Introduction
Over the past decades, the world has seen an increasing frequency in the introduction of invasive alien species (IAS) that significantly impacted plant health and agricultural productivity (e.g. Faulkner et al., Reference Faulkner, Robertson, Rouget and Wilson2016; Mastrangelo et al., Reference Mastrangelo, Paulo, Bergamo, Morais, Silva, Bezerra-Silva and Azeredo-Espin2014; Pozebon et al., Reference Pozebon, Marques, Padilha, O´ Neal, Valmorbida, Bevilaqua and Arnemann2020; Sandvik et al., Reference Sandvik, Olsen, Töpper and Hilmo2022; Turbelin et al., Reference Turbelin, Diagne, Hudgins, Moodley, Kourantidou, Novoa and Courchamp2022). Examples of these included the silverleaf whitefly Bemisia tabaci species complex (De Barro et al., Reference De Barro, Liu, Boykin and Dinsdale2011), Thrips tabaci (Balan et al., Reference Balan, Ramasamy, Hande, Gawande and Krishna Kumar2018), Frankliniella occidentalis (Reitz et al., Reference Reitz, Gao, Kirk, Hoddle, Leiss and Funderburk2020), Helicoverpa armigera (Czepak et al., Reference Czepak, Albernaz, Vivan, Guimarães and Carvalhais2013; Jones et al., Reference Jones, Parry, Tay, Reynolds and Chapman2019; Tay et al., Reference Tay, Soria, Walsh, Thomazoni, Silvie, Behere and Downes2013), Spodoptera frugiperda (Goergen et al., Reference Goergen, Kumar, Sankung, Togola and Tamò2016; Tay et al., Reference Tay, Court, Hoffmann and Polaszek2022a), and the tomato leafminer Phthorimaea absoluta (synonym Tuta absoluta; Chang and Metz, Reference Chang and Metz2021), highlighting the need to better understand, through genomics, mechanisms associated with the introduction and adaptation of these transboundary plant pests to improve global agricultural biosecurity preparedness and developing appropriate management solutions.
The dispersal of propagules from geographically distant locations is a major factor underpinning success of IAS introductions, enabling establishment potentials through higher levels of genetic diversity as compared with a single founder introduction event. These dispersal mechanisms and pathways, in turn, may manifest quantitatively (i.e. the number of individuals introduced to a location where it is not native) and qualitatively (i.e. different parental lineages), many of which are intrinsically linked to human-mediated dispersal routes, such as commercial pathways. The determination of genetic variability associated with species introduction is strongly influenced by the intensity and diversity of source populations. Considering these factors, variations in insect behaviour can result in distinct patterns of genetic diversity (Wilson et al., Reference Wilson, Dormontt, Prentis, Lowe and Richardson2009), although the choice of genetic markers used will also impact on detection of genetic patterns. Consequently, recently introduced invasive species such as H. armigera in Brazil (Czepak et al., Reference Czepak, Albernaz, Vivan, Guimarães and Carvalhais2013; Tay et al., Reference Tay, Soria, Walsh, Thomazoni, Silvie, Behere and Downes2013) where females randomly lay individual eggs, have been shown to exhibit higher genetic diversity in Brazil and its neighbouring countries (Anderson et al., Reference Anderson, Tay, McGaughran, Gordon and Walsh2016; Arnemann et al., Reference Arnemann, Tay, Walsh, Brier, Gordon, Hickmann, Ugalde and Guedes2016, Reference Arnemann, Roxburgh, Walsh, Guedes, Gordon, Smagghe and Tay2019; Tay et al., Reference Tay, Soria, Walsh, Thomazoni, Silvie, Behere and Downes2013) as compared to those that follow natural dispersal pathways, whose expansion is induced by climatic factors (Battisti et al., Reference Battisti, Larsson and Roques2017; Robinet et al., Reference Robinet, Laparie and Rousselet2015).
Various AIS (e.g. P. absoluta, Czepak et al., Reference Czepak, Nunes, Oliveira, Silva, PBSO, Roy, Magalhaes and Marques Filho2018; S. frugiperda, Tay et al., Reference Tay, Meagher, Czepak and Groot2023) have been of particular global concern due to their economic impact (Adelino et al., Reference Adelino, Heringer, Diagne, Courchamp, Faria and Zenni2021; IPBES, 2023) and their increasing introduction frequencies (Mastrangelo et al., Reference Mastrangelo, Paulo, Bergamo, Morais, Silva, Bezerra-Silva and Azeredo-Espin2014; Pozebon et al., Reference Pozebon, Ugalde, Smagghe, Tay, Karut, Copa Bazán, Vitorio, Peralta, Saluso, Ramírez-Paredes, Murúa, Guedes and Arnemann2022). The economic cost of these invasive species, including P. absoluta, that has global footprints, remains poorly understood. For example, the combined economic losses due to P. abosulta for small-scale farmers in four African countries (Ethiopia, Kenya, Tanzania, and Uganda), range between US$70 and US$80 million annually, and contributed to a total of US$1.1 billion when considering also other major invasive species, representing approximately 2.2% of the agricultural Gross Domestic Product (GDP) of the region (Pratt et al., Reference Pratt, Constantine and Murphy2017).
The larvae of P. absoluta are polyphagous pests of solanaceous plants (e.g. egg plants Solanum melongena; potato S. tuberosum) (Smith et al., Reference Smith, Dubois, Mallogo, Njau, Tua and Srinivasan2018; EPPO, 2010), with S. lycopersicum (tomato) being its main host crop. The widespread detection of P. absoluta therefore raises significant concerns on tomato production (Hogea, Reference Hogea2020). Management of P. absoluta is challenging due to reported insecticide resistance (Haddi et al., Reference Haddi, Berger, Bielza, Rapisarda, Williamson, Moores and Bass2017; Silva et al., Reference Silva, Silva, Silva, Campos, Filho and de Siqueira2021), and to its behaviour and high reproductive capacity, which involves overlapping generations and high-field infestation rates, facilitated by its flight dispersal ability, and the propensity to be accidentally introduced via anthropogenic-related activities, causing tomato productivity losses that can reach up to 100% (Biondi et al., Reference Biondi, Guedes, Wan and Desneux2018). Phthorimaea absoluta females lay individual eggs randomly, preferentially on leaves, new shoots, and sepals (less on fruits). Adults can mate up to four times with implications to establishment success of bridgehead invasive populations (Guillemaud et al., Reference Guillemaud, Ciosi, Lombaert and Estoup2011). Multiply-mated female bridgehead founders undertaking range expansion have the potential to increase propagule genetic diversity even if no males were yet present in the new invasive range.
Native to South America and representing one of the main threats to tomato crops (Guedes and Picanço, Reference Guedes and Picanço2012), P. absoluta was identified in 1917 and recorded as widely distributed in Peru (Rojas, Reference Rojas1981; USDA-APHIS, 2023). In 1980, it was reported in southern Brazil (Muszinski et al., Reference Muszinski, Lavendowski and Maschio1982) and brought with it concerns that included higher insecticide application frequencies with associated control costs, as well as issues relating to environmental contaminations with human health implications (Kim et al., Reference Kim, Kabir and Jahan2017; Tudi et al., Reference Tudi, Li, Li, Wang, Lyu, Yang and Connell2022). In 2006, P. absoluta was detected in Spain (Urbaneja et al., Reference Urbaneja, Vercher, Navarro-Llopis, Porcuna Coto and García-Marí2007), followed by reports from other European countries (e.g. France, Italy, Switzerland, Malta, Portugal, Greece, the Netherlands), and the United Kingdom (Potting et al., Reference Potting, Van Der Gaag, Loomans, Van Der Straten, Anderson and Macleod2009). First reports of P. absoluta in northern Africa (e.g., Morocco, Algeria, Tunisia, Libya, Egypt) were between 2007 and 2008 (Mansour et al., Reference Mansour, Brévault, Chailleux, Cherif, Grissa-Lebdi, Haddi and Biondi2018), and in East and South African countries (e.g., Kenya, Malawi, South Africa) between 2014 and 2016 (e.g. IPPC, 2016; Visser et al., Reference Visser, Uys, Nieuwenhuis and Pieterse2017). In Asia, it was detected in Turkey in 2009 (Kilic, Reference Kilic2010) with an assumed rapid dissemination to reach central and southwestern parts of the continent (Biondi et al., Reference Biondi, Guedes, Wan and Desneux2018). P. absoluta was reported in China (e.g. Yili, Xinjinag, Yunnan) from 2017 onward (Li et al., Reference Li, Fu, Guo, Wang and Lu2022; Zhang et al., Reference Zhang, Wang, Gao, Liu, Zhang, Fu and Wan2020), and confirmed in South Korea in 2024 (Lee et al., Reference Lee, Jeong, Lee and Paik2024).
Response to challenges of rapid and widespread invasive agricultural pest population establishments often involves insecticide applications that can be both excessive, inappropriate, and inaccurate (Desneux et al., Reference Desneux, Luna, Guillemaud and Urbaneja2011; Guenaoui et al., Reference Guenaoui, Dahliz, Bensaad and Ouezzani2013; Palumbo et al., Reference Palumbo, Perring, Millar and Reed2016; Potting, Reference Potting2013; Wan and Yang, Reference Wan and Yang2016), thereby contributing to the evolution of insecticide resistance and secondary pest outbreaks (Aynalem, Reference Aynalem2022; Biondi et al., Reference Biondi, Guedes, Wan and Desneux2018; Khan et al., Reference Khan, Uddin, Rizwan, Khan, Farooq, Shah and Ali2020). The spread and successful population establishment of invasive pests may also be facilitated by the presence of resistance alleles that contributes to reduced insecticide effectiveness, as already reported in P. absoluta in South America (Silva et al., Reference Silva, Assis, Ribeiro and Siqueira2016a, Reference Silva, Silva, Campos, Silva, Ribeiro and Siqueira2016b, Reference Silva, Berger, Bass, Williamson, Moura, Ribeiro and Siqueira2016c), Europe (Roditakis et al., Reference Roditakis, Vasakis, Grispou, Stavrakaki, Nauen, Gravouil and Bassi2015), Asia (Zhu et al., Reference Zhu, Chen, Liu, Lin, Liang, Nauen, Li and Gao2024; Zibaee et al., Reference Zibaee, Mahmood, Esmaeily, Bandani and Kristensen2018), and Africa (Bala et al., Reference Bala, Mukhtar, Saka, Abdullahi and Ibrahim2019; Ong’onge et al., Reference Ong’onge, Ajene, Runo, Sokame and Khamis2023). Therefore, identifying resistant genotypes underpinning resistance phenotypes in P. absoluta populations will be crucial for understanding its propagation processes and for the designing of management strategies.
Low levels of nucleotide variation in the partial mitochondrial DNA cytochrome oxidase subunit I (mtCOI/cox1) gene were previously reported in native (e.g. South America; Cifuentes et al., Reference Cifuentes, Chynoweth and Bielza2011) and invasive P. absoluta populations from the Mediterranean Basin (Cifuentes et al., Reference Cifuentes, Chynoweth and Bielza2011), India, and Nepal (Shashank et al., Reference Shashank, Twinkle, Chandrashekar, Meshram, Suroshe and Bajracharya2018). Similar findings were reported also in other significant transboundary pest species such as S. frugiperda (Cock et al., Reference Cock, Beseh, Buddie, Cafá and Crozier2017; Goergen et al., Reference Goergen, Kumar, Sankung, Togola and Tamò2016) and Oryctes rhinoceros (Tay et al., Reference Tay, Marshall, Popa-Baez, Dulla, Blas, Sambiran and Hoffmann2024), emphasizing the general similar nucleotide characteristics for the mtCOI barcoding partial gene region in insects/arthropods.
An over reliance of the partial mtCOI gene region to infer population founding histories is known to result in conflicting population genetic signatures, especially when compared with full mitochondrial genome-derived population signatures (e.g. Elfekih et al., Reference Elfekih, Etter, Tay, Fumagalli, Gordon, Johnson and De Barro2018; Tay et al., Reference Tay, Rane and Padovan2022c, Reference Tay, Marshall, Popa-Baez, Dulla, Blas, Sambiran and Hoffmann2024). The circular arthropod mitochondrial genomes are typically high in A–T compositions (e.g. Pozebon et al., Reference Pozebon, Ugalde, Gansemans, Smagghe, Van Nieuwerburgh, Stürmer, Tay WT and Arnemann2023; De Souza et al., Reference De Souza, Tay, Czepak, Elfekih and Walsh2016; Kunz et al., Reference Kunz, Tay, Court, Elfekih, Gordon, Evans and De Barro2019a), and due to its overall non-recombinant nature (i.e. maternally inherited; Crozier, Reference Crozier1990) and relatively small size (i.e. typically ranged between 15kb and 28kp; Behere et al., Reference Behere, Tay, Firake, Kunz, Burange and Ramamurthy2019; Piper et al., Reference Piper, Van Helden, Court and Tay2016; Tay et al., Reference Tay, Rane, James, Gordon, Downes, Kim and Walsh2022b; Morgan et al., Reference Morgan, Wang, Chen, Moctezuma, Burgos, Le and Huang2022), enabled ease of genome assembly that also afforded significant advantages for more comprehensive inference of population establishment history. Kunz et al. (Reference Kunz, Tay, Elfekih, Gordon and De Barro2019b) and Vyskocilová et al. (Reference Vyskocilová, Tay, van Brunschot, Seal and Colvin2018) also showed that at both inter- and intra-specific levels, amino acid conservation enabled detections of pseudogenes and therefore elimination of poorly annotated mtCOI barcode gene to significantly improve on species identification, while quality control of assembled mitogenomes enabled assembly errors to be identified through comparative genome analyses (e.g. see Walsh et al., Reference Walsh, Perera, Anderson, Gordon, Czepak, McGaughran and Tay2019).
Based on simple sequence repeat markers (SSR) analysis, Guillemaud et al. (Reference Guillemaud, Blin, Le Goff, Desneux, Reyes, Tabone and Lombaert2015) concluded that the invasive Mediterranean P. absoluta populations likely originated from central Chile, while the shared mtCOI genetic similarity linked the species’ invasion patterns from southern and central Europe to the Indian sub-continent and to northern and sub-Saharan Africa (Cifuentes et al., Reference Cifuentes, Chynoweth and Bielza2011; Fiaboe et al., Reference Fiaboe, Agboka, Agboyi, Koffi, Ofoe, Kpadonou, Agnamba, Assogba, Adjevi, Zanou and Fening2021; see also review by Santana et al., Reference Santana, Kumar, Da and Picanço2019). To-date, very few P. absoluta mitogenome resources are available, including one from Spain (Zhang et al., Reference Zhang, Yang and Zhang2019), two from China (i.e. one each from Xinjiang and Yunnan; Li et al., Reference Li, Fu, Guo, Wang and Lu2022), and one from Kenya (Ajene et al., Reference Ajene, Heya and Khamis2024). Li et al. (Reference Li, Fu, Guo, Wang and Lu2022) concluded that multiple mtDNA markers would be needed to increase population genetic diversity estimate accuracy of P. absoluta. This is especially apposite to better understand and assess the scenarios on the African continent that accounts for approximately 18% of the world’s human population, is highly vulnerable to climate change, and where severe food insecurity may result due to invasive pests.
Here, we report three new mitochondrial genome resources for P. absoluta from Malawi and South Africa, and demonstrate the benefit of utilising a whole-genome sequencing (WGS) approach to concurrently characterise known resistance genes and alleles, using the organophosphate (OP) and pyrethroid resistance/DDT resistance genes, i.e. Acetylcholinesterase (AChE-1) and Para-type voltage-gated sodium channel (VGSC), respectively, as examples. We further compare P. abosuta mitogenome gene variability of Spain, Kenya, and China, for re-assessments of mitogenome assembly quality. We also review P. absoluta published partial mtCOI gene sequences to assess the marker’s suitability for understanding both P. absoluta and general arthropods’ invasion biology.
Material and methods
Samples for WGS
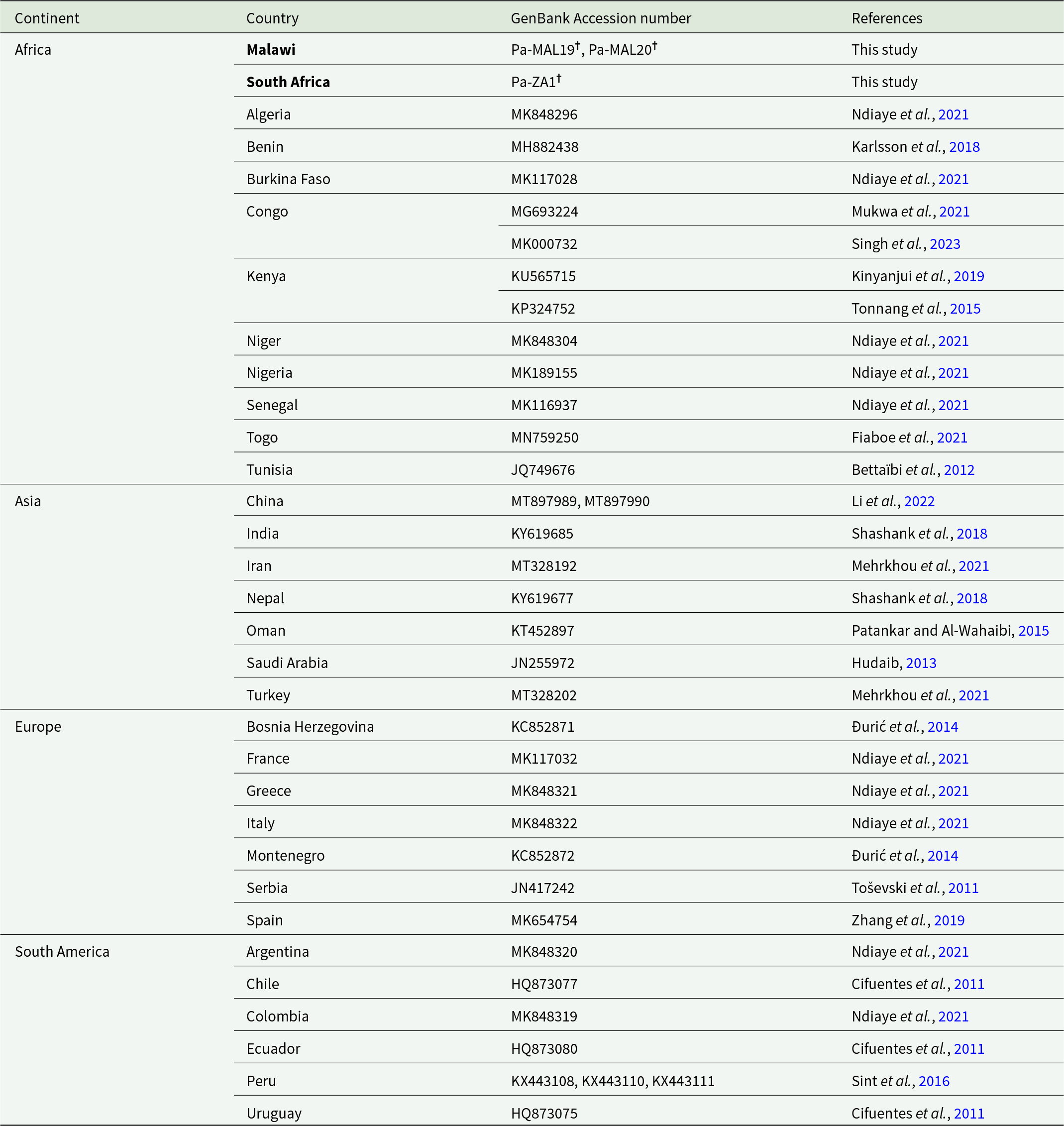
Two P. absoluta individuals were collected from Malawi in East Africa (sample code: MAL19, MAL20) and one individual from South Africa (sample code: ZA1) (Table 1). The Malawian P. absoluta individuals were collected from tomato plant hosts from Thyolo District (−16.054, 35.062) in Malawi (elevation 1,045 m) on 12 June 2018. The South African P. absoluta individual was from tomato near Baltimore (Limpopo province, South Africa) (−23.217337, 28.304087). Samples were collected as early instar larvae and were stored in 100% ethanol until used in DNA extraction. DNA of each individual was extracted using the QIAGEN DNeasy Blood & Tissue DNA extraction Kit (Dusseldorf, Germany) following manufacturer’s recommended protocol. Extracted genomic DNA was quantified using the Qubit 2.0 Fluorometer (Life Technologies). Illumina DNA libraries were prepared as per the manufacturer’s protocols at the Biomolecular Resource Facility at the John Curtin School of Medical Research, Australian National University, Canberra, Australia. WGS of these libraries was performed on an Illumina NextSeq 2000 sequencer, P1 flowcell, (2 x 300bp, PE).
Samples of Phthorimaea absoluta analysed in this study for their partial mtCOI gene. The draft mitogenomes of the African samples (indicated by ‘†’) can be downloaded from CSIRO’s data access portal (Magalhaes et al., Reference Magalhaes, Czepak, van Niekerk, du Plessis, Court and Tay2024), the partial gene region used in this analysis ranged between nucleotide positions 1,572-2,142 of the MAL19 mitogenome (i.e., 570 bp) (see also Figure 1)
Mitogenome assembly and annotation
Mitogenome assembly was performed in Geneious Prime (v11.1.5 from Biomatters Ltd, Auckland, NZ), using the Geneious Mapper program with default settings (Algorithm: Auto; Scoring matrix: 200PAM/K = 2; Gap open penalty: 1.53; Offset value: 0.123). Initial assembly of a complete mitochondrial genome was undertaken for sample MAL19, commencing by mapping the MAL19 whole-genome sequence data against a reference partial mtCOI gene (GenBank MT897989). We build the full mitogenome of MAL19 by genome assembly walking out from the initial assembled partial cox1 gene. Assembled partial mitogenome contig versions were improved iteratively until a draft assembled circularised mitogenome for MAL19 was obtained. Ambiguous nucleotides and sub-optimally assembled regions were checked manually to ensure resolutions. Annotation of the assembled mitogenome was performed using Mitos (Bernt et al., Reference Bernt, Donath, Jühling, Externbrink, Florentz, G, Middendorf and Stadler2013) selecting genetic code 5 (invertebrate mitochondrial genome). Mitos annotated bedfile (.bed) was imported into Geneious to assist with fine tuning of the MAL19 final version of mitogenome (Pa-MAL19; Magalhaes et al., Reference Magalhaes, Czepak, van Niekerk, du Plessis, Court and Tay2024). Fine tuning of mitogenomes involved identifying a ‘Methionine’ (Met) start codon and a termination codon for each of the 13 protein-coding gene (PCG). To infer the stop codon for the 13 PCGs, gene length was extended until the first stop codon was identified, irrespective of whether the predicted PCG over-lapped a transfer RNA (tRNA) gene or not. This approach differed from those used to predict stop codons of the 13 PCGs of the Chinese (Yunnan, Xinjiang) and Spanish T. absoulta mitogenomes (Li et al., Reference Li, Fu, Guo, Wang and Lu2022; Zhang et al., Reference Zhang, Yang and Zhang2019), where the stop codon ended with single T in the cox1, cox2, nad5, and nad1 genes.
The fully assembled and annotated MAL19 mitogenome (Pa-MAL19; Magalhaes et al., Reference Magalhaes, Czepak, van Niekerk, du Plessis, Court and Tay2024) was then used as the reference genome for the assembly of MAL20 (Pa-MAL20; Magalhaes et al., Reference Magalhaes, Czepak, van Niekerk, du Plessis, Court and Tay2024) and ZA1 (Pa-ZA1; Magalhaes et al., Reference Magalhaes, Czepak, van Niekerk, du Plessis, Court and Tay2024) draft mitogenomes. The published mitogenomes of P. absoluta individuals from Spain (GenBank accession number MK654754), China (Xinjiang, GenBank accession number MT897989; Yunnan, GenBank accession number MT897990), and Kenya (SRR22312267; Ajene et al., Reference Ajene, Heya and Khamis2024) were downloaded from GenBank for comparison against our African individuals. The Helicoverpa species’ mitogenome resources reported in Walsh et al. (Reference Walsh, Perera, Anderson, Gordon, Czepak, McGaughran and Tay2019) were downloaded to enable comparative genome analysis assessments of P. absoluta mitogenomes assembly quality via amino acid residue compositions and consideration of their expected biochemical properties.
Assembly and identification of OP and pyrethroid resistance genes
To assess and characterise the resistance genes Acetycholineesterase 1 (AChE-1/ace-1) responsible for OP and the Para-type VGSC gene responsible for pyrethroid and DDT resistance, we downloaded the Tuta absoluta strain TA1 ace-1 mRNA partial coding sequence KU985167 from GenBank for use as reference sequence to assist with assembly of the AChE-1 gene in our Malawi and South African individuals. Similarly, we downloaded the T. absoluta partial coding sequence KY767010 from GenBank to assist with assembly of the pyrethroid/DDT resistance gene ‘VGSC’ of our African P. absoluta individuals. In both AChE-1 and VGSC, resistance to OP and to pyrethroid, respectively, have been identified in various lepidopteran pest species (e.g., S. frugiperda; H. armigera; P. abosulta) as due to nucleotide base substitution resulting in amino acid changes (Table S1). For P. absoluta, the known nucleotide substitutions for KU985167 and KY767010 are compared and as provided in Carvalho et al. (Reference Carvalho, Omoto, Field, Williamson and Bass2013) (Table S1).
Molecular diagnostics
To provide confidence that the Malawian and South African larvae sequenced were P. absoluta and to better understand the implications between utilisation of partial mtCOI gene versus full mitogenome PCGs in understanding the species’ invasion biology, we undertook molecular diagnostics and pairwise nucleotide comparison of published and our assembled mtCOI partial gene sequences. We selected representative sequences encompassing various worldwide locations from GenBank (Table 1) for alignment using the MAFFT Align tool (Katoh and Standley, Reference Katoh and Standley2013) in the Geneious software and trimmed all to 570 bp to ensure no missing data in all sequences prior to downstream analyses.
Quality control of partial mtCOI (cox1) gene sequences
The Staden sequence assembly package’s Pregap and Gap4 programs (Staden et al., Reference Staden, Beal and Bonfield2000) were employed for contig assembly and quality checking of the selected published mtCOI partial sequences (Table 1). Identified base substitutions in clean sequences were translated into amino acids and the likelihood of possible incorrect base-calling in these published sequences were assessed through amino acid residue biochemical properties (Kunz et al., Reference Kunz, Tay, Elfekih, Gordon and De Barro2019b). Presence of premature stop codons that may indicate candidate nuclear mitochondrial (NuMt) sequence was also assessed through amino acid translation. Sequence alignment also enable possible insertion/deletion (INDEL) mutations to be identified.
Basic nucleotide statistics and haplotype network construction
Pairwise nucleotide analysis was conducted to compare all 13 PCGs of the published mitogenomes and mitogenomes of the three African P. absoluta samples reported in this study. Each full length PCG was extracted with the MAL19 PCG gene lengths used as standards for pairwise gene alignment using MAFFT with default parameters (Auto algorithm, scoring matrix: 200PAM/k = 2, Gap open penalty: 1.53, Offset value: 0.123). Once aligned, the genetic variability from the 13 PCGs were recorded as a base substitution table (Table 2) to enable a miotogenome haplotype network to be constructed (Fig. 2). For the haplotype network analysis, SNP data were converted to the Phylip format prior to being analysed using the TCS v1.23 program (Clement et al., Reference Clement, Posada and Crandall2000), with the connection limit for the haplotype network set at 15 steps. This approach allowed for the visualization and analysis of the samples’ network structure, highlighting evolutionary patterns and genetic relationships among them (Fig. 2).
Summary of base substitutions of the Malawian (MAL19, MAL20) and South African (ZA1) P. absoluta samples and including published mitochondrial genomes of Spain (MK654754) and China (MT897989, MT897990) P. absoluta samples. Base substitutions were identified in eight mtDNA PCGs and for the 16S rRNA gene
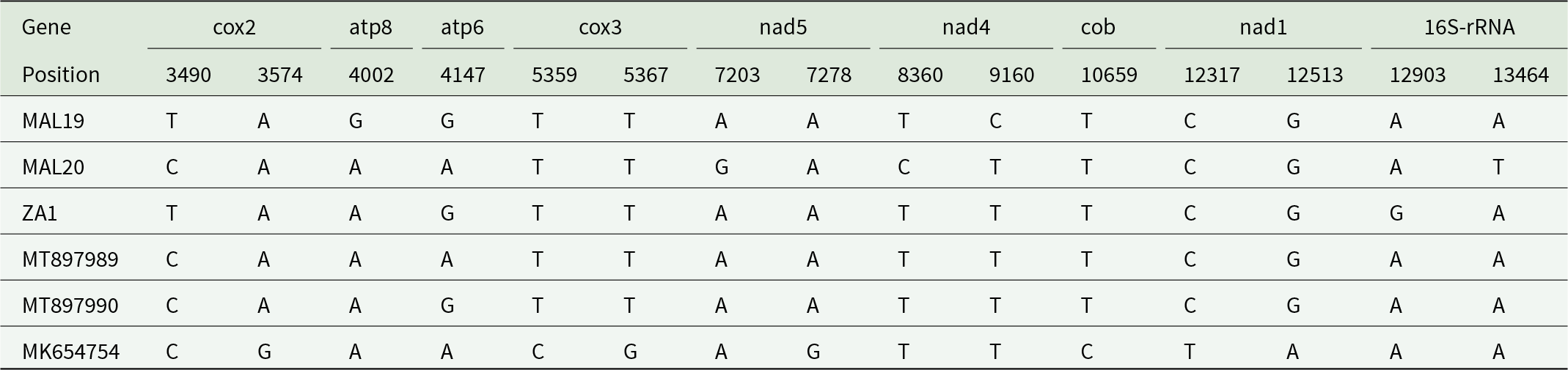
Note: No base substitutions were present in the full mtCOI gene of these six full mitogenomes compared. An INDEL was detected in the 16S-rRNA gene of ZA1 at alignment position 13,930 (ZA1 has a ‘T’ which was not detected in the other five mitogenomes).
Results
The mitogenome organisation and diversity of Phthorimaea absoluta
A total of 107 manual mitogenome assembly iterations was needed to generate our final draft mitogenome for MAL19. All three African P. absoluta mitochondrial genomes (MAL19, MAL20, ZA1) exhibited the expected circular genome structure, as shown for the MAL19 mitogenome (Fig. 1), with draft mitogenome lengths of 15,290 bp for both MAL19 and MAL20, and 15,292 bp for ZA1. The draft mitogenomes of MAL19, MAL20, and ZA1 all contained 13 PCGs, 2 ribosomal RNA (rRNA) genes, 22 tRNA genes, and a control region. Among the 37 genes, 23 were encoded on the heavy strand (H-), while four PCGs (nad5, nad4, nad4l, and nad1), eight tRNAs (trnC-tgc, trnY-tac, trnF-ttc, trnH-cac, trnP-cca, trnL1-cta, trnV-gta, and trnQ-caa), two rRNAs were encoded on the light strand (L-), and an approximately 320 bp non-coding region that is also rich in A + T nucleotide composition (91.9%). No nucleotide base substitutions were detected for six of the mitochondrial genes (i.e. nad2, cox1, atp6, nad3, nad4l, nad6), and one to two base substitutions observed among the cox2, atp8, atp6, nad5, nad4, and 16S rRNA genes (Table 2) between the published and our African P. absoluta individuals.
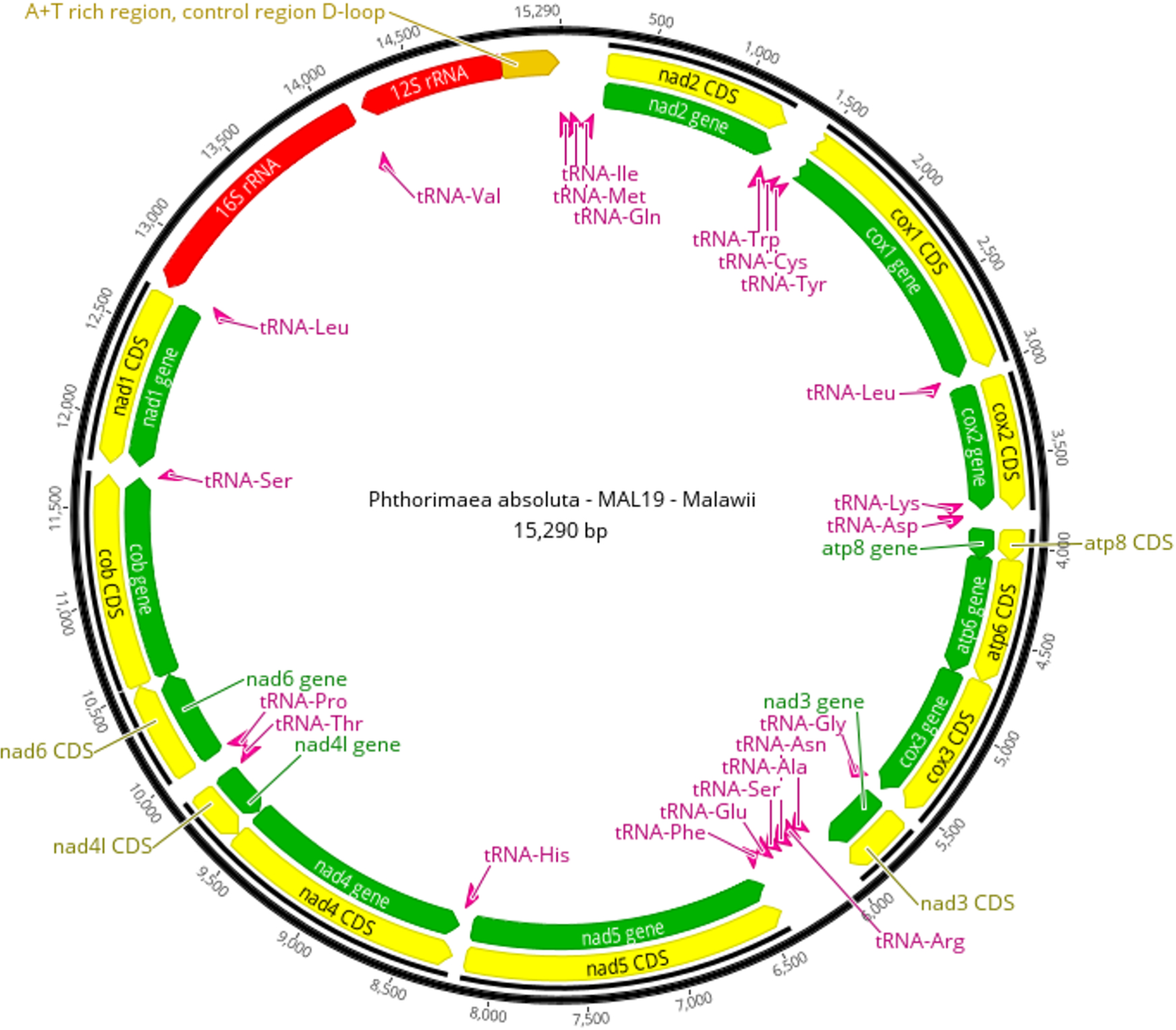
Circular mitochondrial genome (mitogenome) of the phthorimaea absoluta individual MAL19 (Pa-MAL19; Magalhaes et al., Reference Magalhaes, Czepak, van Niekerk, du Plessis, Court and Tay2024) from Malawi, showing 13 protein-coding genes (green), 22 tRNA genes (pink), 2 rRNA genes (red), and an A + T rich control region (Orange).
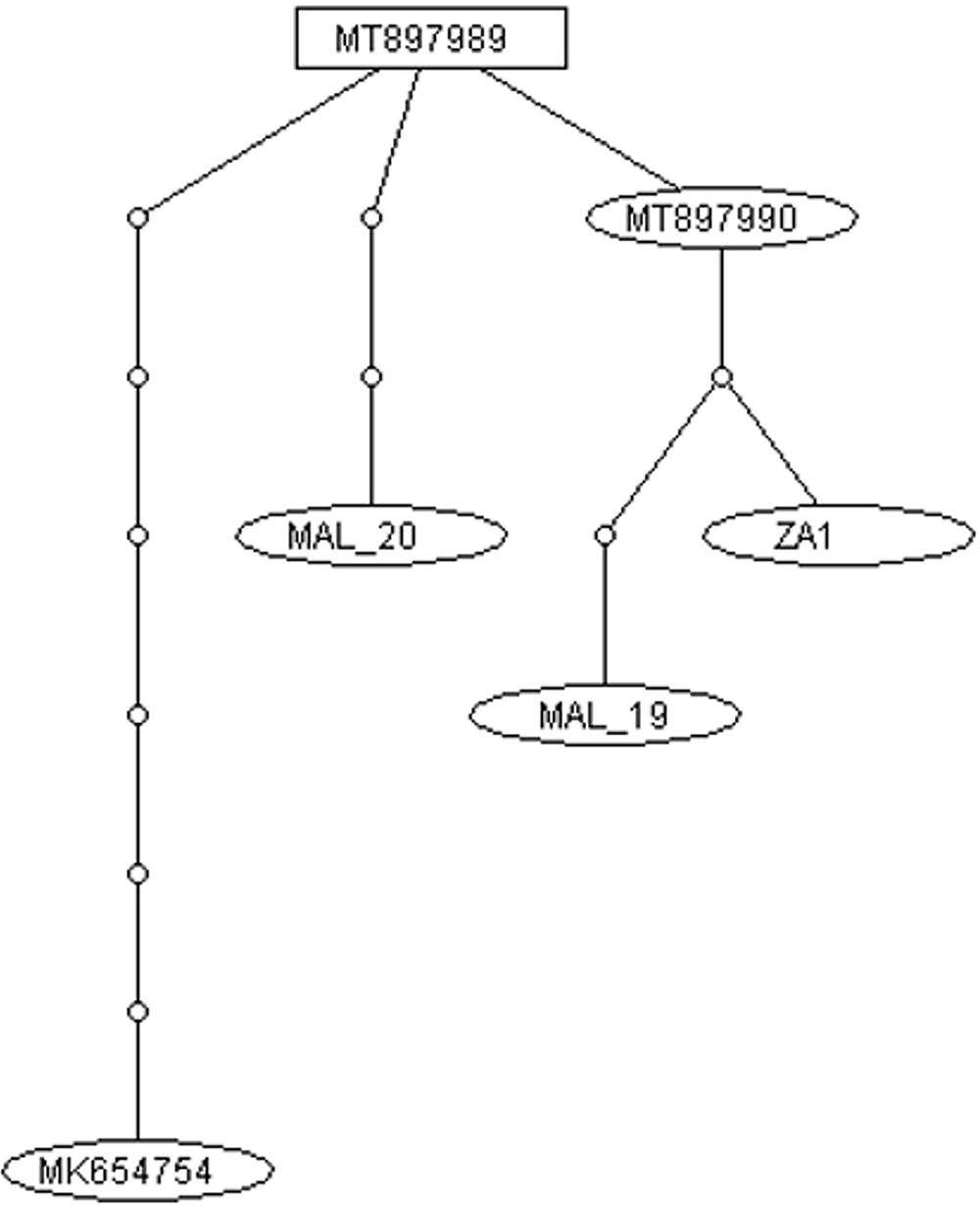
Phthorimaea absoluta draft mitogenome haplotype network. Number of base changes between haplotypes are indicated by open circles.
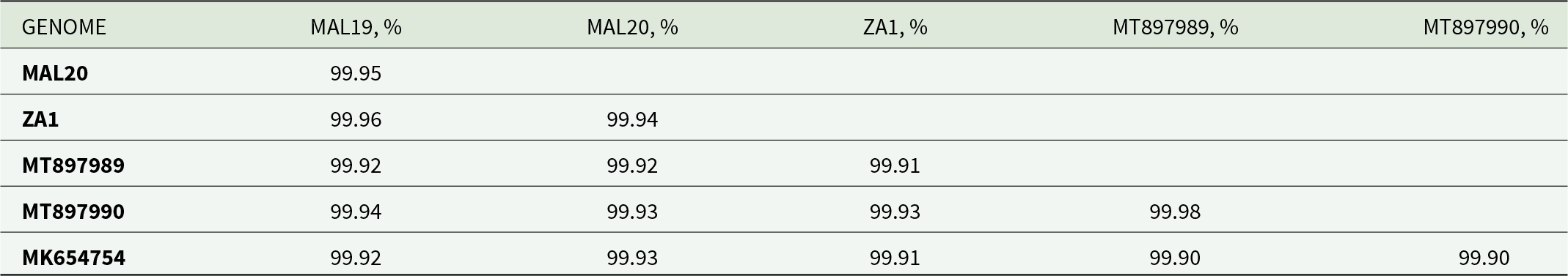
The nucleotide composition of the draft P. absoluta mitogenome based on MAL19 was as follows: 39.3% A, 40.6% T, 8.1% G, and 12.0% C, respectively, and exhibiting an overall high A + T bias (79.8%). Of the 13 coding genes, one started with the CGA codon (cox1), six with the ATG codon (cox2, atp6, nad4, nad4l, cob, cox3), four with the ATT codon (atp8, nad5, nad3, nad2), and two with the ATA codon (nad6, nad1), and all 13 PCGs terminate with the TAA codon (Table 3). The mitogenomes between the African, European, and Asian P. absoluta showed high degree of similarity (i.e. between 99.90 and 99.98%; Table 4), while 17 intergenic sequences and four overlapping sequences were also identified (Table 3).
Annotation of P. absoluta mitochondrial genomes between MAL19 from Malawi (Pa-MAL19; Magalhaes et al., Reference Magalhaes, Czepak, van Niekerk, du Plessis, Court and Tay2024) and published mitogenomes from China (Xinjiang, GenBank MT897989; Yunnan, GenBank MT897990) by Li et al. (Reference Li, Fu, Guo, Wang and Lu2022) and Spain (GenBank MK654754) by Zhang et al. (Reference Zhang, Yang and Zhang2019)
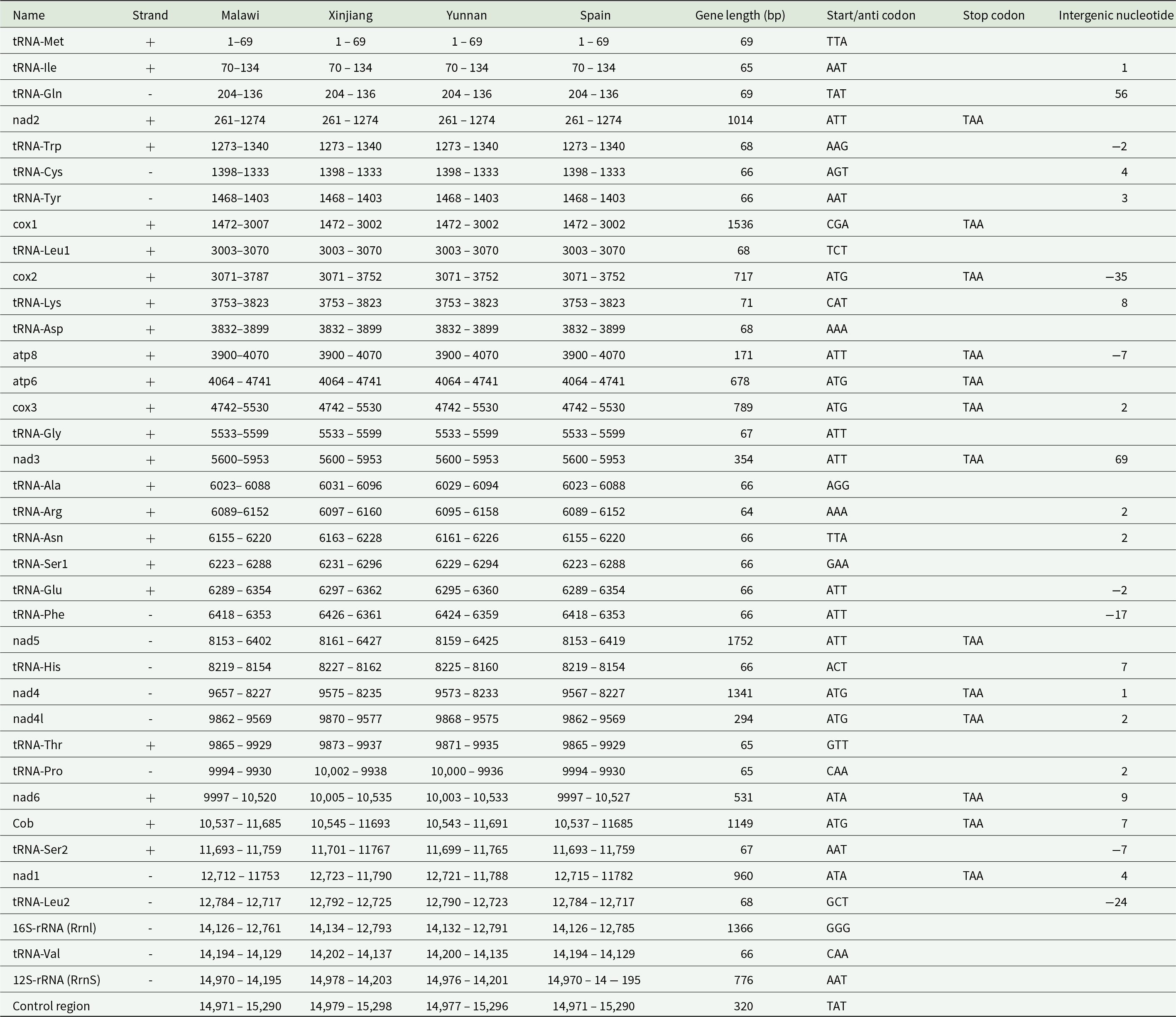
Note: Detailed annotation of P. absoluta mitochondrial genomes for MAL19, MAL20, and ZA1 are also provided in Magalhaes et al. (Reference Magalhaes, Czepak, van Niekerk, du Plessis, Court and Tay2024). Predicted secondary structures of transfer RNAs (tRNAs) are provided in Fig. S1.
Comparison of pairwise nucleotide distances (p-dist) among the 13 concatenated (11,286bp) PCGs from the mitogenomes of Phthorimaea absoluta individuals from Malawi (MAL19, MAL20), South Africa (ZA1), Spain (MK654754), and China (MT897989, MT897990)
Through comparisons of gene lengths within individuals from Africa, from the mitogenomes, it was observed that the genes exhibited similar lengths, except for the 16S-rRNA (Rrnl) region, where the ZA1 individual showed a slightly longer length (1,367 bp) as compared with MAL19 and MAL20 (1,366 bp). Previously, Li et al. (Reference Li, Fu, Guo, Wang and Lu2022) reported differences only in the lengths of intergenic spacers between two Chinese individuals and a Spanish individual. However, in the current study, discrepancies in gene lengths were identified between our African individuals and the others due to the different approaches for determining start and stop codon positions for the PCGs. These differences were particularly notable in the cox1, cox2, nad5, nad1, and 16S-rRNA (Rrnl) regions, where African individuals exhibited longer lengths (Table 3).
Assessments of mitogenome assembly quality prior to calculating intraspecific nucleotide variability between the reported P. absoluta mitogenomes and our African P. absoluta mitogenomes was undertaken (Table S2). Across the seven mitogenomes now being reported, we identified 44 base substitutions that resulted in 27 synonymous and 17 non-synonymous amino acid changes in the 13 PCGs. Of the 17 non-synonymous amino acid changes, one occurred in MAL19 at nucleotide positions (nt) 8358-8360 (within the nad4 gene) and involved a change from Glutamic Acid (E; negatively charged side chain) in all P. absoluta to Glycine (G; ‘special case’) in MAL19. Two non-synonymous amino acid residue changes were detected in the Spain specimen (MK654754, Zhang et al., Reference Zhang, Yang and Zhang2019), and involved a change from Valine (V, hydrophobic side chain) to Glycine (cox3, nt5366-5368), and from Serine (S, polar uncharged side chain) to Phenylalanine (F, hydrophobic side chain) at nt12511-12513 (nad1 gene). The remaining unexpected 14 non-synonymous amino acid changes at the intraspecific level were all detected in the Kenyan P. absoluta mitogenome (SRR22312267; Ajene et al., Reference Ajene, Heya and Khamis2024).
Nucleotide base and amino acid residue substitutions at positions corresponding to those of P. absoluta were also compared at both intra- and interspecific levels in five different Helicoverpa species that included also four known subspecies (H. armigera armigera (Haa), H. armigera confera (Hac), H. assulta assulta (Haas), H. assulta afra (Haaf); Hardwick, Reference Hardwick1965; Walsh et al., Reference Walsh, Perera, Anderson, Gordon, Czepak, McGaughran and Tay2019). At the interspecific level, in the five genetically distinct Helicoverpa species (Hardwick Reference Hardwick1965; Pearce et al., Reference Pearce, Clarke and East2016; Anderson et al., Reference Anderson, Tay, McGaughran, Gordon and Walsh2016; Reference Anderson, Oakeshott, Tay, Gordon, Zwick and Walsh2018), only one non-synonymous amino acid change was observed (i.e. H. armigera (Haa, Hac), H. zea (Hz), H. punctigera (Hp) versus H. assulta (Haas, Haaf), H. geletopoeon (Hg)), and involved a change from Asparagine (N, uncharged side chain) to Glycine (G) at nt10650-10652 (cob gene) (Table S2). Excessive non-synonymous amino acid changes especially at the intraspecific level (e.g. Kenyan P. absoluta SRR22312267 versus all other P. absoluta; Table S2) are not expected for mitogenomes in arthropods (e.g. Kunz et al., Reference Kunz, Tay, Elfekih, Gordon and De Barro2019b; Walsh et al., Reference Walsh, Perera, Anderson, Gordon, Czepak, McGaughran and Tay2019) and the assembly quality of the Kenyan mitogenome will need to be re-checked to ensure accuracies. Base substitutions specific to the Kenyan P. absoluta that did not result in amino acid residue changes were identified (e.g. in cox2, atp6, cox3, nad5, nad4, cob, nad1; Table S2), suggesting that this Kenyan P. absoluta likely belonged to yet another unique (i.e. 8th) maternal lineage. However, suspected mitogenome assembly issues nevertheless precluded calculations and analyses of haplotype network, nucleotide diversity, and intra-specific comparisons of mitogenome characterization involving this Kenyan individual.
Assembly and identification of OP and pyrethroid-resistance genes
We assembled the partial AChE-1 and the partial VGSC gene regions in our P. absoluta samples for characterisation of OP and pyrethroid-resistance alleles, respectively. We successfully identified OP resistance alleles in all three P. absoluta specimens from Malawi and from South Africa. In MAL19 and ZA1, resistance to OP would be expected due to changes to the A201S and the F290V amino acid residues, while for MAL20, base substitution from GCT to TCT is expected to result in OP resistance due to the A201S amino acid change (Table 5). Pyrethroid resistance profiles obtained from the partially assembled VGSC gene identified super-kdr for MAL19 due to the presence of M918T + L1014 amino acid substitutions, and for the first time, amino acid substitution profiles of T929I + L1014F in ZA1 which corresponded to the 1B pyrethroid-resistance profile in Musca domestica, expected to be of greater resistance to pyrethroid insecticides than super-kdr (Roca-Acevedo et al., Reference Roca-Acevedo, Boscaro and Toloza2023). For MAL20, poor coverage prevented characterisation of the L1014F amino acid residue, although the T929I amino acid residue change was detected (Table 5).
Nucleotide substitutions and amino acid residue changes in the Acetylcholine esterase-1 (AChE-1/ace-1) and the Para-type VGSC genes, associated with OP and pyrethroid knockdown resistance (kdr), respectively, in phthorimaea absoluta individuals from Malawi (MAL19, MAL20) and South Africa (ZA1). Missing genome data coverage is indicated by ‘-,’ indetermined amino acid residue is indicated by ‘?.’ Single amino acid residue letter codes used: alanine (A), serine (S), glycine (G), phenylalanine (F), valine (V), methionine (M), threonine (T), isoleucine (I), leucine (L)
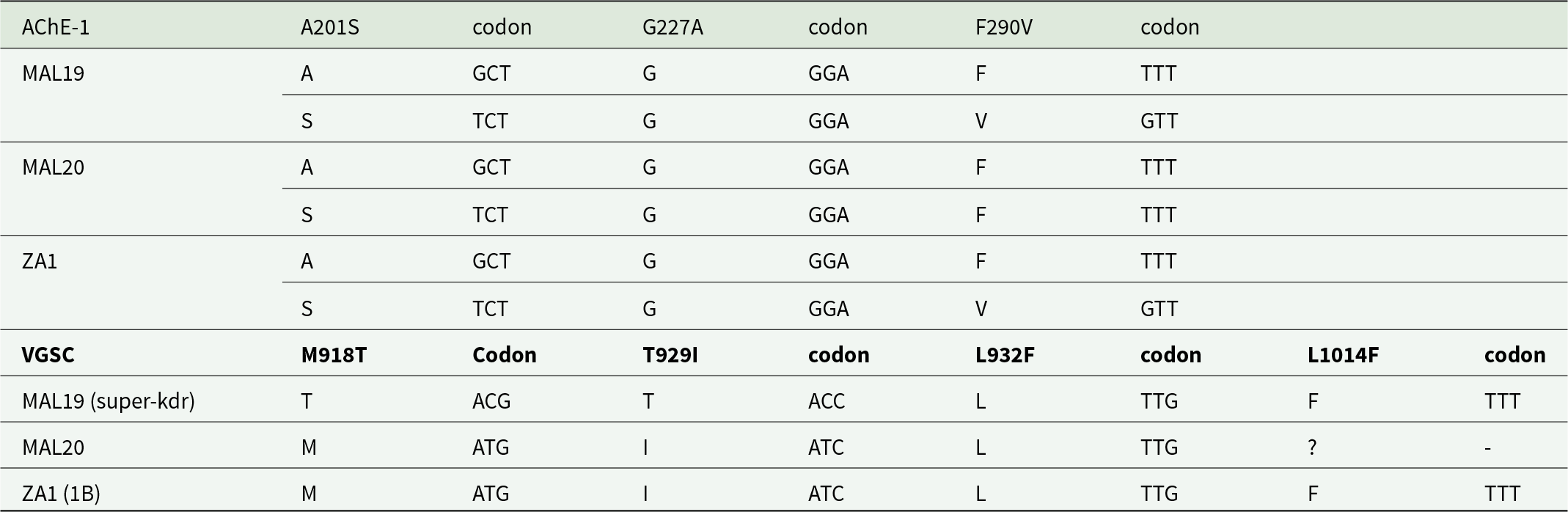
Note: VGSC Kdr (L1014F); super-kdr (M918T + L1014); kdr-his (L1014H); 1B (T929I + L1014F); type N (D600N + M918T + L1014F). General level of protection conferred by these alleles against pyrethroids: kdr-his < kdr < Type N ≤ super-kdr ≤ 1B (see Roca-Acevedo et al., Reference Roca-Acevedo, Boscaro and Toloza2023). Novel concomitant mutations L932F + I936V (Sugiura et al., Reference Sugiura, Kimoto, Itokawa and Kasai2021) for VGSC were not detected in the three P. absoluta individuals.
Mitogenome haplotype network
The network analysis showed that MAL19 and the Spain (MK654754) mitogenomes were most divergent from each other, separate by 11 base substitutions, followed by between MAL19 and MAL20 (7 base substitutions). South African (ZA1) and China P. absoluta shared close maternal evolutionary relationships, being separated by two (i.e. between ZA1 and MT897990) and three (i.e., between ZA1 and MT897989) base substitutions. The two Chinese individuals (MT897989 and MT897990) showed the closest maternal lineage relationship, being separated by only one step. We were unable to confidently assess the Kenyan P. absoluta mitogenome haplotype due to the high frequencies of significant amino acid residue changes at the intraspecific level (Table S2). Raw WGS data should ideally be made available for independent confirmation, and if the mitogenome assembly quality was confirmed, this highly divergent mitogenome haplotype would further support current widespread detections of P. absoluta involved multiple origins of genetically diverse native populations.
Molecular diagnostics and re-assessment of P. absoluta partial mtCOI gene
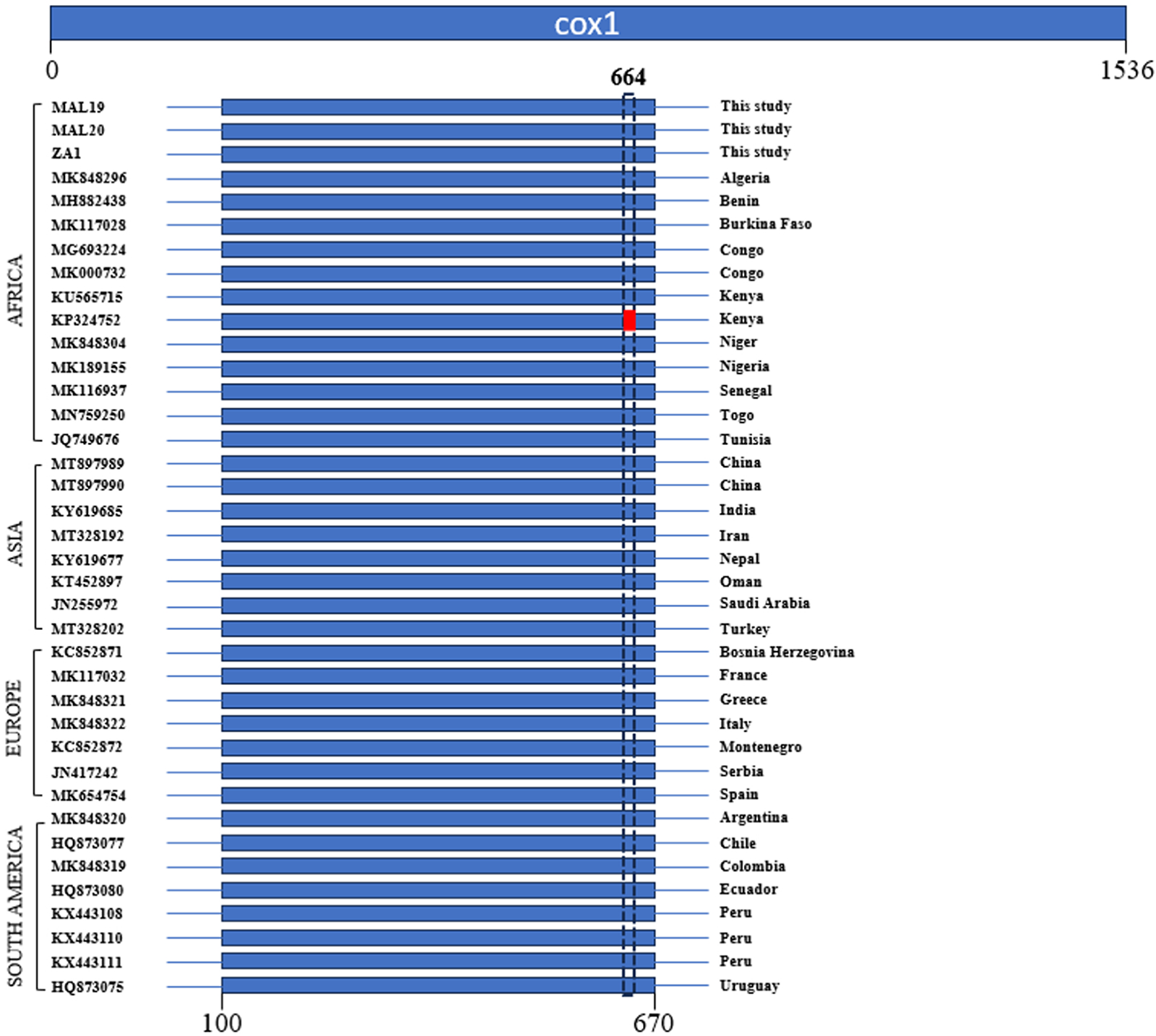
Comparison of the mtCOI partial gene region (i.e. the ‘DNA barcoding’ gene region, Hebert et al., Reference Hebert, Cywinska, Ball and DeWaard2003) typically used in molecular diagnostics supported the P. absoluta species identity for the three African individuals with 100% sequence identity versus partial mtCOI sequences available in GenBank for selected Africa (N = 15), Asia (N = 8), Europe (N = 7), and South America (N = 8) sequences. We further observed that, specifically in the cox1 region, there is evidence of only two maternal lineages (i.e. KP324752 from Kenya versus all others) (Fig. 3). Taken together, and assuming that this base substitution in KP324752 (i.e. a substitution between ‘T’ and ‘G’) to be a potential real SNP (amino acid for all other partial mtCOI gene with ‘TGG’ codon = (Gly), amino acid for KP324752 = ‘GGG’ (Gly), i.e. there is no change to the predicted glycine amino acid residues), there would appear at least seven separate maternal lineages identified in the P. absoluta invasive populations from Europe, Africa, and Asia.
Graphical representation comparing the partial mtCOI gene of Phthorimaea absoluta samples from different parts of the world. A base substitution (G) in the Kenyan partial mtCOI gene (KP324752) that differed from all other partial mtCOI gene at nucleotide position 644 (T) is shown in red rectangle.
Discussion
In this study, we presented draft mitochondrial genomes of two Malawian and one South African P. absoluta, representing the first mitogenome resources from these eastern and southern African countries. When compared with the individual from Spain and two individuals from Xinjiang and Yunnan, China (Tables 2, S2), and the partial mtCOI gene survey from a Kenyan P. absoluta population (Fig. 3), we demonstrated that at least seven different maternal lineages were present in African, European, and Asian invasive P. absoluta populations. We also characterised the AChE-1 and VGSC partial gene regions responsible for OP and pyrethroid resistance in these African P. absoluta individuals. Our results revealed surprising mitogenome haplotypes, as well as OP and pyrethroid resistance diversity in P. absoluta, and highlighted the limitation of using partial mtCOI gene as DNA marker to infer invasion biology of often complex global introduction pathways and incursion frequencies of agriculturally significant AIS.
Until now, genetic analyses of P. absoluta populations have largely relied on the partial mtCOI gene widely used in species diagnostics (Hebert et al., Reference Hebert, Cywinska, Ball and DeWaard2003; but see also, e.g., Guillemaud et al. (Reference Guillemaud, Blin, Le Goff, Desneux, Reyes, Tabone and Lombaert2015); Cifuentes et al. (Reference Cifuentes, Chynoweth and Bielza2011)), and reported high genetic homogeneity among populations (e.g., Flores et al., Reference Flores, Gilardón and Gardenal2003; Cifuentes et al., Reference Cifuentes, Chynoweth and Bielza2011; Shashank et al., Reference Shashank, Twinkle, Chandrashekar, Meshram, Suroshe and Bajracharya2018). A survey of the partial mtCOI gene of multiple native and invasive P. absoluta populations demonstrated this gene region to be lacking in nucleotide polymorphisms (Fig. 3). Here, we identified eight mitochondrial genes where base substitutions were detected that could facilitate the development of alternative DNA markers for both molecular diagnostics and investigation of population diversity of this important Solanaceae pest. Specifically, by combining the cox2, atp6, atp8, and the 16S-rRNA partial genes could enable differentiation of all six maternal lineages. Li et al. (Reference Li, Fu, Guo, Wang and Lu2022) analysed multiple partial mtDNA genes (i.e. cox1, cox2, cox3, atp6, cob, nad1, nad5) to demonstrate that populations between Xinjian and Yunnan were genetically different at the cox2, atp6, nad1, and nad5 partial gene regions. Going forward, there is a need to standardise the set of multiple partial P. absoluta mtDNA gene markers for use in population genetic studies, although adoption of the WGS approach for the characterisation of the mitogenome would be more economic and efficient. Finally, these new P. absoluta mitogenomes will also further contribute to the in-depth understanding of lepidopteran systematics especially for the Gelechiidae family, and between the Gelechioidea superfamily (i.e. for which P. absoluta belonged) and other lepidopteran species as shown by Zhang et al. (Reference Zhang, Yang and Zhang2019) and Chen et al. (Reference Chen, Chen, Liao, Wang, Wang and Huang2022).
Assessment of the Kenyan P. absoluta individual identified exceptionally high amino acid composition changes in this individual as compared with mitogenomes of other P. absoluta individuals (Table S2). Across many of these mitogenome polymorphic sites, amino acid composition conservation was detected not only between P. absoluta individuals (i.e. intraspecific comparison), but also between lepidopteran species (i.e. Helicoverpa species; Table S2) from evolutionary diverged lineages. High amino acid conservation patterns in the partial mtCOI gene region have been shown in the cryptic Bemisia tabaci species complex (Kunz et al., Reference Kunz, Tay, Elfekih, Gordon and De Barro2019b; Vyskocilová et al., Reference Vyskocilová, Tay, van Brunschot, Seal and Colvin2018), and the detection of the high number of intraspecific amino acid residue changes between the Kenyan and other African, Asia, and Europe P. absoluta individuals was therefore unexpected and suggested potential assembly inaccuracies.
With rapidly decreasing WGS cost, investigation of the invasive biology for P. absoluta would benefit from analysis of genome-wide SNP loci and full mitogenomes, as already demonstrated for other invasive Lepidoptera (e.g. S. frugiperda, Tay et al., Reference Tay, Rane, Padovan, Walsh, Elfekish, Downes and Gordon2022d; Rane et al., Reference Rane, Walsh, Lenancker, Gock, Dao, Nguyen and Tay2023; and H. armigera, Anderson et al., Reference Anderson, Tay, McGaughran, Gordon and Walsh2016; Zhang et al., Reference Zhang, Zhang, Tay, Robin, Shi, Guan and Wu2022) that could also enable characterisation of known resistance genes (e.g. Guan et al., Reference Guan, Zhang, Shen, Wang, Padovan, Walsh and Wu2021; Walsh et al., Reference Walsh, Joussen, Tian, McGaughran, Anderson, Qiu and Heckel2018). There is as yet no P. absoluta population genomic studies based on genome-wide loci although Guillemaud et al. (Reference Guillemaud, Blin, Le Goff, Desneux, Reyes, Tabone and Lombaert2015) inferred population genetics of P. absoluta using low numbers of SSR loci, and concluded that there was an overall absence of genetic sub-structure across the Mediterranean Basin populations that very probably corresponded to a single introduction in Africa or Spain, followed by a rapid expansion and little noticeable demographic bottleneck effect, but acknowledging also the recognised difficulties of scoring SSR loci in the Lepidoptera due to high null allele frequencies.
The presence of null alleles and other related factors such as allele drop out and loci not in Hardy Weinberg equilibrium, could be due to association with transposable elements (e.g. Gordon et al., Reference Gordon, Tay, Collinge, Williams, Batterham, Goldsmith and Marec2009; Tay et al., Reference Tay, Behere, Batterham and Heckel2010) and/or due to Wahlund effect and could contribute to poor lepidopteran population genetic result interpretations. A closer examination of the P. absoluta structure analysis (Guillemaud et al., Reference Guillemaud, Blin, Le Goff, Desneux, Reyes, Tabone and Lombaert2015) identified potential genetic diversity that could have been missed especially in the Gre_lef (Greece), Fra_bal/Fra_ber (France), and the Cyp_emp (Cyprius) populations which differed from, e.g., Isr_wga (Israel), Mar_lar (Morocco), and Spa_alm (Spain) populations. Increasing the number of SSR and/or adopting the EPIC-PCR loci approach (e.g. Behere et al., Reference Behere, Tay, Russell, Kranthi and Batterham2013; Tay et al., Reference Tay, Behere, Heckel, Lee and Batterham2008), and avoiding potential marker-association with mobile elements, including using of multiple mtDNA gene markers (e.g. Li et al., Reference Li, Fu, Guo, Wang and Lu2022; Otim et al., Reference Otim, Tay, Walsh, Kanyesigye, Adumo, Abongosi and Agona2018), or by adopting genome-wide SNP loci population genomic approaches (e.g. Elfekih et al., Reference Elfekih, Tay, Polaszek, Gordon, Kunz, Macfayden, Walsh, Vyskocilova, Colvin and De Barro2022), could increase sensitivity to detect potential signatures of population substructure in P. absoluta invasive populations.
Analyses of draft mitogenomes in this study therefore demonstrated that P. absoluta populations in the invasive ranges are genetically diverse and complex, although more mitogenomes from invasive populations are needed. There is also currently no draft mitogenome resources for P. absoluta from native range populations or from South American invasive range such as from Brazil. These diverse maternal lineages increased genetic variability in the invasive populations and underpinned the likelihood of better adaptation to new environments, to better finding suitable colonisation sites, as well as increasing the potential of introducing insecticide resistance alleles (Suinaga et al., Reference Suinaga, Casali, Picanço and Foster2004) that will enable the pest to better survive within agricultural environments (Lockwood et al., Reference Lockwood, Cassey and Blackburn2005). Zhu et al. (Reference Zhu, Chen, Liu, Lin, Liang, Nauen, Li and Gao2024) showed that pyrethroids, organophosphates, and carbamates are likely to be ineffective in controlling P. absoluta and the related P. operculella due to the widespread detection of resistance alleles in invasive populations studied. In this study, we demonstrated that WGS data simultaneously enabled characterisation of both mitogenome and known OP and pyrethroid resistance genes in our limited African samples. The diversity of insecticide resistance gene profiles between P. absoluta populations from the many invaded countries remained to be adequately characterised, although targeted PCR tests have been developed and carried out for both AChE-1 and kdr alleles, e.g. in Kenyan populations (Haddi et al., Reference Haddi, Berger, Bielza, Cifuentes, Field, Gorman and Bass2012; Ong’onge et al., Reference Ong’onge, Ajene, Runo, Sokame and Khamis2023).
Our characterisation of the pyrethroid resistance gene VGSC identified the ‘1B’ (T929I + L1014F) kdr resistance allele first described in Musca domestica, and which was shown to have higher resistance to permethrin + piperonyl butoxide (PBO) and deltamethrin + PBO (Kasai et al., Reference Kasai, Sun and Scott2017; Roca-Acevedo et al., Reference Roca-Acevedo, Boscaro and Toloza2023). Co-occurrence of the Isoleucine I and the Phenylalanine amino acids in T929I and in L101 F, respectively, in P. absoluta is likely in various populations due to the L101F resistance amino acid being fixed in survey populations, and the T929I amino acid change detected in approximately 50% of the tested individuals (Haddi et al., Reference Haddi, Berger, Bielza, Cifuentes, Field, Gorman, Rapisarda, Williamson and Bass2021). The detection of the M. domestica ‘1B’ kdr phenotype in South Africa is nevertheless demonstrated for the first time, although the level of resistance to pyrethroids in this 1B mutational alleles as compared with other kdr mutation alleles remained to be investigated. A review of currently known insecticide resistances in P. absoluta is provided (Table S3).
Diversity in AChE-1 OP resistance alleles in the invasive lepidopteran pest S. frugiperda has been linked to multiple independent introductions across the world (e.g. see Boaventura et al., Reference Boaventura, Ulrich, Lueke, Bolzan, Okuma, Gutbrod and Nauen2020; Tay et al., Reference Tay, Rane and Padovan2022c) and different selective pressures on these pest populations. With the intensity and variety of source populations playing a major role in determining the genetic diversity of the invasive populations (Wilson et al., Reference Wilson, Dormontt, Prentis, Lowe and Richardson2009), it is therefore unsurprising that evidence from both mitogenomes and insecticide resistance gene diversity (e.g. this study; Ong’onge et al., Reference Ong’onge, Ajene, Runo, Sokame and Khamis2023; Zhu et al., Reference Zhu, Chen, Liu, Lin, Liang, Nauen, Li and Gao2024) supported complex global introduction history of P. abosluta. A whole genome sequencing approach incorporating well-designed and well-coordinated insecticide/biopesticide bioassay experiments is now the logical next step to enable development of environmentally responsible future management solutions for P. absoluta at a global scale.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0007485325000252.
Acknowledgements
V.S.M. expresses gratitude for the support provided by CSIRO Health and Biosecurity during his time as a visiting student for the molecular characterization of P. absoluta populations. We thank Rahul Rane and Tom Walsh (CSIRO Applied Genomics Initiative (AGI)) for support with preparing and coordinating the P. absoluta genome sequencing effort. This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior – Brasil (CAPES) – Finance Code 001 and project 3B02782 from the North-West University, South Africa. The authors express their gratitude to Dr Donald L. Kachigamba from the Department of Agricultural Research Services (DARS), Bvumbwe Research Station, Malawi, who provided the Malawian P. absoluta specimens.
Competing Interests
The authors declared no competing interests.










