Introduction
Milk consumption is critical for the survival, growth and development of mammal neonates, and mammary health is important for milk production (Garwolinska et al. Reference Garwolinska, Namiesnik, Kot-Wasik and Hewelt-Belka2018; Li et al. Reference Li, Wang, Li, Han, Mao and Wang2019). Oxidative stress is a common factor that impairs mammary health and the function of milk synthesis (Li et al. Reference Li, Wang, Li, Han, Mao and Wang2019; Spitzer et al. Reference Spitzer, Tian, Choudhary and Zhao2020). Oxidative stress occurs when the scavenging capacity of reactive oxidants is lower than their production, resulting in excessive accumulation of oxidative damages in the mammary gland which leads to decreased milk production and milk quality (Jakubczyk et al. Reference Jakubczyk, Dec, Kaldunska, Kawczuga, Kochman and Janda2020; Spitzer et al. Reference Spitzer, Tian, Choudhary and Zhao2020). Studies showed that some nature substances have great potential to combat oxidative stress (Xu et al. Reference Xu, Hu, Wang and Cui2019, Reference Xu, Ke, Li, Xie, Su, Xie, Mo and Chen2021). Quercetin, a kind of flavonoid substance found in vegetables, fruits, tea, wine and other foods, is one of examples (Polera et al. Reference Polera, Badolato, Perri, Carullo and Aiello2019; Xu et al. Reference Xu, Hu, Wang and Cui2019). In drug research and development, quercetin has been found to have great potential for its antioxidant activity (Song et al. Reference Song, Wang and Gao2020; Xu et al. Reference Xu, Hu, Wang and Cui2019). Studies showed that quercetin improved the expression of antioxidant enzymes in lipopolysaccharide (LPS) -challenged human aortic endothelial cells (Li et al. Reference Li, Zhang and Frei2016), and quercetin protected human umbilical vein endothelial cells against H2O2-induced oxidative damages (Tian et al. Reference Tian, Yang, Lu and Peng2019), suggesting its antioxidant capacity in cardiovascular diseases. However, whether quercetin has protective effects on oxidative stress in the mammary gland has not been well studied.
It is believed that nuclear factor E2-related factor 2 (Nrf2) signaling pathway is the main regulatory pathway of cellular antioxidant defense system (Shaw and Chattopadhyay Reference Shaw and Chattopadhyay2020). When oxidative stress occurs, Nrf2 pathway is activated and regulates the activity of antioxidant enzymes and the expression of stress-resistant proteins to maintain redox homeostasis (Jin et al. Reference Jin, Wang, Liu, Hu, Zhao and Liu2016; Shaw and Chattopadhyay Reference Shaw and Chattopadhyay2020). Severe oxidative stress leads to cell proliferation obstruction, autophagy and apoptosis. The mitogen-activated protein kinase (MAPK) pathway plays an important role in cell survival and death and thus may be closely related to oxidative stress (Kumar et al. Reference Kumar, Singh, Rana and Rana2021; Liu et al. Reference Liu, Shen, Zhao, Sun, Wang, Long, He, Lin, Wu and Wei2020a; Luo et al. Reference Luo, Xu, Sho, Zhang, Xu, Yao and Xu2019). The MAPK family mainly consists of three subtypes: p38 MAPK, extracellular regulated protein kinase (ERK) and c-Jun N-terminal kinase (JNK) (Jin et al. Reference Jin, Wang, Liu, Hu, Zhao and Liu2016). It has been reported that three subtypes of MAPK pathway had positive effects on the redox imbalance in melanocytes and sertoli cells (Li et al. Reference Li, Tang, Su and Li2020a; Liu et al. Reference Liu, Shen, Zhao, Sun, Wang, Long, He, Lin, Wu and Wei2020a). However, it is not clear whether quercetin alleviates oxidative stress via Nrf2 and MAPK pathways in the mammary gland. Therefore, the aim of this study was to investigate the effect and underlying molecular mechanisms of quercetin against redox imbalance in mammary epithelial cells. Our study provided evidence to support the development and utilization of quercetin as an antioxidant additive in lactating animals.
Materials and methods
Reagent
Quercetin (Q4951) was purchased from Sigma Chemical (St. Louis, MO, USA) with the purity ≥95%. Hydrogen peroxide (H2O2, 323381-500 mL), bovine insulin (I6634) and epidermal growth factor (E4127) were also purchased from Sigma Chemical. Fetal bovine serum (No. 900-108) was purchased from Gemini Bio (Calabasas, California, USA). Nrf2 inhibitor (ML385), p38 MAPK inhibitor (SB203580), ERK inhibitor (PD98059) and JNK inhibitor (SP600125) were purchased from Selleck Chemicals (Houston, TX, USA).
Cell culture
The HC11 mammary epithelial cells, derived from female mouse mammary gland, were obtained from American type culture collection (ATCC) (Manassas, Virginia, USA; CRL-3062) and cultured in RPMI 1640 supplemented with 10% fetal bovine serum, bovine insulin (5 µg/mL), epidermal growth factor (10 ng/mL), penicillin (100 U/mL) and streptomycin (0.1 mg/mL) in a humid incubator with 5% CO2 and 95% air at 37°C.
Cell treatment
After HC11 cells adhered to the 96-well plates and grew to 40% confluence, the culture medium was changed to serum-free medium added with H2O2 (50–1000 μΜ) or serum-containing medium added with quercetin (5–25 μΜ) for 24 h to determine the changes of cell proliferation and lactate dehydrogenase (LDH) release. The powder of quercetin was dissolved in dimethyl sulfoxide (DMSO) (D2650, Sigma) to obtain 25 mM and 20 mM storage solution, respectively, based on the highest concentrations required for individual experiment. The inhibitor was dissolved in DMSO to produce a storage solution of 1000 times the concentration of the working solution. At each use, the storage solution of quercetin and inhibitors were diluted with the medium used in the experiment. In experiments with quercetin treatment, 0.1% DMSO was added in the control group. In experiments, after the HC11 cells adhered to the 6-well plates and grew to 40% confluence, HC11 cells were pretreated with 20 μΜ quercetin for 2 h, followed by treatment of 100 μΜ H2O2 for 24 h to determine the effects of quercetin pretreatment on H2O2-induced redox imbalance and the activation of MAPK and Nrf2 pathways. In the experiments with signaling molecule inhibitors, HC11 cells were pretreated with an individual inhibitor for 1 h, and then treated with 20 μΜ quercetin for 2 h, followed by treatment with 100 μΜ H2O2 for 24 h.
Cell viability and cytotoxicity assay
Cell viability was quantified by Cell Counting Kit-8 (Beyotime, Shanghai, China). The 96-well plates were inoculated with about 8000 cells/well. After HC11 cells adhered and grew to 40% confluence, the cell treatments were carried out as described above. The CCK8 solution was added to each well and incubated for 1 h, and the absorbance at 450 nm was detected by microplate reader (Molecular Devices, California, USA). The cytotoxicity of cells was determined by LDH Cytotoxicity Assay Kit (Beyotime). When 1 h before the predetermined time reached, LDH releasing reagent was added to the “sample maximum enzyme activity control well.” Then, the HC11 cells were cultured for 1 h followed by centrifuging at 400 × g for 5 min. A 90 μL volume of supernatant/each well was transferred to a new 96-well plate, and the absorbance at 490 nm was measured by microplate reader (Molecular Devices). In the cell viability and cytotoxicity experiments, six wells were used for one treatment in each experiment, and the experiment was conducted three times independently.
Cellular reactive oxygen species (ROS) assay
The production of cellular ROS was determined by DCFDA/H2DCFDA Cellular ROS Assay Kit (ab113851) purchased from Abcam (Waltham, MA, USA) using flow cytometry (BD FACSVerse™, BD Biosciences, CA, USA) with BD FAC Suite software (BD Biosciences). According to the manufactures’ instructions, HC11 cells were cultured and treated with 100 μΜ H2O2 for 4 h. Then, the cells were prepared into a single cell suspension and incubated in 10 μΜ 2’,7’–dichlorofluorescin diacetate (DCFDA) working solution at 37°C for 30 min in dark. After that, the cells were gently blowing with a pipette to ensure that single cells were suspended, and then 10,000 cells were collected and analyzed in each group by flow cytometry. The excitation wavelength of fluorescence was 485 nm, and the emission wavelength was 535 nm. The fluorescein Isothiocyanate-protein A (FITC-A) mean value was used to analyze the differences between groups.
Assay of total antioxidant capacity (T-AOC), catalase (CAT) and superoxide dismutase (SOD) enzyme activities
The T-AOC was defined as the ratio of reducing efficiency of the samples to the reducing efficiency of Trolox which determined by Total Antioxidant Capacity Assay Kit. The assays for CAT activity, SOD activity and protein concentration were performed using the Catalase Assay Kit, Total Superoxide Dismutase Assay Kit with WST-8 and BCA Assay Kit (Beyotime), respectively, according to the manufactures’ instructions. The activities of CAT and SOD enzymes and T-AOC were normalized by the protein concentration of the samples.
Western blot analysis
Cells were lysed using Radio Immunoprecipitation Assay (RIPA) lysate buffer with 1 mM phenylmethanesulfonyl fluoride (Beyotime). The lysates were centrifuged at 12,000 × g for 10 min, and the supernatants were collected. The protein concentration was determined using BCA Assay Kit (Beyotime). The western blot analysis used 20 μg protein per well. All protein samples were used within 1 week after extraction. Other procedures of western blot were performed according to the previous description (Chen et al. Reference Chen, Zhao, Ren, Han, Liu, Li and Liu2020). Primary antibodies used for western blot were all rabbit anti-mouse antibodies which obtained from Abcam, HuaAn Biotechnology (Hangzhou, China) and Cell Signaling Technology (MA, USA). More information about the antibodies was listed in Table S1, and the secondary antibody was horseradish peroxidase (HRP)-conjugated goat anti-rabbit immunoglobulin G (IgG) antibody (Bioker, Hangzhou, China). The Super ECL Detection Reagent (Bioker) was used for chemiluminescence using an imager from Shanghai CLINX Science Instruments Co., Ltd (Shanghai, China). The relative intensity of bands was quantified using Fusion FX software (Vilber, France) and normalized by reference protein β-actin in the same sample.
Statistical analysis
Data were presented as mean ± standard error of mean (SEM). One-way analysis of variance (ANOVA) followed by Tukey’s multiple comparison was carried out to determine the differences between treatments using IBM SPSS Statistics 19 (IBM, Armonk, NY, USA). P < 0.05 was considered statistically significant. Graphing was carried out using GraphPad Prism Software version 6.0 (GraphPad Software Inc., La Jolla, CA, USA).
Results
Effects of quercetin on H2O2-induced changes in cell viability and LDH release in HC11 cells
Cell viability was significantly reduced by 50–1000 μΜ H2O2, whereas 100–1000 μΜ H2O2 significantly increased LDH release in HC11 cells (Fig. 1A–B). The cell viability was reduced to about 60% of the control group, and LDH release was increased by about 25% when the cells were treated with 100 μΜ H2O2 for 24 h (Fig. 1A–B). Thus, treatment of cells with 100 μΜ H2O2 for 24 h was used as the cell injury model for subsequent experiments. Treatment of cells with 5–20 μΜ quercetin alone did not affect the cell viability, whereas 25 μΜ quercetin significantly reduced the cell viability (Fig. 1C). In addition, 5–25 μΜ quercetin did not affect the release of LDH from the cells (Fig. 1D). The LDH release induced by H2O2 was significantly reduced by 5–25 μΜ quercetin, and cell viability decreased by H2O2 was improved by 10–25 μΜ quercetin (Fig. 1E–F). As a result, 20 μΜ was the most effective concentration of quercetin in alleviating HC11 cell damage caused by H2O2.
Effects of different concentrations of hydrogen peroxide (H2O2) and quercetin on cell viability and lactate dehydrogenase (LDH) release in HC11 cells. (A) Cell viability and (B) LDH release in HC11 cells after 24 h treatment with different concentrations of H2O2 (0, 50, 100, 250, 500, 750 and 1000 μΜ). (C) Cell viability and (D) LDH release in HC11 cells after 24 h treatment with different concentrations of quercetin (0, 5, 10, 15, 20 and 25 μΜ). (E) Cell viability and (F) LDH release in HC11 cells pretreated with different concentrations of quercetin (0–25 μΜ) for 2 h, followed by treatment of 100 μΜ H2O2 for 24 h. One-way ANOVA followed by Tukey’s multiple comparison was used to determine the differences between the groups. Data represent the mean ± SEM. Data marked with different small letters (a, b, c and d) indicated P < 0.05, while data with the same letter indicated no significant difference.

Effect of quercetin on H2O2-induced redox imbalance
The production of ROS in HC11 cells increased by 100 μΜ H2O2 treatment, and quercetin pretreatment restored that to the control level (Fig. 2A–B). H2O2 treatment reduced the T-AOC in HC11 cells, which was significantly improved by quercetin pretreatment (Fig. 2C). H2O2 treatment significantly reduced the activity of CAT and increased the activity of SOD in the cells (Fig. 2D–E), and quercetin pretreatment significantly improved the activity of CAT reduced by H2O2 (Fig. 2D). H2O2 significantly inhibited the expression of glutamate/cystine reverse transporter light chain (xCT) and thioredoxin reductase 1 (TXNRD1), and the inhibition on the expression of xCT was completely removed by quercetin pretreatment (Fig. 2F–G, I). Quercetin partially restored the expression of TXNRD1 under H2O2 (Fig. 2F, I). Although H2O2 treatment did not affect the expression of glutamate-cysteine ligase modifier subunit (GCLM), quercetin treatment with or without H2O2 upregulated the expression of GCLM compared with the control group (Fig. 2F, H).
The ameliorating effects of quercetin on redox balance in HC11 cells. HC11 cells were pretreated with or without 20 μΜ quercetin for 2 h, followed by treatment with or without 100 μΜ H2O2 for 24 h to determine (A and B) the production of ROS, (C) T-AOC, enzyme activity of (D) CAT and (E) SOD. (F) Representative grayscale photographs of immunoblots and quantitative protein expression of (G) xCT, (H) GCLM and (I) TXNRD1. One-way ANOVA followed by Tukey’s multiple comparison was used to determine the differences between four groups. Data represent mean ± SEM. Data marked with different lowercase letters (a, b and c) indicated P < 0.05, while data with the same letter indicated no significant difference.

Effect of quercetin on the activation of MAPK pathway induced by H2O2
Hydrogen peroxide increased the level of phosphorylation of p38 MAPK and the ratio of p-p38 MAPK/p38 MAPK, and quercetin pretreatment restored the phosphorylation level of p38 MAPK and the ratio of p-p38 MAPK/p38 MAPK to the control levels (Fig. 3A–D). Similarly, H2O2 increased the phosphorylation level of ERK protein and the ratio of p-ERK/ERK and decreased the total protein level of ERK (Fig. 3A, E–G). Quercetin pretreatment restored the phosphorylation level of ERK, the ratio of p-ERK/ERK and the protein expression of ERK to a great extent (Fig. 3A, E–G). Meanwhile, H2O2 increased the ratio of p-JNK/JNK and decreased the total protein level of JNK but did not affect the phosphorylation level of JNK, and quercetin pretreatment upregulated the level of JNK and downregulated the ratio of p-JNK/JNK to the control level in H2O2 treated cells (Fig. 3A, H–J).
Effects of quercetin on the H2O2-induced activation of MAPK pathway in HC11 cells. HC11 cells were pretreated with or without 20 μΜ quercetin for 2 h, followed by treatment with or without 100 μΜ H2O2 for 24 h. (A) Representative grayscale photographs of immunoblots. Quantitative protein expression of (B) p-p38/p38, (C) p38, (D) p-p38, (E) p-ERK/ERK, (F) ERK, (G) p-ERK, (H) p-JNK/JNK, (I) JNK and (J) p-JNK. One-way ANOVA followed by Tukey’s multiple comparison was used to determine the differences between four groups. Data represent the mean ± SEM. Data marked with different lowercase letters (a, b and c) indicated P < 0.05, while data with the same letter indicated no significant difference.

Effect of quercetin on the activation of Nrf2 pathway induced by H2O2
Hydrogen peroxide significantly increased the protein expression of p-NRF2 and the ratio of p-NRF2/NRF2 but did not affect the total protein expression of NRF2 (Fig. 4A–D). Quercetin pretreatment restored the expression of p-NRF2 to the level in control cells, and the ratio of p-NRF2/NRF2 in quercetin pretreatment group was significantly lower than that in H2O2 group (Fig. 4).
Effects of quercetin on the H2O2-induced activation of Nrf2 pathway in HC11 cells. HC11 cells were pretreated with or without 20 μΜ quercetin for 2 h, followed by treatment with or without 100 μΜ H2O2 for 24 h. (A) Representative grayscale photographs of immunoblots. (B) Quantitative presentation of the ratio of p-NRF2/NRF2 protein and protein expression level of (C) NRF2 and (D) p-NRF2. One-way ANOVA followed by Tukey’s multiple comparison was used to determine the differences between four groups. Data represent mean ± SEM. Data marked with different lowercase letters (a and b) indicated P < 0.05, while data with the same letter indicated no significant difference.

Role of MAPK pathway in ameliorating H2O2-induced activation of Nrf2 pathway by quercetin
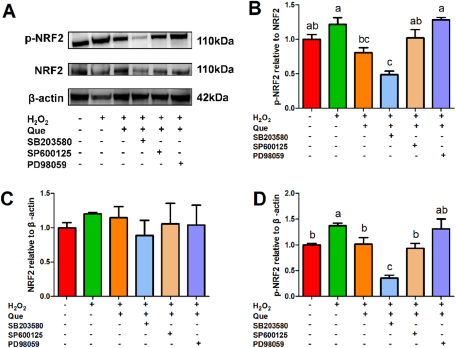
The p38 MAPK inhibitor (SB203580, 25 µM), ERK inhibitor (PD98059, 100 µM), JNK inhibitor (SP600125, 10 µM) and Nrf2 inhibitor (ML385, 5 µM) inhibited the activation of MAPK and Nrf2 pathways, respectively (Figure S1). The ERK inhibitor PD98059 significantly upregulated the ratio of p-NRF2/NRF2 and completely eliminated the recovery effect of quercetin on H2O2-induced activation of Nrf2 pathway in HC11 cells (Fig. 5). The p38 MAPK inhibitor SB203580 significantly reduced the protein expression of p-NRF2, whereas the JNK inhibitor SP600125 did not affect Nrf2 signaling pathway significantly in cells treated with H2O2 (Fig. 5).
Effects of the p38 MAPK inhibitor SB203580 (25 µM), ERK inhibitor PD98059 (100 µM) and JNK inhibitor SP600125 (10 µM) on the quercetin’s ameliorating effect on H2O2-induced activation of Nrf2 pathway in HC11 cells. HC11 cells were pretreated with or without an individual inhibitor for 1 h, then the cells were treated with or without 20 μΜ quercetin for 2 h, followed by treatment with or without 100 μΜ H2O2 for 24 h. (A) Representative grayscale photographs of immunoblots. (B) Quantitative presentation of the ratio of p-NRF2/NRF2 protein and the protein expression level of (C) NRF2 and (D) p-NRF2. One-way ANOVA followed by Tukey’s multiple comparison was used to determine the differences between the groups. Data represent mean ± SEM. Data marked with different lowercase letters (a, b and c) indicated P < 0.05, while data with the same letter indicated no significant difference.
Roles of MAPK and Nrf2 pathways in quercetin’s ameliorating effect on H2O2-induced oxidative stress
All p38 MAPK, ERK and JNK inhibitors significantly reduced the protective effect of quercetin on cell viability (Fig. 6A). There were no differences in cell viability between any group of Nrf2, p38 MAPK and ERK inhibitors and the H2O2 group (Fig. 6A). No inhibitors significantly affected quercetin’s alleviation in the release of LDH induced by H2O2 (Fig. 6B). Nrf2 inhibitor, but not MAPK pathway inhibitors, blocked the protective effect of quercetin on the level of ROS to a certain degree (Fig. 6C–D). In addition, the p38 MAPK, ERK and Nrf2 inhibitors decreased the recovery effect of quercetin on T-AOC of HC11 cells, and the inhibition of ERK pathway reduced the T-AOC to the level of the control group (Fig. 6E). Four inhibitors decreased quercetin’s upregulation of CAT activity to different degrees (Fig. 6F). Nrf2 inhibitor, but not MAPK pathway inhibitors, significantly reduced the upregulation of quercetin on the protein level of xCT to the basal level of H2O2 group (Fig. 6G–H). Furthermore, the Nrf2 inhibitor eliminated the upregulation of quercetin on the expression of GCLM and restored it to the level of H2O2 group, whereas the expression of GCLM in three MAPK pathway inhibitor groups was between the H2O2 group and quercetin group. There was no difference in protein level of GCLM between the Nrf2 inhibitor and p38 MAPK inhibitor groups (Fig. 6G, I).
Effects of the p38 MAPK inhibitor SB203580 (25 µM), ERK inhibitor PD98059 (100 µM), JNK inhibitor SP600125 (10 µM) and Nrf2 inhibitor ML385 (5 µM) on the quercetin’s ameliorating effects on H2O2-induced oxidative stress in HC11 cells. HC11 cells were pretreated with or without an individual inhibitor for 1 h, then the cells were treated with or without 20 μΜ quercetin for 2 h, followed by treatment with or without 100 μΜ H2O2 for 24 h to determine (A) cell proliferation, (B) LDH release, (C and D) the production of ROS, (E) T-AOC and (F) CAT enzyme activity. (G) Representative grayscale photographs of immunoblots. Quantitative protein expression of (H) xCT and (I) GCLM. One-way ANOVA followed by Tukey’s multiple comparison was used to determine the differences between the groups. Data represent mean ± SEM. Data marked with different lowercase letters (a, b, c, d and e) indicated P < 0.05, while data with the same letter indicated no significant difference.

Discussion
Oxidative stress impacts mammary health seriously, resulting in decreased milk yield and milk quality (Li et al. Reference Li, Wang, Li, Han, Mao and Wang2019; Spitzer et al. Reference Spitzer, Tian, Choudhary and Zhao2020). Alleviating oxidative stress by using exogenous substances has been the focus of antioxidation research in the mammary gland (Koch et al. Reference Koch, Zagorska, Marzec and Kukula-Koch2019; Xu et al. Reference Xu, Hu, Wang and Cui2019). Mammary epithelial cells are lactigenous cells and directly related to the health and function of the mammary gland (Mizusawa et al. Reference Mizusawa, Sharmin and Yonekura2019; Yuan et al. Reference Yuan, Zhen, Zhang, Yu, Gao and Ao2019). Therefore, a model of cellular oxidative stress was adapted in HC11 mouse mammary epithelial cells with H2O2 treatment. In this model, the cell viability was gradually decreased by increasing H2O2 concentration from 50 μΜ to 250 μΜ. The 100 μΜ H2O2 reduced cell viability to about 60% of control group and significantly increased the release of LDH from the cells, which was then used for subsequent experiments. Quercetin is a flavonoid extracted from a variety of plants (Polera et al. Reference Polera, Badolato, Perri, Carullo and Aiello2019). Quercetin has shown to have antioxidant properties in previous studies (Song et al. Reference Song, Wang and Gao2020; Xu et al. Reference Xu, Hu, Wang and Cui2019). In our study, HC11 cells were treated with quercetin at commonly used concentrations (Li et al. Reference Li, Zhang and Frei2016; Smith et al. Reference Smith, Fowler, Naftalin and Siow2020) and found that quercetin below 25 μΜ had no negative effect on cell viability and LDH release from the cells. However, 5–20 μΜ quercetin had significant protective effects on the viability of HC11 cells treated with H2O2, and 20 μΜ quercetin showed the maximal effects, which was then used in subsequent experiments to study the underlying mechanisms.
ROS include reductive oxygen-containing free radicals and non-free radical reactive oxidants (Sessa et al. Reference Sessa, Messina, Russo, Salerno, Castruccio, Distefano, Li Volti, Calogero, Cannarella, Mongioi, Condorelli and La Vignera2020). In this study, H2O2 treatment increased ROS production and decreased the T-AOC in HC11 cells, indicating that H2O2 resulted in redox imbalance, which was consistent with previous reports (Jin et al. Reference Jin, Wang, Liu, Hu, Zhao and Liu2016). There are numbers of antioxidant enzymes and non-enzymatic antioxidants in the body, such as CAT, SOD and xCT (Vucetic et al. Reference Vucetic, Cormerais, Parks and Pouysségur2017), which constitute the antioxidant defense system. Quercetin increased T-AOC and the activity of CAT reduced by H2O2, verifying that quercetin restored H2O2-induced redox imbalance in HC11 cells. The xCT is a member of the cystine/glutamate antiporter family and promotes cystine uptakes and protects cells from oxidative stress and cell death (Koppula et al. Reference Koppula, Zhang, Zhuang and Gan2018). In this study, quercetin completely eliminated the reduction of xCT induced by H2O2 and partially restored the level of stress-resistant protein TXNRD1 in HC11 cells. These effects of quercetin on stress-resistant proteins were consistent with previous in vivo studies in pancreatic tissue of type 2 diabetes mice (Li et al. Reference Li, Jiang, Mei, Zhao, Chen, Liu, Tang, Gao and Yao2020b), proving quercetin’s antioxidation function in vivo and in vitro. Moreover, GCLM is the regulatory subunit of GCL which affects the synthesis of glutathione (GSH) (Chen et al. Reference Chen, Xue, Fang, Chen, Chen, Yang, Shen, Chen, Zhang and Ling2019; Nishizawa et al. Reference Nishizawa, Matsumoto, Shindo, Saigusa, Kato, Suzuki, Sato, Ishii, Shimokawa and Igarashi2020). The mRNA abundance of GCLM was found to be downregulated after H2O2 treatment in human retinal pigment epithelial cells (ARPE-19 cells) (Yuan et al. Reference Yuan, Du, He, Zhang and He2020). In the present study, the protein level of GCLM was not changed by H2O2, but it was induced by quercetin treatment, indicating that GCLM may still be one of the important molecules involved in quercetin protection in HC11 cells.
The MAPK signaling pathway is a key pathway that regulates cell proliferation and respond to exogenous stimuli, and MAPK signaling molecules mainly include p38 MAPK, ERK and JNK subtypes (Lanna et al. Reference Lanna, Gomes, Muller-Durovic, McDonnell, Escors, Gilroy, Lee, Karin and Akbar2017; Santarpia et al. Reference Santarpia, Lippman and El-Naggar2012). Studies on cardiac fibroblasts have shown that quercetin reduces the phosphorylation of ERK, p38 and JNK induced by free radicals, but there have only a few studies which investigated the relationship between quercetin and MAPK pathway in mammary epithelial cells (Min et al. Reference Min, Yangchun, Yuquan and Changying2019). In this study, H2O2 activated the p38 MAPK, ERK and JNK molecules by phosphorylation, and quercetin restored the activation of these molecules, revealing that MAPK pathway might play a role in the protective effects of quercetin in mammary epithelial cells. Similarly, quercetin decreased p-NRF2/NRF2 ratio in cells treated with H2O2, proving that Nrf2 pathway may also likely to be involved in quercetin’s protective role to oxidative stress. These results are consistent with the reports that quercetin is a Nrf2-interacting nutrient in improving Alzheimer’s disease, insulin resistance and lung injury (Bousquet et al. Reference Bousquet, Cristol, Czarlewski, Czarlewski, Anto, Martineau, Haahtela, Fonseca, Laccarino, Blain, Fiocchi, Canonica, Fonseca, Vidal, Choi, Kim, Le Moing, Reynes, Sheikh, Akdis and Zuberbier2020; Zaplatic et al. Reference Zaplatic, Bule, Shah, Uddin and Niaz2019).
To investigate the molecular mechanisms of quercetin protection against oxidative stress in HC11 cells, the specific inhibitors of Nrf2 (ML385), p38 MAPK (SB203580), ERK (PD98059) and JNK (SP600125) molecule (Jin et al. Reference Jin, Wang, Liu, Hu, Zhao and Liu2016; Martin-Acosta and Xiao Reference Martin-Acosta and Xiao2021; Zarrin et al. Reference Zarrin, Bao, Lupardus and Vucic2021) were used to explore the role of each of these pathways in quercetin’s function. The inhibition of ERK upregulated the ratio of p-NRF2/NRF2 and eliminated the recovery effect of quercetin on Nrf2 signaling pathway, whereas p38 MAPK inhibition reduced the protein level of p-NRF2, indicating that quercetin may protect HC11 cells by activating ERK-Nrf2 and p38 MAPK-Nrf2 pathways simultaneously. The relationship between MAPK and Nrf2 signaling pathways has not been consistent in different studies (Jin et al. Reference Jin, Wang, Liu, Hu, Zhao and Liu2016; Li et al. Reference Li, Tang, Su and Li2020a; Liu et al. Reference Liu, Shen, Zhao, Sun, Wang, Long, He, Lin, Wu and Wei2020a). For example, ERK inhibition decreased, whereas p38 MAPK inhibition increased the activation state of Nrf2 signaling pathway under resveratrol treatment (Jin et al. Reference Jin, Wang, Liu, Hu, Zhao and Liu2016). All three MAPK pathway inhibitors activated the expression of Nrf2 in particulate matter (PM)2.5-induced spermatogenesis dysfunction (Liu et al. Reference Liu, Shen, Zhao, Sun, Wang, Long, He, Lin, Wu and Wei2020a). Thus, the roles of MAPK and Nrf2 signaling pathways may be cell-specific or rely on the biological activity of different antioxidants.
Yao et al. (Reference Yao, Nussler, Liu, Hao, Song, Schirmeier and Nussler2007) found that the inhibitors of p38 MAPK and ERK blocked the protective effects of quercetin on alcoholic liver injury, whereas JNK inhibitor failed to do that. In this study, the Nrf2, p38 MAPK and ERK inhibitors completely blocked the effect of quercetin on cell proliferation, but the JNK inhibitor only blocked partially. Our observation was similar to the result of Yao et al. (Reference Yao, Nussler, Liu, Hao, Song, Schirmeier and Nussler2007) in alcoholic liver injury, suggesting that the antioxidation mechanisms of quercetin might be at least partially determined by the structure and characteristics of quercetin itself. Besides, the level of ROS is an indicator of cellular oxidation stress (Bottje Reference Bottje2019). Previous reports showed that p38 MAPK inhibitor reduced the elevation of ROS induced by ochratoxin A, and Nrf2 knockdown upregulated ROS level in rat hippocampal neurons, but the effects of Nrf2 and MAPK inhibitors on ROS of HC11 cells were not clear (Dai et al. Reference Dai, Zhang, Zhang and Yan2018; Han et al. Reference Han, Zhang, Wang, Liu, Jiang, Liu, Wang, Yang and Wei2019). In this study, Nrf2 inhibitor, but not MAPK inhibitors, partially blocked the downregulation effect of quercetin on ROS production. However, due to the presence of ERK-Nrf2 and p38 MAPK-Nrf2 pathways, MAPK pathway may also indirectly affect ROS production at other time points. Moreover, our observations indicated that p38 MAPK, ERK and Nrf2 pathways are all necessary in the enhancement of T-AOC by quercetin in HC11 cells, whereas JNK pathway is not. But interestingly, the JNK inhibitor had the greatest effect on CAT activity among four inhibitors. The changes of T-AOC and CAT activity were not completely consistent after 24 h of H2O2 treatment, suggesting that T-AOC in HC11 cells was determined by many antioxidant substances other than CAT enzyme, as other reports in the liver (Jia et al. Reference Jia, Peng, Zhao, Cheng, Zhou, Chai, Zeng, Pan and Xu2019; Liu et al. Reference Liu, Chang, Yu and Xu2020b).
Several studies have confirmed that xCT is a downstream gene of the Nrf2-antioxidant response element pathway (Xie et al. Reference Xie, Cai, Zhao, Li and Tian2020; Xie et al. Reference Xie, Cai, Zhao, Li and Tian2020). Repression of the activation of Nrf2-antioxidant response element pathway by si-RNA or overexpression of Nrf2’s anchor protein KEAP1 inhibited the expression of xCT (Ali et al. Reference Ali, Mohammad, Mujahed, Jonson-Videsäter, Nore, Paul and Lehmann2016; Habib et al. Reference Habib, Linher-Melville, Lin and Singh2015). In this study, the Nrf2 inhibitor completely blocked H2O2-induced protein expression of xCT, which was consistent with other studies in vivo and in vitro (Ali et al. Reference Ali, Mohammad, Mujahed, Jonson-Videsäter, Nore, Paul and Lehmann2016; Xie et al. Reference Xie, Cai, Zhao, Li and Tian2020). However, three MAPK inhibitors did not affect the expression of xCT. Hence, the effect of quercetin on xCT in HC11 cells was mainly dependent on the Nrf2 pathway. Ishikado et al. found that different si-RNAs of Nrf2 inhibited the expression of GCLM in liver and vascular endothelial cells, and the inhibitor or si-RNA of p38 MAPK also had the inhibitory effects on GCLM (Gu et al. Reference Gu, Tao, Xu, Han, Qi, Xu, Yin and Peng2016; Ishikado et al. Reference Ishikado, Sono, Matsumoto, Robida-Stubbs, Okuno, Goto, King, Keith Blackwell and Makino2013; Ji et al. Reference Ji, Sheng, Zheng, Shi and Wang2015). In HC11 cells, the recovery effect of quercetin on the protein level of GCLM was largely dependent on Nrf2 and p38 MAPK pathways and also partially relied on JNK and ERK pathways, proving that the level of GCLM may be affected by two pathways simultaneously.
Conclusion
In summary, our study showed that quercetin protected HC11 cells against H2O2-induced redox imbalance by regulating cell viability, ROS production, enzyme activity of CAT and the expression of stress-resistant proteins via MAPK and Nrf2 signaling pathways.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/anr.2024.2.
Author contributions
Yongxin Li: conceptualization, investigation, visualization and writing – original draft. Ning Han: methodology and investigation. Pengfei Hou: investigation. Feng-Qi Zhao: writing – review and editing. Hongyun Liu: funding acquisition, writing – review and editing, and supervision.
Financial support
This research was supported by grants from the National Natural Science Foundations of China (32072756) and the China Agricultural Research System (Grant No. CARS-36). The funders had no role in the design, analysis or writing of this article.
Conflicts of interest
The authors declare no conflicts of interest.







