INTRODUCTION
Unsafe food is related to several kinds of diseases, ranging from diarrhoeal syndromes to various forms of cancer [Reference Greger1, 2]. In 2005, it was estimated that foodborne or waterborne diarrhoeal diseases were responsible for 2·2 million deaths per year worldwide, 1·9 million of which were children [2].
Salmonella spp. is one of the most common and widely distributed foodborne pathogens in the European Union (EU), with 108 614 laboratory-confirmed cases reported in 2009. Although its relative importance has been decreasing since 2006, S. Enteritidis is still the most frequently reported serovar (52·3% of cases), followed by S. Typhimurium (23·3%), and a wide range of other serovars of public health significance [3].
Identifying the main sources of an illness is a crucial step for the prioritization of control measures [Reference Kuchenmüller4]. This process is called source attribution, and it can be achieved by a variety of methods, one of which is the microbial subtyping approach [Reference Pires5, 6]. The principle of this approach is to compare the occurrence of subtypes in animals or food sources with the same subtypes in humans, provided that subtypes are heterogeneously distributed among the sources. Human infections caused by source-specific subtypes are attributed to the corresponding sources. Infections caused by subtypes found in several reservoirs are distributed relatively to the prevalence of the specific types.
The approach requires an integrated foodborne disease surveillance programme that collects isolates from the major food-animal reservoirs, sporadic human cases, outbreaks and travel-related cases [Reference Hald7]. For that reason, no EU-wide source attribution study has yet been conducted, as a unified European animal-and-human health database does not exist. However, since 2003, efforts have been made in the EU to standardize the reporting of pathogens and diseases in humans and animals, including the conduction of studies to estimate the Member State (MS)-level baseline prevalence of Salmonella in animals of the food chain [8–11]. Another initiative was the harmonization of the monitoring of Salmonella in laying hens [12], broilers [13] and turkeys [14], the last two implemented after the activities described in this paper were conducted. Data from harmonized animal programmes, other production animals and humans are summarized yearly in the European Union Summary Report on Trends and Zoonoses (EUSR) [3].
Based on a review of microbial subtyping-based source attribution studies [Reference Hald7, Reference Pires and Hald15–Reference Pires17], the ‘perfect’ dataset for a EU-wide model would include, for each MS: (1) the number of reported salmonellosis cases in humans, originating from a nationally representative surveillance system in which cases are all laboratory-confirmed and subtyped to an appropriate discriminatory level; (2) information on whether the person had been travelling abroad 1 week prior to symptoms onset; (3) number of outbreak cases and identified outbreak sources; (4) for all major animal sources of human salmonellosis in Europe, the Salmonella prevalence using the same subtyping methods used for humans; and (5) the amount of an animal product originating from one country which is consumed in another country. Serotyping, combined with phage-type data and further differentiation based on antimicrobial resistance profiling, is currently considered the ideal level of subtyping for those models, as it better discriminates common subtypes (e.g. S. Enteritidis and S. Typhimurium) among similar sources, compared to using only serovars [Reference de Knegt18].
This paper describes the data obtained from sources available in 2010 to be used in an EU-wide source attribution microbial subtyping model and the data management steps taken to produce a sufficiently detailed and homogenous dataset containing Salmonella serovar information from humans and animal-food reservoirs. Limitations of the data available are presented, along with the solutions applied to solve them.
METHODS
Data sources
The European Surveillance System (TESSy)
This is a system for collection, validation, analysis and dissemination of data from the EU and European Economic Area (EEA), administered by the European Centre for Disease Prevention and Control (ECDC) which has been functioning since 2008 [19]. Countries report their data on communicable diseases to the system, which also records information on outbreaks and the possibility of infection during international travel. Reporting of specific serovars is mandatory, but countries may report isolates as ‘unknown’, and further subtyping is only done on a voluntary basis. TESSy replaced the data collection systems for the Data Surveillance Network, which collected national data individually [19], therefore data from 2006 and 2007 on Salmonella exist in the system, but not in a completely standardized manner.
The European Union Summary Report on Trends and Sources of Zoonoses, Zoonotic Agents and Foodborne Outbreaks (EUSR)
This report is prepared by the European Food Safety Authority (EFSA) and ECDC. Data on zoonoses and zoonotic agents in animals, foodstuffs and animal feed are reported annually by MS to EFSA and summarized in the EUSR. Serovar reporting follows the same requirements as for humans.
Baseline studies on the prevalence of Salmonella in animal populations in the European Union (BS)
To provide the scientific basis for setting prevalence targets for reduction of Salmonella in commercial and breeding farms, EU-wide studies on the baseline prevalence of Salmonella were conducted on laying hen flocks (2004–2005) [8], broiler flocks (2005–2006) [11], slaughter pigs (2006–2007) [9], fattening and breeding turkeys (2006–2007) [10], broiler carcasses (2008) [20] and breeder pigs (2008) [21]. The studies took place in a 4-year period, and varied in participation due to new EU members in 2004 and 2007, and to the occasional participation of EEA countries. However, the studies still constitute the most uniformly collected and analysed data on Salmonella at the EU-level, allowing valid comparisons between MS.
The Statistical Office of the European Union (EUROSTAT) [22]
EUROSTAT was established in 1953 to provide statistics at the European level to allow comparisons between countries and regions. It collects data on the value and quantity of food and slaughter animals traded among EU MS or with third-party countries. Although EU legislation ensures that the statistics provided to EUROSTAT are based on legal texts and harmonized definitions and procedures [22], information is provided directly by MS, and so data availability and quality are subject to variations in national focus and cultural differences.
Data collected from the sources described were stored and analysed in SAS Enterprise Guide, SAS/STAT® User's Guide, version 8 (SAS Institute Inc., USA).
Reported cases of human salmonellosis
Number and serovar distribution of sporadic human cases reported to TESSy from 2007 to 2009 were provided by ECDC through EFSA. Outbreak-related cases were provided by EFSA as reported by MSs. The total number of reported cases included sporadic, travel and outbreak-related infections. MS with a level of serovar detail insufficient for source attribution were requested to provide additional data, if available. Such national datasets were provided by Poland and Portugal. The MS providing data on sporadic and outbreak cases are summarized in Table 1.
Availability of data from the different datasets by country
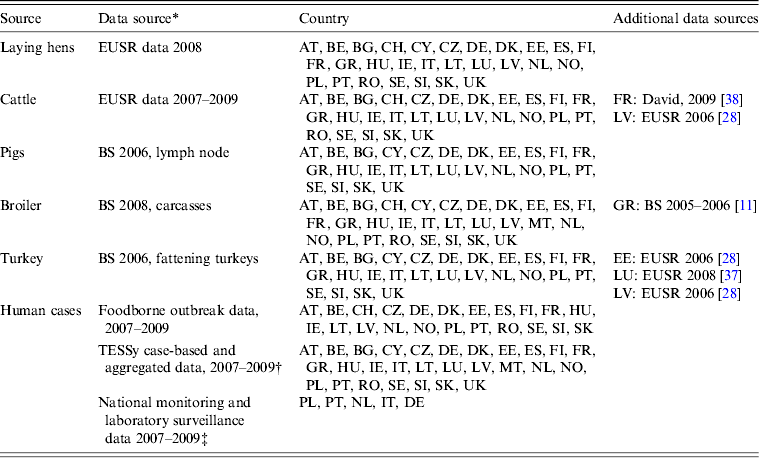
Austria (AT); Belgium (BE); Bulgaria (BG); Cyprus (CY); Czech Republic (CZ); Denmark (DK); Estonia (EE); Finland (FI); France (FR); Germany (DE); Greece (GR); Hungary (HU); Ireland (IE); Italy (IT); Latvia (LV); Lithuania (LT); Luxembourg (LU); Malta (MT); Norway (NO); Poland (PL); Portugal (PT); Romania (RO); Slovakia (SK); Slovenia (SI); Spain (ES); Sweden (SE); Switzerland (CH); The Netherlands (NL); United Kingdom (UK)
* If data were missing from a specific source in a country, used surrogate data sources are indicated.
† Bulgaria reported human cases, but no serovar information was available.
‡ Obtained through direct contact with Member States.
Challenge 1: Underreporting
One issue arising from the use of surveillance data is the underreporting of cases. It is generally understood that the real number of illnesses occurring in the population is larger than the number that are reported to the surveillance system [Reference Wheeler23]. This is explained by the percentage of: (1) people who seek medical care when sick; (2) people who provide clinical specimens when requested; (3) specimens which are tested; (4) sensitivity of the laboratory tests and (5) positive results that get reported [Reference Wheeler23]. Therefore, the true burden of human salmonellosis may be considerably larger than the officially reported incidence. The level of underreporting is expected to vary between countries, depending on differences in organization and effectiveness of local surveillance systems [19, Reference Havelaar24].
Proposed solution
Havelaar et al. used data from a Swedish travel database and the Salmonella incidence from a Dutch population-based study to estimate a set of multipliers for correction of underreporting in 31 European countries [19, Reference Havelaar24]. These underreporting factors (UF) were based on the proportion of cases of salmonellosis that were reported in Sweden upon returning from the Netherlands, and represent an estimation of the number of cases that should have been reported for each case that entered the system. The use of these multipliers is expected to have an impact on the most important sources estimated at EU-level. As the adjustment for underreporting is only done after the attribution process, the corrected numbers are not shown here, but can be found in de Knegt (2013) [Reference de Knegt18].
Challenge 2: Incomplete travel-related information
Travel information, derived from data reported as ‘probable country of infection’, was recorded as ‘travel-related’, ‘domestic’ or ‘unknown’. The proportion of travellers and the amount of information provided is expected to vary among MS, depending on local habits and surveillance priorities.
Proposed solution
The Hald model and its adaptations [Reference Hald7, Reference Pires and Hald15] use the observed proportion of travel cases that were properly discriminated to redistribute cases with no information to the ‘travel-related’ and ‘domestic’ categories, and the same approach could potentially be used in the EU model. In case there is not enough information available for redistribution, cases which did not specifically report a travel history should be considered domestic.
Challenge 3: Incomplete or missing serovar identification
Cases in which serovar identification is missing or incomplete can be summarized as: (a) classification up to genus or species level, e.g. Salmonella spp. or Salmonella enterica; (b) classification up to subspecies level, e.g. Salmonella enterica enterica or Salmonella enterica subspecies I; (c) classification using serogroups based on the O-antigen, e.g. B, C1–C2, O:4 or O:33; (d) main serovars properly specified and the rest aggregated as ‘others’; (e) serovar field left blank or completed as ‘unknown’.
Proposed solutions
Isolates not classified up to serovar level should be reassigned to specific serovars according to proportions observed in previous studies, in the same dataset or in other references, depending on the data availability in each case.
Isolates identified up to genus or species level, left blank or completed as ‘unknown’ should be reassigned to all serovars observed in the country (e.g. if S. Enteritidis accounts for 60% of all serotyped isolates in a country, and 10 isolates in the same country are not properly identified, six of them must be reassigned to S. Enteritidis). Isolates identified up to subspecies level should likewise be reassigned to all serovars in the country, but with proportions calculated using only isolates of S. enterica enterica as the total.
Isolates classified as serogroups should be distributed among serovars pertaining to those groups, in accordance with the Kauffman–White–Le Minor scheme [25] (e.g. if S. Typhimurium accounts for 40% of all isolates in a country, but for 80% of units from serovars belonging to group B, and 10 isolates are only identified as ‘group B’, eight of those should be reassigned to S. Typhimurium).
Isolates classified as ‘others’ are assumed to belong to serovars not described in the current dataset, but nonetheless present in the country. In this case, the reference used for reassignment of proportions is the World Health Organization Global Foodborne Infections Network (GFN) Country Databank (CDB) [26], which contains the 15 most commonly identified Salmonella serovars among human and non-human sources in 84 countries (e.g. a country reports 30 isolates to TESSy: 10 S. Enteritidis, 10 S. Typhimurium, and 10 ‘others’. The CDB shows 80% S. Enteritidis, 10% S. Typhimurium, 7% S. Infantis and 3% S. Hadar for this country, so, according to this reference, S. Infantis and S. Hadar correspond to 70% and 30% of the non-described serovars. The 10 isolates should be redistributed as seven S. Infantis and three S. Hadar, assuming that S. Typhimurium and S. Enteritidis are not included in the ‘others’ group).
Challenge 4: Underreporting and incomplete identification of serovars in outbreak data
For outbreaks of foodborne salmonellosis, the same datasets used for EUSRs 2007–2009 [3, 27, 28] were provided by EFSA. Not all countries report outbreak cases, and not all reported cases have complete serovar information.
Proposed solutions
The same underreporting multipliers used for sporadic cases cannot be applied to outbreaks, as cases belonging to an outbreak are likely to have a different probability of being reported, and some serovars may generate outbreaks more frequently than others [Reference Pires and Hald15]. Based on that, countries which report sporadic cases but no outbreak cases are assumed to have no foodborne Salmonella outbreaks in the period. Outbreak-related cases for which a serovar is not fully identified should be reassigned using the proportions observed in the same outbreak dataset.
Salmonella in livestock and food
Challenge 5: Heterogenous availability of data from MS and animal sources
The EU BS prevalence of Salmonella in the sources was the preferred data source. Due to the admission of new EU members and the voluntary character of BS participation, data were not available for all MS and sources. However, these datasets were considered the most representative of the given reservoirs, since no harmonized EU monitoring in pigs and turkeys was currently in place. In addition, the broiler carcass study was considered to provide more recent data than BS on broiler flocks, and with a better detailing of the serovar distribution compared to the existing EU monitoring data. The laying-hen BS was conducted between 2004 and 2005 [8], and it is expected that the implementation of harmonized monitoring [12] has resulted in significant changes in Salmonella serovar prevalences in this reservoir in many MS. No data from BS or EU-harmonized monitoring exist for cattle.
Proposed solutions
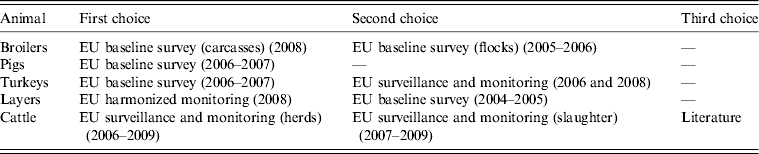
In order to use the most recent data possible, data that are missing from BS should be supplied with surveillance and monitoring data found in the EUSR. When not enough surveillance or monitoring data at the herd/flock level are available for a source or MS, slaughter samples should be surveyed and their quality as substitutes assessed. The order of priority for selecting which animal-food data to include in the model is shown in Table 2.
Order of priority for selection of animal-food data to include in the model
Challenge 6: Incomplete or missing serovar identification
The expected situations in which serovar identification is missing or incomplete are the same as for human data. For BS data, no reference for reassigning serogroups or incomplete serovar identification was available.
Proposed solutions
The criteria for reassigning non-identified or partially identified serovars should be the same as for the human data. Proportions found in the laying-hen BS [8] should be used for re-allocation of laying-hen monitoring data. In datasets where there are no records identified as ‘others’, units should be redistributed according to the proportions found in properly identified serovars in the same dataset.
Food production and trade data
Food production data were derived by EFSA from the EUROSTAT databases on production and slaughtered animals for food consumption [22]. Consumption calculations were based on production and country-to-country trade data. This was done so the attribution model can account for the amount of food present in a given country that originated from other countries, and use the country- and food-specific serovar prevalences for the attribution [Reference de Knegt18]. The domestically produced amount available for consumption of a food source in a MS was estimated as domestic production minus export, whereas the amount of imported food available for consumption in MS A originating from MS B was estimated as import minus re-export (when relevant). For this study, extra EU food trade was not considered [Reference de Knegt18].
Challenge 7: Missing data
Information on poultry for meat production was not available for Belgium in 2007 and 2008. Egg production data was lacking for several countries, and data for most food sources and most years were missing for Cyprus. Data on the export of the food sources to other MS included in this study were available for all considered countries, with the exception of the amount of eggs exported from Cyprus.
Proposed solutions
Missing data on annual quantities of poultry meat products sold per MS, with differentiation between boilers, turkeys and other poultry species are available in the 2009 Annual Report of the Association of Poultry Processors and Poultry Trade in the EU Countries (AVEC) [29]. For all sources, countries with information missing for a year should have the missing value estimated based on the percentage of increase or decrease between available years; when data from only one year are available, that value should be used as surrogate for the missing years.
Challenge 8: Negative estimated amounts available for consumption
Due to differences in numbers reported in the production, import and export datasets, the calculations of the amount of a food source available for consumption in a country in some cases results in negative numbers, meaning that the volume exported is larger than the domestic production.
Proposed solution
In order to ensure that MS will still have nationally produced food available in their own country, re-exporting of imported products should be considered possible.
Challenge 9: Validation of the estimation of consumption data based on trade data
The underlying assumptions for this estimation were that EUROSTAT data were complete and consistent, and that all food available for consumption is actually consumed, in a way that these data reflect the real flow of foodstuffs and consequent exposure in the countries. According to an assessment performed by EFSA [30], the information recorded in those datasets does not fully support these assumptions. The assessment showed the existence and non-reporting of triangular trade, misclassification of food products and problems in the conversion of currency/weight units. Moreover, we expect that in several situations, data for missing years needs to be estimated or supplied with surrogate data (e.g. AVEC data), resulting in a highly manipulated dataset that may not represent reality.
Proposed solution
The data management can be validated by comparing the resulting consumption dataset with consumption data available from the WHO Global Environment Monitoring System Food Consumption Cluster Diets [31]. As the WHO data only offer the broad category ‘poultry’, broilers and turkeys should be summed. Relative proportions of consumption of poultry, pork and eggs must be calculated, so a proportional similarity index (PSI/Czekanowsky index) can be used to compare those proportions between the two groups in each country. The PSI is an estimate of the area of intersection between two frequency distributions [Reference Rosef32], calculated as
It is traditionally used for calculating niche overlap and resource availability in population ecology [Reference Feinsinger, Spears and Poole33] or proportions of identified bacterial strains in epidemiology [Reference Mullner34, Reference Mullner35], but it was considered that each of the relative proportions in the three sources corresponds to the area under a probability curve, and so the same measure could be applied. A PSI of 1 means a complete overlap, or 100% similarity. An ‘overall PSI’ for the whole dataset was calculated by averaging the country PSI values.
RESULTS
Human data
The percentage of records with incomplete identification and that had to be reassigned varied from zero in Portugal to 84% in Romania (Table 3). The most common reason for reassignment was records reported in aggregated form, i.e. with several serovars categorized as ‘others’, and the next reason was isolates reported as ‘unknown’, followed by isolates only classified as serogroup (Table 3). Besides the predicted identification problems, a specific issue regarding S. Typhimurium was found: one of the defining characteristics of S. Typhimurium is the two phases of the H-antigens: ‘i’ and ‘1,2’, which is why the antigenic formula for this serovar is written as ‘1,4,[Reference Pires5],12:i:1,2’ [27]. However, S. Typhimurium-like variants with only the first phase of the H-antigen (e.g. 1,4,[Reference Pires5],12:i:- or 1,4,[Reference Pires5],12:-:-) have been reported, and are referred to as ‘S. Typhimurium-like strains’ or ‘monophasic S. Typhimurium’ [36]. For our purposes, those isolates were reassigned to S. Typhimurium, which is supported by an EFSA Biohazard Panel assessment [36].
Number and percentage of reassigned records in humans

For explanation of country abbreviations see Table 1.
* Salmonella spp, Salmonella enterica, Salmonella not typed, Salmonella untyped.
† Salmonella enterica enterica, subspecies I.
‡ B, C, D, E, D1, C1, C2–C3, D1, E1.
§ ‘Others’, ‘other serovars’.
¶ ‘Unknown’.
Bulgaria, Cyprus, Greece, Italy, Luxembourg, Malta and the UK did not report outbreak cases. Nearly 47% of outbreak cases reported by France had to be reassigned, as the isolates were reported as ‘Salmonella spp’. For Latvia, the proportion was 39% (Table 4). Switzerland reported outbreaks, but no sporadic cases (Table 1).
Number and percentage of reassigned records in foodborne Salmonella outbreaks
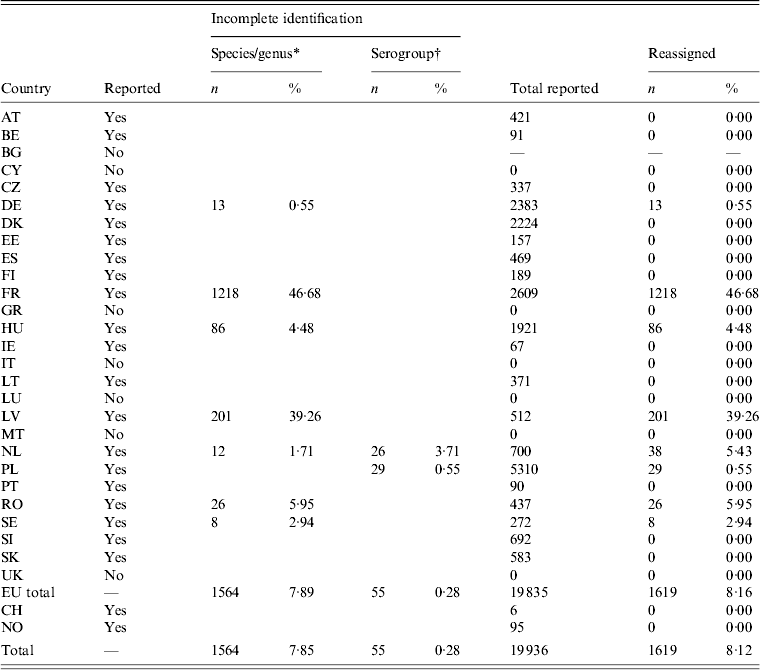
For explanation of country abbreviations see Table 1.
* Salmonella enterica enterica, subspecies I.
† B, C, D, E, D1, C1, C2–C3, D1, E1.
Travel information (Table 5) was reported as ‘unknown’ for 100% of isolates in France, Romania and Slovenia. Full travel information was provided by Austria, Belgium, the Czech Republic, Estonia, Spain, Hungary, The Netherlands and Slovakia. The remaining MS had variable proportions of cases reported as ‘travel-related’, ‘domestic’ or ‘unknown’. Therefore, the proposed ‘informed redistribution’ was not possible, as a large number of countries did not report any travel cases. As a consequence, all records with missing or unknown travel information from countries with serovar details of sporadic cases were considered domestically acquired in the reporting country.
Number of cases reported in the original datasets as travel-related, domestic or unknown and the total used in the model, assuming that any case not specifically mentioned as travel-related was domestic

For explanation of country abbreviations see Table 1.
Table 6 shows the relative occurrence of the 11 most important zoonotic serovars in the last 5 years in sporadic and outbreak cases [3, 28]. S. Enteritidis and S. Typhimurium were the most frequently observed in sporadic cases, along with S. Infantis, S. Newport, S. Kentucky, S. Virchow, S. Derby and S. Agona. The most commonly observed serovars in outbreaks were also S. Enteritidis and S. Typhimurium. As expected, outbreaks may present serovars not normally found in a specific country. That is particularly true in countries with a small number of sporadic cases and good Salmonella control in domestic products, e.g. Finland or Sweden.
Total isolates and relative proportions of the most frequent serovars in total reported (R) and outbreak (O) cases in humans in the EU and Norway, 2007–2009
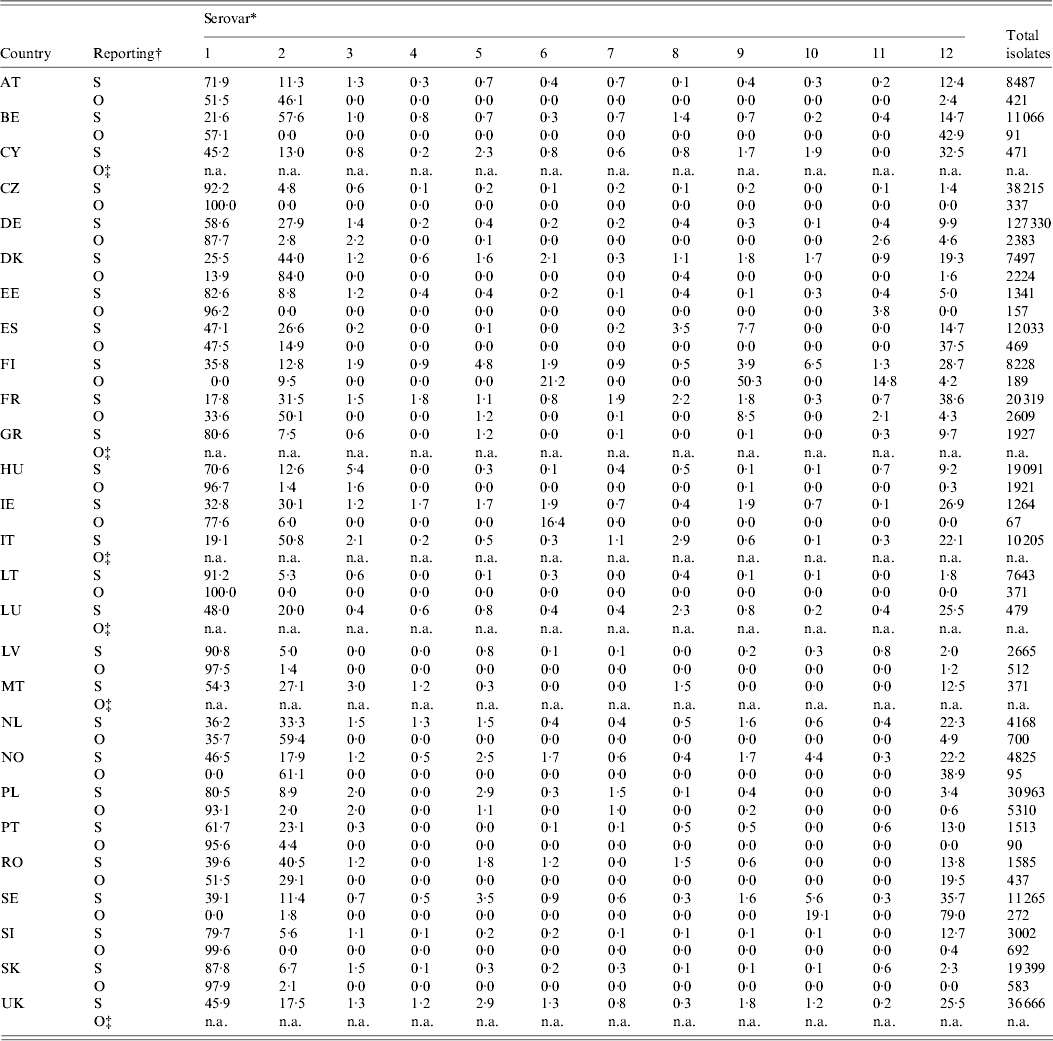
For explanation of country abbreviations see Table 1.
* 1, S. Enteritidis; 2, S. Typhimurium; 3, S. Infantis; 4, S. Kentucky; 5, S. Virchow; 6, S. Agona; 7, S. Hadar; 8, S. Derby; 9, S. Newport; 10, S. Stanley; 11, S. Bovismorbificans; 12, others; n.a., not available.
† S = sporadic cases; O = outbreak cases.
‡ The country did not report any outbreak cases.
Animal-food data
Data was available from 28 countries (Table 1). Laying-hen data from the EUSR 2008 [28] were preferred over BS data, as this was the first year of EU-harmonized reporting for this reservoir, Greece did not participate in the broiler carcasses study [20], being supplied with data from the broiler flocks BS [11]. Malta and Romania did not participate in the slaughter pigs BS [9], and no surrogate data was available for those countries. For turkeys, BS data from fattening flocks were chosen [10], with the exception of Estonia, Latvia, Luxembourg and Romania, which were not part of the study. Data for Estonia, Latvia and Luxembourg were retrieved from EUSR 2006 and 2008 [28, 37]. No surrogate data was available for Romania. Non-harmonized surveillance data on cattle, including carcass samples at slaughter, were retrieved from EUSR 2007, 2008 and 2009 [3, 27, 28], with 2009 data being preferred to the other years. Cattle data for France was retrieved from a PhD thesis [Reference David38]. For this reservoir, no data from Cyprus or Malta were identified, and for some countries only one year of data was available. Belgium and the UK only reported positive samples for cattle, resulting in 100% positivity in those countries. Small samples were observed for broilers in Luxembourg, laying hens in Lithuania and Luxembourg and turkeys in Estonia, Luxembourg and Latvia. The amount and percentage of reassigned records in the total positives are given in Table 7.
Number and percentage of records reassigned to serovars in animal reservoirs
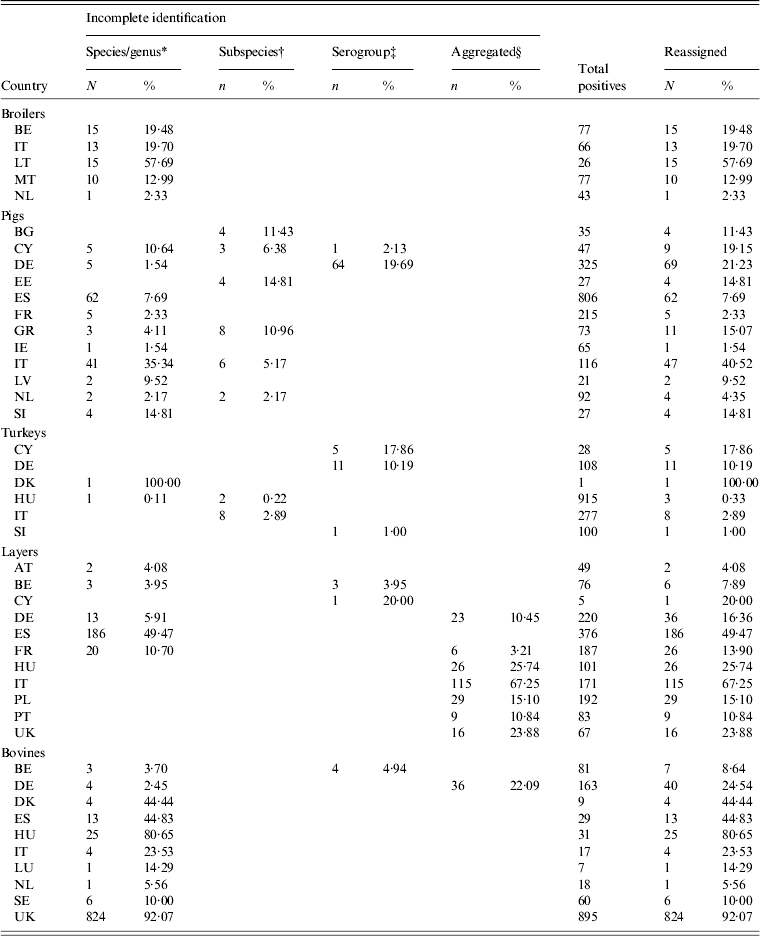
For explanation of country abbreviations see Table 1.
* Salmonella spp., Salmonella enterica, Salmonella not typed, Salmonella untyped.
† Salmonella enterica enterica, subspecies I.
‡ B, C, D, E, D1, C1, C2–C3, D1, E1.
§ ‘Others’, ‘other serovars’.
Serovar predominance varied between countries in all animal sources. Considering the relative occurrence of serovars and number of countries in which they predominated, S. Infantis and S. Enteritidis were the main serovars observed in broilers, S. Typhimurium and S. Derby in pigs, S. Typhimurium, S. Bredeney and S. Hadar in turkeys and S. Enteritidis and S. Infantis in layers. S. Dublin and S. Typhimurium were the main serovars in cattle, but the data was considered too heterogeneous and frail to be representative. The top-ten serovars for broilers, pigs, turkeys and layers are given in Table 8.
Relative proportions of the top-10 Salmonella serovars found in broiler carcasses, pig lymph nodes, turkey flocks and laying hen flocks in the chosen datasets
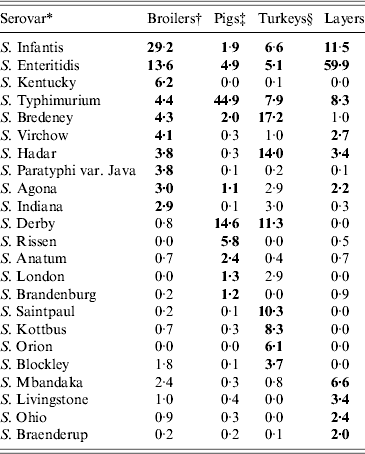
* Combined list of the top ten serovars in all BS. Bold values show the top ten serovars for each animal reservoir.
† [20]. Participating countries: AT, BE, BG, CH, CY, CZ, DE, DK, EE, ES, FI, FR, GR, HU, IE, IT, LT, LU, LV, MT, NL, NO, PL, PT, RO, SE, SI, SK, UK.
‡ [9]. Participating countries: AT, BE, BG, CY, CZ, DE, DK, EE, ES, FI, FR, GR, HU, IE, IT, LT, LU, LV, NL, NO, PL, PT, SE, SI, SK, UK.
§ [10]. Participating countries: AT, BE, BG, CY, CZ, DE, DK, EE, ES, FI, FR, GR, HU, IE, IT, LT, LU, LV, NL, NO, PL, PT, SE, SI, SK, UK.
¶ [27] Participating countries: AT, BE, BG, CH, CY, CZ, DE, DK, EE, ES, FI, FR, GR, HU, IE, IT, LT, LU, LV, NL, NO, PL, PT, RO, SE, SI, SK, UK.
Trade and consumption data
Availability of data on the annual quantities of poultry, pork, bovine meat and eggs produced varied by year and MS. All MS reported imports from other MS for all food products in the study period. The resulting surrogate consumption dataset was considered valid, as shown by the results of the data validation (Table 9). The individual PSI values were > 0·8 in most countries, indicating more than 80% similarity between the estimated data and the reference values. The one exception was Cyprus, with only 42% similarity, which is expected to have an impact on the attribution estimates for this country. The overall PSI was 0·91, indicating that the dataset as a whole can be used without considerable bias.
Comparison of the relative proportion of consumption of pork, poultry meat and table eggs in the WHO GEMS/Food data and the surrogate values calculated from EUROSTAT data
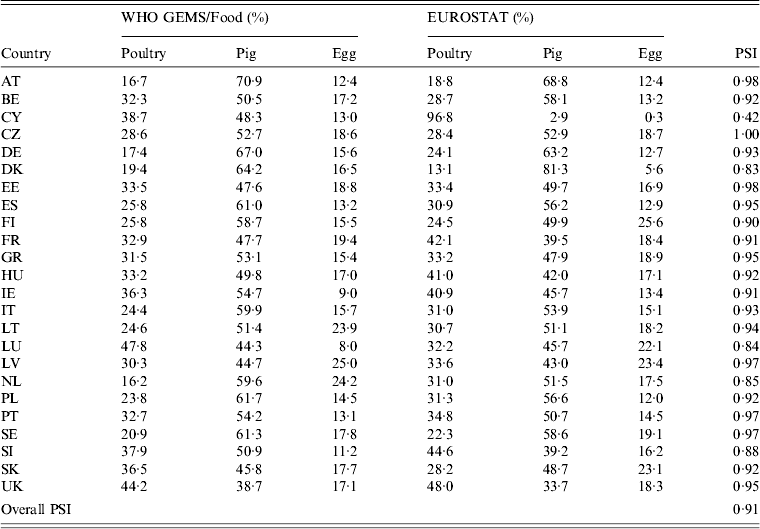
For explanation of country abbreviations see Table 1.
PSI, Proportional similarity index.
Final dataset for the source attribution model
Based on data availability and quality, 24 countries were included in the model: Austria, Belgium, Cyprus, Czech Republic, Denmark, Estonia, Finland, France, Germany, Greece, Hungary, Ireland, Italy, Latvia, Lithuania, Luxembourg, Poland, Portugal, Slovakia, Slovenia, Spain, Sweden, The Netherlands and UK. Countries initially analysed and excluded were Bulgaria, which provided 100% of human cases without serovar details; Romania, which only participated in one BS, did not have enough surrogate data to be retrieved from the EUSR, and reported 84% of human cases without serovar information; Norway and Switzerland, which do not report to EUROSTAT, the latter also does not report to TESSy.
Based on the availability of EU-wide homogeneous data or good-quality surrogates, food-animal sources included were broilers, pigs, turkeys and laying hens (as the animal reservoirs for chicken meat, pork, turkey meat and eggs). Due to better completeness and availability, the resulting trade data from 2009 was used as consumption data for those sources. Cattle data were in general poor, and for some MS consisted of clinical isolates only. The use of herd information from 2007–2008 or slaughterhouse carcass samples was not sufficient to obtain a representative dataset for this source.
Twenty-two serovars were selected to be specifically addressed, based on their presence and importance in humans and chosen animal reservoirs: S. Agona, S. Anatum, S. Bovismorbificans, S. Braenderup, S. Brandenburg, S. Bredeney, S. Derby, S. Enteritidis, S. Hadar, S. Heidelberg, S. Infantis, S. Kentucky, S. Kottbus, S. Livingstone, S. London, S. Mbandaka, S. Montevideo, S. Newport, S. Rissen, S. Saintpaul, S. Typhimurium and S. Virchow. Albeit important in humans in most of the 24 countries, S. Stanley was not isolated from any of the selected reservoirs, while S. Dublin and S. Ohio became irrelevant after cattle were excluded. The building of the final Salmonella dataset (trade data not included) is shown in Figure 1.
The final Salmonella dataset (not including trade data). * For abbreviations see Table 1. FBO, Foodborne outbreaks.

DISCUSSION
This study presented the officially reported data available for use in an EU Salmonella source attribution model based on microbial subtyping [Reference de Knegt18]. Challenges associated with the use of these data were presented, and solutions were proposed. The data available were retrieved from multiple sources and had varied levels of quality and completeness. Although TESSy and EFSA collect and organize the data at the EU-level in a harmonized way, the primary information is collected in different countries, which have their own individual approaches and methods for data collection and management. Non-EU countries such as Switzerland and Norway also contribute to data heterogeneity, as they participate in some studies and report partial data, for example, to the EUROSTAT production database, but not to the trade database. This variability made several data management steps necessary.
The variability observed in the number of reported human Salmonella infections reflects true differences in the burden of salmonellosis across countries, but also differences in foodborne disease surveillance systems in MS and different levels of underreporting. The loss of data at various points along the surveillance chain from patient to official statistics is recognized in all countries [Reference Wheeler23], and multiplying factors [Reference Havelaar24] were used to compensate this loss. Limitations and assumptions connected to the use of those factors should be discussed, as they were calculated based on Swedish cases [Reference Havelaar24], coming from a system where underreporting is also expected to occur. By using the infection rates in returning travellers to calculate incidences for the local population in the countries visited, it was assumed that the eating habits and other exposures of Swedish travellers are the same as for the locals, also disregarding local levels of acquired immunity and differences in circulating strains. Considerations must also be made regarding the use of a Dutch study as a reference to estimate the underreporting in other countries, and a full discussion of the limitations can be found in Havelaar et al. [Reference Havelaar24]. Despite these limitations, the UF-adjusted numbers are still a better reflection of reality than the raw reported data, and this adjustment is expected to affect the relative importance attributed to the different sources by the model at the EU level, as it affects the contribution of each country to the total burden of salmonellosis in the EU.
Information about travelling within or outside Europe was not available in a representative manner, and it was not possible to estimate additional ‘extra’ intra-EU travellers because the proportion of reported cases with missing travel information was 100% in some countries. Thus, it had to be assumed that all reported cases with missing travel information were domestically acquired, which is expected to overestimate domestic cases, since travel information as reported to TESSy is often incomplete and may not reflect the true relationship between travel and domestic cases [3].
Concerning animal data, the panel of participating MS varied with each BS, as countries have the right to refuse participation in EU-wide baseline studies. The admittance of new MS to the EU also generates different lists of reporting countries for each animal source, as data were collected in different years. The resulting data gaps were, when possible, filled with information from EUSR. There are currently no EU-wide studies on the baseline prevalence of Salmonella in cattle and no harmonized monitoring in place, which is the main reason why this reservoir was excluded. However, this is not expected to have a large impact on the model, as national attribution studies have suggested that the contribution from the cattle reservoir in general is low compared to the other sources [Reference Pires and Hald15].
Serovar information was also heterogeneous both in humans and animals. Countries were approached directly for more complete datasets, and records were reassigned based on the serovar distributions observed in available data or external reference datasets (e.g. WHO GFN/CDB). One limitation of this approach is that any emergence of new serovars or other profile fluctuations may be lost, particularly when a whole year of typing is missing and the records are reassigned based on data from previous years. This is also a problem for outbreak cases, as two MS had nearly 50% of reassigned records, while others had the reference proportions calculated from a small number of reported cases. Therefore, serovar reassignment is considered a large source of uncertainty around the final data, and it is proposed that future models use a stochastic approach for reassigning, allowing this uncertainty to be expressed and quantified.
The consumption dataset presented a special challenge, as it had to be based on estimates from surrogate trade data, and an evaluation of the quality of EUROSTAT data revealed major inconsistencies in the intra-EU trade statistics [30]. However, according to comparison with WHO GEMS/Food, this approach produced valid results, as 19/24 countries had a PSI of ⩾0·9 and three had a value of >0·8, suggesting that the consumption profiles composed using EUROSTAT data are highly similar to the GEMS/Food profiles for most countries. An exception was noted for Cyprus, which is likely to be a reflection of the large proportion of extrapolated data, and which may have an effect on the attribution outcomes for that country. Nonetheless, the dataset as a whole showed 91% similarity.
In conclusion, as long as a thorough data evaluation is performed and specific countries and reservoirs with insufficiently representative data are excluded, public surveillance and monitoring data from multiple countries can potentially be used for scientific purposes, particularly for microbial subtyping-based source attribution methods. This could be a first step for the conduction of source attribution studies in countries or regions where no country-wide baseline studies have been conducted, but where programmes for Salmonella monitoring in food or surveillance in humans are currently up and running.
ACKNOWLEDGEMENTS
We acknowledge Timour Koupeev from Vose Risk Consulting for the collaboration on the management of the EUROSTAT data.
The staff of EFSA's Task Force of Zoonoses Data Collection is acknowledged for providing the original datasets necessary to conduct this study. The views or positions expressed in this publication do not necessarily represent in legal terms the official position of the European Food Safety Authority. The European Food Safety Authority assumes no responsibility or liability for any errors or inaccuracies that may appear.
The views and opinions of the authors expressed herein do not necessarily state or reflect those of the ECDC. The accuracy of the authors' statistical analysis and the findings they report are not the responsibility of ECDC. ECDC is not responsible for conclusions or opinions drawn from the data provided. ECDC is not responsible for the correctness of the data and for data management, data merging and data collation after provision of the data. ECDC shall not be held liable for improper or incorrect use of the data.
DECLARATION OF INTEREST
The initial version of the EU model was developed with partial funding from contract CT/EFSA/Zoonoses/2010/02 (contract value 45000 Euros) between EFSA and the DTU National Food Institute, in relation to Question no. EFSA-Q-2010-00685.












