1. Introduction
Heading date (HD) is one of the most important agronomic traits in rice (Xing et al., Reference Xing, Xu, Hua, Tan and Sun2001; Guo et al., Reference Guo, Luo, Xing, Xu, Wang, Shi, Mei, Zhong and Ying2002) and closely associated with yield, quality and tolerance to stresses. Previous studies have shown that HD is a quantitative trait subject to the influences of environmental factors. Photoperiod and temperature are the major environmental factors affecting HD. Different areas and cropping seasons require varieties with different HD. Varieties with HD less affected by environmental factors have wider adaption, and thus can be planted in a larger geographical region. A better understanding of the genetic control of HD in rice will greatly enhance breeding efficiency by adopting the most effective breeding method and techniques. QTL mapping studies in the last two decades have reported 734 HD QTL distributed on all 12 rice chromosomes (http://www.gramene.org). Eight of the QTLs with large effects have been cloned using positional cloning (Yamamoto et al., Reference Yamamoto, Lin, Sasaki and Yano2000; Yano et al., Reference Yano, Katayose, Ashikari, Yamanouchi, Monna, Fuse, Baba, Yamamoto, Umehara, Nagamura and Sasaki2000; Takahashi et al., Reference Takahashi, Shomura, Sasaki and Yano2001; Kojima et al., Reference Kojima, Takahashi, Kobayashi, Monna, Sasaki, Araki and Yano2002; Doi et al., Reference Doi, Izawa, Fuse, Yamanouchi, Kubo, Shimatani, Yano and Yoshimura2004; Matsubara et al., Reference Matsubara, Yamanouchi, Wang, Minobe, Izawa and Yano2008, Reference Matsubara, Yamanouchi, Nonoue, Sugimoto, Wang, Minobe and Yano2011; Wei et al., Reference Wei, Jiang, Xu, Zhang, Lu, Zhang and Wan2008; Xue et al., Reference Xue, Xing, Weng, Zhao, Tang, Wang, Zhou, Yu, Xu, Li and Zhang2008; Yan et al., Reference Yan, Wang, Chen, Zhou, Li, Wang, Ding, Zhang, Yu, Xing and Zhang2011).
There is little evidence that the identified HD QTLs have been effectively exploited for breeding (Wei et al., Reference Wei, Liu, Xu, Jiang, Zhang, Wang, Zhai and Wan2010). One of the reasons is that most of the reported HD QTLs were identified using only one mapping population tested in a single environment. It is understandable that many studies have reported inconsistent results. Indeed, just like the phenotypic performance of a complex trait is affected by environmental factors, the expression of a QTL for the complex trait is also strongly dependent on environmental factors (Hittalmani et al., Reference Hittalmani, Huang, Courtois, Venuprasad, Shashidhar, Zhuang, Zheng, Liu, Wang, Sidhu, Srivantaneeyakul, Singh, Bagali, Prasanna, McLaren and Khush2003; Li et al., Reference Li, Yu, Lafitte, Huang, Courtois, Hittalmani, Vijayakumar, Liu, Wang, Shashidhar, Zhuang, Zheng, Singh, Sidhu, Srivantaneeyakul and Khush2003). Moreover, the expression of a QTL is also dependent on the genetic background it resides (Mei et al., Reference Mei, Xu, Li, Yu, Guo, Wang, Ying and Luo2006; Chevin & Hospital, Reference Chevin and Hospital2008; Wang et al., Reference Wang, Cheng, Sun, Zhou, Zhu, Xu, Xu and Li2009). The identification of QTLs with stable expression across genetic backgrounds and environments is a key for marker-assisted manipulation of complex traits in breeding (Ribaut & Hoisington, Reference Ribaut and Hoisington1998; Ye & Smith, Reference Ye and Smith2010).
We made an attempt to identify stable HD QTL using two sets of BC2F6 reciprocal introgression lines (ILs) in 2 years and a set of recombinant inbred lines (RILs) in 1 year. The objectives are to detect QTL effects under different genetic backgrounds, identify stable QTL and analyse pyramiding effects of different main-effect QTL.
2. Materials and methods
(i) Plant materials
A high-yielding japonica rice cultivar–Xiushui09 from China was crossed with a drought tolerant indica breeding line–IR2061-520-6-9 (herein abbreviated as IR2061) from International Rice Research Institute (IRRI). The F1 plants were simultaneously backcrossed to the two parents to develop two BC1F1 populations of 100 plants. The BC1F1 plants were used as the male parent to backcross with the two parents to produce two BC2F1 populations. The BC progenies were consecutively selfed for seven generations without any selection to obtain BC2F6 populations. Ultimately, two sets of reciprocal ILs, each comprising of 240 BC2F6 lines, were developed in 2006. Using the two sets of reciprocal ILs, genetic background effect on salt-tolerant QTL detection was revealed (Cheng et al., 2012). In this study, herein we designated the ILs in Xiushui09 and IR2061 backgrounds as XS-ILs and IR-ILs, respectively. In addition, 240 F8 RILs derived from the same cross of Xiushui09 and IR2061 were developed by single seed decent method.
(ii) Field trails and phenotyping
The two sets of reciprocal ILs and their parents were evaluated in the experimental station of Institute of Crop Science, Chinese Academy of Agricultural Sciences at Sanya (18.3°N, 109.3°E) and Hainan province in 2007 and 2009. A randomized complete block design with two replicates was used. Five-leaf seedlings were transplanted into the field on 25th November. Plot size was two 10 plants-rows with 25 cm row space and 17 cm between plant spaces. During the flowering period, eight representative plants per plot were recorded for HD. HD was recorded as days from the time of sowing to that of the first panicle flowering in 50% of the plants in each plot. The RILs were tested in the same site at Sanya, Hainan province in 2009 using the same design and agronomic practices.
(iii) Genotyping, linkage map construction and QTL analysis
Ten grams of fresh leaf tissues were bulk harvested for DNA extraction using the Cetyltriethylammonium bromide (CTAB) method with minor modifications. The two sets of reciprocal IL populations and the RIL population were assayed with a set of same 142 well-distributed simple sequence repeat (SSR) markers. Linkage maps were constructed using the MapManager QTX15 software (Manly & Olson, Reference Manly and Olson1999).
Inclusive interval mapping (ICIM) implemented in the Icimapping 3.2 software was used to conduct QTL mapping. ICIM consists of two steps (Li et al., Reference Li, Ye and Wang2007). In the first step, marker selection is conducted through stepwise regression by considering all marker information simultaneously. Phenotypic values are then adjusted by all markers retained in the regression equation, except the two markers flanking the current mapping interval. In the second step, the adjusted phenotypic values are used in one-dimensional scanning. In this study, the two probabilities for entering and removing variables in the first step were set at 0·001 and 0·002, respectively. The empirical LOD threshold was set at 2·5 for claiming a significant QTL based on 1000 runs of randomly shuffling the trait values in the second step (Churchill & Doerge, Reference Churchill and Doerge1994). HD data collected from different years were analysed separately. QTLs with overlapping of 95% confidence interval under different conditions were considered to be the same one.
(iv) Analysis of QTL pyramiding effect
To analyse pyramiding effects of introgressed donor alleles and allele combinations at the detected QTL on HD in the XS-ILs, all lines were grouped according to the allele and allele combinations using the markers closest to the QTL in the XS-ILs. Duncan's multiple range test (DMRT) was employed to examine the significant differences among different QTL genotype groups.
3. Results
(i) Linkage maps of the three mapping populations
The linkage map constructed for the XS-ILs population spanned 1665·5 cM with an average distance of 11·7 cM between adjacent markers. The linkage map constructed for the IR-ILs population spanned 1528·7 cM with an average distance of 10·8 cM. The marker distributions on chromosomes and orders were the same for the XS-ILs and IR-ILs. The linkage map constructed for the RILs population spanned a total of 1867·8 cM with an average distance of 13·2 cM.
On an average, the introgressed donor genome was 6·1% ranging from 1·3 to 26·4% in the 240 XS-ILs, whereas it was 7·2% ranging from 0·42 to 39·4% in the 240 IR-ILs. The peak values of the reciprocal sets of ILs separated without any overlapping between frequency distributions of the Xiushui09 genome (Fig. 1).
Frequency distribution of Xiushui09 genome in the reciprocal IL and RIL populations derived from the cross of Xiushui09/IR2061.

(ii) Phenotypic variations of the parents and different populations
Xiushui09 flowered significantly earlier than IR2061 with the average HD being 93·6 and 100·2 days in 2007, and 79·2 and 89·4 days in 2009, respectively (Table 1). Transgressive segregations were observed for both of the ILs populations in 2 years. The RILs population also showed a significant transgressive segregation in HD in 2009 (Table 1). The phenotypic variance in the RIL population was roughly two times more than in the IL populations.
Phenotypic performance of HD in the reciprocal ILs and RILs and their parents under different conditions

* , ** Represent significant differences at levels of P < 0·05 and 0·01.
(iii) QTL detection
A total of nine QTLs explaining 3·0–33·7% of the total phenotypic variance were identified in the XS-ILs in 2 years (Table 2 and Fig. 2). They were on chromosomes 3, 4, 6, 7, 8, 10 and 12. Five QTLs (QHd3, QHd7, QHd10a, QHd10b and QHd12) were identified in 2007 and the introgressed IR2061 alleles at QHd7 and QHd10b promoted flowering, whereas those of the other three QTLs (QHd3, QHd10a and QHd12) delayed HD. In 2009, eight QTLs (QHd3a, QHd4a, QHd6a, QHd6b, QHd7, QHd8, QHd10b and QHd12) were detected. The IR2061 alleles at all QTLs except QHd7 and QHd10b delayed HD. Among them, QHd3, QHd7, QHd10b and QHd12 were detected in both of the 2 years.
QTL for HD detected in the reciprocal ILs (XS-ILs and IR-ILs) and the RILs derived from the cross of Xiushui09/IR2061 in years 2007 and 2009.

QTL affecting HD detected in the reciprocal ILs and RILs under different conditions

a Genetic distance of the left marker.
b Genetic distance of the putative QTL apart from the left marker.
c Underlined markers are those closer to the true QTL positions.
d A represents the additive effect, estimated as the substitution effect of Xiushui09 allele by IR2061 allele in XS-ILs and RILs, while the substitution effect of IR2061 allele by Xiushui09 allele in IR-ILs.
Five QTLs on chromosomes 4, 5, 6, 7 and 10 were identified in the IR-ILs in 2 years. They explained 3·6–25·9% of the total phenotypic variance (Table 2 and Fig. 2). Four QTLs (QHd5, QHd6b, QHd7 and QHd10b) were detected in 2007 and the introgressed Xiushui09 alleles at the first two loci promoted HD, whereas those at the latter two loci delayed HD. Four QTLs (QHd4b, QHd6b, QHd7 and QHd10b) were detected in 2009 and the introgressed Xiushui09 alleles at the first two loci delayed HD, whereas those at the latter two loci promoted HD. Three QTLs (QHd6b, QHd7 and QHd10b) were simultaneously identified in 2 years.
Seven QTLs (QHd2, QHd3, QHd4b, QHd6a, QHd7, QHd10a and QHd10b) were detected in RIL population in 2009 (Table 2 and Fig. 2). They were distributed on chromosomes 2, 3, 4, 6, 7 and 10 and accounted for 3·0–11·0% of the total phenotypic variance. QHd10b had the largest additive effect and accounted for 11·0% of the total phenotypic variance. The HD-delaying alleles at all loci except QHd2, QHd7 and QHd10b were associated with IR2061 (Table 2 and Fig. 2).
Four QTLs (QHd3, QHd6a, QHd7 and QHd10b) detected in XS-ILs were commonly detected in RILs with same directions of gene effect in 2009. Similarly, four QTLs (QHd4b, QHd6a, QHd7 and QHd10b) detected in IR2061 ILs were also detected in the RILs with same directions of gene effect. However, only two (QHd7 and QHd10b) or three QTLs (QHd6a, QHd7 and QHd10b) were commonly identified in the two reciprocal IL populations in 2007 and 2009. QHd6a, QHd7 and QHd10b were detected in all the three populations, indicating that these three QTLs were less affected by the differences in genetic backgrounds.
(iv) Pyramiding effect of QHd7 and QHd10b
QHd7 and QHd10b detected in different populations in 2 years were used to investigate the effect of pyramiding. The IR2061 alleles at QHd7 and QHd10b increased HD. All XS-ILs were divided into four groups based on the introgressed IR2061 alleles or allele combinations at the above two QTLs (Table 3). In 2007 and 2009, the averages of HD of 192 lines without introgressed IR2061 alleles at QHd7 and QHd10b (group 1) were 94·7 days and 74·5 days, respectively, which was similar to the recurrent parent, Xiushui09. Twelve and 17 lines with introgressed IR2061 alleles at QHd7 and QHd10b (groups 2 and 3) had significant shorter HD than group 1. The average HD of the five lines with introgressed IR2061 alleles at QHd7 and QHd10b (group 4) was significantly shorter than those of groups 2 and 3, suggesting that the effects of the two QTLs could be accumulated by pyramiding.
Effects on HD of introgression of IR2061 alleles into Xiushui09 background
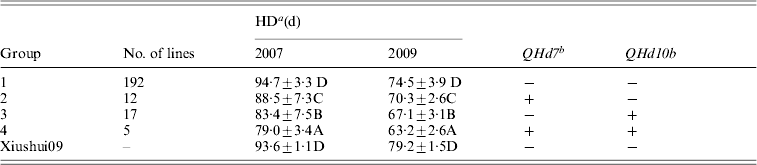
a Different letters represent significant levels at P ⩽ 0·05 based on DMRT.
b ‘ + ’ indicates lines with the favourable allele from the donor parent, IR2061 while ‘ − ’ without allele introgression from the donor parent.
To further confirm the two HD QTLs and their pyramiding effect in a similar genetic background, three ILs, XS-28 (QHd10b), XS-112 (QHd7) and XS-211 (QHd7 and QHd10b), were tested in different years. The introgressed donor genome (IR2061) of the three ILs was 4·0, 11·9 and 6·6%, respectively (Fig. 3). In 2007, the HDs of XS-28 and XS-112 were 78·5 days and 87 days, respectively, which were significantly shorter than 93·6 days of the recurrent parent, Xiushui09. Similarly, in 2009, the HDs of XS-28 and XS-112 were 68·5 days and 69·5 days, respectively, which were significantly shorter than 79·2 days of the recurrent parent, Xiushui09. The HD of the two-QTL pyramided line, XS-211, was 75·5 days (65·5 days) in 2007 (2009), which was significantly shorter than those of XS-28 and XS-112 (single-QTL ILs).
Graphical genotype and performance of HD of near iso-genic ILs with QTL QHd7, QHd10b and the two pyramided QTLs, respectively, in Xiushui09 background. A stands for Xiushui09, B, C and D for the ILs, XS-28, XS-112 and XS-211 with QHd7, QHd10b and the two pyramided QTLs, respectively and E for IR2061. The grey bars indicate homozygous for the Xiushui09 genome, whereas the black bars indicate introgressing homozygous IR2061 genome. Data in the right two columns are HD of each line recorded in years 2007 and 2009. Different letters represent significant levels at P ⩽ 0·05 based on DMRT.

4. Discussion
So far, a wide range of segregating populations derived from bi-parental crosses, including RILs (Xiao et al., Reference Xiao, Li, Yuan and Tanksley1996; Li et al., Reference Li, Arif, Zhong, Fu, Xu, Domingo-Rey, Ali, Vijayakumar, Yu and Khush2006), doubled haploid (DH) lines (Hittalmani et al., Reference Hittalmani, Huang, Courtois, Venuprasad, Shashidhar, Zhuang, Zheng, Liu, Wang, Sidhu, Srivantaneeyakul, Singh, Bagali, Prasanna, McLaren and Khush2003; Mu et al., Reference Mu, Li, Li, Zhang, Wu, Li and Wang2003) and F2 populations (Tang et al., Reference Tang, Ma, Teng, Yan, Wu, Dai and Li2007) have been used for mapping QTLs. Abundant evidences in molecular quantitative genetics indicate that QTL expression is strongly dependent on genetic background and environmental factors (Li et al., Reference Li, Yu, Lafitte, Huang, Courtois, Hittalmani, Vijayakumar, Liu, Wang, Shashidhar, Zhuang, Zheng, Singh, Sidhu, Srivantaneeyakul and Khush2003; Wang et al., Reference Wang, Cheng, Sun, Zhou, Zhu, Xu, Xu and Li2009; Cheng et al., Reference Cheng, Wang, Meng, Hu, Cui, Sun, Zhu, Ali, Xu and Li2012). The use of reciprocal ILs tested in 2 years in this study, allowed studying environmental and genetic background effects. Moreover, we also compared the results from the two reciprocal ILs populations with that of a random RILs population. In 2007, a total of seven QTLs for HD were identified in the two reciprocal ILs populations, two QTLs (28·6%) were detected in both genetic backgrounds. In 2009, 3 (33·3%) of the nine QTLs were commonly detected in the two ILs populations. QHd7 and QHd10b were detected and confirmed under different backgrounds and environments, indicating that they were stable. Four (57·1%) and five (50·0%) QTLs detected using the IR-ILs and XS-ILs populations were also identified using the RILs population. Three QTLs (25·0%) were detected in all three populations. Obviously, the number of commonly identified QTL between different mapping populations largely depends on the degree of the overlap between their genomes. In most cases, the genetic backgrounds are very different between mapping and breeding populations. Identification of QTL with large effects should be followed by extensive validation in multiple backgrounds across environments before utilization in breeding through QTL pyramiding or other marker-assisted selection (MAS) methods (Ye & Smith, Reference Ye and Smith2010). QTL mapping and breeding are better conducted in the same genetic background or population (Tanksley & Nelson, Reference Tanksley and Nelson1996). In this context, ILs have advantages compared with other mapping population. In our study, QHd6a, QHd7 and QHd10b could stably express in different genetic backgrounds and environments, indicating that they may be useful for breeding by MAS. However, it should be pointed out that the three QTLs were detected in the three populations which derived only from the same two parents under short-day condition. These QTLs still need to be further validated in other genetic backgrounds and in various photoperiod conditions. This is particularly true for breeding for rice-production region in north or central China.
Among the three stable QTLs for HD, QHd6a was mapped near to RM136 (9683415 bp) on chromosome 6. Hd1, a previously reported gene for promoting heading under short-day conditions, was located in the region with physical distance from 9335536 to 9337360 bp on chromosome 6 (Yano et al., Reference Yano, Katayose, Ashikari, Yamanouchi, Monna, Fuse, Baba, Yamamoto, Umehara, Nagamura and Sasaki2000). Hence, QHd6a is mostly related to Hd1 and the allele from Xishui09 promoted heading in the natural short-day condition. QHd10b was mapped on chromosome 10 and located in the region of RM271–RM258 (16637171–16934494 bp). The closer marker RM271 was also linked with Ehd1 gene (16723434–16899784 bp) for the promotion of heading under short-day conditions as reported by Doi et al. (Reference Doi, Izawa, Fuse, Yamanouchi, Kubo, Shimatani, Yano and Yoshimura2004). So Hd10b was probably related to Ehd1. QHd7 was mapped near to the marker RM501 (8006020 bp) on chromosome 7. So far, a total of 83 QTL for HD has been reported on chromosome 7 (http://www.gramene.org). Most of them were located in three regions on chromosome 7. The first one is on the upper chromosome 7 where Li et al. (Reference Li, Yu, Lafitte, Huang, Courtois, Hittalmani, Vijayakumar, Liu, Wang, Shashidhar, Zhuang, Zheng, Singh, Sidhu, Srivantaneeyakul and Khush2003) detected a QTL for HD in the region of RZ488–RG477 (4572812 bp–6778514 bp) across eight different environments. The second one was in the middle of chromosome 7 where the Hd4 and Ghd7 (9154038 bp) were detected by Lin et al. (Reference Lin, Liang, Sasaki and Yano2003) and Xing et al. (Reference Xing, Xu, Hua, Tan and Sun2001), respectively. The third one was located at the end of chromosome 7 where the gene Hd2 (29434231 bp) conferring photoperiod responses was identified by Yano et al. (Reference Yano, Harushima, Nagamura, Kurata, Minobe and Sasaki1997). QHd7 was located near to the Ghd7. The gene Ghd7 had significant effect on HD (15 d) under long-day condition but no significant effect under short-day condition, whereas the QHd7 was consistently identified in the three populations under short-day condition in Sanya of Hainan province (18.1_N, 109.5_E). Therefore, QHd7 was different from Ghd7.
To develop cultivars with HD required by the target production environments is one of the most important objectives of rice breeding. For instance, in Hainan province many of the current cultivars flower too late. The reduction of HD is particularly important for many tropic regions where triple cropping seasons per year are practiced. The IR2061 alleles at QHd7 and QHd10b identified in this study on average shortened HD by 4·4 and 6·2 days in 2 years in XS-ILs, respectively. The experiment with ILs (XS-28 and XS-112) with only a small portion of the donor (IR2061) genome in XS background confirmed that that the introgressed IR2061 alleles at QHd10b and at QHd7 significantly reduced HD, confirmed the effects of the two main-effect QTLs. The ILs containing both of the IR2061 alleles of QHd7 and QHd10b had significantly shorter HD than most of the ILs with IR2061 alleles of only one of the two QTLs, indicating that QTL pyramiding is an efficient method in breeding for HD to the target environment. Likewise, in north and central China, early maturing varieties are required for rice production due to the short summer and long-day condition. The success of rice production in such areas depends on raising relative photoperiod insensitivity varieties with short basic vegetative growth period (Okumoto et al., Reference Okumoto, Ichitani, Inoue and Tanisaka1996). Two of the three stable QTLs identified under short-day condition in this study, QHd6a and QHd10b are very close to Hd1 and Ehd1, respectively, and could be useful for this purpose if further studies in various photoperiod conditions could confirm their effectiveness and stability.
It might be worth pointing out that the two testing seasons had similar day length but very different temperatures. A big snowstorm occurred in late winter of 2008 and reduced the temperature greatly. The low temperature significantly delayed flowering time in 2009 than in 2007. This might contribute to the observed fewer number of QTL detected across years. Nevertheless, the three QTLs with large effects, which are most useful in MAS, were detected across 2 years.
In conclusion, expressions of HD QTL could be strongly affected by genetic background and environmental factors. Among the QTLs detected in this study, QHd6a, QHd7 and QHd10b could stably express in the three populations with different complexities of the genetic backgrounds and two testing environments and could be exploited for breeding for earliness (short HD). The combined effect QHd7 and QHd10b was highly significant larger than their individual effects, suggesting that QTL pyramiding will be an effective method for the development of lines with adequate HD.
The authors thank two anonymous reviewers for their invaluable comments and suggestions on the manuscript. This work was supported by the ‘948’ Project (2011-G2B) from the Chinese Ministry of Agriculture, the ‘863’ Project (2012AA101101) and the International Cooperative Project (S2012ZR0160) from the Chinese Ministry of Science and Technology.
5. Declaration of Interest
None.







