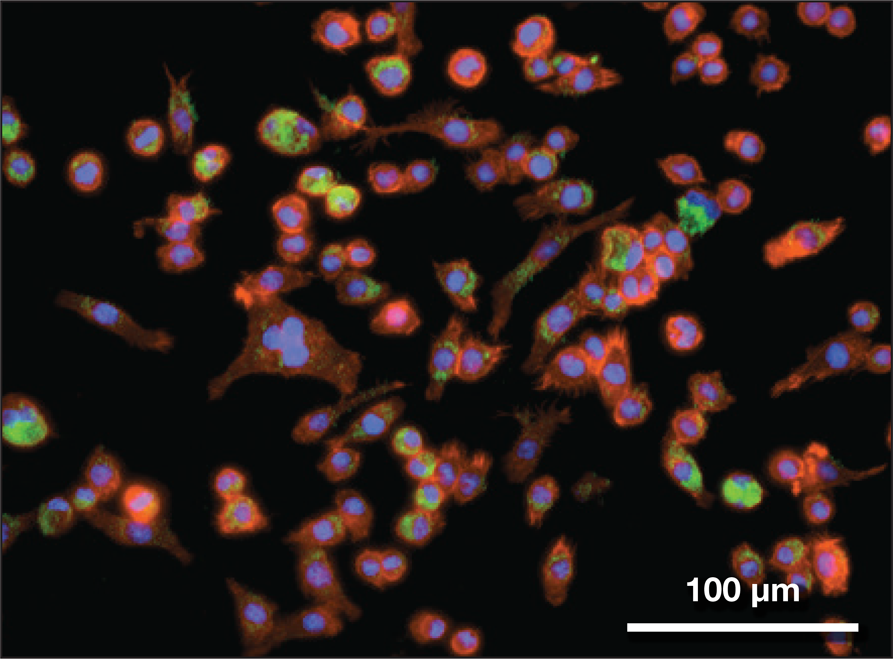
Inhaled nanoparticles (green) taken up by immune cells (aveolar macrophages) like those that are present in the lung. Credit: Bader Jarai and Catherine Fromen, Fromen Research Group, University of Delaware.
A second strategy for ameliorating COVID-19, Fromen said, is not to target the virus itself but to intervene in the human immune response that causes the potentially fatal respiratory problems. “Part of the challenge with COVID-19 is that this virus promotes a delayed and improper immune response,” she said. “The lag allows the virus to replicate with less host interruption. At some point, the body recognizes things are out of control and rushes to catch up, which leads to intensive production of cell-signaling molecules called cytokines. These signals lead to recruitment of more inflammatory cells, which can cause tissue damage and generate more cytokines, leading to an uncontrolled runaway response.” This is called a cytokine storm, and it precedes Acute Respiratory Distress Syndrome (ARDS) and, sometimes, death. “Your own immune system's response to the virus is essentially what leads to death, not the virus itself,” Fromen said.
Although antivirals might help to stop matters reaching this point, she said, often by the time people have come into the hospital it is because of symptoms caused by the immune response. It is too late then to slow down the virus; what is needed instead is a way to stop the immune system from getting out of control. Currently that might involve agents that block inflammatory cytokines, such as dexamethasone or the so-called monoclonal antibody agent Tocilizumab.
Fromen's research group has looked at how nanoparticle therapies might do that job instead. In one study in mice with acute lung injury, her group showed that micro- and nanoparticles functionalized with carboxylate groups and directly injected into the bloodstream could act as a distraction for the white blood cells called neutrophils (reported in ACS Nano, doi:10.1021/acsnano.7b03190). These are the “first responders” to acute inflammation and are therefore the key cell type involved in ARDS. When the particles were injected, the neutrophils, rather than migrating to the sites of lung inflammation, became diverted into mopping them up and carrying them to the liver for disposal.
Jacob Brenner at the University of Pennsylvania has also explored this approach, screening a range of protein-based nanoparticles to find ones that specifically elicit the attention of neutrophils in the lung (posted in bioRxiv, doi:10.1101/2020.04.15.037564). His research group found that particles made from certain proteins, such as polymer-cross-linked and denatured (amorphous) albumin or liposomes with tangled proteins on their surface, will show this effect, the specificity seeming to come more from the nature of the supramolecular organization of the proteins than from their chemical composition. By labeling the nanoparticles with an indium radioisotope that acts as an imaging agent via gamma-ray emission, they could diagnose acute lung injury this way. And some of the nanoparticles could lower the ability of neutrophils to stick to the blood vessels of inflamed lung tissue and cause further swelling.
Fromen and her colleagues are also looking at inhalable particulates that can adsorb fluid or cytokines in the lung and prevent the cytokine storm. By tuning the properties of particles, such as size and charge, she said it might be possible to avoid their clearance via the mucus layer, and instead allow the particles to diffuse through this layer and enter the bloodstream, carrying drug molecules (such as anti-inflammatory agents) with them. She thinks that vaccines might be more effectively delivered this way too. “Delivery of the vaccine directly to the lung allows for distribution and activation of lung-specific immune cells, which leads to better protection of the lung,” she said. Webster said that the peptide nanoparticles his group has developed also show signs of reducing lung inflammation in animal tests, and so seem able to ameliorate the cytokine storm too.
These nanoparticle approaches present many materials-related challenges, Fromen said, such as precise functionalization of the particles and the tailoring of aerodynamic properties. “The materials design components are critical in which cells are targeted and what information is delivered to the immune cells,” she said. The strategy of using inhalable particles as modulators of lung immune cells is relatively new and so cannot draw on existing methods for preclinical testing. But she said that “there are considerable benefits to direct inhalable lung delivery, especially that it would enable fighting the virus directly at the site of infection,” as well as engaging the unique immune cells that reside in the lung itself.
Cells on chips
Developing antiviral therapeutic agents requires techniques for rapidly screening candidate drugs, as well as learning more about the mechanisms of viral infection. A project involving researchers at Cornell and Stanford Universities and the University of Cambridge, and funded by the US Defense Advanced Research Projects Agency is seeking to develop an electronic biosensor that might assist both of those efforts by detecting the presence of the coronavirus with high sensitivity.
The device incorporates a small piece of a cell membrane onto an electronic chip. The Cornell team, led by biomolecular engineer Susan Daniel, has figured out how to detach a portion of the lipid membrane (a vesicle) from human cells—containing all the native membrane proteins—and fuse it with an array of electrodes and transistors produced by Alberto Salleo's team at Stanford and made from conducting polymers, which can measure the conductivity across the membrane. The researchers at Cambridge are then studying how this conductivity changes when outside particles such as viruses stick to and perhaps disrupt the membrane (Figure 6).
(a) Organic electronic devices for detecting viruses as they attach to fragments of cell membrane on a chip. (b) When a virus attaches, its lipid coat merges with the membrane and causes a change in ion-channel conductivity, which can be monitored by the conducting-polymer circuit. (c) The on-chip membranes themselves begin as “blebs”—vesicle-like membrane compartments—that detach from living cells with their complement of membrane proteins included. Credits: Device photo by A. Salleo team at Stanford; schematic by S. Daniel team at Cornell; blebbing by R. Owens team at Cambridge/BioRender software.

“A major advantage of our system is that because the vesicle was produced from cells, it is like having a biopsy of the cell,” said Róisín Owens, a biochemist on the Cambridge team. “We have all the material that would be present including proteins and lipids, but none of the inconvenience of using live cells.” The collaboration has shown that the membrane on a chip preserves all the molecular orientation and functionality of the original cell membrane of mammalian kidney cells (Langmuir, doi:10.1021/acs.langmuir.0c00804), and that devices like this can detect the action of chemical agents known to alter ion-channel conductivity (ACS Nano, doi:10.1021/acsnano.0c01330)—something that would previously have required conductivity measurements on individual cells.
The collaboration has studied this approach before for other pathogens, such as the influenza virus. “When the global pandemic struck, we quickly thought how we could shift our focus to looking at SARS-CoV-2,” Owens said. Because of the hazards of the coronavirus, however, the researchers use disabled virus particles that have the same shell (including the proteins responsible for cell adhesion) but none of the nucleic acid needed for replication. The researchers hope soon to be able to detect agents that disrupt the lipid coat of the coronavirus, or that bind to the spike protein.
Finding the right materials for the device is critical, Owens said. They are using a conducting polymer—a mixture of thiophene and sulfonated polystyrene derivatives, denoted PEDOT:PSS—for the electrodes and transistors. “Up until now, researchers have created membranes on hard, rigid materials such as glass or silicon,” said Owens. “But the membrane functionality [then] suffers because of a stiffness mismatch.” The soft polymers can cushion the downward-facing portions of the membrane proteins while allowing them enough mobility to accomplish their function. An added advantage, said Owens, is that the devices can be transparent, allowing for optical imaging.
Given the previous successful demonstration of virus fusion with influenza, she added, “we are poised to have a preliminary device within a couple of months after starting the research.”
Lab life in lockdown
Despite all this encouraging activity, scientists worldwide have been struggling with the fact that, at the very moment their work is most urgently needed for the COVID-19 crisis, their ability to conduct it is most compromised. Not only have many research establishments been shut down, but supply chains for reagents are disrupted. Even laboratories that had dispensation to continue operating in lockdown have been beset with problems due to the absence of administrative staff.
“I think many new technologies and ideas right now from universities are being kept from realization since they cannot do the fundamental studies needed to attract industry's interest,” Webster said. Fortunately, that problem is not bedeviling his own efforts, since his team had already established a partnership with a startup company that can culture the coronavirus and test the materials.
But the constraints have not had entirely negative effects. “The lockdown allowed us time to really look at our data and try to understand them better,” said Owens, who has been working from home since the UK lockdown began. “We have done some really nice modelling of our data to get even more out of it. And we have a killer list of experiments drawn up, where we have honed all of the necessary controls and so on, because we have had extra time to think about it.” She hopes that once laboratory work can resume, they can hit the ground running.
There are plenty of lessons to be learned from this crisis—and an urgent need to learn them. “We really need to make sure that we are better prepared for the next time something similar occurs,” said DeCoste. “There will surely be new research in the area of virucidal materials and coatings in the near future, but it is going to be very important to do that research keeping in mind how these advances can either be integrated into existing technologies, or how they can be mass-produced.” He thinks that modular, distributed production platforms such as additive manufacturing (e.g., 3D printing) will be valuable in this regard.
Addressing the pandemic, then, is not just about finding new materials solutions, but finding ways to ensure that they can be produced in volume and at speed. Because society will surely need to do so again.