INTRODUCTION
Bartonella spp. are vector-borne, haemotropic, Gram-negative, rod-shaped bacteria in the family Bartonellaceae which can cause long-lasting intra-erythrocytic bacteraemia [Reference Chomel1]. Bartonella invade erythrocytes, endothelial cells, CD34+ progenitor cells, and dendritic cells of their hosts and subsequently induce persistent infection [Reference Chomel1, Reference Dehio2]. Bartonella bacteria have been isolated or detected increasingly for the last two decades [Reference Boulouis3, Reference Kaiser4] in a wide range of mammals, including humans, and in various arthropods [Reference Chomel and Kasten5, Reference Saisongkorh6]. At least 30 Bartonella spp. and subspecies have been identified [Reference Biswas and Rolain7] which are mainly transmitted by haematophagous arthropods such as fleas, ticks, lice and possibly mites and flies [Reference Boulouis3]. Domestic animals and wildlife represent major reservoirs of Bartonella spp., especially in relation to human infection [Reference Billeter8]. The distribution of Bartonella spp. varies between countries and their domestic and wildlife reservoir species [Reference Boulouis3]. Bartonella infections have been reported more frequently in humans [Reference Znazen9] and domestic animals in warm and humid climates [Reference Chomel10, Reference Jameson11] and tropical countries [Reference Saisongkorh6, Reference Chomel12–Reference Suksawat15].
Domestic dogs can be infected with several Bartonella spp. or subspecies, including B. vinsonii subsp. berkhoffii, B. henselae, B. vinsonii subsp. arupensis, B. clarridgeiae, B. washoensis, B. elizabethae, B. quintana, B. koehlerae, B. bovis, B. rochalimae, and several new candidate species, such as B. volans-like and a strain similar to B. bovis called HMD [Reference Bai14, Reference Diniz16, Reference Pérez17]. In infected dogs, the clinical signs are similar to those seen in human patients [Reference Biswas and Rolain7, Reference Breitschwerdt18]. Therefore dogs represent excellent epidemiological sentinels for human exposure yet their role as reservoirs requires further investigation [Reference Henn19].
Several studies have shown a high prevalence of antibodies in stray dogs but very low rates in pet dogs in various parts of the world, including the USA [Reference Henn20–Reference Pappalardo22], Europe [Reference Honadel21, Reference Solano-Gallego23, Reference Zobba24] and North Africa [Reference Henn25]. Seropositivity was shown to be associated with outdoor activities, heavy tick or flea infestation, and rural environment [Reference Pappalardo22].
The aims of this study were to determine the seroprevalence of antibodies against B. clarridgeiae, B. henselae and B. vinsonii subsp. berkhoffii in stray dogs living in four tropical countries and to detect and identify by molecular techniques the presence of Bartonella DNA in the subset of seropositive and seronegative dogs.
MATERIALS AND METHODS
Dog samples
Whole blood (1–2 ml) or serum samples (0·5–1 ml) used for this study were initially collected from a convenient sample of stray dogs from four tropical countries, two in the Americas (Colombia, Brazil), and two from Asia (Sri Lanka, Vietnam), for studies investigating Toxoplasma gondii infection in domestic dogs [Reference Dubey26–Reference Dubey29].
Dogs of different breeds and age groups from Colombia (Bogota), Brazil (São Paulo) and Sri Lanka (Peradeniya) were caught by the respective municipalities and euthanized by intravenous injection of sodium thiopental [Reference Dubey26, Reference Dubey27, Reference Dubey29]. For the dogs from Vietnam (provinces in southern Vietnam, near Ho Chi Min City: Bin Phuoc (10 dogs), Dong Nai (five dogs) and Long An (five dogs), each dog (aged 1–4 years) was from a different home and the houses were at least 2 km apart. These dogs, not considered as pets but raised to be sold for human consumption, were purchased from individual homes and euthanized by a veterinarian with an overdose of sodium thiopental (30–40 mg/kg i.v.) [Reference Dubey28].
Serology
Antibodies against B. vinsonii subsp. berkhoffii, B. clarridgeiae, and B. henselae in stray domestic dog samples were detected using an indirect immunofluorescent antibody assay (IFA). These three antigens were selected, as B. henselae and B. vinsonii subsp. berkhoffii have been frequently detected in dogs as well as B. clarridgeiae, which is also a good substitute for detection of B. rochalimae [Reference Namekata30, Reference Schaefer31] The IFA procedure was similar to a procedure previously described [Reference Henn19], with the following modifications. A 90% confluent tissue culture flask (containing MDCK cells) was inoculated with a 4-day-old culture of B. vinsonii subsp. berkhoffii (ATCC 51672) resuspended in 0·5 ml saline. Similarly, flasks containing Vero tissue cultures were inoculated with either B. clarridgeiae ATCC 51734 or a mixture of B. henselae ATCC 49882 and B. henselae U4 (University of California, Davis, strain). Serum samples added to the test wells were screened at 1:64 dilution in PBS with 5% milk. Slides were incubated at 37 °C for 30 min, followed by three washes in PBS. Fluorescein-conjugated goat anti-dog immunoglobulin G (IgG; ICN Biomedicals Inc., USA) was diluted in PBS (1:1400 for B. vinsonii subsp. berkhoffii, 1:3600 for B. clarridgeiae, and 1:2800 for B. henselae) with 5% milk containing 0·001% Evans Blue, and 20 μl of the dilution was applied to each well. The slides were incubated at 37 °C for 30 min and again washed in PBS three times. The intensity of bacillus-specific fluorescence was scored subjectively from 1 to 4. Samples with a fluorescence score of ⩾2 at a dilution of 1:64 were reported as positive and final titration was performed (last dilution with a score ⩾2). The same two readers performed a double-blind reading of each slide. Negative and positive control samples were included on each slide.
Bartonella PCR procedures
To allow for specific identification of Bartonella isolates, DNA was extracted from the blood of seropositive dogs (dogs for which only serum was available were excluded) and a sampling of seronegative dogs (three dogs from Sri Lanka, three dogs from Brazil, four dogs from Colombia) and analysed by PCR, gel electrophoresis, and DNA sequencing of the 16S–23S intergenic spacer (ITS) region [Reference Rolain32], the citrate synthase gltA [Reference Norman33] gene and the rpoB gene [Reference Renesto34]. The DNA was extracted as described previously [Reference Chang35] using a Qiagen DNeasy blood and tissue kit (Qiagen Sciences, USA). The primers used for the ITS region were 325s 5′-CTTCAGATGATGATCCCAAGCCTTYTGGCG-3′ and 1100as 5′-GAACCGACGACCCCCT GCTTGCAAAGCA-3′ [Reference Diniz36]. The run conditions were 95 °C for 2 min and then 54 cycles of 94 °C for 15 s, 66 °C for 15 s, and 72 °C for 18 s, followed by a final step at 72 °C for 1 min. The primers used for the gltA gene were Bhcs.781p 5′-GGGGACCAGCTCATGGTGG-3′ and Bhcs.1137n 5′-AATGCAAAAAGAACAGTAAACA-3′ [Reference Norman33]. Run conditions were 95 °C for 10 min and then 40 cycles of 94 °C for 30 s, 57 °C for 1 min, and 72 °C for 2 min, followed by a final step at 72 °C for 5 min. The primers used for the rpoB gene were primers 1615s 5′- ATYACYCATAARCGYCGTCTTTCTGCTCTTGG-3′ [Reference Diniz36] and 2300R 5′- GGATCTAAATCTTCYGTYGCACGRATACG-3′ [Reference Renesto34]. Following a first denaturation step of 94 °C for 10 min, a three-step cycle of 94 °C for 30 s, 53 °C for 30 s, and 72 °C for 1 min was repeated 45 times. The PCR programme was ended by a single 5-min extension step at 72 °C [Reference Renesto34].
An approximately 400-bp fragment of the gltA gene, 670-bp fragment of the ITS region and 700-bp fragment of the rpoB gene were amplified and then verified by gel electrophoresis. Amplified PCR products were stained with GelRedTM (Phenix Research Products, USA) and visualized by ultraviolet light after electrophoresis on 2% agarose gels (SeaKem LE agarose; Cambrex Bio Science Rockland Inc., USA).
Bartonella DNA sequencing and alignment
PCR products from the gltA and rpoB genes and ITS region were used for DNA sequencing. Products were purified with the QIAquick PCR purification kit (Qiagen Sciences, USA), and the sequencing of both DNA strands was done using a fluorescence-based automated sequencing system (Davis Sequencing, USA).
A consensus sequence for each amplification product was obtained after raw sequences were imported into MEGA 5 [Reference Tamura37]. Blastn (http://ncbi.nih.gov/BLAST/) was utilized to compare sequences with entries in GenBank. MEGA 5 was then used to align sequence variants of each gene with one another and with relevant sequences available through GenBank.
Statistical tests
Exact χ2 contingency table analysis was performed to evaluate any differences in seroprevalence by geographical region and antigen response, using StatXact 8 (Cytel Corporation, USA). A P value <0·05 was considered statistically significant. Epi Info version 3.5.1 was used to calculate P values, χ2, and Fisher's exact test.
RESULTS
A total of 455 urban stray dogs or rural owned dogs from four tropical countries on two continents (South America, Asia) were enrolled in the study, including 118 (25·9%) dogs from Brazil (São Paulo), 258 (56·7%) from Colombia (Bogota), 59 (13·0%) from Sri Lanka (Peradeniya) and 20 (4·4%) from Vietnam. Overall, 376 (82·6%) dogs were from South America and 79 (17·4%) from Asia.
Serology
Bartonella spp. antibodies at a titre of at least 1:64 were detected by IFA in 38 (8·3%) of these 455 dogs, including 26 (10·1%) dogs from Colombia, nine (7·6%) dogs from Brazil, and three (5·1%) dogs from Sri Lanka (Table 1). None of the 20 dogs from Vietnam had detectable Bartonella antibodies. Overall, 30 (6·6%) dogs were seropositive for B. henselae, 25 (5·5%) for B. clarridgeiae, and 21 (4·6%) for B. vinsonii subsp. berkhoffii (Table 1). In Colombia, the antibody titres ranged from 64 to 4096 with a mode of 256 for the three antigens. In Brazil, the titres ranged from 64 to 512, with dogs seropositive for B. clarridgeiae having the highest titres (256–512). Dogs from Sri Lanka had titres ranging from 64 to 1024.
Number of dogs seropositive (+) and by respective final titres for Bartonella henselae, B. clarridgeiae and B. vinsonii subsp. berkhoffii by country of origin
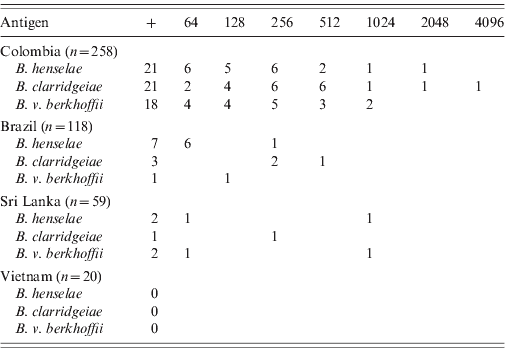
Many dogs were seropositive to more than one antigen. Overall, 53·8% (14/26) of the seropositive dogs from Colombia and 25% (1/4) of the seropositive dogs from Sri Lanka were positive for the three antigens. Of the seropositive dogs, a smaller number of dogs were seropositive for two antigens, including four dogs from Colombia and one dog from Brazil which were seropositive for B. henselae and B. clarridgeiae, and one dog each from Brazil and Colombia which were seropositive for B. henselae and B. vinsonii subsp. berkhoffii. Finally, one dog from Colombia was seropositive for B. clarridgeiae and B. vinsonii subsp. berkhoffii.
Statistical analysis
A significant association between seropositivity for all three Bartonella spp. and country of origin was found [χ2 = 8·16, 2 degreses of freedom (d.f.), P = 0·017]. Similarly, there was a significant association between seropositivity for B. clarridgeiae (χ2 = 9·50, 2 d.f., P = 0·009), B. vinsonii subsp. berkhoffii (χ2 = 9·41, 2 d.f., P = 0·009) and B. henselae (χ2 = 6·11, 2 d.f., P = 0·047) and country of origin. Dogs from Colombia were more likely to be seropositive for B. vinsonii subsp. berkhoffii (Fisher's exact test: P = 0·005) and for B. clarridgeiae (Fisher's exact test: P = 0·014) than dogs from Brazil.
Molecular testing
Whole blood for DNA extraction was available for 26 (63%) of the 41 seropositive dogs, including the four seropositive dogs from Sri Lanka and 22/28 seropositive dogs from Colombia. DNA was also extracted from the whole blood of 10 seronegative dogs (Colombia: four samples, Brazil: three samples, Sri Lanka: three samples) and were all negative by PCR. Of these 26 DNA extracts, four had suspect bands on PCR gel electrophoresis. After purification and sequencing in both directions, the Bartonella strains could be identified as B. rochalimae for a dog (dog 1) from Bogota, Colombia and B. vinsonii subsp. berkhoffii for another dog (dog 2) from Bogota (Figs 1–3). The two other samples were from dogs from Sri Lanka and both were identified as identical to strain HMD, previously described in dogs from southern Italy and Greece [Reference Diniz16]. For one of the two Sri Lanka dogs, a possible mixed Bartonella infection was detected, but the second sequence could not be interpreted.
Dendogram of Bartonella strains identified by partial sequencing of the gltA gene. The tree shown is based on the neighbour-joining method. Bootstrap values are based on 1000 replicates. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) is shown next to the branches. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. There were a total of 272 positions in the final dataset. Evolutionary analyses were conducted in MEGA 5.

Dendogram of Bartonella strains identified by partial sequencing of the 16S–23S genic interspacer. The tree shown is based on the neighbour-joining method. Bootstrap values are based on 1000 replicates. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) is shown next to the branches. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. All positions containing gaps and missing data were eliminated. There were a total of 160 positions in the final dataset. Evolutionary analyses were conducted in MEGA 5.

Dendogram of Bartonella strains identified by partial sequencing of the rpoB gene. The tree shown is based on the neighbour-joining method. Bootstrap values are based on 1000 replicates. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) is shown next to the branches. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. All positions containing gaps and missing data were eliminated. There were a total of 570 positions in the final dataset. Evolutionary analyses were conducted in MEGA 5.

The B. rochalimae strain (dog 1) was confirmed by PCR and sequencing on all three genes or interspacer segment (Figs 1–3). The HMD strain from the dog with mixed infection (dog 4) was confirmed by both PCR and sequencing of the ITS region and the rpoB gene (Figs 2 and 3). The gltA sequence could not be interpreted because of the mixed infection. The B. vinsonii subsp. berkhoffii (dog 2) and the other HMD strain (dog 3) were both confirmed only on the sequence of the gltA gene (Fig. 1).
DISCUSSION
We report the investigation of Bartonella seroprevalence in stray and domestic dogs from four different countries within the tropics, including two countries (Colombia, Sri Lanka) for which no information on Bartonella infection in dogs has been previously reported. Furthermore, we were able to identify from extracted DNA belonging to a subset of seropositive dogs the Bartonella spp. involved in these infections. Rural dogs from Vietnam were surprisingly all seronegative, as Bartonella infection is common in dogs from neighbouring countries, such as Thailand, where 38% of 49 dogs presented at the veterinary teaching hospital of Kasetsart, Bangkok were seropositive for B. vinsonii subsp. berkhoffii [Reference Suksawat15] and 31·3% (60/192) of dogs were bacteraemic from Bangkok and Khon Kaen [Reference Bai14]. However, the limited sample size for each province (5–10 dogs) may explain these negative results. Of note, approximately 7% of the tested dogs from Sri Lanka were Bartonella seropositive, including two dogs which were PCR positive. Sequencing indicated that these strains were identical to a strain previously identified in shelter dogs from southern Italy and sick dogs from northern Greece [Reference Diniz16]. This is the first report of presence of this new Bartonella sp., similar to B. bovis, from dogs from Asia.
Prevalence of Bartonella antibodies was low in stray dogs in São Paulo, Brazil. One dog was seropositive for B. vinsonii subsp. berkhoffii (0·8%) and seven (5·9%) were seropositive for B. henselae, which are values close to those previously reported for 197 sick dogs seen at the São Paulo State University Veterinary Teaching Hospital (2% for B. vinsonii subsp. Berkhoffii, 1·5% for B. henselae) [Reference Diniz36]. Unfortunately, none of the nine seropositive dogs could be tested by PCR, due to a lack of whole blood.
By contrast, the highest seroprevalence for the three antigens was identified in stray dogs from Bogota, Colombia, including a much higher prevalence for B. clarridgeiae antibodies than for the dogs from Brazil or Sri Lanka. Of the two Colombian dogs that were PCR positive, one was infected with B. vinsonii subsp. berkhoffii and the other with B. rochalimae, which are Bartonella spp. previously isolated or detected in dogs or their fleas in the Americas, including California [Reference Henn19, Reference Henn38], Chile [Reference Pérez-Martínez39], and Peru [Reference Diniz40], as well as from Europe [Reference Diniz16]. Several dogs were detected to be seropositive for B. clarridgeiae, which is a species closely related to B. rochalimae [Reference Henn38]. Using B. clarridgeiae as a substitute for the detection of B. rochalimae has been reported previously [Reference Namekata30, Reference Schaefer31]; therefore, it is likely that most of these dogs could have been infected with B. rochalimae rather than B. clarridgeiae, as shown by molecular analysis.
In conclusion, we report the first detection of Bartonella infection in stray dogs from Colombia, South America and Sri Lanka in the Indian subcontinent as well as the first detection of HMD strain from Asia. In addition, we confirmed the low prevalence of Bartonella infection in stray dogs from São Paulo, Brazil and the lack of infection in rural dogs from Vietnam.
DECLARATION OF INTEREST
None.





