Introduction
To understand human physiology and pathology, the scientific community in its beginnings used dogs, chickens, rabbits, rodents, pigs, cows, sheep and non-human primates (Ref. Reference Franco1). It is indisputable that, for the time, the use of these animals was necessary and allowed gaining the wide basic knowledge fundamental for the dissemination of several therapies that currently exist.
However, despite their historical contribution, animal models have limitations inherent in interspecific differences, especially regarding immunological response, gene expression, metabolism and signalling pathways, which often compromises their predictive ability for human outcomes (Refs. Reference van der Worp, Howells, Sena, Porritt, Rewell, O’Collins and Macleod2, Reference Hartung3). It is estimated that more than 90% of drugs that demonstrate efficacy in animal models fail in human clinical trials due to unexpected toxicity or lack of efficacy (Ref. Reference Mak, Evaniew and Ghert4). These discrepancies highlight the need for systems more representative of human biology.
Not only studies evoking treatments but also those that aimed to improve the ways of studying went ahead. In this scenario, after some time, alternative models were thought and many continue to be developed to reduce the use of animals, based on the concept of the ‘three Rs’ (replacement, reduction and refinement) created during the 1950s by Russel and Burch (Refs. Reference Russell and Burch5, Reference Doke and Dhawale6).
Nevertheless, the use of these approaches has not fully replaced animal use yet. Animal testing is still required when it comes to understanding systemic situations, as even with excellent alternative models, the interaction between physiological systems has not been fully replaced to understand the homoeostasis of an organism (Ref. Reference Barré-Sinoussi and Montagutelli7). Examples of these situations are studies involving immunological diseases, sepsis and pharmacological action of drugs or chemical substances. In these cases, a scenario emerges that highlights the need to search for new methods that can reverse this situation, drawing attention to the bioethical issue. In addition, the United Nations (UN) established 17 Sustainable Development Goals (SDGs) in 2015, where the promotion of innovation is cited (SDG 9: build resilient infrastructures, promote inclusive and sustainable industrialization and foster innovation).
In this unfilled gap is noticeably visible the emergence of the need to promote more discussions to save animals. It is perceived that there is a worldwide concern, as pharmaceutical companies have been developing cruelty-free products and a growing number of people have adhered to veganism (Ref. Reference Wang, Zhao and Song8). According to recent surveys, approximately 1% of Americans and 2% of British adults identify themselves as vegan (Refs. Reference Jones9, Reference Mathieu and Ritchie10). Furthermore, in a study addressing public opinion regarding animal use in medical training, Merkley et al. (2018) found that the majority of the interviewed agree that if effective alternative models exist to study a determined condition, they should be used instead of animals for medical training. A supermajority of these participants agreed that it is unethical or morally wrong to use animals for this reason when non-animal methods are available (Ref. Reference Merkley, Pippin and Joffe11).
Regulatory and ethical advances
Following this time frame, several movements around the world have gained strength to refine ethical issues. In Brazil, a law was created in 2008 to manage animal use: the Arouca Law (11.794/2008). It has established procedures for the scientific use of animals and created the National Council for the Control of Animal Experiments (CONCEA), the highest Brazilian agency in the area. The CONCEA validates alternative methods for testing medicines, cosmetics and health products, among other substances, aiming to replace the use of animals in cosmetic tests. Moreover, each in vivo study needs to be approved by a local committee before starting. Not rare, the committee returns the study to the researcher for adjustments, aiming to minimize the number of animals used and their suffering. Similar to Brazil, other countries have their laws and guidelines with the same goal.
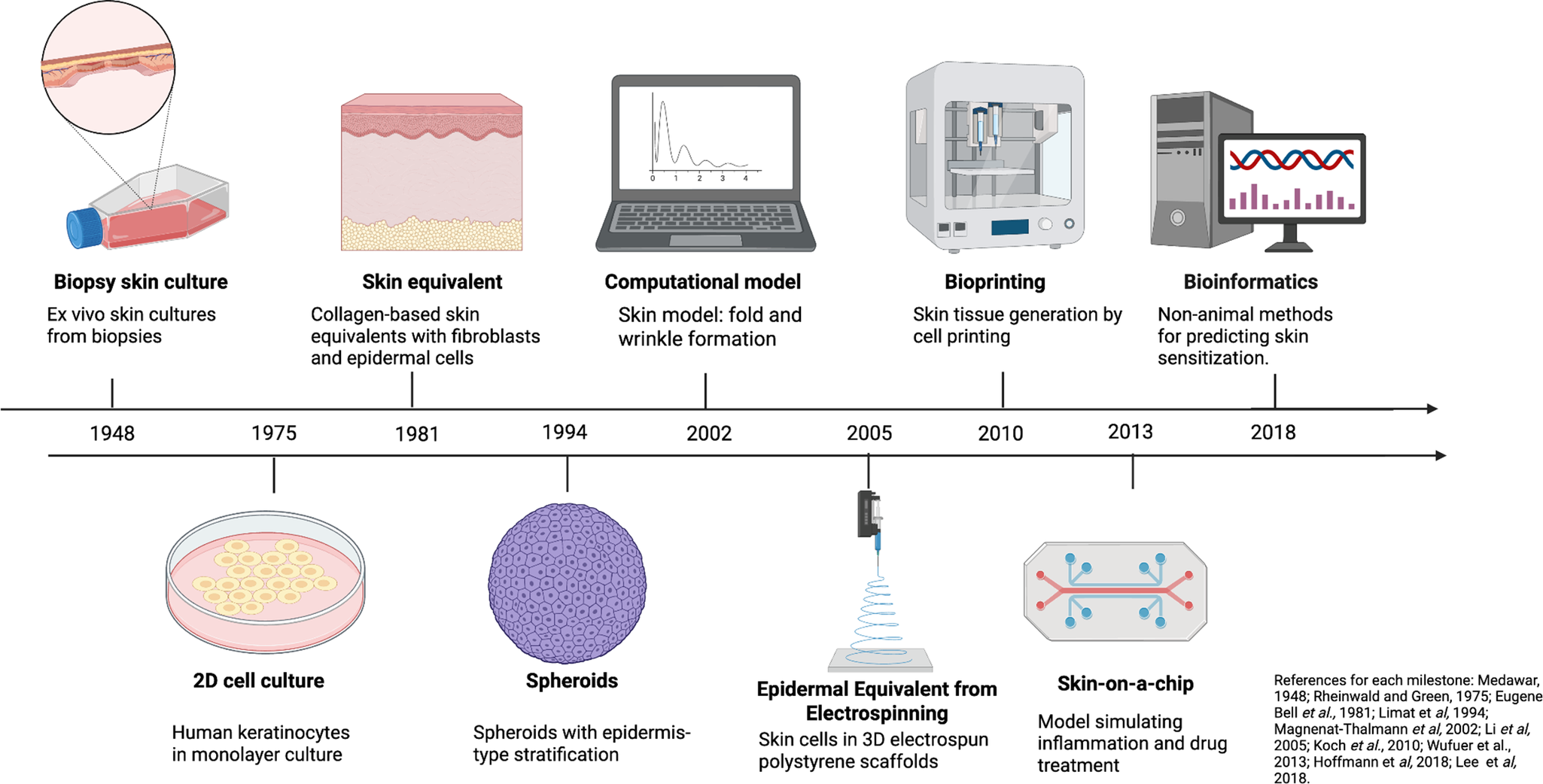
In an attempt to avoid the use of animals, the development and enforcement of regulations have played a fundamental role in defining ethical standards and research practices regarding the use of animals in science worldwide, as well as in stimulating the development of alternative models and improving the existing ones. In this context, one field that has benefited significantly is dermatology. Considering that dermatology involves the testing of cosmetic substances, most alternative models were developed to meet this need. Alternative in vitro and in silico models are emerging as a new paradigm, offering tools capable of reproducing aspects of human physiology that animals cannot accurately simulate. While traditional in vivo models allow the assessment of systemic responses and interactions between organs, they lack cellular and molecular precision. On the other hand, in vitro models, derived from human tissues or cells, provide direct observation of specific human processes – such as gene regulation, protein expression and metabolism – eliminating interspecific variability and increasing the reproducibility of results (Figure 1) (Refs. Reference Rheinwald and Green12–Reference Gangwal and Lavecchia20).
Timeline showing the emergence of alternative methods. The figure summarizes the chronological development of key in vitro and in silico models, from early biopsy and 2D monolayer cultures to advanced 3D bioprinting, skin-on-a-chip and computational systems. Created with BioRender.com.

In the European Union (EU), animal tests for finished cosmetics and cosmetic ingredients were banned in 2004 and 2009, respectively (Ref. 21). In this context, the Scientific Committee on Consumer Safety (SCCS) was created in the EU as an independent committee of experts from different countries. One of its responsibilities is to establish guidelines for cosmetic industries to develop studies for the safety evaluation of cosmetics and relevant toxicological tools for the safety evaluation of cosmetic ingredients as alternatives to animal testing, such as corrosivity and irritation, skin sensitization, dermal/percutaneous absorption, repeated dose toxicity, reproductive toxicity, mutagenicity/genotoxicity, carcinogenicity, toxicokinetics studies, photo-induced toxicity and human data (Ref. 22). The main challenge of overcoming in vitro results is to predict the real benefits of cosmetic products in contact with all the systems that involve the human body as a whole (Ref. Reference Eberlin, da Silva, Facchini, da Silva, Pinheiro, Eberlin and Pinheiro23).
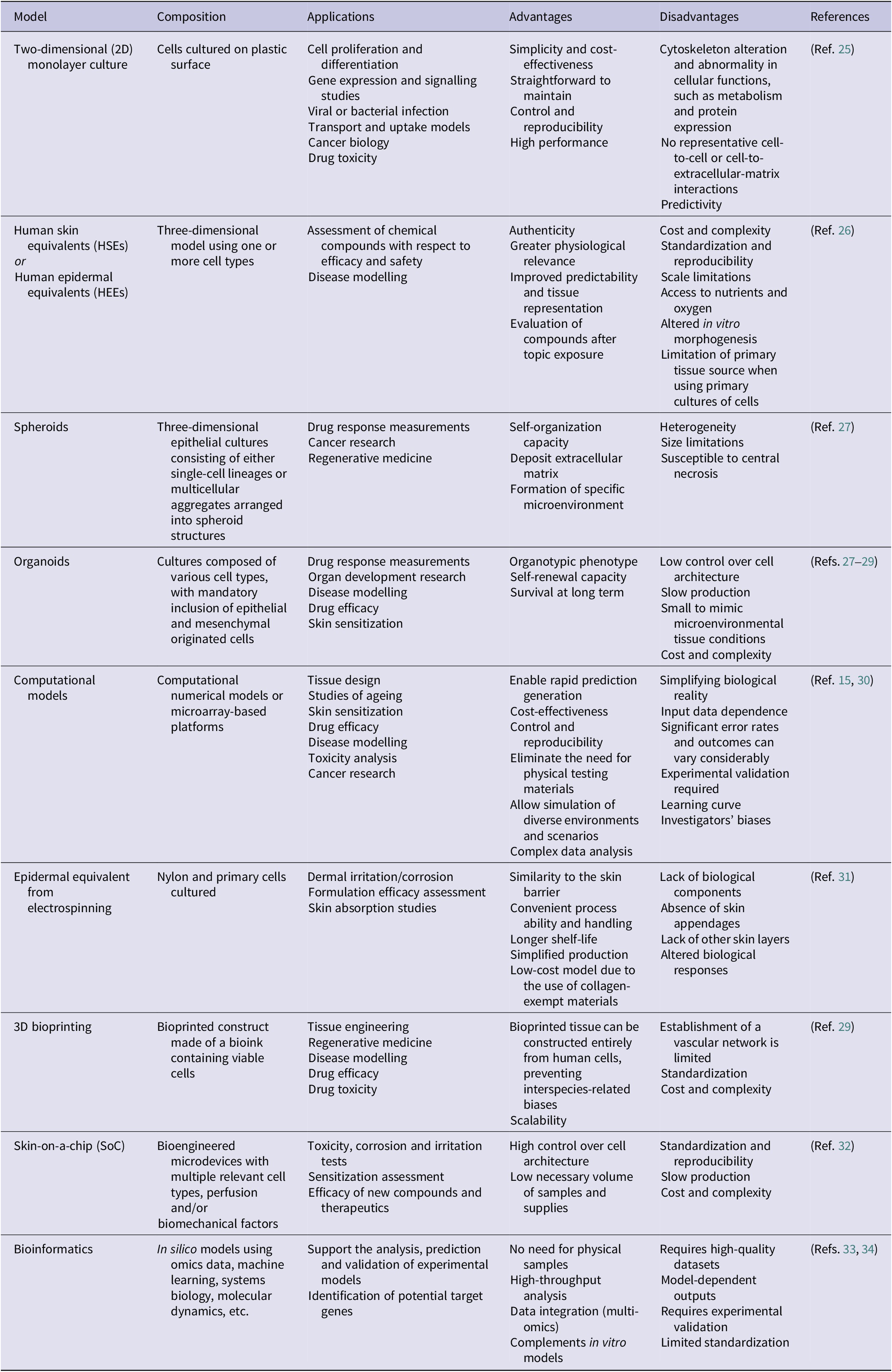
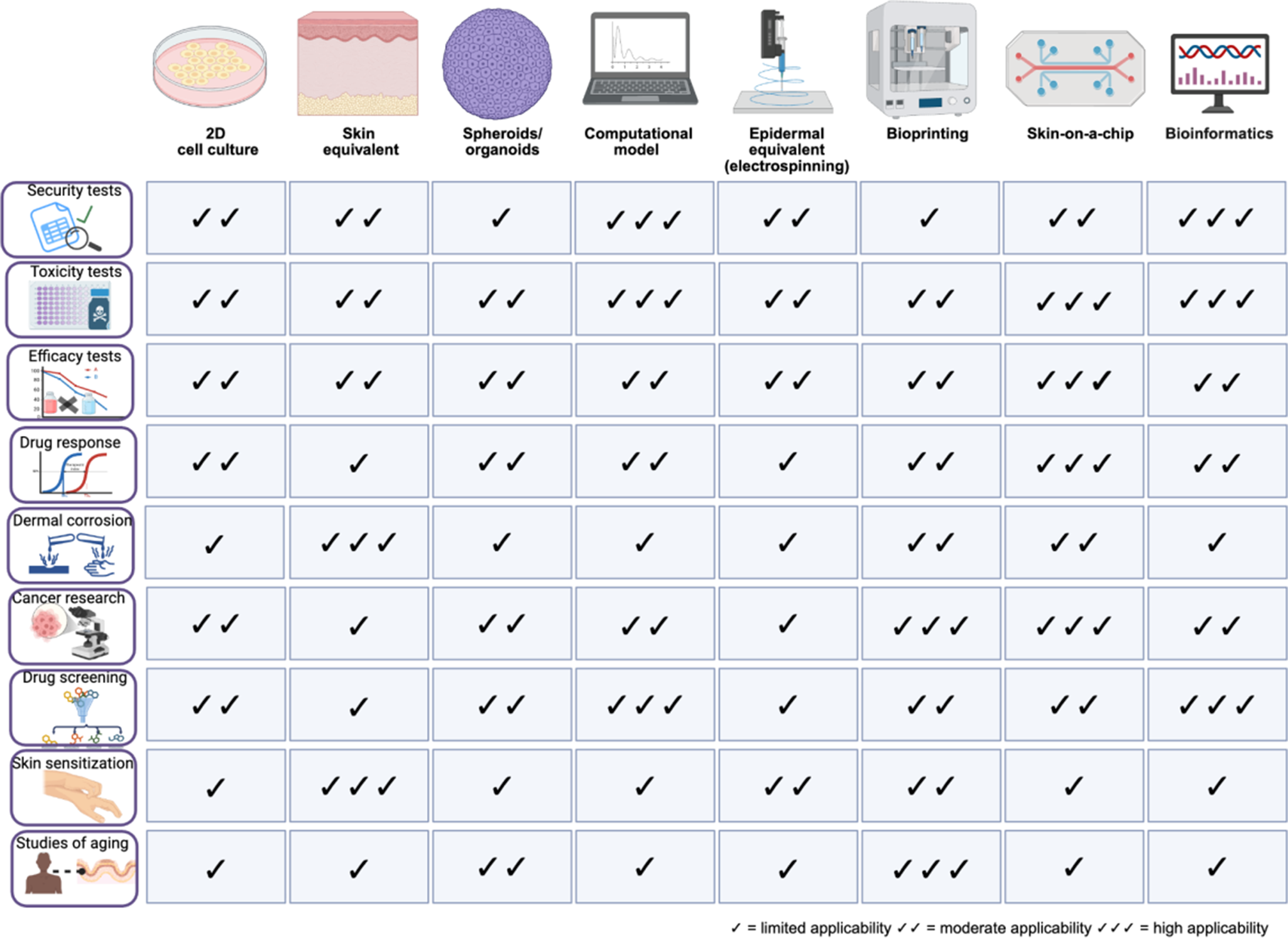
The alternative methods to dermatology research have their own advantages and disadvantages (Table 1). A common feature among them is the use as proof of concept, where a large funnel is generated and only substances and/or therapies with pharmacological activity or those with biological potential will proceed to animal testing (Ref. Reference Denayer, Stöhr and Van Roy24). The majority of these alternative methods are cell-based in vitro models. Figure 2 summarizes the main alternative dermatological models and their relative applicability to different biological purposes, highlighting differences in suitability, maturity and physiological relevance of each model.
Main applications, advantages and disadvantages of alternative dermatological methods in substitution for animal tests

Alternative methods and their correlation with different biological purposes. Created in BioRender. Wink, M. (2026) https://BioRender.com/6v061nr.

Alternative in vitro models in dermatological research
Two-dimensional (2D) monolayer cultures, spheroids and organoids
Two-dimensional (2D) models are the most commonly used. In adherent 2D cultures, cells grow as an attached monolayer, and the advantages of these cultures are associated with simple and cost-effective maintenance. Unfortunately, adherent cultures have disadvantages as the cultured cells do not mimic the natural tissue structures and, after tissue isolation under 2D conditions, the morphology of the cells is altered as well as the mode of cell division (Ref. Reference Kapałczyńska, Kolenda, Przybyła, Zajączkowska, Teresiak, Filas, Ibbs, Bliźniak, Łuczewski and Lamperska35). Three-dimensional (3D) models that mimic skin tissue have been developed to overcome the limitations of predictivity of 2D in vitro models. 3D models showing the interactions between different cell types of tumour and stroma are difficult to generate; thus, to better mimic the in vivo situation, different 3D cell culture approaches with several degrees of complexity have been developed. Spheroids have formats that are very useful for basic and applied melanoma research, track cellular behaviour in a cell-type-specific manner and recapitulate different characteristics of early melanoma stages. They can be used to evaluate novel drugs and combination therapies (Ref. Reference Vörsmann, Groeber, Walles, Busch, Beissert, Walczak and Kulms36). Organoids represent an advanced 3D in vitro model that mimics the structural and functional organization of native tissues through the self-organization of stem cells or progenitor cells. In dermatological research, skin organoids are capable of reproducing multiple cell types found in the epidermis and dermis, providing a physiologically relevant platform for studying processes such as differentiation, wound healing and disease modelling. These systems offer a more faithful representation of tissue microarchitecture compared to conventional monolayer cultures, bridging the gap between 2D models and full-thickness engineered skin equivalents. Recent studies have demonstrated their potential for applications in drug response testing, toxicity evaluation and personalized medicine approaches in skin biology (Refs. Reference Mak, Evaniew and Ghert4, Reference Lee, Bӧscke, Tang, Hartman, Heller and Koehler37, Reference Kim, Koo and Knoblich38).
Human skin equivalents (HSEs) and reconstructed human epidermis (RHE)
Techniques for developing 3D skin models demand a high level of innovation and complexity through cell cultures. One of the main goals is the in vitro development of skin equivalents for dermal toxicity evaluation (Ref. Reference do Nascimento Pedrosa, Catarino, Pennacchi, de Moraes Barros and Maria-Engler39). The simplest 3D model is known as reconstructed human epidermis (RHE) and contains an epidermal layer of skin composed only of keratinocytes. EpiSkin™, Epiderm™, SkinEthic™, epiCS™ and LabCyte EPI-MODEL24 are commercially available models of RHE approved by European Centre for the Validation of Alternative Methods (ECVAM) to replace in vivo rabbit skin irritation tests and for skin corrosion tests of cosmetic ingredients; more recently, Skin+ and KeraSkin™ have also been validated as irritation tests (Refs. 40, 41). KeratinoSens™ and LuSens are models constructed with immortalized HaCaT stably transfected with a selectable plasmid, which are used in skin sensitization tests also approved by ECVAM (Ref. 42). Full-thickness skin models or HSEs are more complex models, which consist of an epidermis and dermis (keratinocytes and fibroblasts) and have been used to evaluate drug or treatment efficacy (Refs. Reference Bataillon, Lelièvre, Chapuis, Thillou, Autourde, Durand, Boyera, Rigaudeau, Besné and Pellevoisin43, Reference Mathes, Ruffner and Graf-Hausner44). Recently, 3D vascularized human skin equivalents (vHSEs) that can mimic native skin have been developed; these models reproduce key histological and functional characteristics of real skin, including a multilayered, differentiated epidermis supported by a fibroblast-derived dermal matrix containing endothelial cells organized into a capillary-like microvascular network (Refs. 40, 41). This way, the electrospinning technique has been used to form polymeric mats that, together with porous structures, allow the seeding of cells such as keratinocytes and fibroblasts, to produce an equivalent skin (Ref. Reference Camarena, Matsuyama, Maria-Engler and Catalani31).
3D bioprinting
3D bioprinting is another important tool for dermal toxicity evaluation; it is used to extrude materials and develop scaffolds to target tissues in the required structure (Ref. Reference Ng and Yeong45). It enables the accurate construction based on biomolecules, synthetic/natural hydrogels and cells (Refs. Reference Fayyazbakhsh46, Reference Manita, Garcia-Orue, Santos-Vizcaino, Hernandez and Igartua47). Different bioprinting methods have been used to fabricate skin tissue models, such as extrusion-based, laser-assisted and microvalve-based bioprinting (Ref. Reference Ng and Yeong45). Authenticity, scalability and reproducibility of the tissues are the main advantages of bioprinted skin, when compared to conventional constructs (Ref. Reference Millás, Lago, Vasquez-Pinto, Massaguer and Maria-Engler26). 3D skin bioprinting has enabled the construction of a complete skin with a hypodermal layer (Ref. Reference Moakes, Senior, Robinson, Chipara, Atansov, Naylor, Metcalfe, Smith and Grover48) and can be applied in penetration and absorption tests of cosmetic ingredients. However, there are challenges to overcome, such as producing sensitive, dry, oily skin with different textures, pigmented with different shades, and appendages, hair follicles, microvessels and immune cells (Refs. Reference Millás, Lago, Vasquez-Pinto, Massaguer and Maria-Engler26–Reference Olejnik, Semba, Kulpa, Dańczak-Pazdrowska, Rybka and Gornowicz-Porowska49). 3D bioprinting is a research target of global cosmetic leaders and still requires standardization and regulatory approvals.
Skin-on-a-chip (SoC)
Limitations on traditional 2D cultures and 3D organ models have stimulated the use of technologies such as microfabrication in the last decade. In this context, organs-on-chips have been developed. They consist of complex tissue-like structures within microfluidic chips that allow the dynamic culture of cells inside, in order to model or mimic the physiology of a tissue or organ. Skin-on-a-chip (SoC) has made considerable advances in the transport of substances or delivery. Microfluidics maintain the high-throughput capacity of the systems while reducing costs and reagent volumes needed for the experiments (Refs. Reference Zhang, Sito, Mao, He, Zhang and Zhao50, Reference Risueño, Valencia, Jorcano and Velasco51)). In 2D SoC models, skin cells are cultured directly in microfluidic channels to simulate different skin components. Additionally, SoC is a possible platform to study cell–cell interactions, expose cells to mechanical strains or even study immune response (Ref. Reference Zoio and Oliva52). In 2D SoC systems, monolayer cultures differ from conventional 2D cultures because cells are exposed to dynamic nutrient flow, mechanical stimuli and controlled microenvironments, which help preserve morphology and enable the physiological study of cell–cell interactions. However, 3D SoC models are the most accurate to replicate the spatial organization, cell interactions and functional complexity of native skin, surpassing the limitations of 2D cultures and 2D SoC systems (Refs. Reference Zhang, Korolj, Lai and Radisic53–Reference Abdo, Zhao, Okhovatian, Vargas, Wagner, Shakeri, Vosoughi and Radisic55).
Therefore, these models bring human physiology closer than ever, unlike animals, whose physiological parameters (e.g. perfusion rate and oxygen gradients) vary widely. SoC models allow precise control of the cellular microenvironment, reproducing perfusion, mechanical tension and nutrient flow under simulated human conditions (Ref. Reference Rimal, Marquardt, Nevolianis, Djeljadini, Marquez, Huth, Chigrin, Wessling, Baron, Möller and Singh56).
Computational and in silico models
Bioinformatics has emerged as a promising alternative to the use of animals and in vitro experiments as it allows the investigation of biological phenomena using computational tools and biological databases. Through in silico simulations and modelling, it is possible to predict molecular interactions, effects of genetic mutations and drug responses, thus reducing the need for laboratory testing.
The computational models generated from bioinformatics tools are emerging as a new opportunity to reduce not only the use of animal models but also the volume of in vitro experiments, contributing to the formulation of more accurate hypotheses before performing biological experiments due to the large volumes of omics data (transcriptomics, proteomics and metabolomics) generated – to the detriment of limited data of traditional tests.
Furthermore, bioinformatics makes it possible to leverage data already available in public databases, enabling comparative and predictive analyses without the need for new experimental tests. This translates into savings in time and resources, as well as greater reproducibility and control over the variables analysed. Thus, scientific research becomes more sustainable and aligned with ethical principles that seek to replace, reduce and refine the use of animals in experimentation.
Utilizing bioinformatics tools and in silico experiments, data from human cell cultures exposed to different toxicants can be analysed (e.g. (Ref. Reference Grafström, Nymark, Hongisto, Spjuth, Ceder, Willighagen, Hardy, Kaski and Kohonen57)). Interestingly, dermatology was one of the first medical disciplines to welcome and support bioinformatics results, and the term ‘skinomics’ has been proposed to designate specifically the bioinformatics studies in the area. The objective is to provide knowledge of skin biology, improve the function of healthy skin and assist in treating pathological skin conditions through in silico experiments, where computer simulations using databases allow the modelling of an organic phenomenon (Refs. Reference Younis, Shnayder, Blumenberg, Maglaveras and Chouvarda58–Reference Madden, Enoch, Paini and Cronin60). In silico models make predictions concerning absorption, distribution, metabolism and excretion of a chemical from molecular data, and new possibilities and results that can be generated by this approach are promising and exciting (Refs. Reference Younis, Shnayder, Blumenberg, Maglaveras and Chouvarda58–Reference Madden, Enoch, Paini and Cronin60).
More recently, researchers are exploring the uses of artificial intelligence (AI) to improve or supplement current screening processes in melanoma and nonmelanoma skin cancer (Ref. Reference Madden, Enoch, Paini and Cronin60). These methods can be explored in the future for use as strategic alternatives to animal research as well. In the context of dermatological and cosmetic sciences, AI can be trained on databases containing chemical descriptors, gene-expression signatures and cellular responses from reconstructed human skin models to predict irritation, sensitization, penetration and antimicrobial efficacy of novel compounds before physical testing. These approaches not only accelerate early-stage screening but also substantially reduce the number of compounds entering animal experimentation, thereby minimizing animal use and costs (Ref. Reference Tieghi, Moreira-Filho, Martin, Wellnitz, Otoch, Rath, Tropsha, Muratov and Kleinstreuer61).
Despite their clear advantages, AI and other computational approaches still rely on robust, high-quality datasets – often derived from in vivo, in vitro and clinical studies – to ensure the reliability and generalizability of their predictions (Ref. Reference Gangwal and Lavecchia20). In this context, bioinformatics plays a central role in data analysis, model prediction and the refinement of experimental designs, establishing itself as an indispensable tool in modern science. It contributes to the integration of technological innovation with ethical and sustainable research practices (Figure 3). Nonetheless, computational predictions still require experimental validation to confirm their biological relevance and applicability. Therefore, additional time, resources and continued investment will be necessary before such tools can meaningfully reduce, though not fully replace, the use of traditional in vitro and in vivo models.
Flowchart illustrating the progressive transition from basic to advanced animal-free dermatological models and their integration with predictive models.

Conclusion and perspectives
The alternative models cited represent a valuable strategy to mimic biological systems and minimize animal use. A number of models and techniques have been developed for different purposes. 2D monolayer cultures remain indispensable for hypothesis generation and screening studies due to their simplicity and cost-effectiveness. However, by design, they lack the 3D architecture and microenvironment of native skin. As a result, 2D systems cannot simulate complex tissue responses and should be complemented by advanced 3D models, microphysiological platforms and in silico approaches when higher physiological relevance is required.
Despite their potential, advanced methods are not yet widely adopted across research settings. It is notable that more developed countries are already ahead in the use of these methodologies. However, these innovative solutions must reach all countries, enabling them to refine and rethink the use of animals in dermal pharmacology and toxicology research and progressively in other fields.
Encouragingly, the latest ECVAM status report indicates that in 2022, approximately 8.5 million animals were used for scientific purposes across the EU, representing a 5% reduction compared to 2018, coinciding with a rapid expansion of in vitro and in silico biotechnologies (Ref. Reference Zuang, Baccaro, Barroso, Berggren, Bopp, Bridio, Capeloa, Carpi, Casati, Chinchio, Corvi, Deceuninck, Franco, Gastaldello, Katsanou, Langezaal, Malinowska, Mennecozzi, Milcamps, Piergiovanni, Prieto-Peraita, Selfa, Valsesia, Whelan, Wittwehr and Worth62). These trends demonstrate that while complete replacement remains challenging, significant advances are being achieved.
Altogether, we can conclude that there is no unique methodology capable of replacing in vivo experiments. It is necessary that each researcher evaluate the scientific question that should be addressed and then select and develop the best in silico and in vitro protocols or their combination. After that, in vivo experimentation needs to be done for the questions that remain to be answered at the systemic level – for example, modulation of the immune system, pharmacokinetics, pharmacodynamics, chronic toxicity – until new possibilities and technologies arise.
Data availability statement
No datasets were generated or analysed during the study.
Funding statement
Schematic representations were created with BioRender.com. M.R.W. is a recipient of productivity research fellowship (PQ1D) from Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq). This study was supported by CNPq MS-SCTIE-DECIT-DGITIS-CGCIS/CNPq n° 26/2020 (442586/2020–5) and INCT 3D (406436/2022–3); Fundação de Amparo à Pesquisa do Estado do Rio Grande do Sul (FAPERGS) – RITES (22/2551–0000385-0) and ARD/ARC (25/2551–0000943-0). Science Suite Inc. dba BioRender (‘BioRender’) has granted Márcia Wink permission to use this completed graphic in accordance with BioRender’s Terms of Service and Academic License Terms (‘License Terms’). Subscription Type: Academic Individual. Type of Publication: Open-access. Publisher Name: Expert Reviews in Molecular Medicine. Effective Date: January 30, 2026.
Competing interests
The authors have no conflicts of interest to declare.