Introduction
Schizophrenia is a chronic psychiatric disorder affecting approximately 1% of the global population, reducing lifespan by 20 years (Kahn et al., Reference Kahn, Sommer, Murray, Meyer-Lindenberg, Weinberger, Cannon and Insel2015). It involves generalized cognitive deficits across multiple domains (Dickinson, Ragland, Gold, & Gur, Reference Dickinson, Ragland, Gold and Gur2008; Fioravanti, Bianchi, & Cinti, Reference Fioravanti, Bianchi and Cinti2012; Le Roy, Reference Le Roy and Prouteau2011), severely impairing functional outcomes (McCutcheon, Keefe, & McGuire, Reference McCutcheon, Keefe and McGuire2023; Prouteau & Verdoux, Reference Prouteau, Verdoux and Prouteau2011). Cognitive deficits are present, although milder, at prodromal stages of the schizophrenia spectrum, i.e. in individuals at ‘ultra-high risk’ (UHR) of psychosis (Fusar-Poli et al., Reference Fusar-Poli, Deste, Smieskova, Barlati, Yung, Howes and Borgwardt2012, Reference Fusar-Poli, Borgwardt, Bechdolf, Addington, Riecher-Rössler, Schultze-Lutter and Yung2013). Around 25% of UHR convert to chronic psychosis within 3 years of follow-up (Salazar de Pablo et al., Reference Salazar de Pablo, Radua, Pereira, Bonoldi, Arienti, Besana and Fusar-Poli2021), and these converters exhibit more severe baseline cognitive deficits than non-converters (Fusar-Poli et al., Reference Fusar-Poli, Borgwardt, Bechdolf, Addington, Riecher-Rössler, Schultze-Lutter and Yung2013). Unaffected first-degree relatives (FDR) also present poorer cognition than healthy controls (HC) with no family history of schizophrenia (Snitz, MacDonald, & Carter, Reference Snitz, MacDonald and Carter2006). Understanding the mechanisms underlying cognitive deficits is essential for early intervention and prevention of disorder progression. However, cognitive assessments are time-consuming and use varied tests, hindering large cohort collection and cross-study comparisons.
To address this issue, the general factor of intelligence, or g-factor, appeared as a relevant construct. The g-factor measures overall cognitive ability, capturing common core features of cognitive ability across different cognitive tasks (Jensen, Reference Jensen1998; Spearman, Reference Spearman1904). It is considered as the single higher-order factor at the apex of the hierarchical intelligence models built upon the Cattell–Horn–Carroll theory of intelligence (Carroll, Reference Carroll1993, Reference Carroll and Nyborg2003; Cattell & Horn, Reference Cattell and Horn1978; Schneider & McGrew, Reference Schneider, McGrew, Flanagan and Harrison2012). Computationally, the g-factor can be defined as the first component of a principal component analysis (PCA) performed on an individual's cognitive test scores (Jensen, Reference Jensen1998). The g-factors derived from different test batteries have been demonstrated to be highly correlated (Johnson, Bouchard, Krueger, McGue, & Gottesman, Reference Johnson, Bouchard, Krueger, McGue and Gottesman2004, Reference Johnson, te Nijenhuis and Bouchard2008), supporting the existence of a single global intelligence factor and the consistency of its computation, provided the test batteries assess sufficiently diverse cognitive abilities (Dickinson, Goldberg, Gold, Elvevåg, & Weinberger, Reference Dickinson, Goldberg, Gold, Elvevåg and Weinberger2011). The g-factor thus enables comparisons and data pooling across studies with different cognitive assessments, a major stake in the era of big data.
Using a specifically selected neuropsychological battery, the g-factor has been validated as a higher-order cognitive factor in schizophrenia patients (SZ), their unaffected siblings, and HC (Dickinson et al., Reference Dickinson, Goldberg, Gold, Elvevåg and Weinberger2011). However, further investigation is required to ascertain its generalizability to other samples and test batteries, and its potential in discriminating subgroups along the schizophrenia spectrum. The literature would also benefit from a more comprehensive understanding of the sociodemographic, clinical, and genetic contributions to cognition, which could, in turn, inform the development of alternative treatments. Indeed, antipsychotic medication, the primary treatment for psychosis, appeared relatively ineffective in reducing cognitive deficits (Keefe et al., Reference Keefe, Sweeney, Gu, Hamer, Perkins, McEvoy and Lieberman2007; Millan et al., Reference Millan, Andrieux, Bartzokis, Cadenhead, Dazzan, Fusar-Poli and Weinberger2016).
Cognitive phenotypes have a high heritability (Blokland et al., Reference Blokland, Mesholam-Gately, Toulopoulou, del Re, Lam, DeLisi and Petryshen2017; Mallet, Le Strat, Dubertret, & Gorwood, Reference Mallet, Le Strat, Dubertret and Gorwood2020). Polygenic risk scores (PRS) estimate the genetic heritability of traits based on genome-wide association studies (GWAS) (Choi, Mak, & O'Reilly, Reference Choi, Mak and O'Reilly2020). Previous research by our team and others found significant correlations between cognition and several PRS in UHR patients (He et al., Reference He, Jantac Mam-Lam-Fook, Chaignaud, Danset-Alexandre, Iftimovici, Gradels Hauguel and Chaumette2021) and within SZ cases (Richards et al., Reference Richards, Pardiñas, Frizzati, Tansey, Lynham, Holmans and Walters2019), but lacked comparisons with HC and the full schizophrenia spectrum.
The present study had three objectives: first, to compute and validate g-factors derived from different cognitive test batteries as global cognitive scores that can discriminate between the stages of the schizophrenia spectrum; second, to use the g-factors to investigate potential mechanisms underlying cognitive impairment in the schizophrenia spectrum; and third, to determine which test battery produced the most informative g-factor for discriminating schizophrenia spectrum subgroups and correlating with other dimensions. PCA was used to derive 10 g-factors, the correlations of which with intelligence quotient and with one another were examined for validation as global cognitive scores. Analysis of variance (ANOVA) assessed differences across subgroups, namely HC, FDR, UHR, first-episode psychosis (FEP), and SZ. Correlations between the g-factors, sociodemographic, and clinical data were investigated. Linear regressions between the g-factors and the PRS for cognitive performance and for schizophrenia were conducted, and results pooled through meta-analyses to establish global correlations between PRS and cognition.
Materials and methods
Participants
The present retrospective study compiled data from 892 individuals included in six previous studies coordinated by M.-O. K. at GHU Paris Psychiatry and Neuroscience, assessing various aspects, including the influence of genetics or environmental factors in psychosis and its transition. Baseline visits were selected for interventional and longitudinal studies to prevent bias. Data were collected between 2005 and 2021. Further details on the individual studies can be found in previous publications (Gay et al., Reference Gay, Plaze, Oppenheim, Mouchet-Mages, Gaillard, Olié and Cachia2013; Krebs et al., Reference Krebs, Magaud, Willard, Elkhazen, Chauchot, Gut and Kazes2014; Magaud et al., Reference Magaud, Morvan, Rampazzo, Alexandre, Willard, Gaillard and Krebs2014; Martinez et al., Reference Martinez, Alexandre, Mam-Lam-Fook, Bendjemaa, Gaillard, Garel and Krebs2017). Study abbreviations are detailed in the ‘Acknowledgments’ section.
Inclusion criteria for the present analyses were an age over 15 years old, and diagnoses within the schizophrenia spectrum: 338 UHR identified using the Comprehensive Assessment of At-Risk Mental State (Krebs et al., Reference Krebs, Magaud, Willard, Elkhazen, Chauchot, Gut and Kazes2014) (ICAAR, START, PrEPP studies), 100 FEP (ICAAR and PRIMEPI studies), and 167 SZ diagnosed with DSM-IV criteria after the Diagnosis Interview for Genetic Studies 3.0 (Nurnberger et al., Reference Nurnberger, Blehar, Kaufmann, York-Cooler, Simpson, Harkavy-Friedman and Reich1994) (AUSZ, PrEPP, PRIMEPI, and PsyDEV studies). Additionally, 124 HC with no personal or familial psychiatric history up to second-degree relatives were included (AUSZ, START, and PsyDEV studies), and 163 FDR (PsyDEV study). Exclusion criteria included severe somatic or neurological disorders, intelligence quotient below 70, severe substance use disorders for more than 5 years, or involuntary hospitalization. Free and informed written consent was obtained from all participants or their legal representatives. The studies adhered to the French regulatory framework and received ethics committees approvals.
Cognitive data
The French version of the Wechsler Adult Intelligence Scale (WAIS) was administered in four studies: PrEPP, ICAAR, and AUSZ studies used the 3rd edition (WAIS-III) (Wechsler, Reference Wechsler1997) with 199 UHR, 35 FEP, 53 SZ, and 25 HC; START study used the 4th edition (WAIS-IV) (Wechsler, Reference Wechsler2008) with 44 UHR and 7 HC. The WAIS is commonly used to derive the g-factor, hence our focus on this scale and its subtests.
Due to time constraints in larger studies, a shorter cognitive assessment, referred to as ‘Minicog’, was used in five studies to reduce participant burden. This assessment takes approximately 30 min, compared to 90 min for the WAIS. The ICAAR and AUSZ studies used the complete Minicog (‘Minicog 6’) with 106 UHR, 29 FEP, 34 SZ, and 21 HC. The ICAAR, AUSZ, START, PsyDEV, and PRIMEPI studies used the partial Minicog (‘Minicog 3’) with 157 UHR, 77 FEP, 93 SZ, 43 HC, and 92 FDR. The Minicog comprised: Verbal Fluency Test (Cardebat, Doyon, Puel, Goulet, & Joanette, Reference Cardebat, Doyon, Puel, Goulet and Joanette1990); Trail Making Test A & B (Reitan, Reference Reitan1956); Proverbs, i.e. item N5 from the Positive and Negative Syndrome Scale (PANSS) (Lançon, Reine, Llorca, & Auquier, Reference Lançon, Reine, Llorca and Auquier1999); WAIS Similarities; Wechsler Memory Scale-IV Coupled Words subtest (Wechsler, Reference Wechsler2010); and the French adaptation of the National Adult Reading Test (Mackinnon, Ritchie, & Mulligan, Reference Mackinnon, Ritchie and Mulligan1999). The partial Minicog included the first three of these tests.
Computation of the g-factors
The g-factors were derived from 10 combinations of neuropsychological tests. Seven g-factors were derived from the WAIS: g WAIS global was derived from the Verbal and Performance intelligence quotients of the WAIS-III (n = 323 individuals); g WAIS indices was derived from the four indices of the WAIS-IV, i.e. Verbal Comprehension, Working Memory, Perceptual Reasoning, and Processing Speed (n = 51); g WAIS-III was derived from the nine main subtests of the WAIS-III (n = 312); g WAIS-IV was derived from the 10 main subtests of the WAIS-IV (n = 51); g WAIS 8 was derived from the eight subtests that were common to both WAIS-III and WAIS-IV (n = 363); g WAIS 4 was derived from four subtests representing each index of the WAIS-IV (Gignac, Reference Gignac2015), i.e. Similarities, Digit Span, Matrix Reasoning, and Symbol Search (n = 51); and g WAIS 2 was derived from Coding and Information, identified as the most valid and reliable dyad for estimating the Full-Scale Intelligence Quotient (FSIQ) (Girard, Axelrod, Patel, & Crawford, Reference Girard, Axelrod, Patel and Crawford2015) (n = 366). It is important to note the distinction between the g-factor and FSIQ, although the WAIS is based on the Cattell–Horn–Carroll theory, as are most modern intelligence tests. The FSIQ is a composite score derived from standardized intelligence subtests measuring an individual's overall cognitive ability compared to a normative sample, whereas the g-factor is a theoretical construct derived from statistical analysis and representing underlying cognitive ability across various cognitive tasks. Two g-factors were derived from the Minicog: g Minicog 6 derived from the complete Minicog (n = 190) and g Minicog 3 derived from the partial Minicog (n = 462). Finally, one g-factor was computed by pooling data from the WAIS and the Minicog, using WAIS FSIQ and the scores of the complete Minicog, excluding WAIS Similarities to avoid redundancy (n = 239). In total, at least one g-factor could be computed for 598 individuals.
Whole-genome genotyping data
Both cognitive and genotyping data were available for 220 individuals (40 HC, 104 UHR, 5 FEP, 71 SZ). Despite the small sample size for FEP, their data were retained as our objective was to investigate the relationship between cognition and genetics across the schizophrenia spectrum as a whole, rather than within each subgroup. Genotyping was conducted using the high-throughput genome-wide Illumina Infinium PsychArray-24 v1.3 BeadChip (Illumina, California, USA). Summary statistics from recent GWAS related to cognitive performance (Lee et al., Reference Lee, Wedow, Okbay, Kong, Maghzian, Zacher and Cesarini2018) and schizophrenia (Schizophrenia Working Group of the PGC et al., Reference Trubetskoy, Pardiñas, Qi, Panagiotaropoulou, Awasthi, Bigdeli and O'Donovan2022) were used as base data.
Computation of the PRS
Quality control on raw genotyping data was performed using PLINK (v1.9) (Chang et al., Reference Chang, Chow, Tellier, Vattikuti, Purcell and Lee2015), following standard procedures (Choi et al., Reference Choi, Mak and O'Reilly2020; He et al., Reference He, Jantac Mam-Lam-Fook, Chaignaud, Danset-Alexandre, Iftimovici, Gradels Hauguel and Chaumette2021; Marees et al., Reference Marees, de Kluiver, Stringer, Vorspan, Curis, Marie-Claire and Derks2018). Single-nucleotide polymorphisms (SNP) with a missingness rate exceeding 2% and Hardy–Weinberg equilibrium p values below 10–6 were removed. Pruning retained only independent SNP. A sex check excluded one SZ patient with inconsistent gender assignment. Imputation was performed with the Sanger Imputation Service (McCarthy et al., Reference McCarthy, Abecasis, Cardon, Goldstein, Little, Ioannidis and Hirschhorn2008) to enhance statistical power (He et al., Reference He, Jantac Mam-Lam-Fook, Chaignaud, Danset-Alexandre, Iftimovici, Gradels Hauguel and Chaumette2021; Porcu, Sanna, Fuchsberger, & Fritsche, Reference Porcu, Sanna, Fuchsberger and Fritsche2013), using EAGLE2 for pre-phasing and PBWT for imputation on a reference panel combining 1000 Genomes Phase 3 and UK10K. Imputed SNP with information scores below 0.8, missingness rates above 2%, and deviations from Hardy–Weinberg equilibrium were filtered out. The remaining SNP were pruned. Relatedness analysis excluded five FDR or second-degree relatives, using a cut-off coefficient of 0.125. Population genetic stratification was assessed with a PCA using Peddy (Pedersen & Quinlan, Reference Pedersen and Quinlan2017) and excluded 19 ethnic outliers, retaining only individuals of European ancestry.
Following quality control, 195 individuals remained (38 HC, 88 UHR – of whom 85 were also included in a previous analysis from our team focused on UHR only [He et al., Reference He, Jantac Mam-Lam-Fook, Chaignaud, Danset-Alexandre, Iftimovici, Gradels Hauguel and Chaumette2021] – 5 FEP, 64 SZ). A total of 131 individuals had a g WAIS global, 130 a g WAIS-III, 143 a g WAIS 8, 144 a g WAIS 2, 109 a g Minicog 6, 178 a g Minicog 3, and 127 a g Mix. Thirteen individuals had a g WAIS indices, a g WAIS-IV, and a g WAIS 4, but those samples were excluded from PRS calculations due to the small sample size.
PRS were computed using PRSice-2 (v2.3.3) (Choi & O'Reilly, Reference Choi and O'Reilly2019), filtering out SNP with information scores below 0.8 or minor allele frequencies below 5%. Analyses were conducted for each PRS and the seven g-factor samples at 14 p value thresholds (He et al., Reference He, Jantac Mam-Lam-Fook, Chaignaud, Danset-Alexandre, Iftimovici, Gradels Hauguel and Chaumette2021), retaining scores computed at the most predictive threshold for each PRS.
Psychopathological and neurodevelopmental data
Patients completed the PANSS (Kay, Fiszbein, & Opler, Reference Kay, Fiszbein and Opler1987; Lançon et al., Reference Lançon, Reine, Llorca and Auquier1999) that assessed positive and negative symptoms, and general psychopathology; the Brief Psychiatric Rating Scale-Expanded (BPRS-E) (Dingemans, Linszen, Lenior, & Smeets, Reference Dingemans, Linszen, Lenior and Smeets1995; Mouaffak et al., Reference Mouaffak, Morvan, Bannour, Chayet, Bourdel, Thepaut and Krebs2010) that assessed general psychiatric symptoms; the Developmental Disorders Screening Scale (DDSS) (Martinez, Reference Martinez2017) that assessed developmental disorders in childhood; and the Neurological Soft Signs Examination (NSS) (Krebs, Gut-Fayand, Bourdel, Dischamp, & Olié, Reference Krebs, Gut-Fayand, Bourdel, Dischamp and Olié2000) that assessed subtle integrative neurological anomalies.
Statistical analyses
Statistical analyses were conducted using R (R Core Team, 2022) with a significance level of 0.05. Descriptive statistics, one-way ANOVA, and pairwise comparisons were performed on sociodemographic, cognitive, and clinical data using the compareGroups package (Subirana, Sanz, & Vila, Reference Subirana, Sanz and Vila2014).
Bartlett's sphericity tests from the psych package (Revelle, Reference Revelle2022) were used for data redundancy analysis, and PCA on 10 neuropsychological test battery scores using FactoMineR (Lê, Josse, & Husson, Reference Lê, Josse and Husson2008) provided the g-factors. Spearman's correlations between the g-factors and FSIQ, corrected for multiple comparisons using Benjamini–Hochberg (BH) method (Benjamini & Hochberg, Reference Benjamini and Hochberg1995), were computed using stats (R Core Team, 2022) and psych (Revelle, Reference Revelle2022) packages, and visualized using corrplot (Wei & Simko, Reference Wei and Simko2021)
One-way between-subjects ANOVA assessed subgroup differences in g-factors using the stats package (R Core Team, 2022), with post hoc comparisons based on estimated marginal means from the emmeans package (Lenth, Reference Lenth2022), corrected using the BH method. Effect sizes were measured with the parameter dm, analogous to Cohen's d (Bögge, Colás-Blanco, & Piolino, Reference Bögge, Colás-Blanco and Piolino2022; Cohen, Reference Cohen1988).
Spearman's correlations between the g-factors, sociodemographic, and clinical data, corrected using the BH method, were analyzed. Linear regressions of standardized g-factors against standardized PRS were conducted using stats (R Core Team, 2022), adjusting for sex, age, and 10 top principal components from population stratification. All g-factors and PRS were scaled to have a mean of 0 and a standard deviation of 1 to standardize the regression estimates as β coefficients (Richards et al., Reference Richards, Pardiñas, Frizzati, Tansey, Lynham, Holmans and Walters2019). Effect sizes were converted from β coefficients to correlations using esc (Lüdecke, Reference Lüdecke2019), and pooled for each PRS through meta-analyses based on the generic inverse variance method with a fixed-effects model, using meta (Balduzzi, Rücker, & Schwarzer, Reference Balduzzi, Rücker and Schwarzer2019). Between-sample heterogeneity was quantified using Higgins and Thompson's I 2 statistic (Higgins & Thompson, Reference Higgins and Thompson2002).
Graphs were created using ggplot2 (Wickham, Reference Wickham2009), ggpubr (Kassambara, Reference Kassambara2020), ggsignif (Ahlmann-Eltze & Patil, Reference Ahlmann-Eltze and Patil2021), and ggExtra (Attali & Baker, Reference Attali and Baker2022), and forest plots using meta (Balduzzi et al., Reference Balduzzi, Rücker and Schwarzer2019).
Results
Sample description
Descriptive statistics, ANOVA p values, and significant pairwise differences, for the sociodemographic, cognitive, and clinical data across the schizophrenia spectrum are provided in online Supplementary Table S1.
Regarding sociodemographic data, sex ratios were balanced for HC and FDR, but skewed toward males for UHR, FEP, and SZ (approximately 70% male). Mean age differed significantly between groups, reflecting the disease severity progression over time. There was also a bias for FDR, who were significantly older than other groups, given that they were mostly patients' parents. Only HC had significantly more years of education than all other groups (all p < 0.001).
Regarding cognitive data, most WAIS scores showed no significant difference between subgroups. Conversely, most tests in the Minicog battery presented significant differences between HC and patients, as well as between patient groups.
Regarding clinical scores, PANSS, BPRS, and NSS scores were found to worsen with disease severity. No group differences were observed for DDSS. Chlorpromazine equivalent doses significantly increased with disease severity.
Computation of the g-factors and comparison across the schizophrenia spectrum
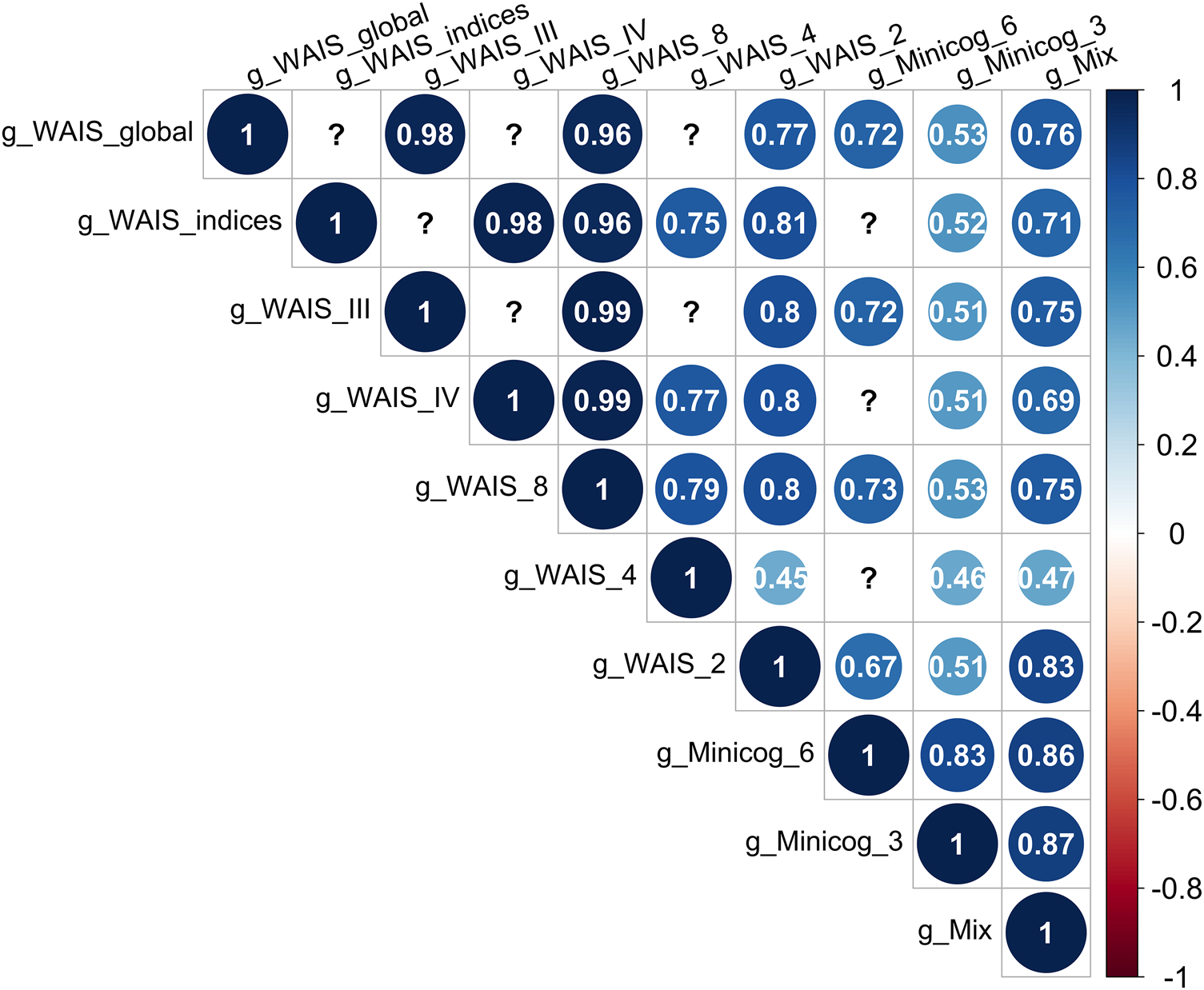
Bartlett's sphericity tests were significant, justifying the use of PCA on the 10 test combinations (Table 1). The g-factors explained between 36% and 81% of the total variance in cognitive scores. Pairwise correlations between each of the g-factors and WAIS FSIQ were significant and strong (Spearman's r between 0.55 and 0.99), as were correlations among g-factors (r > 0.50 for each pairwise correlation) (Fig. 1).
Combinations of cognitive tests used for the computation of 10 g-factors, proportion of total variance explained by each g-factor and its correlation with WAIS FSIQ
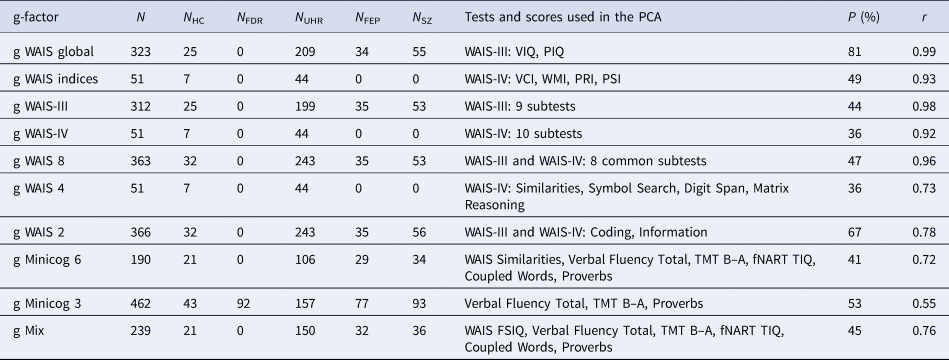
N, total sample size; N HC, number of healthy controls; N FDR, number of first-degree relatives; N UHR, number of patients at ultra-high risk of psychosis; N FEP, number of patients who had a first-episode psychosis; N SZ, number of patients with schizophrenia; PCA, principal component analysis; P: proportion of total variance explained by the g-factor; r, Spearman's correlation between the g-factor and WAIS (Wechsler Adult Intelligent Scale) full-scale intelligence quotient (FSIQ).
VIQ, verbal IQ; PIQ, performance IQ; VCI, Verbal Comprehension index; WMI, Working Memory index; PRI, Perceptual Reasoning index; PSI, Processing Speed index; TMT, Trail Making Test; fNART, French adaptation of the National Adult Reading Test.
Correlations between the g-factors derived from different cognitive test combinations. All correlations are significant at a p value threshold of 0.05. The question marks correspond to correlations which could not be computed because the two g-factors were not available on the same sample of individuals.
The ANOVA performed for each g-factor revealed significant group differences for seven out of the 10 g-factors, i.e. g WAIS global (F (3,319) = 4.65, p = 0.003, η G2 = 0.04), g WAIS-III (F (3,308) = 3.99, p = 0.008, η G2 = 0.04), g WAIS 8 (F (3,359) = 5.75, p = 0.001, η G2 = 0.05), g WAIS 2 (F (3,362) = 5.37, p = 0.001, η G2 = 0.04), g Minicog 6 (F (3,186) = 10.30, p < 0.001, η G2 = 0.14), g Minicog 3 (F (4,457) = 39.08, p < 0.001, η G2 = 0.26), and g Mix (F (3,235) = 17.98, p < 0.001, η G2 = 0.19) (Fig. 2).
Comparison of the g-factors across the schizophrenia spectrum. HC, healthy controls; FDR, first-degree relatives; UHR, patients at ultra-high risk of psychosis; FEP, patients who had a first-episode psychosis; SZ, patients with schizophrenia. p value significance: *p < 0.05; **p < 0.01; ***p < 0.001.
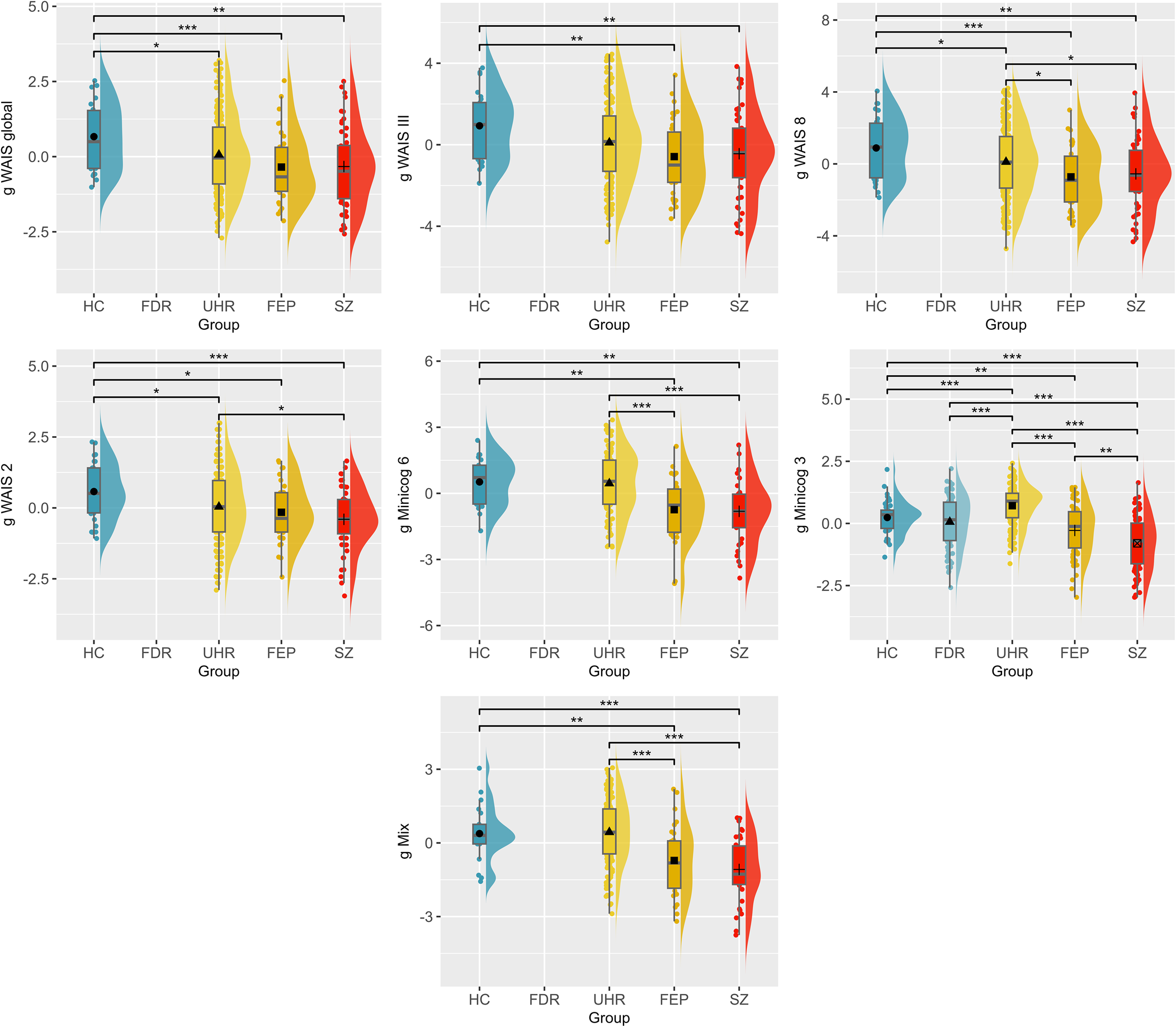
All seven g-factors were significantly higher for HC compared to FEP (g WAIS global: p = 0.01, dm = 0.81; g WAIS-III: p = 0.01, dm = 0.77; g WAIS 8: p = 0.002, dm = 0.85; g WAIS 2: p = 0.02, dm = 0.64; g Minicog 6: p = 0.01, dm = 0.86; g Minicog 3: p = 0.01, dm = 0.51; g Mix: p = 0.01, dm = 0.80), and to SZ (g WAIS global: p = 0.01, dm = 0.79; g WAIS-III: p = 0.01, dm = 0.70; g WAIS 8: p = 0.002, dm = 0.76; g WAIS 2: p = 0.001, dm = 0.86; g Minicog 6: p = 0.001, dm = 0.98; g Minicog 3: p < 0.0001, dm = 1.15; g Mix: p < 0.001, dm = 1.20). Four g-factors also differed significantly between HC and UHR (g WAIS global: p = 0.048, dm = 0.48; g WAIS 8: p = 0.04, dm = 0.41; g WAIS 2: p = 0.02, dm = 0.47; g Minicog 3: p = 0.02, dm = −0.44). The g Minicog 3 further differentiated FDR from UHR (p < 0.0001, dm = −0.69) and SZ (p < 0.0001, dm = 0.90), with better cognitive scores in the FDR group relative to SZ, but worse scores compared to UHR.
Regarding comparisons between patient groups, five g-factors distinguished UHR and SZ (g WAIS 8: p = 0.03, dm = 0.35; g WAIS 2: p = 0.02, dm = 0.39; g Minicog 6: p < 0.001, dm = 0.89; g Minicog 3: p < 0.0001, dm = 1.58; g Mix: p < 0.0001, dm = 1.22). Four g-factors discriminated UHR from FEP (g WAIS 8: p = 0.03, dm = 0.44; g Minicog 6: p = 0.001, dm = 0.77; g Minicog 3: p < 0.0001, dm = 0.95; g Mix: p < 0.001, dm = 0.82). Additionally, g Minicog 3 differentiated FEP and SZ (p < 0.001, dm = 0.64).
All g-factors showed higher scores for HC and decreased with the more advanced stages of the disease.
Relationships between cognition and sociodemography, psychopathology, neurodevelopmental load, medication, and genetics
No significant correlations were observed between three g-factors (g WAIS indices, g WAIS-IV, and g WAIS 4) and sociodemographic and clinical data. The results presented in this section will therefore focus on the seven other g-factors, i.e. g WAIS global, g WAIS III, g WAIS 8, g WAIS 2, g Minicog 6, g Minicog 3, and g Mix (see online Supplementary Table S2 for details).
Regarding sociodemographic data, Spearman's correlations indicated that all seven g-factors were significantly positively correlated with educational attainment. Only g Minicog 3 was negatively correlated with age.
Regarding psychopathological data, four out of the seven g-factors (i.e. g WAIS 2, g Minicog 6, g Minicog 3, and g Mix) were negatively correlated with PANSS Total score; three g-factors were negatively correlated with PANSS Positive (i.e. g WAIS 8, g Minicog 3, and g Mix); three were negatively correlated with PANSS Negative (i.e. g Minicog 6, g Minicog 3, and g Mix), and six were negatively correlated with PANSS General Psychopathology score (all seven g-factors except g WAIS 8). Only g Minicog 3 was negatively correlated with BPRS Total score.
Regarding neurodevelopmental data, six g-factors (all seven g-factors except for g Minicog 6) were negatively correlated with NSS Total score; one g-factor (g Minicog 8) was negatively correlated with NSS Motor Coordination factor; six were negatively correlated with NSS Motor Integration factor (all seven g-factors except for g WAIS 2); all seven g-factors were negatively correlated with NSS Sensory Integration factor; and one g-factor (g Minicog 3) was negatively correlated with NSS Involuntary Movements factor. No g-factor was significantly correlated with NSS Lateralization factor nor with DDSS scores.
Regarding medication, all seven g-factors were negatively correlated with chlorpromazine equivalent dose.
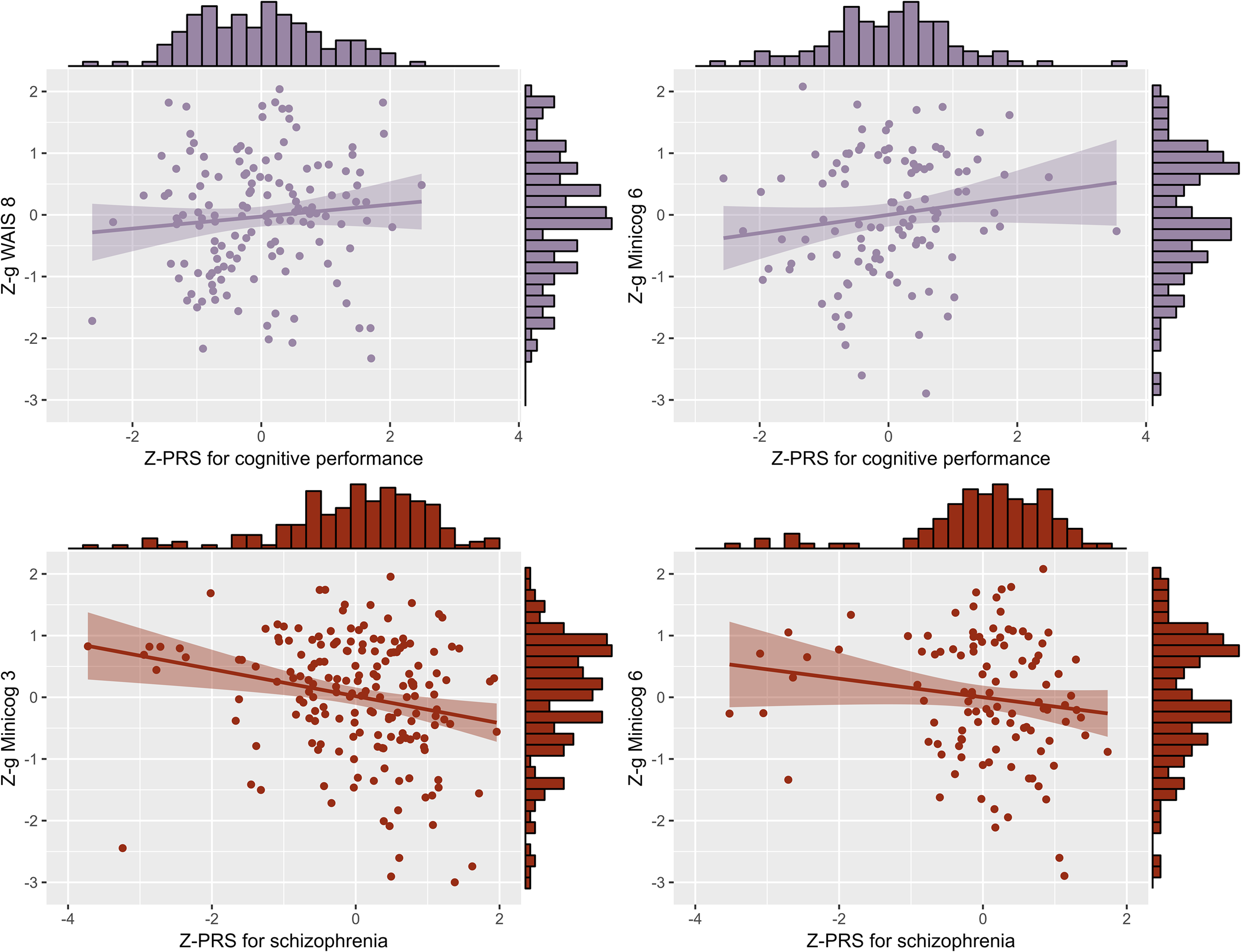
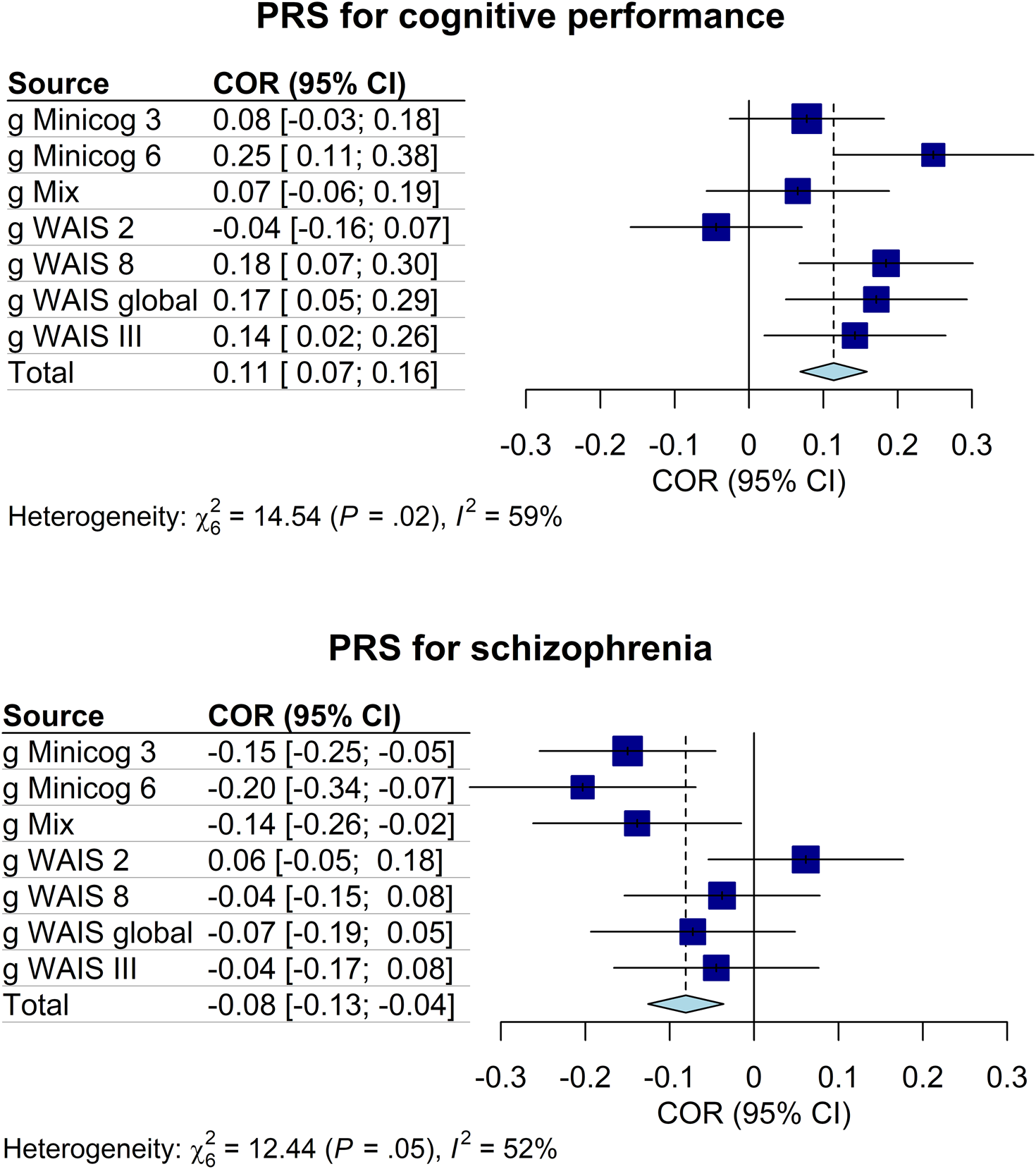
Regarding genetics, linear regressions of the g-factors against the PRS indicated that the PRS for cognitive performance had a significant positive influence on g WAIS 8 (P T = 0.3, standardized estimate β = 0.19, standard error [s.e.] = 0.09, p = 0.04, proportion of g-factor variance explained by the PRS PRS.R2 = 0.03) and on g Minicog 6 (P T = 1 × 10−5, β = 0.25, s.e. = 0.10, p = 0.01, PRS.R2 = 0.05). The PRS for schizophrenia had a significant negative influence on g Minicog 6 (P T = 1 × 10−5, β = −0.20, s.e. = 0.10, p = 0.03, PRS.R2 = 0.04), and on g Minicog 3 (P T = 1 × 10−5, β = −0.15, s.e. = 0.07, p = 0.04, PRS.R2 = 0.02) (Fig. 3 and online Supplementary Table S3). The meta-analysis of regression results revealed a significant positive correlation between the PRS for cognitive performance and the g-factor (pooled effect size r pooled = 0.11, p < 0.0001, I 2 = 58.7%), and a significant negative correlation between the PRS for schizophrenia and the g-factor (r pooled = −0.08, p < 0.001, I 2 = 51.8%) (Fig. 4 and online Supplementary Table S4). Upon examination of the g-factors individually, those derived from WAIS scores and the complete Minicog demonstrated a stronger correlation with the PRS for cognitive performance, compared to those derived from the partial Minicog, from only two WAIS subtests, or g Mix. Conversely, the g-factors derived from WAIS scores exhibited weaker correlations with the PRS for schizophrenia compared to those derived from Minicog scores and g Mix.
Significant linear regression models of the standardized g-factors against the standardized PRS for cognitive performance and for schizophrenia. Z-g, standardized g-factor; Z-PRS, standardized polygenic risk score.
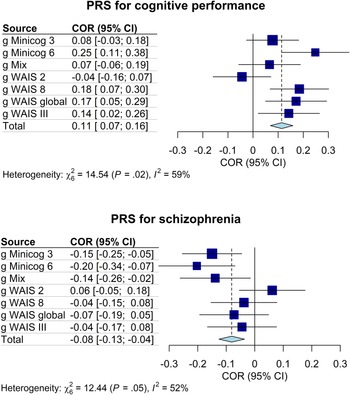
Forest plots of the correlations between the g-factors and the PRS for cognitive performance and for schizophrenia. COR, effect size as correlation; CI, confidence interval; I 2, Higgins and Thompson's between-sample heterogeneity index. Error bars represent the 95% confidence intervals of the means.
Discussion
In this study, we established and validated g-factors derived from various cognitive test batteries, providing global cognitive scores sensitive to subgroups within the schizophrenia spectrum. The g-factors were used to investigate underlying mechanisms of cognitive impairment in the spectrum. Results indicated that educational attainment and the PRS for cognitive performance were positively associated with cognition, whereas general schizophrenia psychopathology, neurodevelopmental load, medication, and the PRS for schizophrenia were negatively associated with cognitive scores across the spectrum.
The g-factors derived from 10 distinct batteries strongly correlated with FSIQ, thereby confirming their association with global intelligence (Canivez & Watkins, Reference Canivez and Watkins2010). Despite differing methodologies, both the g-factor and FSIQ quantify general cognitive ability, serving distinct purposes in psychological assessment and research. The g-factors explained a large proportion of the variance in neuropsychological scores and were overall positively correlated with the PRS for cognitive performance, providing further evidence of their validity as global cognitive scores. Moreover, high correlations among the g-factors supported Spearman's principle of indifference of the indicator (Dickinson et al., Reference Dickinson, Goldberg, Gold, Elvevåg and Weinberger2011; Johnson et al., Reference Johnson, Bouchard, Krueger, McGue and Gottesman2004, Reference Johnson, te Nijenhuis and Bouchard2008; Spearman, Reference Spearman1904).
The following discussion will focus on seven g-factors, excluding g WAIS indices, g WAIS-IV, and g WAIS 4, which were only available for two unbalanced subgroups (7 HC and 44 UHR) and a limited sample size that did not pass our post hoc power analyses. Overall, the averaged g-factors showed a gradual decline across the schizophrenia spectrum. Three g-factors presented significantly lower scores for UHR than HC: although the effect sizes were small, they confirm that mild cognitive deficits are present at the prodromal stage (Fusar-Poli et al., Reference Fusar-Poli, Deste, Smieskova, Barlati, Yung, Howes and Borgwardt2012, Reference Fusar-Poli, Borgwardt, Bechdolf, Addington, Riecher-Rössler, Schultze-Lutter and Yung2013). All seven g-factors were lower on average in the more advanced stages of schizophrenia compared to prodromal stages, with medium-to-large effect sizes, but the difference was smaller between FEP and SZ. These findings indicate that cognitive impairment increases from prodromal to FEP stages, followed by a relative stabilization after the first episode. Notably, one g-factor, g Minicog 3, was on average lower for HC and FDR than UHR, potentially linked to age-related cognitive decline in the FDR, who were significantly older; other studies have also shown that some cognitive scores of FDR were more closely aligned with those of FEP than of HC (Scoriels et al., Reference Scoriels, Salek, Goodby, Grainger, Dean, West and Jones2015). The g-factors enabled overall to discriminate between the stages of the schizophrenia spectrum when WAIS scores did not. As the g-factors capture the fundamental cognitive ability shared across tasks, they may have reduced within-group variability, thereby facilitating the detection of between-group variability. In contrast, the IQ profiles within each subgroup of the schizophrenia spectrum might have been heterogenous (Magaud et al., Reference Magaud, Morvan, Rampazzo, Alexandre, Willard, Gaillard and Krebs2014), resulting in high within-group variability.
Correlations between the g-factors, and sociodemographic and clinical data revealed that educational attainment was associated with higher cognition, while psychopathological symptoms, neurodevelopmental load, and medication negatively affected cognitive performance. Specifically, six g-factors were negatively correlated with PANSS General Psychopathology score, whereas only one was negatively correlated with BPRS Total score, highlighting the specific relationship between cognitive functioning and schizophrenia psychopathology (Chavez-Baldini et al., Reference Chavez-Baldini, Nieman, Keestra, Lok, Mocking, de Koning and Denys2023), but not with general psychiatric psychopathology. However, only three g-factors were correlated with PANSS Positive and Negative scores, in accordance with previous research indicating that cognitive alterations are relatively independent of schizophrenia positive and negative symptoms and constitute a distinctive dimension of the disease (Bell, Lysaker, Milstein, & Beam-Goulet, Reference Bell, Lysaker, Milstein and Beam-Goulet1994; Capatina, Miclutia, & Toma, Reference Capatina, Miclutia and Toma2017; de Gracia Dominguez, Viechtbauer, Simons, van Os, & Krabbendam, Reference de Gracia Dominguez, Viechtbauer, Simons, van Os and Krabbendam2009). Six g-factors were also negatively correlated with NSS Total score and Motor Integration factor, and seven g-factors were negatively correlated with NSS Sensory Integration factor, which is consistent with models of a neurodevelopmental origin of cognitive impairment in schizophrenia (Bora, Reference Bora2015). Finally, seven g-factors were negatively correlated with chlorpromazine equivalent dose, which corroborates previous findings that higher doses of medication were associated with poorer cognition in schizophrenia (Haddad, Salameh, Sacre, Clément, & Calvet, Reference Haddad, Salameh, Sacre, Clément and Calvet2023; Sweeney, Keilp, Haas, Hill, & Weiden, Reference Sweeney, Keilp, Haas, Hill and Weiden1991).
The genetic contribution to cognitive impairment in the schizophrenia spectrum was also investigated. The g-factors were positively correlated with the PRS for cognitive performance and negatively correlated with the PRS for schizophrenia, when considering the entire spectrum without differentiating the subgroups, suggesting that genetic factors related to both cognitive performance and schizophrenia influence cognition in the schizophrenia spectrum. These results were congruent with those previously found by our team (He et al., Reference He, Jantac Mam-Lam-Fook, Chaignaud, Danset-Alexandre, Iftimovici, Gradels Hauguel and Chaumette2021) and another study (Hubbard et al., Reference Hubbard, Tansey, Rai, Jones, Ripke, Chambert and Zammit2016), with the difference that those previous studies explored correlations between FSIQ and the PRS for schizophrenia in UHR patients and in children from a population-based birth cohort, respectively. Therefore, our findings demonstrate the consistency between results based on the FSIQ and on the g-factor, and they generalize the relationship between cognition and the PRS for schizophrenia to the whole schizophrenia spectrum. Prior research has also shown that cognitive phenotypes partially share genetic etiology with schizophrenia liability (Blokland et al., Reference Blokland, Mesholam-Gately, Toulopoulou, del Re, Lam, DeLisi and Petryshen2017; Kendler, Ohlsson, Sundquist, & Sundquist, Reference Kendler, Ohlsson, Sundquist and Sundquist2015; Koch, Caldwell, & Fuchs, Reference Koch, Caldwell and Fuchs2013; Legge et al., Reference Legge, Cardno, Allardyce, Dennison, Hubbard, Pardiñas and Walters2021; Mallet et al., Reference Mallet, Le Strat, Dubertret and Gorwood2020; Mistry, Harrison, Smith, Escott-Price, & Zammit, Reference Mistry, Harrison, Smith, Escott-Price and Zammit2018), although cognition in schizophrenia appears to be explained to a larger extent by alleles associated with cognition in the global population than risk alleles for schizophrenia per se (Mallet et al., Reference Mallet, Le Strat, Dubertret and Gorwood2020; Richards et al., Reference Richards, Pardiñas, Frizzati, Tansey, Lynham, Holmans and Walters2019). When considered individually, the g-factors derived from WAIS scores were more correlated with the genetics linked with cognitive performance than those derived from Minicog scores and g WAIS 2, which was expected since the WAIS is designed primarily to evaluate global intelligence. Conversely, the g-factors derived from Minicog scores were more correlated with genetics linked to schizophrenia, which can be explained by their stronger correlations with psychopathology.
Ultimately, the g-factor derived from the partial Minicog appeared as the most informative one, as it enabled discriminating between subgroups of the schizophrenia spectrum, and was correlated with sociodemography, psychopathology, neurodevelopmental load, and schizophrenia genetics.
Several limitations should be considered. First, the sample sizes for each subgroup were relatively small, especially for genetic analyses. Second, the observed decline in the g-factor across disease stages represents an average population and does not account for within-group variability, limiting its ability to discriminate between individuals due to substantial cognitive heterogeneity. Additionally, the cross-sectional design of the study warrants longitudinal investigations to elucidate cognitive evolution across stages within individual trajectories, given the varied trajectories of cognitive development observed within the schizophrenia spectrum (Reckziegel et al., Reference Reckziegel, Czepielewski, Hasse-Sousa, Martins, de Britto, Lapa and Gama2022). Third, the correlational nature of the analyses implied associations rather than causality, and the global approach did not discern associations between cognition and other factors within each subgroup of the spectrum separately.
Nevertheless, the study aimed to demonstrate that the g-factor could constitute a relevant indicator of the cognitive state of patients, sensitive to schizophrenia spectrum stages, and provide insights into potential mechanisms underlying cognitive impairment. Using the g-factor as a global cognitive score enabled data pooling across studies with different cognitive assessments, enhancing statistical power. Other methods have been proposed for computing global cognitive ability while pooling data across different studies, such as composite scores based on the mean of all standardized cognitive scores (Anda et al., Reference Anda, Brønnick, Johannessen, Joa, Kroken, Johnsen and Løberg2019). While composite scores, which include all available data and are closer to original test scores, are better able to capture specific deficits and thus more relevant for clinical assessment and treatment, the reductionist approach of PCA simplifies cognitive data by eliminating redundancy and heterogeneity that may hinder between-group comparisons. Moreover, despite the g-factor being an abstract construct that may not directly correspond to specific cognitive processes, it is supported by a robust theoretical background and empirical evidence; conversely, composite scores lack a theoretical foundation, limiting their interpretability and comparisons across studies. Future research on the determinants of cognitive impairment should include the g-factor in larger samples and causal analyses, and investigate subgroups with distinct cognitive trajectories to address the heterogeneity among SZ comprehensively.
Conclusion
The g-factors derived from diverse cognitive test batteries proved to be valid global cognitive scores across the schizophrenia spectrum, effectively distinguishing between different disease stages on average. The g-factors were used to investigate underlying mechanisms of cognitive impairment within the schizophrenia spectrum. Educational attainment and genetics related to cognitive performance were associated with better cognition, whereas general psychopathology of schizophrenia, neurodevelopmental load, antipsychotic medication, and genetic risk for schizophrenia were linked to poorer cognition. The progressive nature of cognitive deficits across the schizophrenia spectrum, coupled with the positive correlation with educational attainment, provide compelling arguments in favor of early cognitive training interventions to mitigate cognitive decline associated with schizophrenia, and thus prevent the alteration of functional outcomes.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0033291724002538
Acknowledgments
We would like to sincerely thank the participants who volunteered for the studies, and clinicians from the GHU Paris Psychiatry and Neuroscience involved in the data collection for the different studies (PrEPP: ‘Programme d’Évaluation et de Prévention des Psychoses’, PRIMEPI: ‘Étude des facteurs prédictifs d’évolution lors d'un PREMier ÉPIsode psychotique’, ICAAR: ‘Influence du Cannabis sur l’émergence de symptômes psychopathologiques des Adolescents et jeunes Adultes présentant un état mental à Risque’, AUSZ: ‘Continuum AUtisme et SchiZophrénie: recherche de marqueurs phénotypiques cliniques, cognitifs, biologiques et en imagerie’, START: ‘Étude sur les marqueurs de Stress et efficacité des Thérapies comportementales et cognitives chez les sujets jeunes présentant un état mental À Risque’, PsyDEV: ‘Étude familiale et génétique des aspects DÉVeloppementaux des maladies PSYchiatriques’): Emilie Bonnard, Olivier Canceil, Mylène Charre, Claire Daban-Huard, Edouard Duchesnay, Ariel Frajerman, Mathilde Kazes, Oussama Kebir, Antoine Lambert, Valeria Lucarini, Albane Mansoux, Marion Plaze, Fabrice Rivollier, and Guillaume Tanguy.
Author contributions
L. S. and M.-O. K. supervised the work. M.-O. K. was the principal investigator of all the included studies. G. M., J. B.-D., F. M., C. D.-A., M. G., C. J., and N. B. collected the data for the PrEPP, PRIMEPI, ICAAR, AUSZ, START, and PsyDEV studies. E. K., A. I., and D. Y. extracted and tidied the data. D. Y. performed the data analysis, with the help of Q. H. and B. C. regarding the genetic data analyses. D. Y. and L. S. interpreted the data. D. Y. drafted the manuscript. D. Y., L. S., B. C., A. I., Q. H., and M.-O. K. reviewed and edited the manuscript. All authors approved the submitted version.
Funding statement
This work was supported by grants from the French government (recipient: M.-O. K.): AUSZ (ANR-2010-NEUR-002-01b), ‘Investissements d'Avenir’ PsyCARE (ANR-18-RHUS-0014); from the French Health Ministry: START (PHRC AOM 13-681), ICAAR (PHRC AOM 07-118); and from the Fondation de France: AUSZ (Eng 2010-15142).
Competing interests
None.
Ethical standards
The authors assert that all procedures contributing to this work comply with the ethical standards of the relevant national and institutional committees on human experimentation and with the Helsinki Declaration of 1975, as revised in 2008.