Introduction
Fungi are eukaryotic organisms essential to ecosystems, serving as decomposers of organic material and playing a vital role in nutrient recycling (Treseder and Lennon, Reference Treseder and Lennon2015) and bioremediation (Deshmukh et al., Reference Deshmukh2016; El-Gendi et al., Reference El-Gendi2022). Saprophytic fungi, for instance, are used in industry as cellular factories for the sustainable production of organic acids, alcohols, and other fine chemicals. For example, Penicillium chrysogenum is used in the production of penicillin (Houbraken et al., Reference Houbraken2012), whereas Aspergillus niger is used to generate organic acids such as citric (Behera, Reference Behera2020) and itaconic acid (Steiger et al., Reference Steiger2013). Fungi are also excellent sources of antifungals. For example, Echinocandins, a class of antifungal drugs to treat invasive fungal infections, were first isolated from fungi fermentation metabolites (Jiang et al., Reference Jiang2024). More recently, Ibrexafungerp, an antifungal to treat vulvovaginal candidiasis, is a derivative of enfumafungin that was discovered through natural compound screening of endophytic fungi (Vanreppelen et al., Reference Vanreppelen2023). Fungi also serve as important model organisms to study the biochemistry, genetics, and cell biology of higher-order eukaryotes. A review by McCluskey and Baker (Reference McCluskey and Baker2017) provides an excellent summary of the historical use of filamentous fungi as model organisms from the early genetic era to modern omics approaches.
Studies on the cell walls and plasma membranes of various fungal species have also led to specific industrial or biotechnological processes, particularly to produce natural products (Liu et al., Reference Liu2023). In these cases, the remodelling of the cell wall can be beneficial. For example, in Aspergillus nidulans, deletion of the α-1,3-glucan synthase genes agsB and agsA to produce ΔagsB and ΔagsAΔagsB variants improves fermentation processes by yielding more hyphal cells. Similarly, studies on Aspergillus oryzae mutants deficient in both galactosaminogalactan (GAG) and α-1,3-glucan improve the production of low-molecular-weight chemicals and proteins (Yoshimi et al., Reference Yoshimi2022). Similar to cell wall modifications, lipid membrane engineering provides an efficient and sustainable strategy to produce industrial chemicals and biofuels. For example, genetic modification of oleaginous yeasts by controlling the degree of lipid saturation or the length of the lipid chain influences membrane fluidity and integrity, which results in increased lipid titre (Mota et al., Reference Mota2022).
While most of the over 1.5 million known fungal species are beneficial, a few hundred are pathogenic and cause plant and animal diseases (Köhler et al., Reference Köhler2015). In the context of human health, invasive fungal infections (IFIs) are an increasing public health threat and cause over 1.6 million human deaths annually (Brown et al., Reference Brown2012). In 2022, the World Health Organisation, for the first time, released a list of fungi pathogens posing a threat to human health and modern medicine (Parums, Reference Parums2022). Fungal pathogens also cause billions of dollars of loss to agricultural industries by infecting livestock such as beef cattle, dairy cows, chickens, and sheep (Savary et al., Reference Savary2012; Mitchell et al., Reference Mitchell2016; Gupta and Meshram, Reference Gupta and Meshram2018). In plants, fungal pathogens are one of the dominant causative agents of plant diseases (Doehlemann et al., Reference Doehlemann2017) and can cause crop losses as high as 20% of total yield, worth billions, negatively impacting individual farmers and global food security (Avery et al., Reference Avery2019; Case et al., Reference Case2022). Thus, new antifungal agents are urgently needed to prevent and treat pathogenic fungi in animals and plants.
The main challenge to developing human antifungal agents is the similarity between mammalian and fungal cells, as both are eukaryotic and heterotrophic organisms. The core difference between mammalian and fungal cells is the presence of a cell wall and, to a lesser extent, differences in the lipid composition of the plasma membrane. Not surprisingly, most antifungal drugs work by directly interacting with or inhibiting the cell wall synthesis or the distinct plasma membrane components (Ghannoum and Rice, Reference Ghannoum and Rice1999). A review by Cortés et al. (Reference Cortés2019) highlighted (1–3)-β-D-glucan, mannan, Glycosylphosphatidylinositol (GPI)-linked proteins, chitin and chitosan, and specific cell wall enzymes such as (1–3)-α-D-glucan synthases and (1–3)-β-D-glucan remodelling glucanosyltransferases as the targets for the development of novel antifungal compounds or designing diagnostic tools for IFIs. The fungal cell wall is also implicated in antifungal drug resistance (Lee et al., Reference Lee2012; Walker et al., Reference Walker2013; Derkacz et al., Reference Derkacz2022; Revie et al., Reference Revie2022). For example, studies have shown that high concentrations of echinocandin drugs, used to treat infections caused by Candida spp. or Aspergillus fumigatus, can trigger ‘paradoxical growth’ due to increased chitin content (Lee et al., Reference Lee2012; Walker et al., Reference Walker2013). This paradoxical growth results in cell wall reinforcement, driven by changes to the signalling pathways related to cell wall integrity (Revie et al., Reference Revie2022).
Studying the composition, structure, and properties of fungal cell walls and membranes is relevant for both basic science and various applications. Key beneficial applications of fungi include developing antifungal drugs, understanding antifungal drug resistance, genetic studies, and various industrial processes. This review examines biophysical techniques (Table 1) used to study fungal cellular systems and their interactions with antifungal drugs. We begin with an overview of cell wall and membrane components in key fungal species and then use selected studies to illustrate how biophysical methods have enhanced our understanding of cell envelope heterogeneity and dynamics. The review then discusses model systems for studying drug-membrane interactions, the characterisation of relevant properties, and the challenges of bridging the gap between simple models and complex cellular systems.
Overview of commonly used biophysical techniques applied to study fungal cell walls and membranes, summarising the key capabilities, limitations, and challenges

The structure and composition of the cell wall vary among species
The cell wall, along with the cell membrane, acts as a semipermeable and selective barrier that controls the distribution of water, ions, and small nutrient molecules between the extracellular and intracellular medium (Garcia-Rubio et al., Reference Garcia-Rubio2020). The cell wall provides fungal cells with structural support and protection against osmotic and mechanical stress. Its heterogeneous and dynamic structure allows for the plasticity needed during the cell cycle, enabling cell division and proliferation. The relative abundance of cell wall components varies among species and also dictates the immunogenicity of the cell and the interactions with its host environment (Snarr et al., Reference Snarr2017).
The cell wall is an intricately structured matrix predominantly composed of polysaccharides. The main components are chitin, chitosan, glucans (β- and α-linked), mannans, and glycoproteins (Garcia-Rubio et al., Reference Garcia-Rubio2020). The inner wall, made up of β-glucans and chitin, is responsible for shape and mechanical strength. The glucans are the most abundant polysaccharides and varied components in the cell wall. Their glucose moieties are either linear or branched, networked via α or β linkages (or both), forming amorphous or microfibrillar structures. The branched β-1,6-glucan connects to chitin at the non-reducing ends through β-1,4 or β-1,2 linkages, and the outer layer glycoproteins through a GPI residue in the yeast cell walls (Kollár et al., Reference Kollár1997). The β-1,3-glucan and chitin intrachain of hydrogen bonds assemble into fibrous microfibrils, forming an exoskeleton that surrounds the cell (Gow et al., Reference Gow2017). Chitin and its deacylated form, chitosan, which is mostly found in Cryptococcus neoformans (Cr. neoformans), are the cell wall components closest to the plasma membrane (Figure 1). Chitin and chitosan are linear polymers of N-acetylglucosamine that are packed in a highly ordered manner to form crystalline, fibrous structures that are exceptionally rigid and serve as a structural backbone of the cell wall (Feofilova, Reference Feofilova2010; Kothe and Freihorst, Reference Kothe and Freihorst2016). Chitin exists in diverse polymer lengths and degrees of deacetylation, which produce respective immune responses (Tsigos et al., Reference Tsigos1999; Da Silva et al., Reference Da Silva2009). Figure 1 shows a scalar model of the inner and outer cell walls, which form a 120 nm layer (60 nm each), and the plasma membrane of Candida and Saccharomyces spp. based on recent work summarised by Gow and co-workers (Gow et al., Reference Gow2017; Gow and Lenardon, Reference Gow and Lenardon2023).
Structural composition and organisation of the most commonly studied and clinically relevant fungal spp — Saccharomyces, Candida, Cryptococcus, and Aspergillus (conidium and hyphae). Candida and Saccharomyces spp. share a very similar cell wall structure comprised of a flexible, three-dimensional network of branched β-1,3-glucan attached to β-1,6-glucan and chitin through their non-reducing ends. The major differences in the cell wall structure species are the relative quantities of β-glucans, Chitin, mannan-linked protein (mannoproteins), and the cell wall proteomes. A. fumigatus exists as either conidia or hyphae. Aspergillus conidia have a hydrophobin rodlet and a melanin outer layer. The hyphae contain α-1,3-glucan and galactosaminoglycan (GAG) in the outer cell wall and fewer glycosylated proteins. Cr. neoformans has a capsular outer layer of glucuronoxylomannan (GXM) and less galactoxylomannan (GalXM). The capsule is connected to α-1,3- glucan via peptides or other glucans to anchor to the cell wall. The chitin of Cry. neoformans is mostly deacetylated to chitosan. Cr. neoformans also have a melanin layer, speculated to be integrated into the polysaccharides around the chitin/chitosan layers. However, its exact position is uncertain. All the fungal species have similar but varying quantities of inner walls comprised of a β-1,3-glucan and chitin core. Figure adapted from Gow and Lenardon, Nature Review Microbiology, 2023.

While the inner-layer component is shared among most fungal species, the outer wall varies significantly across species (Figure 1). The outer wall variations comprise polymers and glycosylated proteins that determine the permeability and adhesiveness of the cell wall, including polysaccharides such as mannans (Candida yeasts), galactomannans and galactosaminoglycan (A. fumigatus hyphae), glucuronoxylomannan and galactoxylomannan (Cryptococcus), β-glucans, and melanin (Aspergillus conidium). Even within a given species, cell wall components vary. For example, Candida auris (Corynebacterium auris) has a distinctive mannan structure with mannose-α-1-phosphate side chain (Dakalbab et al., Reference Dakalbab2024) not present in other Candida spp. These species-specific components of the outer cell wall provide chemical and immunological heterogeneity, critical for the interaction of fungal cells with their host environment (Gow and Lenardon, Reference Gow and Lenardon2023). Embedded into the cell wall layers are also various glycoproteins implicated in pathogenicity (Erwig and Gow, Reference Erwig and Gow2016; Moon et al., Reference Moon2025).
Biophysical characterisation of the fungal cell wall
Before the recent advances in microscopy, methods for characterising the cell wall were mostly based on extracting or chemically treating the corresponding components, separating them using chromatography, and identifying them using spectroscopy techniques. While this enabled identification of components, it does not provide information on their three-dimensional arrangements and dynamic nature, which is critical to the cell wall and its response to external factors. The advancement of biophysical techniques and their application to fungal cells, particularly in microscopy, spectroscopy, and electron tomography, has facilitated an improved detection, quantification, and probing of sugars and lipids, feasibly even in live and intact cells.
Traditionally, cell wall components have been chemically identified and quantified based on the relative abundance of individual components after extraction from cells through chemical treatments and/or after enzymatic digestion. Most often, after mechanical milling, the resulting cell homogenate undergoes differential centrifugation to separate the cell wall and membrane components from the cytosolic components (Pitarch et al., Reference Pitarch2008). The polysaccharides are further separated for analysis using chemical fractionation (Wang et al., Reference Wang2022), typically yielding amorphous alkali-soluble and fibrillar alkali-insoluble components that require further purification. Traditionally, analytical techniques such as high-performance liquid chromatography (HPLC), ultra performance liquid chromatography (UPLC), gas chromatography–mass spectrometry (GC–MS), Fourier transform infrared spectroscopy (FTIR), and nuclear magnetic resonance (NMR) were then used to identify and quantify the different components.
The chromatography (gas or liquid) technique provides the ability to separate, identify, and quantify analytes, allowing for a complete characterisation of the monosaccharide contents of fungal cell wall polysaccharides. In gas chromatography, the monosaccharides hydrolysed from the polysaccharides are subsequently derivatised into O-methyl-trimethylsilyl (TMS) derivatives. The detection of these derivatives is based on the difference in the boiling point of each TMS derivative, which is determined using a temperature gradient developed in the gas chromatograph. MS is used as a complementary technique to determine the monosaccharide linkages of partially methylated alditol acetate derivatives or partially hydrolysed polysaccharides (Millette et al., Reference Millette2023). For liquid chromatography, monosaccharides are separated based on their polarity and identified by a fluorescent ultraviolet (UV) tag. The total polysaccharide content of biomass can be quantitatively assessed by comparing the concentration of eluted samples to a standard at a specified retention time. For example, the total contents of glucosamine and N-acyl glucosamine from the biomass of zygomycetes have been quantitatively analysed with HPLC (Zamani et al., Reference Zamani2008; Mohammadi et al., Reference Mohammadi2012). Similarly, three monosaccharides (glucose, mannose, and xylose) of a polysaccharide extracted from golden needle mushrooms have also been characterised with HPLC (Wu et al., Reference Wu2010). HPLC is often combined with other characterisation techniques such as NMR and FTIR for extensive qualitative and quantitative analysis (Prieto et al., Reference Prieto2007; Corradi da Silva et al., Reference Corradi da Silva2008; Aimanianda et al., Reference Aimanianda2009).
IR spectroscopy relies on unique, characteristic bands to determine the presence of a component in a complex mixture of cell wall components. For example, chitin spectra have bands matching 970 and 810; near 3265, 3105, 1655, 1620, 1550, 953, and 392, and a distinctive 1590 cm−1 for chitosan. The 1370, 1120, 1040, and 900 cm−1 show the presence of mannan, chitin, and glucans, respectively. As these bands can be affected by impurities such as proteins, lipids, or other cell components, IR is particularly useful to evaluate the efficacy of an extraction method. The efficiency of this technique is, however, dependent on the nature of the components in the sample, the relative proportion of these components, and the degree of crystallinity. For example, detecting β -glucan in a mixture with mannan and α-glucans in a mixture with cellulose does not give a very accurate result. It is also limited to determining the degree of branching of branched polymers containing more than one type of linkage, but it is efficient in identifying the nature of major types of linkages (Michell and Sourfield, Reference Michell and Sourfield1970). While analytical techniques were critical in the early studies of identifying and quantifying sugars and lipids in fungal cells, these techniques cannot provide any information on how the individual components assemble into the complex and heterogeneous structure or how individual components contribute to the plasticity of the cell membrane or cell wall.
Solid-state nuclear magnetic resonance (ssNMR) spectroscopy and single-cell technologies such as atomic force microscopy (AFM) are non-destructive and allow cell wall components to be studied in situ (Table 1). For instance, a recent study applied ssNMR and magic angle spinning dynamic nuclear polarisation (MAS-DNP) on intact A. fumigatus cells to provide the first atomic-level description of polymer rigidity and physical packing of the cell wall component in mould species. The results showed that chitin and α-1,3-glucan form a rigid-phase hydrophobic scaffold base that is engulfed by a flexible mobile-phase hydrated matrix of distinctly linked β-glucans, overlaid by a dynamic layer of glycoproteins and α-1,3-glucan (Kang et al., Reference Kang2018) (Figure 2). Compared to previous studies, this updated model of the A. fumigatus cell wall identified which of the components determine cell wall rigidity. Based on new hydration and mobility data, the authors showed that chitin and α-1,3-glucan form the rigid scaffold, a role that was previously attributed to β-1,3-glucan. The study also detailed the many intermolecular interactions between the main carbohydrates and their roles in the structure and rigidity of the cell wall. This detailed model of the A. fumigatus cell wall provides critical reference information for future studies focused on how external stresses or drugs affect the cell wall. The molecular-level detail will also be critical to building in-silico models of fungal cell walls for computational studies.
Diagram showing the rigid and mobile inner domain and the mobile outer domain of the A. fumigatus cell wall as determined by ssNMR and magic angle spinning dynamic nuclear polarisation. Intermolecular interactions between the carbohydrate components are indicated by dashed circles and were determined from NMR restraints. Average hydration levels (in %) and average 1H-T1ρ relaxation times (in milliseconds) are indicated. Figure taken from Kang et al, Nature Communications, 2021, published under the Creative Commons Attribution 4.0 https://creativecommons.org/licenses/by/4.0/.

In a subsequent paper (Chakraborty et al., Reference Chakraborty2021), the same authors used ssNMR and functional genomics to investigate the role of the four major structural polysaccharides, either α-1,3-glucan, chitin, galactomannan (GM) or GAG, in the morphology of the A. fumigatus cell wall. For this, the authors compared the polysaccharide composition, the spatial organisation and dynamics of cell wall components in the wild-type (parental) A. fumigatus strain and its mutant strains lacking either α-1,3-glucan, chitin, GM, or GAG following the homogeneous labelling of culture samples with 13C-glucose and 15N-NaNO3 of the hyphal cell walls of the parental α-1,3-glucan and or chitin. The rigid phase molecules, which are made up of chitin and α-1,3-glucan, were identified by the dipolar-based 1H-13C cross-polarisation, while the mobile phase molecules comprising hydrated glycoproteins and α-1,3-glucan were also shown by the short recycle delay in 13C direct polarisation. The study confirms previously reported compensatory responses to the lack of a specific cell wall component due to gene deletion. For example, the lack of α-1,3-glucan and chitin caused an increase in β-1,3-glucan and GAG contents, respectively. Using a combination of alkali-solubility and information from 13C–T1 relaxation times, the authors were able to construct a model of the mobile and rigid phase of the A. fumigatus cell wall and provided new insight into the formidable plasticity of the fungal cell wall and the capacity of the pathogen to adapt to the lack of essential cell wall components.
ssNMR has also been used to provide pivotal insights into the accumulation of melanin within the fungal cell wall, revealing its structural integration and its role in antifungal resistance. Incorporation of melanin is previously known to be facilitated by interactions with cell wall polysaccharides, forming a matrix that relies predominantly on chitin (Banks et al., Reference Banks2005). ssNMR analyses, however, support a composite scaffold composed of both chitin and α-1,3-glucans that accommodates the aromatic nature of melanin pigments, enhancing the hydrophobicity of the cell wall, thereby limiting permeability and contributing to antifungal resilience (Kang et al., Reference Kang2018). These insights highlight the unique capacity of ssNMR to explore biopolymer dynamics and hydration in situ, offering exceptional resolution of cell wall architecture. Recent advances are addressing the challenges of resonance assignments posed by the high structural complexity and heterogeneity of cell wall polysaccharides (Zhao et al., Reference Zhao2022). In a combined ssNMR, transmission electron microscopy (TEM), and fluorescence/DIC study, the ssNMR analysis showed that deletion of Aah1 and Aah3 enzymes from S. pombe cells led to a notable loss of α-1,3-glucans across rigid and mobile wall domains, along with a marked increase and polymorphic expansion of β-1,3-glucans. These molecular changes were accompanied by depletion of the galactomannan-rich outer layer and redistribution of β-1,6-glucan, indicating extensive cell wall remodelling. Consistently, TEM and fluorescence/DIC microscopy confirmed cell wall thickening, enhanced septal β-glucan staining, and pronounced defects in cell morphology and separation (Jacob et al., Reference Jacob2025). Beyond revealing structural features inaccessible to conventional chemical methods and a possibility to study the cell wall remodelling resulting in wall-related enzymatic manipulation, ssNMR also provides the possibility to understand molecular mechanisms of cell remodelling and influences of melanisation (in Cr. neoformans) under physiological and environmental stresses, a key defensive adaptation linked to fungal drug resistance in situ. A structural-mechanistic understanding of this phenomenon could lead to new therapeutic and biotechnological advantages in fungi studies.
AFM has emerged as a significant technique to observe living microbial cells at nanometre resolution and characterise the topography of cell surfaces in near-native environments (Dufrêne, Reference Dufrêne2014). While AFM suffers from poor temporal resolution, the method enables the mechanical manipulation of surface components and the measurement of mechanical properties in live cells under physiologically relevant conditions. For example, Gibbs et al. (Reference Gibbs2021) determined the polarised growth zones and their mechanical properties of Schizosaccharomyces pombe (S. pombe). A nonuniform ridge in the cell wall was observed to have an uneven accumulation of cell wall constituents, indicating cell wall remodelling and a cell morphological transition during cytokinesis. Additionally, regions of the cell poles were found to exhibit more elastic cell wall properties, albeit with decreased cellular rigidity, per the relative biophysical characteristics of the S. pombe cell body and cell pole. A review by Dufrêne (Reference Dufrêne2010) summarises several applications and the advancement of AFM for studying the fungal cell wall. Likewise, Bhat et al. (Reference Bhat2021) reviewed the use of AFM to understand processes related to pathogenicity, including quantifying host-pathogen interactions and pathogen adhesion, understanding the roles of fungal and bacterial cell walls in pathogenicity, characterising viral infectivity, and the coupling of AFM with optical microscopy for elucidating microbial pathogenicity. AFM can also be combined with IR or Raman spectroscopy, enabling chemical characterisation of components (Quilès et al., Reference Quilès2017; Barroso-Solares et al., Reference Barroso-Solares2025). A combined approach of AFM and ssNMR could provide a more detailed insight into the activity and spatial regulation of cell wall synthases during cytokinesis in fission yeast. This approach has the potential to elucidate the synthesis and organisation of nascent cell wall material at the division site, offering a mechanistic understanding of how polarised cell proliferation drives septum formation.
Cryo-electron microscopy (Cryo-EM) is extensively used in structural biology, and recent advancements in its detectors and image analysis have significantly improved the resolution of macromolecules (Latgé and Wang, Reference Latgé and Wang2022). While Cryo-EM has been used to study fungal proteins, the technique remains underexplored in the study of fungal cell walls and membranes. The benefits of the technique for studying cell walls were highlighted in a study by Matias and Beveridge (Reference Matias and Beveridge2006) that focused on the composition of the bacterial cell wall of Staphylococcus aureus (S. aureus). For the first time, the study found robust evidence of a periplasmic gap in S. aureus by identifying a bipartite bacterial cell wall composed of a lower density of 16-nm inner wall zone and a larger density of 19-nm outer wall zone. Cryo-EM has also been useful in studying the topography of the plant cell wall (Sarkar et al., Reference Sarkar2014) and in understanding the cell wall destruction by the supramolecular assemblies of the antibiotic teixobactin in B. subtilis (Hurst et al., Reference Hurst2021). These studies demonstrate the applicability of Cryo-EM to studying cell wall structures that, in the future, might be extended to fungal cells.
Cryo-electron tomography (Cryo-ET) also offers a powerful window into providing an ultrastructure of the fungal cell wall by visualising the 3D organisation of the cytoskeletal elements. With Aspergillus nidulans, a Cryo-ET review shows a central hub for targeted delivery of vesicles into the cell wall components, guided by intricate networks of actin filaments and microtubules (Hohmann-Marriott et al., Reference Hohmann-Marriott2006). The review also highlights how Cyro-ET uncovers the nanoscale choreography behind fungal morphogenesis and cell wall architecture. In another study using an optimised 3D ET on C. albicans, researchers used high-resolution imaging paired with advanced computer vision algorithms to build a nanoscale model that accurately mapped the thickness and composition of each layer in the fungal cell wall (Lenardon et al., Reference Lenardon2020). This approach has given a detailed architecture of both inner and outer cell wall layers (as previously described). The combination of ET imaging and computational modelling enabled precise visualisation of these features, advancing our understanding of fungal cell wall organisation on a nanometre scale.
Interactions of the cell wall with antifungal drugs
The fungal cell wall is the first point of contact for antifungal drugs. Current antifungal drugs target either the membrane itself (polyenes), membrane proteins that synthesise ergosterol (azoles), cell wall components (echinocandins), or cell-internal components (flucytosine, griseofulvin). Consequently, these drugs have to permeate the cell wall, and imaging techniques, combined with biochemical and molecular biology experiments, have been used to show the molecular details of this permeation process. Imaging techniques have been useful to understand how antifungal drugs affect the morphology of the fungal cell walls. Images of cell wall indentation, shrinkages, grooves, cell elongation, and atypical cell structures are observed in exposure to antifungal drugs. Besides spatial resolution, the temporal resolution often poses a challenge when studying the response of the cell surface to antifungal drugs.
In a seminal study, Walker et al. (Reference Walker2018) used TEM to demonstrate that AmBiosome, the liposomal formulation of polyene antifungal drug Amphotericin B, moves through the cell walls of both C. albicans and Cr neoformans as an intact liposome despite their outer layer differences and liposomal large size (60 to 80 nm) (Figure 3). Interestingly, liposomes devoid of the drug very rarely entered the inner β-glucan–chitin layer, thus highlighting the cell wall’s remarkable sensitivity in responding to its environment. In another TEM study, Thevissen et al. (Reference Thevissen2012) revealed that the plant defensin RsAFP2 induces mechanical stress on the fungal cell wall. The study highlights the morphological alterations on the cell wall of fungal cells by the peptide with a signature of an unusual budding profile showing broadened neck size, altered cell shape, and atypical cell wall structure that likely contribute to cell death. A study by Grela et al. (Reference Grela2023) examined the synergistic effect of bovine serum albumin (BSA) with AmB on C. albicans. Grela et al.’s work combined scanning electron microscopy (SEM), fluorescence microscopy and spectroscopy with biochemical analysis to show that the synergy is likely based on BSA increasing AmB uptake across the cell wall. The authors showed that treating C. albicans cells with the AmB-BSA complex alters cell morphology, including shrinkages, indentations, and broadening with deformed cell walls (Figure 4a), while fluorescence lifetime imaging microscopy revealed that the AmB-BSA complex is more efficient in crossing the cell wall barrier than AmB alone (Figure 4b). The study also establishes that the cell wall acts as a barrier and defensive mechanism of the C. albicans cells, which traps a portion of the AmB molecules, thereby reducing the therapeutic effect. The exact cell wall component and AmB interaction remain unknown. The presence of extracellular vesicles, containing ergosterol in the cell wall, and melanin (as shown in Cr neoformans (Ikeda et al., Reference Ikeda2003)) are viable agent involved in the AmB interaction and the entrapment in the cell wall.
Transmission electron microscopy images display intact sphere-shaped and electron-dense AmBisome (~60 nm) as shown by arrows at the inner and outer cell walls (a, b, c, d, and e) and the surface of the plasma membrane (f) of C. albicans. AmBisome migration through the cell wall to the plasma membrane is indicated by AmBisome particles at the plasma membrane surface with no AmBisome accumulation at the outer cell wall or the base of the microfibrillar layer. Figure taken from Walker et al., mBio, 2018 published under the Creative Commons Attribution 4.0 https://creativecommons.org/licenses/by/4.0/.

Scanning electron microscopy (SEM) and fluorescence lifetime images of C. albicans cells showing the effects of AmB, BSA, and AmB-BSA complex. a) SEM images of C. albicans incubated with PBS, DMSO, and BSA (upper row), AmB (mid-row), and AmB-BSA complex (lower row). b) Fluorescence lifetime-based images of cells before and after exposure to AmB and AmB–BSA (upper row), followed by images based on fluorescence anisotropy values, and images based on the amplitude of the short-lifetime fluorescence component (0.56 ns) ascribed to AmB (third row). The fluorescence lifespan analysis of the cells displayed in the upper panels was plotted as shown at the bottom. The components of fluorescence lifespan were determined as follows: 0.56 ns assigned to AmB (blue), whereas the cell autofluorescence was 2.8 ns (green) and 8.8 ns (red). Figure made from images obtained from Grela et al., The Journal of Physical Chemistry, 2023, published under the Creative Commons Attribution 4.0 https://creativecommons.org/licenses/by/4.0/.

AFM is also routinely used to determine the interactions of antifungal compounds. Awassa et al. (Reference Awassa2024) demonstrated that zinc-based nanoparticles elicit their antifungal activity via ‘sticky surface contact’, which causes significant cell wall damage and, consequently, cell death. Analysis of the adhesion forces between the nanoparticles and AFM-tipless cantilever attached with C. albicans through AFM-based single-particle and single-cell force spectroscopy measured higher frequencies and higher force magnitudes that correlated to the strong adhesive interactions of the nanoparticle, leading to severe cell wall damage compared to the negative controls. AFM imaging and force curves have also been used to study the interaction of caspofungin with C. albicans. The drug-induced cell wall remodelling, characterised by increased chitin and decreased β-glucan content, does not alter the overall morphology and cell division. The changes affected the adhesive properties (known property exhibited by C. albicans cells with the involvement of β-glucan (Tsai et al., Reference Tsai2011)) of the cell surfaces, which are known to be related to pathogenicity (Formosa et al., Reference Formosa2013).
Fluorescence microscopy enables live-cell imaging of specific structural details and molecular processes in real-time. In studies of fungal cells, fluorescently tagged antifungals with conventional microscopy have enabled a better understanding of how drug uptake, temporal subcellular localisation, and membrane perturbation influence virulence factors, drug efficacy and resistance (Jospe-Kaufman and Fridman, Reference Jospe-Kaufman and Fridman2025). However, conventional microscopy resolution is limited by light diffraction, preventing visualisation of nanoscale cellular processes. Super-resolution fluorescence microscopy, such as structured illumination microscopy, super-resolution through radial fluctuations (SRRF), stimulated emission depletion (STED), photoactivated localisation microscopy (PALM) and stochastic optical reconstruction microscopy (STORM), overcomes this constraint, with each applying unique approaches to visualise subcellular structures at nanoscale resolution (Huang et al., Reference Huang2009; Sahl et al., Reference Sahl2017). For instance, in C. albicans, outer mannoproteins can mask β-glucan, reduce immune detection and limit access of glucan-targeting drugs like caspofungin. When wall damage or signalling defects unmask β-glucan, exposure increases, enhancing immune responses and drug susceptibility (Hameed et al., Reference Hameed2021; Wagner et al., Reference Wagner2023). While conventional optical microscopy can show overall β-glucan exposure, it cannot precisely resolve its spatial geometry or the structural changes associated with the unmasking. Using dSTORM, β-glucan organisation in the cell wall of C. albicans before and after antifungal treatment has been resolved. The study demonstrates that super-resolution imaging can reveal the detailed cellular architecture of fungal cells, offering new insights into their pathogenic mechanisms. However, super-resolution microscopy is, in some cases, constrained by potential phototoxic effects in live cell imaging (Prakash et al., Reference Prakash2025).
ssNMR is another non-invasive, whole-cell technique that is mostly complementary to imaging methods. Safeer et al. used 1H-detected fast MAS ssNMR spectroscopy, a method with increased spectral resolution compared to other ssNMR methods, to refine their previous models of proteins in the Sericopelma commune cell wall and demonstrated that components in the outer and inner layers of the cell wall respond differently to the treatment with metal ions or antifungal peptides (Safeer et al., Reference Safeer2023). In another ssNMR study, C. albicans and C. auris share a similar basic wall architecture yet respond differently to caspofungin. The cell wall remodelling caused by caspofungin treatment showed thickened and chitin–glucan reorganisation in C. albicans, whereas C. auris, particularly resistant strains, showed minimal remodelling alongside increased core dehydration, highlighting the role of species-specific wall architecture in antifungal drug interactions and resistance (Dickwella Widanage et al., Reference Dickwella Widanage2025). Another ssNMR study by Bishoyi et al. (Reference Bishoyi2025) shows that the antifungal peptide cathelicidin-2, which acts via a membrane-altering mechanism, first localises in the cell wall of A. fumigatus, where the peptide showed specific interactions with certain cell wall components. In the mobile domain of the cell wall, the peptide interacts with galactosaminogalactan and polysaccharides and causes an increase in the water content in the rigid domain, thus causing reorganisation and an overall increase in the cell wall water content. Earlier fluorescent microscopy studies of the peptides against C. albicans showed no evidence of cell wall interactions, but the peptides were indicated to localise in the membrane with intracellular effects (Ordonez et al., Reference Ordonez2014). For C. albicans and A. fumigatus, having varying cell wall outer components, the interaction of the peptide in the cell wall could also vary, but further ssNMR study on the cell wall interaction of C. albicans with the peptide is required to ascertain this hypothesis. A review by Lima et al. (Reference Lima2019) highlights the fungal species’ mutable responses to antifungal interactions and resistance, which can be attributed to the differences in the cell wall components and synthesis.
Other structural characterisation techniques, such as ultra-small-angle and small-angle X-ray scattering (USAXS/SAXS), have been applied to whole bacterial cell studies. USAXS/ SAXS can provide comprehensive information on structural elements, such as the thickness and lattice parameters of the cell surface layers on intact, hydrated cells, non-destructively, and the changes caused to the cell when exposed to different stresses, such as pH, oxidation, and surfactants (Sekot et al., Reference Sekot2013; Duarte et al., Reference Duarte2022). In Duarte et al. (Reference Duarte2022) study, the model fit to the scattering data showed changes in the overall Gram-positive bacterium, Staphylococcus epidermidis, size and cell wall thickness induced by the different stresses, including pH and hydrogen peroxide. As fungal cell and wall structural changes are linked to drug efficacy and resistance, this approach, combined with microscopy and ssNMR, is viable to provide a comprehensive approach to understanding how antifungal molecules and stressors affect fungal cell and wall structures.
Structure and composition of the fungal plasma cell membrane
Like in other eukaryotic cells, the plasma membrane of fungal cells consists of a phospholipid bilayer with integral membrane proteins. The main components of the lipid bilayer comprise 50–75% glycerophospholipids of the total lipid content, along with varying amounts of sphingolipids and sterols (Sant et al., Reference Sant2016). The most dominant glycerophospholipids as determined in Saccharomyces cerevisiae (S. cerevisiae) include zwitterionic phosphatidylethanolamine (PE) (23%) and phosphatidylcholine (PC) (22%) lipids, and negatively charged lipids such as phosphatidylserine (PS) (2.8%), phosphatidic acid (PA) (13%), phosphatidylinositol (PI) (32%), phosphatidylglycerol (PG) (<0.50%), lyso-glycerophospholipids (Lyso-GPL) (5.5%), and cardiolipin (1.5%) of total mol% of phospholipids (Figure 5). Compared to mammalian plasma membranes, the fungal plasma membrane is enriched in PA and PI lipids, providing them with a higher surface charge (Ejsing et al., Reference Ejsing2009; Klug and Daum, Reference Klug and Daum2014).
Structures of major lipid species found in the fungal cell membrane. Neutral and charged glycerophospholipids account for 50–75% of lipids, most of which are also found in other eukaryotes. Ergosterol is the main sterol found in fungal membranes and is unique to fungal species. While the glycosphingolipid is found in most eukaryotes, many of the inositol-phosphoryl-ceramide lipids are unique to fungi.

Sphingolipids found in fungal cells include the neutral sphingolipids glucosylceramide (GlcCer) and galactosylceramide (GalCer), and the negatively charged inositol-phosphoryl-ceramide, mannosyl-inositol-phosphoryl-ceramide, and mannosyl-diinositol-phosphoryl-ceramide (M(IP)2C) (Mota Fernandes and Del Poeta, Reference Mota Fernandes and Del Poeta2020) (Figure 5). In contrast to the glycerophospholipids, the acyl chains of sphingolipids are usually saturated, resulting in a higher lipid packing and membrane thickness (Ferraz et al., Reference Ferraz2021). Besides changing the structure of the bilayer, sphingolipids are essential for signal transduction, endocytosis, stress responses, and virulence (Heung et al., Reference Heung2004).
The main sterol found in fungal cells is ergosterol (Figure 5). Equivalent to cholesterol in mammalian cells, ergosterol regulates membrane fluidity and permeability and is also involved in the sorting of proteins and the activity of membrane-associated enzymes (Choy et al., Reference Choy2023). Sterols and sphingolipids aggregate into dense microdomains that influence the mechanical characteristics of the plasma membrane. The changes in fungal membrane lipid microdomain change the membrane in response to changes in environmental conditions, also influencing fungal pathogenicity (Rella et al., Reference Rella2016). Variations in the relative abundance of phosphatidylinositol (PI), sphingolipids, and sterols across fungal species have resulted in strong pathogen selectivity, particularly for mammalian cells (Rautenbach et al., Reference Rautenbach2016). However, the role of specific lipids in pathogenicity and the membrane biophysical properties is still poorly understood. More extensive studies about characterising the lipid composition and its effect on the biophysical properties, the virulence processes, and virulence factors in pathogenic fungi are needed. Insights from such studies may open new strategies for developing new therapeutic options (Rella et al., Reference Rella2016). The main challenges to dissect the roles of specific lipids in pathogenicity include the identification of lipids in the outer vs inner leaflet of the plasma membrane and detecting and quantifying minor and/or transient lipid species through lipidomics. The lack of fluorescent probes to label and thus track specific lipid classes or species in live cells, and the rapid nature of lipid remodelling during infection, make it challenging to pinpoint cause-and-effect as well.
Biophysical characterisation of the fungal plasma membrane
The plasma membrane characterisation has also largely depended on solvent extraction, chromatographic separation, and spectroscopic identification of lipid species. Limitations in this approach to understanding the three-dimensional organisation and dynamics of membranes have restricted insight into their functional and adaptive roles. Advances in quantitative microscopy now enable direct studies of membrane structure and dynamics within live intact cells. Such approaches provide, to an extent, spatially resolved insights into the lipid organisation under near-physiological conditions.
The fungal plasma membrane is complex and dynamic, and characterising its composition, structure and physicochemical properties requires a multipronged approach. Contemporary protocols, thus, apply a joint approach involving biochemical and genomic approaches to manipulate cells with biophysical characterisation techniques (Lombardo et al., Reference Lombardo2020; Zhang et al., Reference Zhang2021). Lipid profiling requires extraction, and the earliest membrane extraction methods were developed by Folch and Bligh & Dyer in the 1950s (Folch et al., Reference Folch1951; Bligh and Dyer, Reference Bligh and Dyer1959), based on chloroform and methanol. Due to safety and environmental concerns, cleaner solvent methods have been developed that provide a comparative yield (Saini et al., Reference Saini2021). The critical hurdles to overcome in lipid extraction are the ability to obtain a high yield and absolute separation of non-lipid constituents from the homogenate. In particular, the latter can impact the results of lipid profiling by MS. For example, in identifying lipid variations due to metabolic changes linked to cellular homeostasis and disease aetiology, impure samples can lead to underestimation and misinterpretation of the lipid species. To overcome these hurdles, numerous extraction strategies have been established (Saini et al., Reference Saini2021). The common laboratory methods include osmotic shock extraction, enzyme-assisted extraction, supercritical CO2 extraction, microwave irradiation-assisted extraction, methyl-tert-butyl ether (MTBE) extraction, and ultrasonic extraction. For efficiency, rapid, and pure lipid recovery for MS lipid profiling, MTBE extraction is recommended (Matyash et al., Reference Matyash2008; Ghorasaini et al., Reference Ghorasaini2021).
Lipid molecular species in lipid crude extracts from cell membranes can be characterised using conventional or non-conventional approaches. The conventional approaches include chromatographic and gravimetric techniques, while the non-conventional approaches include spectroscopic and colourimetric techniques. Chromatographic quantification is either thin-layer chromatography, gas chromatography, or HPLC. For the spectroscopic qualitative analysis and structure elucidation, IR, NMR, Raman, fluorescence, and dielectric approaches are widely used. Both conventional and non-conventional approaches have been extensively reviewed by Patel et al. (Reference Patel2019).
Lipid profiling has been essential for identifying disease biomarkers, biological and evolutionary studies, understanding cell metabolic processes, and mechanisms of drug efficacy and resistance. In a non-targeted fungal lipidomics study, UPLC–Orbitrap MS analysis of the antifungal mechanism of encapsulated nanoemulsions of eugenol, carvacrol, and cinnamaldehyde against Penicillium digitatum, showed endogenous lipid profile changes. The study enabled a comprehensive profiling of plasma-membrane lipid remodelling in response to the activities of the antifungal compounds. It also provided insight into the functional perturbation of fungal lipid homeostasis and supported the identification of lipids associated with nanoemulsion-mediated antifungal activity (Yang et al., Reference Yang2022). Using a high-throughput mass spectrometry-based lipidomics approach, Singh et al. (Reference Singh2010) profiled the lipidomes of multiple Candida spp. Major phospholipid classes were identified to be present in broadly comparable levels across the Candida spp, but each exhibits a distinct molecular lipid-species signature. In particular, all the major phospholipids, such as PC, PE, PI, PS, PA, and PG, showed relatively comparable quantities but different molecular species. A comparative lipidomics of drug-resistant and drug-susceptible C. auris strains using an electrospray ionisation tandem mass spectrometry approach also showed changes among lipid species associated with the drug-resistant strains. The drug-resistant strain indicated higher levels of phosphoglycerides, an increased acyl-chain unsaturation index, accumulated 16:1- and 16:0-diacylglycerols, reduced 18:1- and 18:0-triacylglycerols, and higher levels of free ergosterol. Consistent with these strain-level differences, principal component analysis of this result against previously reported Candida spp lipidomes identified a distinct lipid signature for C. auris (Shahi et al., Reference Shahi2020). For a human pathogenic Cryptococcus spp study, a combined lipid profiling approach (Phosphoglycerides by 31P-NMR, sphingolipids by LC-ESI-MS/MS, and sterols by GC–MS) also showed strain- and species-specific differences in the lipid compositions, as well as variations in acyl-chain length and saturation. The distinct lipidomic signatures reveal that the pathogenic Cryptococcus strains and species undergo lipid compositional shifts that are predicted to modulate membrane organisation, stress adaptation, and virulence-associated growth under host-relevant conditions (Singh et al., Reference Singh2017). These studies highlight and emphasise that species-specific membrane organisation constitutes a defining molecular feature that can be linked to the adaptive capacity and drug resistance of fungal cells. Complementary whole-cell fungal biophysical studies to show these observations of lipidomic variations to membrane properties, such as lipid packing and domain formation, which influence membrane fluidity, are required to distinguish between species-specific membrane virulence mechanisms and molecule interactions.
With the growing recognition of the role of fungal plasma membrane lipid changes in virulence and biomolecular interactions, the use of MTBE and other organic solvents in lipidomic studies poses environmental risks. Novel or optimisation of greener lipid extraction methods are needed to achieve high lipid recovery and yield a highly precise and sensitive approach for more reliable lipidomics studies. Current challenges in fungal lipidomics include unique identification of structural variants and minor, fungal-specific lipids due to a lack of reference data.
Interaction of antifungal compounds with fungal cell membranes
Identical to all other biological membranes, the fungal cell membrane is a complex and intricately balanced environment. The cell relies on an intact lipid bilayer to keep the semipermeable nature of the membrane and thus control gradients of ions, protons, and nutrients across cellular compartments and the intra- and extracellular environment. Disruption of lipid bilayer integrity results in cellular stress and, in severe cases, cell death. Membrane disruption is one of the mechanisms by which AmB kills fungal cells (Kamiński, Reference Kamiński2014; Cavassin et al., Reference Cavassin2021) and also underpins the mechanism of action of many antifungal peptides (Fernández de Ullivarri et al., Reference Fernández de Ullivarri2020).
Studies of fungal cell membrane permeability typically involve treating fungal cells with an antifungal agent and assessing permeability using fluorescence dye staining or by labelling the membrane or a cytoplasmic peptide for imaging or fluorescence measurement. For example, the antifungal effect of curcumin on the C. albicans cell membrane was investigated using fluorescence spectroscopy with the lipophilic dye DiBAC43. Because DiBAC43 can enter cells only when membrane permeability increases, this approach helped establish that the antifungal mechanism involves membrane disruption or other forms of physical membrane damage (Lee and Lee, Reference Lee and Lee2014). Similarly, fluorescence microscopy experiments use membrane-impermeant fluorescent dyes such as propidium iodide (PI), SYto-9, and Sytox Green (SG) to assess membrane integrity (Gonzalez et al., Reference Gonzalez2002; Muñoz et al., Reference Muñoz2006). Lee and Kim (Reference Lee and Kim2022) used confocal laser scanning and fluorescence microscopy with PI to study myricetin-treated C. albicans cells for evidence of potential antifungal effects. Based on cell debris observed in the microscopy images, the authors concluded that myricetin damages the cell wall. In contrast, AmB-treated cells showed red fluorescence with an intact shape, indicating an exclusive defect of the cell membranes. Antifungal effects can also be induced by destabilising membranes via altering fluidity. Rodríguez-Moraga et al. (Reference Rodríguez-Moraga2023) used molecular dynamics (MD) simulations to study the effects of rhamnolipids, antifungal compounds naturally produced by Pseudomonas and Burkholderia bacteria. Rhamnolipids partition into the bilayer, where they form strong interactions with ergosterol, creating local microdomains and overall increasing membrane fluidity and reducing dipole potential, consistent with the biosurfactant properties of rhamnolipids. The simulations also showed that the rhamnolipids inserted more readily into PE-containing membranes compared to the PC lipids study, highlighting the importance of lipid composition when studying antifungal compounds.
SEM/TEM on the whole cell can show a structural change in the plasma membrane induced by the antifungal agents. For example, cell morphologies visualised with TEM illustrated that the antifungal peptides C4, cathelicidin LL-37, human defensin HNP-1, and plant defensin HsAFP-1 deformed intracellular organelles and internalised cell membranes but conserved cell shape. The disruption of cell membranes and cytosolic organelles, but not the cell wall of C. albicans, implies that these antifungal peptides affect the plasma membrane and cytosolic organelles (Menzel et al., Reference Menzel2017). In a combined microscopy, SEM and TEM study, Fusarium oxysporum cells treated with the antifungal peptide P852 or AmB, showed images of uneven cell surface, shrunken cytoplasm membrane detached from the cell wall and irregular organelle structures. These peptide-induced changes in membrane structure are indicative of a membrane-disruptive mechanism similar to that of AmB (Han et al., Reference Han2019).
Advanced high-resolution imaging techniques provide a nanoscale approach to understanding the fungal plasma membranes and their response to antifungal molecules. For instance, STED microscopy and lipid-rich structures labelling with Nile Red, a nanoscale architecture of live C. albicans, has been achieved with ~85 nm lateral resolution. The approach resolved subcellular structures, such as the plasma membrane, mitochondria, lipid droplets, and organelle interactions, and allowed tracking of lipid droplet dynamics within the cell over time (Baxter et al., Reference Baxter2025). While this approach is robust, technique-compatible, membrane lipid-specific labels or dyes are required to reduce phototoxicity and achieve high spatial resolution to lipid molecules. The specifications of these labels also need to cover the ability to quantify or show lipid variations and changes that occur during membrane-drug interaction, as shown by lipidomic analyses. There is a lack of sensors or dyes that bind specific lipid head groups (Butler et al., Reference Butler2024).
Model membranes as mimics of fungal membranes
While microscopy and spectroscopy are useful to determine the effect of an antifungal compound on the membrane permeability in live cells, these techniques are limited in explaining how they affect specific membrane properties and thus cannot elucidate the structural details of a compound’s mechanism of action or the role of specific lipids in the drug-induced membrane damage. The dynamic nature and the complex lipid compositions of the plasma membrane of living cells make mechanistic studies of peptide-membrane interactions very challenging (Muro et al., Reference Muro2014; Andersson et al., Reference Andersson2018). Model membranes are thus routinely used in biophysical studies to understand the effects of potential drugs or drug-like compounds on cellular membranes (Andrade et al., Reference Andrade2021; Luchini and Vitiello, Reference Luchini and Vitiello2021).
Model membrane systems are also used for biophysical techniques to quantify binding affinities or study membrane disruption with a particular focus on the effect of lipid composition on these properties, something that is challenging or not possible in living cells due to the complexity of the cell membrane. For instance, high-resolution binding kinetics measurements, such as surface plasmon resonance (SPR) spectroscopy and quartz crystal microbalance, commonly use model membranes to quantitatively examine membrane-peptides or drug-like interactions (De Medeiros et al., Reference De Medeiros2014; Wang et al., Reference Wang2015; Jaikishan et al., Reference Jaikishan2024). Besides reducing complexity and being ideal for broad-spectrum biophysical techniques, model membranes enable control of lipid composition and environmental conditions such as ion concentrations, pH, and temperature. Consequently, model membranes are ideal for isolating the role of specific lipids or environmental conditions on properties such as bilayer morphology, fluidity, permeability, domain formation or surface charge, as well as the effect of small molecules on these properties. Compared to bacterial or mammalian plasma membranes, fungal plasma membranes are understudied. Limited information is available on which lipid composition is most suitable to mimic the properties of fungal membranes or how lipid composition affects membrane–drug interactions. Recent advances and insights from fungal lipidomics will help address these questions. Another challenge is that some fungal-specific lipids, in particular sphingolipids, are not commercially available.
Numerous membrane model systems have been developed to mimic biological membranes to reproduce the essential structural and physicochemical properties of the phospholipid bilayer in cell membranes. They are largely classified into liposomes, also called vesicles, monolayers, and supported bilayers (Luchini and Vitiello, Reference Luchini and Vitiello2021). Liposomes are 3D structures that mimic both the inner and outer leaflets of cell membranes, which makes them a suitable model to study drug–membrane interactions. However, most liposomes used in biophysical studies are much smaller than cells, resulting in a much higher curvature than found in the plasma membrane (Andrade et al., Reference Andrade2021). This difference in curvature has to be considered, particularly when studying membrane-disrupting peptides, some of which are known to sense or change membrane curvature (Sani and Separovic, Reference Sani and Separovic2016; Deplazes et al., Reference Deplazes2020a). To study antifungal drug-membrane interactions, liposomes are often composed of phospholipids such as PC or PE (Figure 5) with added ergosterol, as this sterol is unique to fungal cells and the target of specific antifungal drugs. Such liposomes have been used to study the membrane permeability of antifungal AMPs, and the antifungal compound curcumin (Lee and Lee, Reference Lee and Lee2014; Kakar et al., Reference Kakar2022); pore formation of antifungal agents like the lactoferricin B peptide, and AmB, (Choi et al., Reference Choi2013; Lee et al., Reference Lee2014); and ergosterol interaction with antifungal drugs such as natamycin and imidazopyrazoindole (Mohid et al., Reference Mohid2022; Revie et al., Reference Revie2022).
Monolayers are commonly used to mimic a single leaflet of the phospholipid bilayer in a cell membrane. In the context of antifungal drug development, monolayers have been utilised to examine the mechanisms of action of AmB (Ha̧c-Wydro et al., Reference Ha̧c-Wydro2005; Araque-Marín et al., Reference Araque-Marín2022), nystatin (Hac-Wydro et al., Reference Hac-Wydro2007), antimicrobial peptides (Nasir and Besson, Reference Nasir and Besson2012; Zakharova et al., Reference Zakharova2019), and other membrane-altering biomolecules (Nobre et al., Reference Nobre2015). Because in monolayers, phospholipid tails are exposed to air in the air/liquid or air/solid interface, it is often unclear whether the findings from monolayer studies apply to cellular membranes. This challenge is partially resolved by solid-supported model membranes, which are constructed on a solid substrate, usually made of glass, silicon, aluminium oxide, silicon oxide, quartz, sapphire, mica, silver, or gold. The lipid bilayer is either separated from the substrate by a spacer unit or in direct contact with the solid surface. If sufficient spacers are used, supported bilayers can mimic the fluidity of cell membranes. Solid-supported model membranes are particularly effective for studying membrane electrochemistry and changes in ion permeability caused by membrane-altering compounds (Sebaaly et al., Reference Sebaaly2018; Hossain et al., Reference Hossain2021, Reference Hossain2022).
The benefit of the simple model systems in reducing complexity is also their main limitation. Most model membranes are made from a few lipids, usually phospholipids such as PC, PE, PS, or PA, with added ergosterol (Table 2), contrasting the dozens of lipid types present in varying compositions in the biological cell membranes. There are several reasons for this. First, many lipids found in fungal cells, in particular the fungal-specific sphingolipids, are not available as synthetic lipids, and many others are too expensive to be used in the quantities required for making vesicles. Second, until recently, there was limited information on the lipid composition of fungal membranes, in particular, data on the absolute values of lipids as opposed to changes in specific lipids after exposure to drugs or changes in growth conditions. Compared to synthetic lipid model membranes, Carey et al. (Reference Carey2022) suggest that model membranes developed from lipid extracts generate superior models as different lipid species are intrinsically incorporated. It is still unclear whether the drug-lipid interactions that control the variations in antifungal activity seen in cells are accurately replicated in these model membranes (made from synthetic or extracted lipids). One of the issues with lipid extracts is that they are made from cell homogenates, meaning they contain the lipids of not just the plasma membrane but also cell-internal organelles, which significantly differ in their lipid composition. It is not clear whether model membrane made from synthetic lipids and based on detailed lipidomics information of the plasma membrane are better at mimicking drug-membrane interactions than model membranes made from homogenate lipid extracts. Lipids from the plasma membranes can be isolated for MS analysis, but this involves the use of enzymatic digestion. While the resulting lipid fragments can be analysed by MS, this approach means the extracted lipids can no longer be used to prepare model membranes.
Summary of commonly used lipid compositions of models, model applications, and their general common limitations

Fundamental properties that complicate fungal cell membranes but are required for models and antifungal biomolecule interactions
When using membrane models for biophysical studies aimed at developing new antifungal drugs or understanding the mechanisms of existing ones, it is important to consider how the lipid composition affects the morphology and physicochemical properties of the membrane bilayer. Properties such as membrane thickness, lipid bilayer symmetry, fluidity, and surface charge can all influence how a drug interacts with or permeates through the membrane.
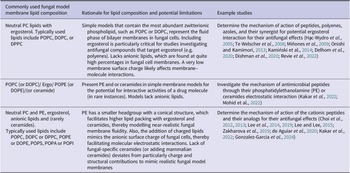
In the context of antifungal drug interaction studies, the simplest model membrane is usually made up of zwitterionic phospholipids (e.g. POPC, DOPC, DPPC) and ergosterol (to mimic fungal membrane) or cholesterol (to mimic the human cell membrane). While PE is abundant in most fungal species, it is less suited for the formation of stable liposomes or bilayers due to its preference for negative curvature. PE is thus often added at a lower mol% than reported from fungal lipidomics data. Charged lipids such as phosphatidylserine (e.g. POPS, DPPS) and sphingolipids are added to mimic the surface charge properties of the fungal membrane. If sphingolipids are added, sphingomyelin or other sphingolipids from mammalian or plant membranes are often used since fungal-specific sphingolipids are not available commercially.
Ergosterol and cholesterol are the dominant sterols in fungal and mammalian cells, respectively. Their structural similarities mean that they have similar effects on membrane properties. Both sterols act as regulators of fluidity through the formation of microdomains in the membrane bilayer and act as signalling molecules. Yet the minor chemical structural differences between ergosterol and cholesterol result in varying effects on membrane fluidity. The effect of sterols, as observed by Arora et al. (Reference Arora2004), using an environmentally sensitive fluorescent membrane probe, diphenylhexatriene (DPH), monitored the organisation and the dynamics of the interior regions of membrane models developed. The fluorescent anisotropy of DPH in vesicle membranes prepared from POPC and DPPC with increasing sterols (ergosterol and cholesterol) concentrations (0–50 mol%) was associated with the rotational diffusion rate, sensitive to the packing of fatty acyl chains. Increasing the sterol concentration reduces the rotational mobility of DPH due to decreased membrane fluidity caused by the sterols. While the effect was more pronounced at higher cholesterol concentrations, ergosterol concentrations above 20% had no further effect on membrane fluidity in the liquid-disordered phase. Similar observations of this difference in cholesterol and ergosterol effect were made from membranes in the gel phase. Further investigation using fluorescence and NMR measurements also revealed that the fluidity effects of ergosterol on liquid-disordered membranes require less than 20 mol% ergosterol concentration (Soubias et al., Reference Soubias2005; Hsueh et al., Reference Hsueh2007; Shrivastava and Chattopadhyay, Reference Shrivastava and Chattopadhyay2007). As the ergosterol concentration increases to approximately 20–35 mol%, POPC chain ordering and the membrane fluidity no longer change (Hsueh et al., Reference Hsueh2007). Similarly, MD simulations showed that both cholesterol and ergosterol increase lipid order of DPPC bilayers but to different extent (Alavizargar et al., Reference Alavizargar2021). Exploiting the atomistic-level insight provided by MD simulations, this study suggests that the higher rigidity of cholesterol compared to ergosterol means the former can align slight better with the saturated tails, causing lipids to be more conformationally restrained thus inducing a strong ordering effect than ergosterol. Based on their data and previous studies, the authors note that the reducing ordering effect of ergosterol means that the sterol alone cannot explain the relatively high membrane order observed in fungal membranes, and it is likely that other factors, such as the saturation of phospholipid tails, are involved. When studying the effect of sterols on phospholipid bilayers it is also important to consider the phospholipid bilayer used. For example, although many studies suggest that cholesterol has a stronger ordering effect than ergosterol, an MD simulation study by Czub et al. showed that, in DMPC, this effect is reversed (Czub and Baginski, Reference Czub and Baginski2006). The authors compared the effects of ergosterol and cholesterol on DMPC bilayers, and while both increased membrane orders, the effect was stronger for ergosterol. These contrasting findings may result from differences in the fluidity of DPPC and DMPC and highlight the complexity of even simple two-lipid membrane systems. The studies also highlight that cholesterol is not a suitable sterol for model membrane systems that aim to mimic fungal cell membranes.
Temperature and pressure have also been shown to exert a significant influence on this sterol effect. In a combined Fourier transform-infrared (FT-IR) spectroscopy and small-angle X-ray scattering (SAXS) study, Pühse et al. (Reference Pühse2008) show that for a binary lipid mixture of DPPC: ergosterol (78:22 mol%), temperature and pressure co-exist to influence the membrane ordering effects of ergosterol. For both the FT-IR and the SAXS data, an increase in temperature increases acyl-chain disorder and decreases lamellar spacing. At the same time, pressure stabilises ordered phases and expands bilayer d-spacing.
The fluidity of biological cell membranes is mostly determined with 1- [4-(trimethylamino) pheny]-6-phenyl-1,3,5-hexatriene (TMA-DPH). TMA-DPH diffuses more uniformly in the bilayer than DPH in the plasma membrane without moving into the cytoplasm. Abe and Hiraki (Reference Abe and Hiraki2009), using TMA-DPH, showed that sterol membrane modifications influence the mechanical properties of membranes. Mutant strains of S. cerevisiae lacking proteins involved in the late stages of ergosterol synthesis were used to show that membrane organisation and the presence of defects (voids) affect membrane stiffness, drug sensitisation, and passive diffusion of molecules across the membrane. A time-resolved anisotropy analysis of ergosterol-depleted mutants demonstrated a decrease in membrane order parameters and an increased rotational diffusion of TMA-DPH in the plasma membrane. These observations emphasised the crucial role of ergosterol in membrane integrity. Another commonly used dye, Laurdan (1-[6-(Dimethylamino) naphthalen-2-yl] dodecane-1-one), is dipole-sensitive to the amount of water in its microenvironment. As changes in fluidity alter the amount of interfacial water, the dye shows distinct emission spectra for liquid-ordered and liquid-disordered phases of membranes. Derkacz et al. (Reference Derkacz2022) used Laurdan to show that overexpression of the ERG11 gene in C. albicans increased ergosterol, which caused an expected decrease in membrane fluidity. This suggests that changes in membrane fluidity can occur for ergosterol-targeting antifungal drugs, which can be evaluated with Laurdan or TMA-DPH (Khmelinskaia et al., Reference Khmelinskaia2020; Suchodolski et al., Reference Suchodolski2021; Shen et al., Reference Shen2024).
The condensing effect of sterols on phospholipids also results in a decreased area per phospholipid and an increase in the thickness of the bilayer (Levine and Wilkins, Reference Levine and Wilkins1971; Henriksen et al., Reference Henriksen2006). In a study using 2H-NMR and micropipette aspiration, both ergosterol and cholesterol were shown to have a significant POPC membrane stiffening effect based on surface compressibility and flexural moduli. The aerial expansion modulus and bending stiffness obtained from the micropipette suction, as well as their correlations with the magnetic moments obtained from the 2H-NMR spectra, indicated that the membrane modification of the sterols on the POPC hydrophobic thickness mainly influences the mechanical properties of the POPC membrane. Although there were differences in the measure of how sterols (ergosterol, lanosterol, and cholesterol) influence and affect membrane thickness, particularly the formation and stability of domains, it is generally accepted that sterols have a universal condensing effect on the membrane and reduce hydrophobic thickness, compressibility, and elasticity of phospholipid bilayers (Henriksen et al., Reference Henriksen2006). Hung et al. (Reference Hung2016) showed a contradicting viewpoint with an X-ray diffraction analysis. The study first found that, unlike cholesterol, mixing ergosterol with POPC or DOPC becomes challenging at high sterol concentrations. POPC-ergosterol mixtures were only monophasic up to 20 mol% ergosterol. For DOPC-ergosterol, this threshold was even lower, 7 mol% ergosterol. While cholesterol showed a strong condensation effect by increasing the membrane thickness from ~36.0 to 43.3 Å, ergosterol had only a slight condensation effect on DMPC; the thickness of the membrane increased slightly by ~2.0 Å, but not on POPC and DOPC membranes. Interestingly, ergosterol rather has a thinning effect on POPC and DOPC bilayers, in contrast to the thickening effect of cholesterol. Compared with previous NMR studies, the interactions of ergosterol and cholesterol with phospholipid chains are different, especially with unsaturated chains. It is, however, often difficult to compare studies. Slight changes make a major difference. For instance, studies on POPC, DOPC, DMPC or DPPC, and even the ions present (e.g. Ca2+ has a stiffening effect on the bilayer while Na+ and K+ usually do not show that effect) are relevant in generating different outcomes (Deplazes et al., Reference Deplazes2020a,Reference Deplazesb; Deplazes et al., Reference Deplazes2021). The choice of lipids and the buffer conditions for mimicking fungal membranes for antifungal drug interaction studies is important to be evaluated and tuned towards more realistic outcomes comparable to the biological membrane.
The outer and inner leaflet composition and the potential for spontaneous phospholipid translocation in the bilayer of the fungal membrane are structurally important in drug interactions. A study by Kohli et al. (Reference Kohli2002) demonstrated that the asymmetric distribution of PE, with a higher concentration in the outer leaflet of the C albicans plasma membrane, is a signature and contributes to azole resistance. For Cr neoformans, the loss of plasma membrane phospholipid asymmetry has been shown to disrupt vesicle trafficking, heighten susceptibility to diverse antifungals, and obliterate its virulence (Shor et al., Reference Shor2016). Most model membranes are symmetric, which deviates from the asymmetric biological membranes, and indicates a major setback to their applications (Zhukov and Popov, Reference Zhukov and Popov2023). The available type of lipid in the outer leaflet to create asymmetry that mimics a biological membrane for interaction mainly depends on controlled lipid translocations between the membrane leaflets. In determining and controlling such compositions and symmetry, fluidity is indispensable due to its major contribution to the ability of lipids to move between leaflets. The model membrane’s bilayer asymmetry results from the spontaneous translocation of phospholipids in the fluid phase, as determined by a neutron reflectometry (NR) study (Gerelli et al., Reference Gerelli2012). Using a similar approach, Florek and co-workers examined the fluidity of simple bilayer membrane models made from different lipid compositions: d-DPPC: h-DPPS, d-DPPC: h-POPS, d-DPPC:(h-DPPC:h-POPS,1:1 mol: mol) and d-DPPC:(h-DPPS: ergosterol, 1:1 mol: mol) and their contribution to asymmetry. The structural parameters obtained establish that the bilayer containing fully saturated phospholipids (DPPC and DPPS) had a comparatively higher asymmetry. The inner leaflet of this model was determined to be made up of 73.7% DPPC and 17.0% DPPS, while the outer leaflet was 1.9% DPPC and 88.8% DPPS. The asymmetric membrane produced offers a higher charge lipid, DPPS, at the outer leaflet, creating a high surface charge for potential membrane-drug electrostatic interaction studies. However, the phospholipids of the biological membrane are a heterogeneous mixture of saturated and unsaturated phospholipids, limiting the application of the DPPC: DPPS model membrane. The membrane model composed of fully saturated d-DPPC, and partially unsaturated h-POPS showed a higher translocation level across the bilayer leaflets. The addition of ergosterol enhanced membrane fluidity in the outer layer, resulting in more lipid rearrangement between the inner and outer leaflets (Florek et al., Reference Florek2018). From the study, a better translocation of lipids for the asymmetric bilayer membrane that represents fungal membrane to capture realistic antifungal-membrane interaction requires an appropriate combination of saturated and unsaturated phospholipids (zwitterionic and charged) with ergosterol.
The fungal cell membrane is also negatively charged, making it electrostatically active for charge particle (antifungal peptide) interactions. Determining cell surface charge is challenging; thus, zeta potential (also known as electrokinetic potential) is utilised to better understand the electrical potential difference at the hydrodynamic sliding surface, defined as the interface between the aqueous liquid and the stationary fluid layer at the cell surface. The surface charge of the cell not only influences the permeability of molecules into the cell but also determines how they adhere to the surfaces of the host to elicit their virulence activities. Investigating the sub-inhibitory concentration of antifungal drugs Ketoconazole, Miconazole, AmB, aculeacin A, and azalomycin F affect the adherence of C. albicans to host surfaces, the obtained correlation between zeta potential and cell adherence indicates that decreased electric repulsive forces enhance cell adherence. These antifungal agents were shown to affect the zeta potential of the cell by altering the cell wall and plasma membrane electrocharacteristics (Miyake et al., Reference Miyake1990). In an antifungal activity study of cyclic temporin L peptide, changes in the zeta potential of C. albicans cells following the addition of the peptide analogues provided insight into the mechanism of action of the peptides. The changes in the zeta potential implied that the peptides aggregated and covered the cell surface through electrostatic interactions (Bellavita et al., Reference Bellavita2022). A similar change in zeta potential was observed in a nanoparticle, nMgO, in activities against fungal phytopathogens. The changes in zeta potential also signified the electrostatic interaction of the positively charged nanoparticle and the negatively charged surface of the fungi cells (Chen et al., Reference Chen2020). de Aguiar et al. (Reference de Aguiar2020) also showed that the anti-biofilm and mechanism of action of an antifungal peptide, crotalicidin Ctn[15–34], results from the peptide incorporation in the lipid bilayer to reduce the anionic surface charge of the cells (Figure 6). This was evidenced in the anionic decrease of zeta potentials of C. albicans membrane-mimicking vesicles (POPC: POPE: POPS: POPI: Erg at the mole ratio of 59: 21: 3: 4: 13) and protoplasts before and after the addition of the peptide. Zhang et al. (Reference Zhang2022) also show that the binding of their designed antifungal peptide analogues, (IIKK)3, to fungal membrane model vesicles (POPC: SOY-PI: Ergosterol (9: 1: 2 by wt) caused a change in zeta potential. Even though zeta potential does not give comprehensive evidence on the mechanism of action of antifungal agents, it provides complementary evidence of whether molecules bind or accumulate on the surface of the cell or the plasma membrane.
Intermittent contact mode AFM images and zeta potentials of C. albicans cells, C. albicans protoplasts and lipid vesicles showing membrane interaction effects of the antifungal peptide Ctn[15-34]. a) Images show the change in the morphology of drug-resistant clinical isolates and wild-type C albicans cell biofilms after exposure to Ctn[15–34]. The presence of vesicles on the cell surface is indicated with white arrows(N–P). b) Changes in zeta potential of C. albicans cells, protoplasts, and vesicles in exposure to Ctn[15-34]. The zeta potential differs in interactive properties of Ctn[15–34] with the membranes of C. albicans cells, protoplasts, and vesicles. Figure made from de Aguir et al., International Journal of Molecular Sciences, 2020, published under the Creative Commons Attribution 4.0 https://creativecommons.org/licenses/by/4.0/.

The critical role of biophysical techniques in developing a less toxic AmB variant
AmB is a lifesaving drug that is often used as a last resort to treat life-threatening fungal infections. Despite >60 years of clinical use, acquired resistance is rare (Cavassin et al., Reference Cavassin2021). Nevertheless, due to its small therapeutic window, AmB shows significant off-target effects and causes severe side effects, including kidney failure. While the liposomal formulation AmBiosome reduces the drug’s toxicity, many low- and middle-income countries, where HIV-related IFIs are prevalent, lack access to AmBiosome and rely on the traditional AmB-deoxycholate formulation (Lancet Global Health, n.d.). Decades of research have (so far) unsuccessfully tried to develop a safer AmB variant. Recent pioneering work in the laboratories of Martin Burke and Chad Rienstra is an excellent illustration of how insights gained from biophysical techniques can be harvested for rational drug design to develop an AmB variant that is renal sparing yet retains the drug’s broad-spectrum fungicidal activity (Maji et al., Reference Maji2023). Their work will hopefully soon translate into a much-needed safer AmB variant.
The mechanism of AmB has been studied extensively to reveal its complex and multipronged antifungal activity. AmB targets ergosterol in fungal cell membranes and kills fungal cells by complex and multipronged mechanisms that include the formation of pores and membrane-associated aggregates resulting in membrane permeabilisation and subsequent cell lysis. For decades, the AmB’s ion channel formation was thought to be the main driver of antifungal activity, yet Burke and co-workers have demonstrated that ergosterol sequestration is sufficient to kill fungal cells (Gray et al., Reference Gray2012). In a subsequent study, the authors used TEM and ssNMR to show that the ergosterol-sequestration results in the formation of large, extramembranous, sponge-like AmB-ergosterol aggregates (Anderson et al., Reference Anderson2014). Characterisation of these aggregates with ssNMR (Lewandowska et al., Reference Lewandowska2021) and extensive medicinal chemistry then resulted in the design of a new AmB variant that accelerates ergosterol extraction in an ergosterol-selective manner, thus increasing the drug’s therapeutic window (Maji et al., Reference Maji2023).
Summary and conclusions
Characterising the fungal cell envelope and its interaction with small molecules at high spatial and temporal resolution remains challenging due to its structural complexity and dynamic nature. Advances in imaging techniques such as SEM, TEM, and AFM, together with structural information from solid-state NMR (ssNMR), have been instrumental in developing models of fungal cell wall architecture and identifying how individual components influence morphology, permeability, and mechanical properties. These approaches have also provided important insights into how environmental stress and antifungal compounds alter cell wall and membrane organisation and how these changes contribute to antifungal resistance. Complementary to cellular studies, model membrane systems have enabled detailed investigation of membrane properties such as fluidity, surface charge, and permeability, as well as the molecular mechanisms of antifungal drugs. However, the extent to which current model systems capture the compositional and functional diversity of fungal plasma membranes remains unclear.
Compared with bacterial and mammalian systems, fungal membranes remain relatively understudied, highlighting the need to bridge the gap between simplified membrane models and cellular complexity. A key priority is to understand better how fungal-specific lipids influence membrane biophysical properties and how these properties affect drug interactions and pathogenic processes. Expanded lipidomics studies will be essential for defining lipid diversity across species and for determining how lipid composition changes in response to environmental stress or antifungal treatment. These data will enable the development of lipidomics-informed model membranes and systematic comparisons with cellular measurements to identify membrane compositions that best reproduce in vivo behaviour. The broader use of asymmetric bilayers will further improve the physiological relevance of these systems.
Another major challenge is understanding the coupling between the plasma membrane and the cell wall. Although typically studied separately, these structures form a mechanically and chemically integrated system. Developing experimental models that allow systematic variation of cell wall components, analogous to lipid variation in membrane models, will be essential for quantifying how individual polysaccharides contribute to properties such as porosity, rigidity, and structural integrity. More broadly, future progress will depend on linking molecular-scale insights from biophysical measurements with cellular phenotypes such as antifungal resistance, stress adaptation, and pathogenicity.
Financial support
Obed Obeng-Gyasi is supported by a University of Queensland Graduate School Scholarship. Evelyne Deplazes declares none.
Competing interests
Evelyne Deplazes and Obed Obeng-Gyasi declare no conflict of interest.