I. INTRODUCTION
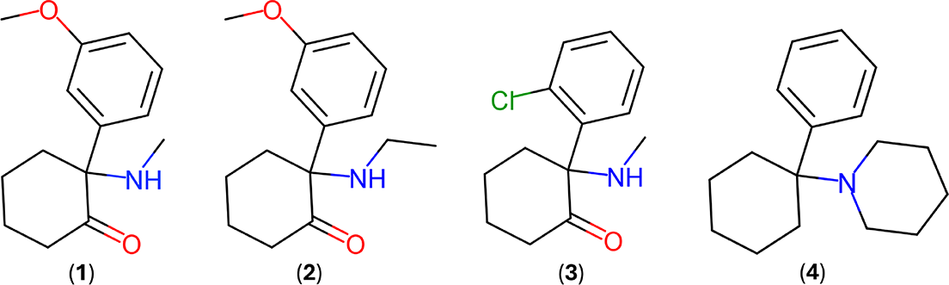
Methoxmetamine, 2-(3-methoxyphenyl)-2-(methylamino)cyclohexanone, C14H19NO2, MMXE (1) and methoxetamine, 2-(ethylamino)-2-(3-methoxyphenyl)cyclohexanone, C15H21NO2, MXE (2) are arylcyclohexylamine derivatives, chemically related to ketamine (3) (C13H16ClNO, 2-(2-chlorophenyl)-2-(methylamino)cyclohexanone) and to the internationally controlled substance phencyclidine (4) (C17H25N, 1-(1-phenylcyclohexyl) piperidine, PCP), a dissociative hallucinogenic drug popularly known as “angel dust” (Morris and Wallach, Reference Morris and Wallach2014; Kaizaki-Mitsumoto et al., Reference Kaizaki-Mitsumoto, Noguchi, Yamaguchi, Odanaka, Matsubayashi, Kumamoto, Fukuhara, Funada, Wada and Numazawa2016). Figure 1 depicts the molecular diagrams of these compounds.
Molecular diagrams of methoxmetamine (1), methoxetamine (2), ketamine (3), and phencyclidine (4).

MMXE and MXE are among the constantly growing number of new psychoactive substances synthesized and introduced in the global market in the last two decades. The term “new psychoactive drugs” has also been used to designate these compounds. Particularly, MXE was the subject of a joint report prepared by the European Monitoring Centre for Drugs and Drug Addiction (EMCDDA) and the European Union Agency for Law Enforcement Cooperation (Europol) published in 2014 (EMCDDA–Europol, 2014). The EMCDDA report contains an assessment of the chemical and pharmacological properties of methoxetamine and its abuse- and dependence-causing potential. The health and social risks associated with this drug, prevalence of use, and the involvement of organized crime in its production and distribution are also presented.
MXE has been sold primarily over the Internet, typically as a whitish powder that can be inhaled, under names such as “Mexxy,” “MXE,” “Special M,” “Legal ketamine,” and “M-ket”. In addition, it has been sold as powder-filled capsules, tablets, and liquids, either in pure form or mixed with other illegal and/or controlled substances such as ketamine, amphetamines, cannabis, morphine, and heroin, or with compounds such as caffeine, with plant material, among many other components (EMCDDA–Europol, 2014). More than 110 non-fatal intoxications and 20 deaths associated with methoxetamine reported by eight EU Member States are contained in the EMCDDA–Europol report.
In the last few years, we have been particularly interested in the structural characterization of active pharmaceutical ingredients and other bioactive compounds using laboratory X-ray powder diffraction data (Dávila-Miliani et al., Reference Dávila-Miliani, Dugarte-Dugarte, Toro, Contreras, Camargo, Henao, Delgado and de Delgado2020; Toro et al., Reference Toro, Dugarte-Dugarte, van de Streek, Henao, Delgado and de Delgado2022; Dugarte-Dugarte et al., Reference Dugarte-Dugarte, Toro, van de Streek, Henao, Fitch, Dejoie, Delgado and de Delgado2023; Dugarte-Dugarte et al., Reference Dugarte-Dugarte, Toro, Henao, de Delgado and Delgado2024). The present work was prompted by our interest in these types of materials and the good room temperature X-ray powder diffraction patterns contained in the Powder Diffraction File™ (Kabekkodu et al., Reference Kabekkodu, Dosen and Blanton2024) for the HCl forms of MMXE and MXE (MMXE·HCl: PDF 00-070-1080, CAS No. 1781829-56-8; MXE·HCl: PDF 00-069-1044, CAS No. 1239908-48-5). A search in the Cambridge Structural Database (CSD) version 5.45 (Groom et al., Reference Groom, Bruno, Lightfoot and Ward2016) did not show reports on their crystal structures. Interestingly, a search using the ketamine fragment resulted in eight ketamine-related hydrochlorides contained in the CSD: APCNON (Stevens et al., Reference Stevens, Treat, Pillai, Schmonsees and Glick1973), KUQPAQ (Gao et al., Reference Gao, Gao, Wu, Li, Yang and Zhang2020), KUQPAQ01 (Hakey et al., Reference Hakey, Ouellette, Zubieta and Korter2008), MUWSUV (Smirnova et al., Reference Smirnova, Zhukhlistova, Tishchenko, Grinenko, Krupitskii and Moshkov1989), OWEYIB (Luelf et al., Reference Luelf, Reiss, Bokel and Urlacher2021), QIMGIF, QIMGOL, and QIMGUR (Jurásek et al., Reference Jurásek, Králík, Rimpelová, Čejka, Setnička, Ruml, Kuchař and Kohout2018b). Supplementary Table S1 contains the crystal data for these compounds, and Supplementary Figure S1 presents their molecular diagrams.
In view of the importance of having the most complete information possible about psychoactive substances for their identification and analysis, given their health and social impact, it was decided to undertake the structure determination of MMXE·HCl and MXE·HCl.
II. EXPERIMENTAL METHODS
The X-ray powder diffraction data for MMXE·HCl and MXE·HCl are contained in entries PDF 00-070-1080 and PDF 00-069-1044, respectively, of the Powder Diffraction File™ (Kabekkodu et al., Reference Kabekkodu, Dosen and Blanton2024). These data were initially reported in the Powder Diffraction Journal, recorded from samples previously synthesized by the corresponding authors (Maixner et al., Reference Maixner, Jurásek, Himl, Kuchař and Babor2017; Jurásek et al., Reference Jurásek, Babor, Huber and Kuchař2018a).
The diffraction pattern for MMXE·HCl was collected at room temperature on a Bruker D8 Discover powder diffractometer with Bragg–Brentano θ/θ geometry using Cu Kα radiation (λ = 1.5418 Å, generator setting: 40 kV, 40 mA). A selected specific energy from the detector was used for filtering Cu Kβ. Data were collected from 5° to 80° (2θ) with a step size of 0.02° (2θ) (Jurásek et al., Reference Jurásek, Babor, Huber and Kuchař2018a).
The diffraction pattern for MXE·HCl was recorded at room temperature with a PANalytical X’Pert3 powder diffractometer with Bragg–Brentano θ/θ geometry using Cu Kα radiation (λ = 1.5418 Å, Ni filter, generator setting: 40 kV, 30 mA). A PIXCEL detector was employed to collect the data from 5° to 70° (2θ) with a step size of 0.013° (2θ) and a counting time of 316.97 s per step (Maixner et al., Reference Maixner, Jurásek, Himl, Kuchař and Babor2017).
III. COMPUTATIONAL STUDIES
A. DFT-D calculations
The crystal structures obtained from laboratory X-ray powder diffraction data were energy-minimized, including the unit-cell parameters, with the program GRACE (Neumann, Reference Neumann2019), which calls VASP (Kresse and Furthmüller, Reference Kresse and Furthmüller1996) for single-point DFT calculations with the PBE functional (Perdew et al., Reference Perdew, Burke and Ernzerhof1996) to which a dispersion correction (Neumann and Perrin, Reference Neumann and Perrin2005) has been added. The method has been extensively validated against about 600 crystal structures, and the upper limit for the root-mean-square Cartesian displacement between the structure from the Rietveld refinement, if correct, and the energy-minimized structure was established to be approximately 0.35 Å (van de Streek and Neumann, Reference van de Streek and Neumann2014). Details of the calculations can be found elsewhere (Neumann and Perrin, Reference Neumann and Perrin2005).
B. Hirshfeld surface analysis
The software CrystalExplorer21 (Spackman et al., Reference Spackman, Turner, McKinnon, Wolff, Grimwood, Jayatilaka and Spackman2021) was used to produce “Fingerprint plots” of the parameter d norm mapped onto the Hirshfeld surface (HS; Spackman and Jayatilaka, Reference Spackman and Jayatilaka2009). These plots are useful for visualizing the atoms involved in intermolecular contacts, the strength of such contacts, and their contribution to the stabilization of the crystal structure.
IV. RESULTS AND DISCUSSION
A. Structure determination
The indexing of the reported MMXE·HCl and MXE·HCl patterns was accomplished with DICVOL14 (Louër and Boultif, Reference Louër and Boultif2014), running under the PreDICT graphical user interface (Blanton et al., Reference Blanton, Papoular and Louër2019). Using the first 20 peaks of the patterns, monoclinic unit cells were obtained in both cases. The de Wolff (Reference de Wolff1968) and Smith–Snyder (Reference Smith and Snyder1979) figures of merit obtained were M 20 = 17.9 and F 20 = 40.9 (0.0132, 37) and M 20 = 26.2 and F 20 = 43.8 (0.0127, 36), respectively. It must be noted that five very weak peaks of the MMXE·HCl pattern were not indexed. They are most likely due to an impurity phase. The unit-cell parameters obtained for both phases were the same as the parameters previously reported (Maixner et al., Reference Maixner, Jurásek, Himl, Kuchař and Babor2017; Jurásek et al., Reference Jurásek, Babor, Huber and Kuchař2018a). A reduced cell search in the CSD (Groom et al., Reference Groom, Bruno, Lightfoot and Ward2016), combined with a chemical elements search having only C, H, N, O, and Cl, for both compounds, yielded no hits.
The fit of the recorded patterns was carried out with the Pawley algorithm by modeling the background, sample displacement errors, absorption correction, surface roughness (Pitschke et al., Reference Pitschke, Hermann and Mattern1993), cell parameters, and peak shape parameters (including anisotropic broadening) using TOPAS-Academic (Coelho, Reference Coelho2018). The intermediate Gaussian–Lorentzian function was employed with a correction for axial divergence as proposed by the program.
In the fit of the MMXE·HCl pattern, a 15-term Chebyshev polynomial was used to model the background. The regions in which the five weak peaks appear (11.940°–12.060°, 19.540°–19.680°, 20.070°–20.180°, 24.180°–24.260°, and 31.580°–31.740°) were excluded. The final Pawley refinement produced a good fit with discrepancy factors Rp = 0.0285, R wp = 0.0383, and GoF = 1.719. The space group P21/n was suggested by the Bayesian extinction symbol algorithm in DASH 4.0.0 (Markvardsen et al., Reference Markvardsen, David, Johnson and Shankland2001) and by DAJUST (Vallcorba et al., Reference Vallcorba, Rius, Frontera, Peral and Miravitlles2012).
For MXE·HCl, a 10-term Chebyshev polynomial was used to model the background. The Pawley refinement produced a good fit to all the diffraction maxima with residuals Rp = 0.0120, R wp = 0.0154, and GoF = 1.669, confirming the correctness of the unit cell and the single-phase nature of the material. DASH 4.0.0 (Markvardsen et al., Reference Markvardsen, David, Johnson and Shankland2001), as well as DAJUST (Vallcorba et al., Reference Vallcorba, Rius, Frontera, Peral and Miravitlles2012), suggested the space group to be P21.
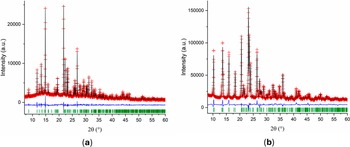
The initial molecular model for MMXE·HCl, introduced as a mol file, was built using the MOPAC2022 software (Moussa and Stewart, Reference Moussa and Stewart2024) with the PM7 method (Stewart, Reference Stewart2013). With this model and the profile parameters obtained from the Pawley fit, the crystal structure was determined using DASH 4.0.0 (David et al., Reference David, Shankland, van de Streek, Pidcock, Motherwell and Cole2006) with χ 2 = 12.213. The refinement of the structure was carried out using TOPAS-Academic (Coelho, Reference Coelho2018). The final Rietveld refinement plot is shown in Figure 2a.
Rietveld refinement plots for (a) MMXE·HCl and (b) MXE·HCl. The red crosses represent the observed data points, and the black line represents the calculated pattern. The blue curve is the normalized error plot, and the green sticks are the peak positions.

For MXE·HCl, the starting molecular model was built from the MMXE molecule in the MMXE·HCl model. The MMXE+ moiety was exported as a .mol file, and then one carbon atom was added to the methylammonium group. The resulting molecule was optimized using MOPAC2022 (Moussa and Stewart, Reference Moussa and Stewart2024) with the PM7 method (Stewart, Reference Stewart2013). With this model, the structure determination was carried out using DASH 4.0.0. The refinement of the structure, carried out using TOPAS-Academic (Coelho, Reference Coelho2018), produced a good fit. Figure 2b shows the final Rietveld refinement plot.
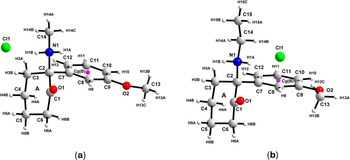
The refinement for MMXE·HCl included an overall scale parameter, the background, the peak shapes (including anisotropic broadening), unit-cell parameters, surface roughness parameter (Pitschke et al., Reference Pitschke, Hermann and Mattern1993), atomic coordinates, four B iso parameters, and a March–Dollase parameter (Dollase, Reference Dollase1986). For MXE·HCl, the same parameters were included in the refinement, with the difference that three B iso parameters were used instead of four. Planes (0, 2, 1) and (1, 0, 0), associated with preferred orientation, were included (see Table I). The bond distances and angles for both structures were restrained based on the values of the DFT-D structure obtained using GRACE. One planar restraint for the molecule (phenyl group) with a standard deviation of 0.01 Å was also added. The isotropic atomic displacement parameters for the hydrogen atoms were fixed to be 1.2 times the parameter of the C or N atom to which they are attached. Figure 3, drawn using DIAMOND v 5.1.0 (Putz and Brandenburg, Reference Putz and Brandenburg2023), depicts the molecular structures of MMXE·HCl and MXE·HCl, showing the atom and ring labeling scheme.
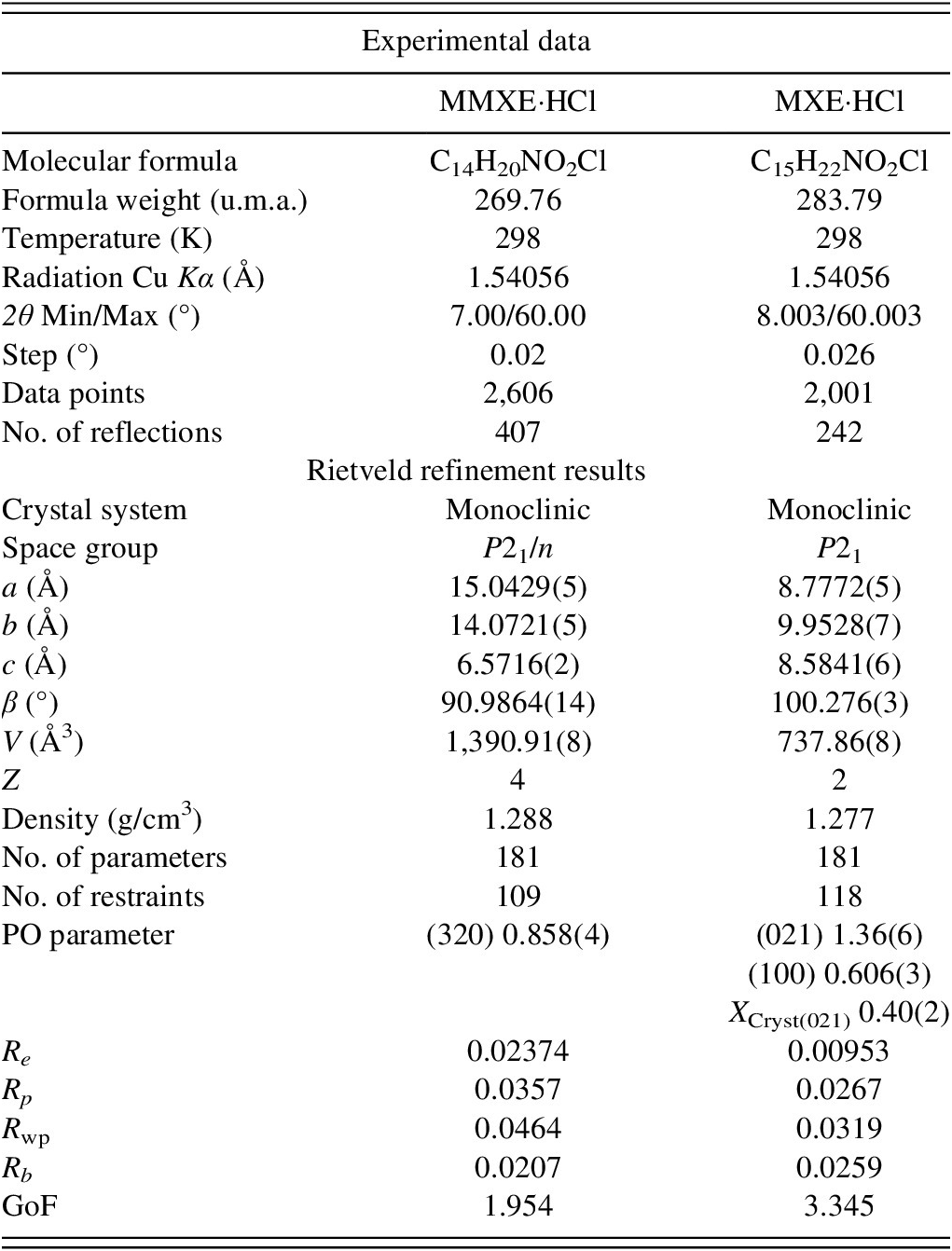
Crystal data, experimental parameters, and refinement results for MMXE·HCl and MXE·HCl

Molecular structure of (a) MMXE·HCl and (b) MXE·HCl with the labeling scheme for atoms and rings.

In total, for MMXE·HCl, 181 parameters were refined with 2,606 data points (407 reflections), 109 restraints, and 4 constraints. The final whole pattern fitting converged with good figures of merit: Re = 0.02374, Rp = 0.0357, R wp = 0.0464, and GoF = 1.954. For MXE·HCl, the powder pattern has 2,001 data points (242 reflections), and 118 restraints and 3 constraints were used to refine 183 parameters. The final refinement converged with good discrepancy factors: Re = 0.00953, Rp = 0.0297, R wp = 0.0351, and GoF = 3.345.
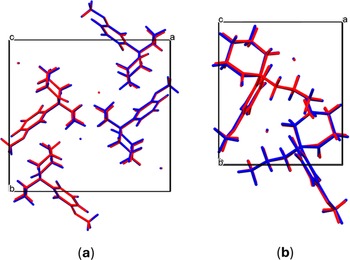
Table I summarizes the crystal data, experimental parameters, and the refinement results obtained for MMXE·HCl and MXE·HCl. The DFT-D calculations of the determined structure led to root mean square Cartesian displacement (RMSCD) values of 0.087 and 0.083 Å for MMXE·HCl and MXE·HCl, respectively. Both values are clearly lower than 0.35 Å, a value that has been recognized as a benchmark to indicate the correctness of the structure (van de Streek and Neumann, Reference van de Streek and Neumann2014). Figure 4 shows the comparison between the determined and the DFT-D-optimized molecular structures for each compound.
Superposition of the experimentally determined (red) and the energy minimized (blue) structure for (a) MMXE·HCl and (b) MXE·HCl.

Figure 5 depicts the superposition of the S-MMXE moiety of MMXE·HCl and the MXE moiety in MXE·HCl determined in the present study. As can be seen, the orientation of the methoxy group in MMXE·HCl is opposite with respect to MXE·HCl.
Superposition of the MMXE+ cation (cyan) and the MXE+ cation (magenta).

B. Molecular structure
The atomic coordinates, the isotropic displacement parameters, and the bond distances and angles are reported in Supplementary Tables S2–S7. In the statistical analysis performed using the Mogul Geometry Check (Bruno et al., Reference Bruno, Cole, Kessler, Luo, Motherwell, Purkis, Smith, Taylor, Cooper, Harris and A. G.2004), all the bond distances and angles in MMXE·HCl and MXE·HCl have Z-score values below 3, indicating that they are within expected values based on the reports contained in the CSD.
The asymmetric units of the MMXE·HCl and MXE·HCl structures contain one cation (MMXE+ and MXE+, respectively) and one chlorine ion (see Figure 3). The molecules are protonated at N1. Atom C2 is a chiral center.
The MMXE·HCl structure is centrosymmetric. Therefore, the two enantiomers, R and S, are present. In contrast, the structure of MXE·HCl is non-centrosymmetric. In this case, in the structure determination from the powder data, the S-enantiomer was used as the input model. It must be noted that the structure refinement of the S- and R-configuration led to the same results. It is well known that X-ray powder diffraction data cannot distinguish between the two configurations.
After the structure determination of the two compounds using the powder diffraction data contained in the PDF-5+ database was carried out in Merida, Venezuela, the authors of the powder data submission were contacted and invited to be co-authors of the publication. Then, they mentioned that a few years ago, they collected single-crystal data and determined the structure of these compounds, but never published the results. The structures determined using powder data agree with the ones obtained from the single crystal studies (Babor et al., Reference Babor, Jurásek and Kuchař2025).
Unfortunately, the MXE·HCl data were collected using Mo Kα radiation, and the absolute structure could not be determined. The statistics of the single crystal data, |E * E − 1| = 0.663, favor a non-centrosymmetric structure, as the expected values are 0.968 for a centrosymmetric structure and 0.736 for a non-centrosymmetric structure. After refinement, for the S-enantiomer, the Flack −0.54(4), Parsons −0.53(5), and the Hooft −0.55(4) parameters were obtained, which are physically unrealistic. These parameters should be close to zero if the structural model has the correct handedness. After inverting the handedness of the molecule, the values of these parameters should be close to 1. This is clearly not observed. When the structural model is inverted, the parameters are 1.54(4), 1.53(5), and 1.55(4) for Flack, Parsons, and Hooft, respectively. The possibility of having a racemic twin was also examined, but the modeling did not produce meaningful results. Even though a Cl atom is present in the molecule, the compound seems to be a weak anomalous scatterer. In addition, the resolution of the Mo Kα dataset is low, and the number of Friedel opposites is also low.
In summary, the structure of MMXE·HCl is correctly described in the centrosymmetric space group P21/n in which the two enantiomers, R and S, are present. On the other hand, the structure of MXE·HCl is non-centrosymmetric. No special precautions were taken during the synthesis to ensure that the product is enantiomerically pure, but nevertheless both the crystal structure from powder diffraction data and the crystal structure from single-crystal data resulted in a space group allowing only molecules of a single chirality. We must conclude that MXE·HCl crystallizes as a conglomerate.
The cyclohexanone ring (A) has a chair conformation in both compounds. The substituents in the ring are in axial (C2–C7) and equatorial (C1–O1 and C2–N1) conformation. Rings A/B (B is the phenyl ring) make an angle of 82.2(3)° in the MMXE+ molecule, whereas the corresponding angle in the MXE+ molecule is 79.0(3)°. The values of the torsion angles O1–C1–C2–N1 and O1–C1–C2–C3 were − 7.4 (8)° and −123.9 (7)° in MMXE+ and −25.0 (7)° and −138.1 (7)° in MXE+, respectively.
The torsion angles C13–O2–C9–C8 and C13–O2–C9–C10 in MMXE·HCl are 173.2(5)° and −9.9(10)°, respectively, and those in MXE·HCl are −177.2(3)° and 68.3(3)°, respectively. These values indicate that the methoxy groups are in opposite positions (see Figure 5). Aside from the orientation of the methoxy group, the MMXE+ and MXE+ cations have similar conformation, with an RMSCD deviation of 0.50 Å and a maximum deviation of 1.74 Å.
C. Intermolecular hydrogen bonds
In the structure of MMXE·HCl, as seen in Figure 6a, a cyclic motif is formed by two chloride ions and two MMXE+ moieties connected through strong N–H···Cl hydrogen bonds (N1–H1A···Cl1, 2.200 (11) Å, 155.5 (10)°; N1–H1B···Cl1, 2.133 (10) Å, 166.0 (11)°). This motif (identified as I) is described by the graph set symbol [R
![]() $ {\displaystyle \begin{array}{c}2\\ {}4\end{array}} $
(8)]
$ {\displaystyle \begin{array}{c}2\\ {}4\end{array}} $
(8)]
![]() $ {\displaystyle \begin{array}{c} Cl1\\ {}N1\end{array}} $
(Etter, Reference Etter1990; Bernstein et al., Reference Bernstein, Davis, Shimoni and Chang1995). Then, this motif is connected with another one via C14–H14A···O1 interactions, forming tapes parallel to the c-axis. The sequence can be represented by [R
$ {\displaystyle \begin{array}{c} Cl1\\ {}N1\end{array}} $
(Etter, Reference Etter1990; Bernstein et al., Reference Bernstein, Davis, Shimoni and Chang1995). Then, this motif is connected with another one via C14–H14A···O1 interactions, forming tapes parallel to the c-axis. The sequence can be represented by [R
![]() $ {\displaystyle \begin{array}{c}4\\ {}6\end{array}} $
(20)]
$ {\displaystyle \begin{array}{c}4\\ {}6\end{array}} $
(20)]
![]() $ {\displaystyle \begin{array}{c}O1, Cl1\\ {}C14,N1\end{array}} $
(II). These tapes are similar to the tapes found in other hydrochloride derivatives related to MMXE and MXE reported in the CSD database, such as APCNON (Stevens et al., Reference Stevens, Treat, Pillai, Schmonsees and Glick1973), KUQPAQ (Gao et al., Reference Gao, Gao, Wu, Li, Yang and Zhang2020), KUQPAQ01 (Hakey et al., Reference Hakey, Ouellette, Zubieta and Korter2008), QIMGIF, QIMGOL, and QIMGUR (Jurásek et al., Reference Jurásek, Králík, Rimpelová, Čejka, Setnička, Ruml, Kuchař and Kohout2018b).
$ {\displaystyle \begin{array}{c}O1, Cl1\\ {}C14,N1\end{array}} $
(II). These tapes are similar to the tapes found in other hydrochloride derivatives related to MMXE and MXE reported in the CSD database, such as APCNON (Stevens et al., Reference Stevens, Treat, Pillai, Schmonsees and Glick1973), KUQPAQ (Gao et al., Reference Gao, Gao, Wu, Li, Yang and Zhang2020), KUQPAQ01 (Hakey et al., Reference Hakey, Ouellette, Zubieta and Korter2008), QIMGIF, QIMGOL, and QIMGUR (Jurásek et al., Reference Jurásek, Králík, Rimpelová, Čejka, Setnička, Ruml, Kuchař and Kohout2018b).
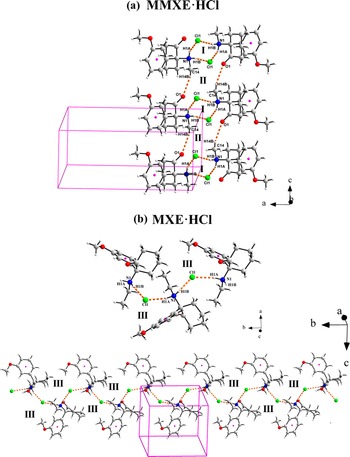
(a) Tapes formed by N1–H···Cl1 (I) and C14–H14B···O1 (II) hydrogen bonds along the c-axis in the structure of MMXE·HCl. (b) Zig-zag chains formed by MXE+ and Cl1− connected via N–H···Cl hydrogen bonds (III) in the structure of MXE·HCl.

In contrast, in the structure of MXE·HCl, there are zig-zag chains parallel to the b-axis, formed by MXE+ cations connected with Cl1− by strong N1–H1A···Cl1 (2.341 (17) Å, 164.6 (13)°) and N1–H1B···Cl1 (2.265 (18) Å, 161.6 (14)°) hydrogen bonds (Figure 6b). The arrangement can be described by the [C
![]() $ {\displaystyle \begin{array}{c}1\\ {}2\end{array}} $
(4)]
$ {\displaystyle \begin{array}{c}1\\ {}2\end{array}} $
(4)]
![]() $ {\displaystyle \begin{array}{c} Cl1\\ {}N1\end{array}} $
graph set symbol (III). This feature is less common in hydrochloride derivatives of ketamine and related compounds. It can be found in the structures of CSD entries KUQPAQ (Gao et al., Reference Gao, Gao, Wu, Li, Yang and Zhang2020), KUQPAQ01 (Hakey et al., Reference Hakey, Ouellette, Zubieta and Korter2008), and OWEYIB (Luelf et al., Reference Luelf, Reiss, Bokel and Urlacher2021) (see Figure 8). Although a cyclic motif is formed in the crystal structure of CSD entry KUQPAQ or KUQPAQ01, a zig-zag chain can also be observed. The cyclic arrangement is completed by the hydrogen bond C–H···Cl1. Furthermore, there are other weaker C–H···O interactions that contribute to the stabilization of the crystal packing. Table II contains the geometries of the hydrogen bonds, as well as π···π and C–H···π interactions of both compounds.
$ {\displaystyle \begin{array}{c} Cl1\\ {}N1\end{array}} $
graph set symbol (III). This feature is less common in hydrochloride derivatives of ketamine and related compounds. It can be found in the structures of CSD entries KUQPAQ (Gao et al., Reference Gao, Gao, Wu, Li, Yang and Zhang2020), KUQPAQ01 (Hakey et al., Reference Hakey, Ouellette, Zubieta and Korter2008), and OWEYIB (Luelf et al., Reference Luelf, Reiss, Bokel and Urlacher2021) (see Figure 8). Although a cyclic motif is formed in the crystal structure of CSD entry KUQPAQ or KUQPAQ01, a zig-zag chain can also be observed. The cyclic arrangement is completed by the hydrogen bond C–H···Cl1. Furthermore, there are other weaker C–H···O interactions that contribute to the stabilization of the crystal packing. Table II contains the geometries of the hydrogen bonds, as well as π···π and C–H···π interactions of both compounds.
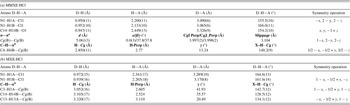
Geometry of hydrogen bonds, as well as π⋯π and C–H⋯π interactions in (a) MMXE·HCl and (b) MXE·HCl

The geometry of the contacts is defined in PLATON (Spek, Reference Spek2020) by the following parameters: a) Cg(B) is the centroid of ring B, defined in Figure 4; d = CgI···CgJ distance; α = dihedral angle between Planes I and J; β = angle between the Cg(I) → Cg(J) vector and the normal to plane I; γ = angle between the Cg(I) → Cg(J) vector and the normal to plane J; CgI Perp = perpendicular distance of Cg(I) on ring J; CgJ_Perp = perpendicular distance of Cg(J) on ring I; Slippage = distance between Cg(I) and perpendicular projection of Cg(J) on ring I. b) H···Cg is the distance of H to Cg, H-Perp = perpendicular distance of H to ring plane J, γ = angle between Cg–H vector and ring J normal, X–H···Cg = X–H–Cg angle.
D. π···π and C–H···π interactions
In addition to hydrogen bonding for MMXE·HCl, one weak π···π interaction and one weak C–H···π interaction provide connectivity between MMXE+ moieties. Figure 7a describes the geometry of the face-to-face offset π···π interaction Cg(B)⋯Cg(B) (5.061(3) Å) related by symmetry operation 1 − x, 2 − y, 2 − z. In addition, a type III (Malone et al., Reference Malone, Murray, Charlton, Docherty and Lavery1997) C4–H4B···π interaction (2.850(11) Å) between H4B and the centroid of Ring B (Cg(B)) of the molecule related by symmetry operation 1/2 − x, −1/2 + y, 3/2 − z was observed. Both interactions are extended in the ab-plane forming layers that stack along the c-axis.
C–H···π and π···π interactions in the structure of (a) MMXE·HCl and (b) MXE·HCl.

Hirshfeld surface mapped onto d norm, shape index, and curvedness for MMXE·HCl (a–f) and MXE·HCl (g–l).

On the other hand, the structure of MXE·HCl exhibits only three type V (Malone et al., Reference Malone, Murray, Charlton, Docherty and Lavery1997) weak C–H···π interactions, as seen in Figure 7b. One side of ring B displays the C3–H3A⋯Cg(B) and the intramolecular C14–H14B⋯Cg(B) contacts (3.052 Å; 3.103 Å, respectively) connecting MXE+ moieties. On the opposite side, the main interaction is the C13–H13A⋯Cg(B) contact (3.320 Å) with an MXE+ molecule related by symmetry operation −x, −1/2 + y, 1 − z. As in the case of MMXE·HCl, these weak interactions produce layers parallel to the ab-plane.
In summary, the structure of MMXE·HCl is characterized by cyclic motifs of N–H···Cl and C–H···O hydrogen bonds between MMXE+ cations and Cl− anions that produce tapes along the c-axis. Weak C–H···π and π···π interactions occur in layers parallel to the ab-plane that stack along the c-axis. For MXE·HCl, zig-zag chains parallel to the b-axis are connected by MXE+ and Cl− moieties via N–H···Cl hydrogen bonds. The weak C–H···π interactions form layers parallel to the ab-plane. These layers stack along the c-axis and are connected to each other through the zig-zag chain of hydrogen bonds.
E. Hirshfeld surface analysis
In the structure of MMXE·HCl and MXE·HCl, the HSs mapped onto d norm (Figure 8a,d for MMXE·HCl and Figure 8g,j for MXE·HCl) show two very intense red spots corresponding to the N–H···Cl hydrogen bonds. Additionally, faint red spots are observed, which are related to weak C–H···O interactions for the MMXE+ and MXE+ cations. The shape index plots (Figure 8b,h) show the red and blue areas corresponding to the C–H···π(phenyl) interactions for both compounds. However, only in MMXE·HCl, there are π···π interactions indicated by the red and blue triangles in the shape index plot and the flat area in the curvedness plot (Figure 8c). The crystal packing is very compact with almost no void space, as the volume of the surface corresponds to 97.6% and 97.7% of the unit cell for MMXE·HCl and MXE·HCl, respectively.
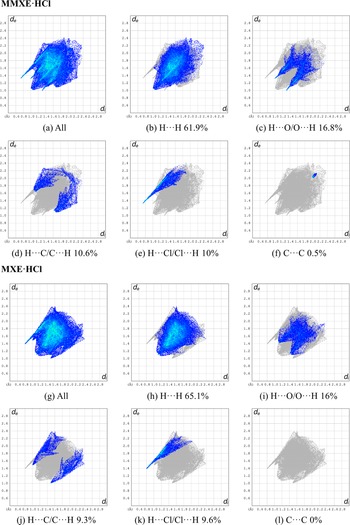
Figure 9 shows the fingerprint plots (Spackman et al., Reference Spackman, Turner, McKinnon, Wolff, Grimwood, Jayatilaka and Spackman2021) for important interactions present in the two structures. The main contribution for MMXE·HCl and MXE·HCl are the H···H (61.9% and 65.1%) and H···O/O···H (16.8% and 16%) contacts. The H···C/C···H and H···Cl/Cl···H interactions for the MMXE·HCl structure represent 10.6% and 10%, respectively, while for the MXE·HCl structure, they are 9.3% and 9.6%, respectively. The weaker C···C contact in the MMXE·HCl structure contributes 0.5%, which is completely absent in MMXE·HCl.
Fingerprint plots for interactions in MMXE·HCl and MXE·HCl.

Fingerprint plots and the contributions of the intermolecular contacts to the HS area for the Ketamine·HCl-related compounds reported in the CSD are also presented in Supplementary Figures S2 and S3.
V. CONCLUSION
The crystal structures of (RS)-methoxmetamine hydrochloride (MMXE·HCl) and (rac)-Methoxetamine hydrochloride (MXE·HCl), two arylcyclohexylamine derivatives chemically related to ketamine, have been determined from laboratory X-ray powder diffraction data, contained in the Powder Diffraction File™. The structures were determined with DASH and refined by the Rietveld method with TOPAS-Academic. They were further confirmed by DFT-D calculations. The crystal structure of MMXE·HCl is based on tapes formed by Cl− ions, which interact via hydrogen bonds with MMXE+ cations. The MMXE+ moieties in the tapes interact with other MMXE+ via π···π and C–H···π interactions. In the packing of MXE·HCl, a zig-zag chain is formed by hydrogen bonding of Cl− ions and MXE+ moieties. Layers are constructed from C–H···π interactions between MXE+ molecules. Fingerprint plots show an important contribution from H···π and H···Cl interactions, which are evident as characteristic red spots in the d norm and shape index representations.
DEPOSITED DATA
Crystallographic Information Framework (CIF) files containing the results of the Rietveld refinement and of the energy-minimization calculations for MMXE·HCl and MXE·HCl were deposited with the ICDD. The data can be requested at pdj@icdd.com. The crystal structure data were also deposited with the Cambridge Crystallographic Data Centre (CCDC 2424756 and 2424757, respectively).
CONFLICTS OF INTERESTS
The authors declare none.
SUPPLEMENTARY MATERIAL
To view supplementary material for this article, please visit http://doi.org/10.1017/S0885715625100948.
FUNDING STATEMENT
The authors thank the support received from the Cambridge Crystallographic Data Centre (CCDC) through the Frank H. Allen International Research & Education Programme (FAIRE), which allows access to the CSD for Universidad de Los Andes (Venezuela). This work used the Extreme Science and Engineering Discovery Environment (XSEDE), which is supported by the National Science Foundation under Grant No. ACI-1548562. J.W.R. acknowledges support for the Canadian Light Source from the Canadian Foundation for Innovation, the Natural Sciences and Engineering Research Council of Canada, the National Research Council Canada, the Canadian Institutes of Health Research, the Government of Saskatchewan, Western Economic Diversification Canada, and the University of Saskatchewan. M.K. and B.J. were supported by the Ministry of the Interior of the Czech Republic (Project No. VK01010212).