INTRODUCTION
Shiga toxin-producing Escherichia coli (STEC) are an important group of foodborne pathogens associated with a broad spectrum of human diseases ranging from mild diarrhoea to haemorrhagic colitis (HC) and haemolytic uraemic syndrome (HUS) [Reference Griffin and Tauxe1–Reference Tarr3]. STEC are zoonotic pathogens which are asymptomatically carried by ruminants, mainly cattle, which are considered their principal reservoir [Reference Blanco4]. Worldwide, the most important STEC serotype reported is O157:H7, owing to its association with severe disease and many outbreaks. However, some non-O157 strains also pose a substantial concern to public health, as they can cause the same clinical complications as O157 and are increasingly more common [Reference Johnson, Thorpe and Sears5, Reference Tarr and Neill6].
Production of one or more Shiga toxins (Stx1 and/or Stx2) is believed to be the most important contributor to HUS development [Reference Karmali7]. Several subtypes of Stx2 have been identified; in particular, Stx2a and Stx2c have been associated with severe human diseases [Reference Persson8]. Moreover, several subtypes of Stx1 have been described, but they all appeared less important for human disease [Reference Siegler9]. Stx genes are present in the genomes of temperate, lambdoid bacteriophages, which appear to regulate Shiga toxin expression as part of their lytic switch [Reference Bolton and Aird10]. Several other virulence factors are also involved in the pathogenicity of STEC. The locus of enterocyte effacement encodes factors responsible for adherence of the bacterium to the enteric cells, like intimin (Eae) [Reference Yu and Kaper11]. The large plasmid of STEC encodes for additional virulence factors, such as enterohaemolysin (Ehx), which acts as a pore-forming cytolysin; the bifunctional catalase-peroxidase (KatP) [Reference Brunder, Schmidt and Karch12]; and the serine protease (EspP), which cleaves the human coagulation factor V [Reference Brunder13].
The most commonly used molecular biology-based method used in epidemiological research of outbreaks and monitoring of the spread of potential pathogens is pulsed-field gel electrophoresis (PFGE), owing to its high discriminatory power and reproducibility [Reference Karama and Gyles14]. Moreover, this method has been standardized for several pathogens such as E. coli O157 to facilitate the subtyping of the pathogens in various laboratories [Reference Swaminathan15].
During 2000–2007, about 48 cases of STEC infections per year were reported in Belgium. Nationally, all suspected STEC isolates from humans and food samples are collected and further verified by the Belgian national VTEC (Verocytotoxin-producing E. coli) reference laboratory (Professor D. Piérard). Despite the long-running investigation of STEC occurrence and characteristics in Belgium since 1990, a comprehensive long-term study on the genetic diversity of STEC isolates, including non-O157 serogroups and isolates from different sources, had not yet been done. In the present study, we used genomic virulence typing and whole genome genetic variation analysis (PCR and PFGE) to examine the virulence potential and genetic relatedness between STEC isolates of serogroups O157, O26, O103, O111, and O145. Second, the influence of the stx genotype, the serotype, and age on the development of HUS were studied. Third, genetic relatedness was verified with epidemiological data in order to delineate the Belgian situation and to evaluate it on the international scene.
MATERIALS AND METHODS
Bacterial isolates
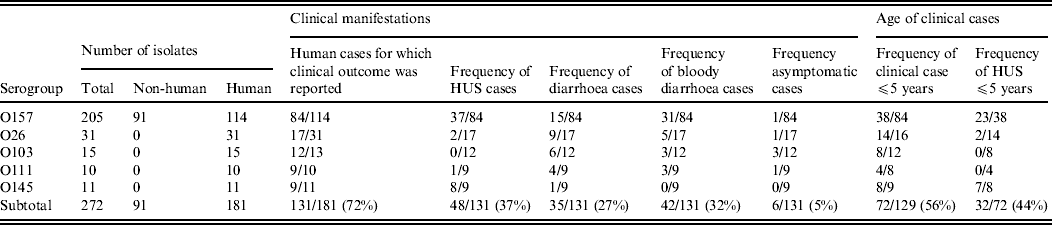
This study included 272 E. coli isolates belonging to serogroups O157 (n = 205), O26 (n = 31), O103 (n = 15), O111 (n = 10), and O145 (n = 11) (Table 1). Isolates were collected by the Belgian national VTEC reference laboratory between 2000 and 2007. The majority (n = 181) of the isolates originated from humans suffering from diarrhoea, bloody diarrhoea, HUS, or asymptomatic infection. Those isolates represented the five serogroups O157, O26, O103, O111, and O145. Clinical manifestation was reported for 131 of the isolates. In addition to the human isolates, 91 isolates belonged to serogroup O157 exclusively, isolates originated from animal sources (two faecal samples from cattle, one faecal sample from a dog, and one dust sample from a cattle barn) or foods (including cattle carcasses (n = 68), beef, minced beef, carpaccio, and raw-milk cheese). Eighty-one of these isolates possessed stx genes. Serogroups were investigated by bacterial agglutination using O antisera for O157, O26, O103, O111, and O145 (Statens Serum Institute, Copenhagen, Denmark).
Origin, clinical manifestation and age associated to the isolates used in this study

HUS, Haemolytic uraemic syndrome.
Detection of stx1, stx2, eae, ehx, espP and katP gene sequences using PCR
The PCR assays for identifying gene sequences were based on literature: for the stx1, stx2, eae, ehx gene sequences, we used the primers and conditions reported by Botteldoorn et al. [Reference Botteldoorn16]. For detection of the katP and espP genes, we used the primers described by Nielsen & Andersen [Reference Nielsen and Andersen17] and the conditions described by Botteldoorn et al. [Reference Botteldoorn16].
Stx2 genotyping
Isolates that gave a positive result for stx2 were tested for the presence of stx2a and stx2c [Reference Wang, Clark and Rodgers18]. Subtypes of stx were denominated according to the subtyping nomenclature established at the 7th International Symposium on Shiga Toxin (Verocytotoxin)-Producing Escherichia coli Infections (Buenos Aires, 10–13 May 2009). Stx2 genes that differed from stx2a and stx2c were considered to be undefined subtypes.
PFGE
PFGE was performed in accordance with the PulseNet-Europe protocol (http://www.pulsenet-europe.org/docs.htm). Genomic DNA was digested by XbaI (Roche Diagnostics, Germany) and analysed in 1% Seakem Gold agarose gels (Lonza, USA) in 0·5 × TBE buffer [45 mm Tris, 45 mm boric acid, 1 mm EDTA (pH 8)] at 14°C using the CHEF Mapper system (Bio-Rad, UK). The runtime was 19 h at 6 V/cm, with initial and final switch times of 2·16 s and 54·17 s, respectively. Gels were stained with ethidium bromide, destained in water, and digitally captured under UV light. Gel images were visually analysed with BioNumerics version 6.5 (Applied Maths, Belgium) using the XbaI-digested DNA from Salmonella enterica Braenderup H9812 as a normalization reference. The similarity between PFGE patterns of the same serogroup was calculated using the Dice coefficient (with an optimization of 1·0% and a position tolerance of 1·0%), and they were grouped together according to their similarities using the unweighted pair-group method with arithmetic mean (UPGMA). Pulsotypes were assigned based on the difference in the presence or absence of at least one band. Pulsogroups were delineated on the basis of 80% similarity according to Dice similarity. Isolates that were not found within a group at 80% similarity, were denominated single isolates. Pulsosubgroups were delineated on the basis of 90% similarity according to Dice similarity.
Statistical analysis
First, univariable logistic regression was performed to determine the association between the presence of a certain stx genotype, a certain genogroup and the age of the patient (⩽5 years vs. >5 years) (risk factors) and the presence of HUS (dependent variable). Next, significant risk factors were tested in a multivariable logistic regression using a backwards stepwise procedure. Statistical analyses were performed using SPSS Statistics v. 20 (SPSS Inc., IBM Corporation, USA). Statistical significance was considered at P < 0·05.
The diversity in isolates of the same serogroup was determined by calculating Simpson's diversity index with 95% confidence intervals as described by Carriço et al. [Reference Carriço19]. Simpson's diversity index accounts for the number and the size of pulsogroups and single isolates for a certain serogroup. A low index indicates that a high number of strains are located within the same group. Agreement between the partition of pulsogroups and pulsosubgroups by PFGE analysis and the virulence profile determined by PCR typing was calculated using the adjusted Wallace index, respectively, with 95% confidence intervals as described previously [Reference Carriço19]. A high adjusted Wallace index is obtained if a virulence property is associated well with a certain pulso(sub)group.
RESULTS
Virulence markers
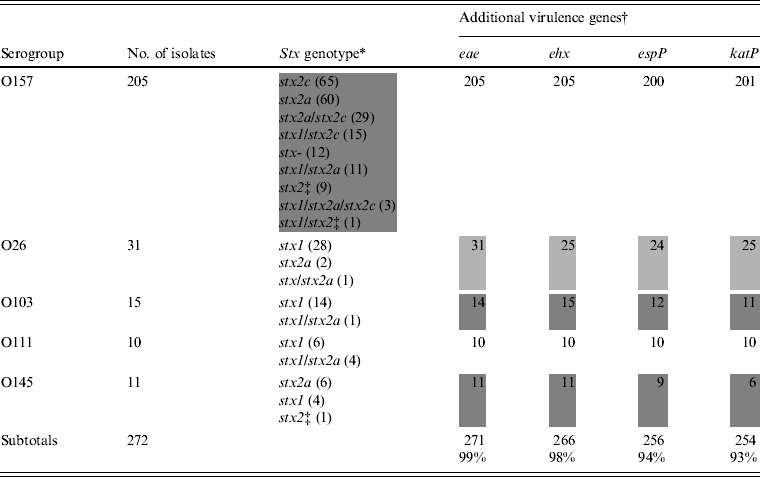
The stx genotype and the presence of additional virulence genes are listed in Table 2. Nine stx genotypes were observed among isolates of O157 (Table 2). Of these, stx2a (60/205 isolates, 29·3%), stx2c (65/205, 31·7%), and stx2a/stx2c (29/205, 14·1%) were the most prominent. Stx genotype stx1 was not observed in the O157 isolates, but the combinations stx1/stx2a, stx1/stx2c, stx1/stx2a/stx2c, and stx1 combined with an undefined subtype of stx2 were present in 12, 15, three, and one isolates, respectively. Nine isolates of O157 harboured a single undefined subtype of stx2 and 12 were stx negative (from cattle carcasses). Isolates of O145 belonged to three stx genotypes, of which stx2a was the most prominent (6/11) (Table 2, shaded area). In the O26, O111 and O103 isolates, the stx genotype stx1 predominated. Eae was found in all isolates except one isolate of O103 (related to a case of diarrhoea). Many combinations of large plasmid-encoded genes (ehx, espP, katP) were observed in isolates of O26, O103 and O145 (Table 2), whereas in all O111 isolates and almost all of O157 these three genes were present.
Virulence properties of STEC O157, O26, O103, O111, and O145 isolates from humans, foods and animals in Belgium between 2000 and 2007

* Subtypes of stx were denominated according to the subtyping nomenclature established at the 7th International Symposium on Shiga Toxin (Verocytotoxin)-Producing Escherichia coli Infections (Buenos Aires, 10–13 May 2009).
† Number of positive isolates.
‡ Undefined subtype of stx2 different from stx2a and stx2c. Heterogeneous results are indicated by grey shading.
PFGE patterns and clonal analysis
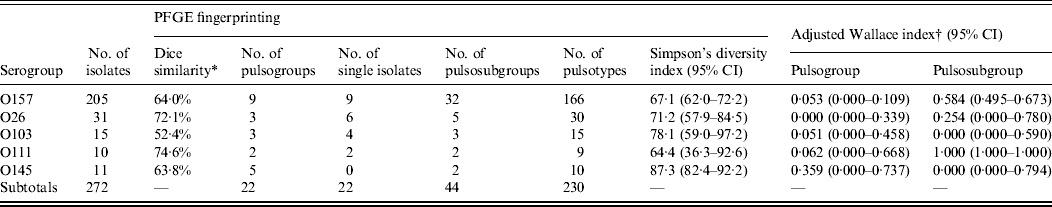
Isolates of serogroup O111 had the highest degree of similarity (74·6%), followed by isolates of serogroups O26 (72·1%), O157 (64·0%), O145 (63·8%), and O103 (52·4%) (Table 3). The diversity of isolates of the same serogroup was determined by Simpson's diversity index. No significant difference between the indices was observed, although O103 and O145 showed the highest diversity index.
Genotypic similarities among STEC O157, O26, O103, O111, and O145 isolates from humans, foods and animals in Belgium between 2000 and 2007

PFGE, Pulsed-field gel electrophoresis; CI, confidence interval.
* Within the analysed isolates of the serogroups.
† Agreement between virulence profile and PFGE typing as described by Carriço et al. [Reference Carriço19].
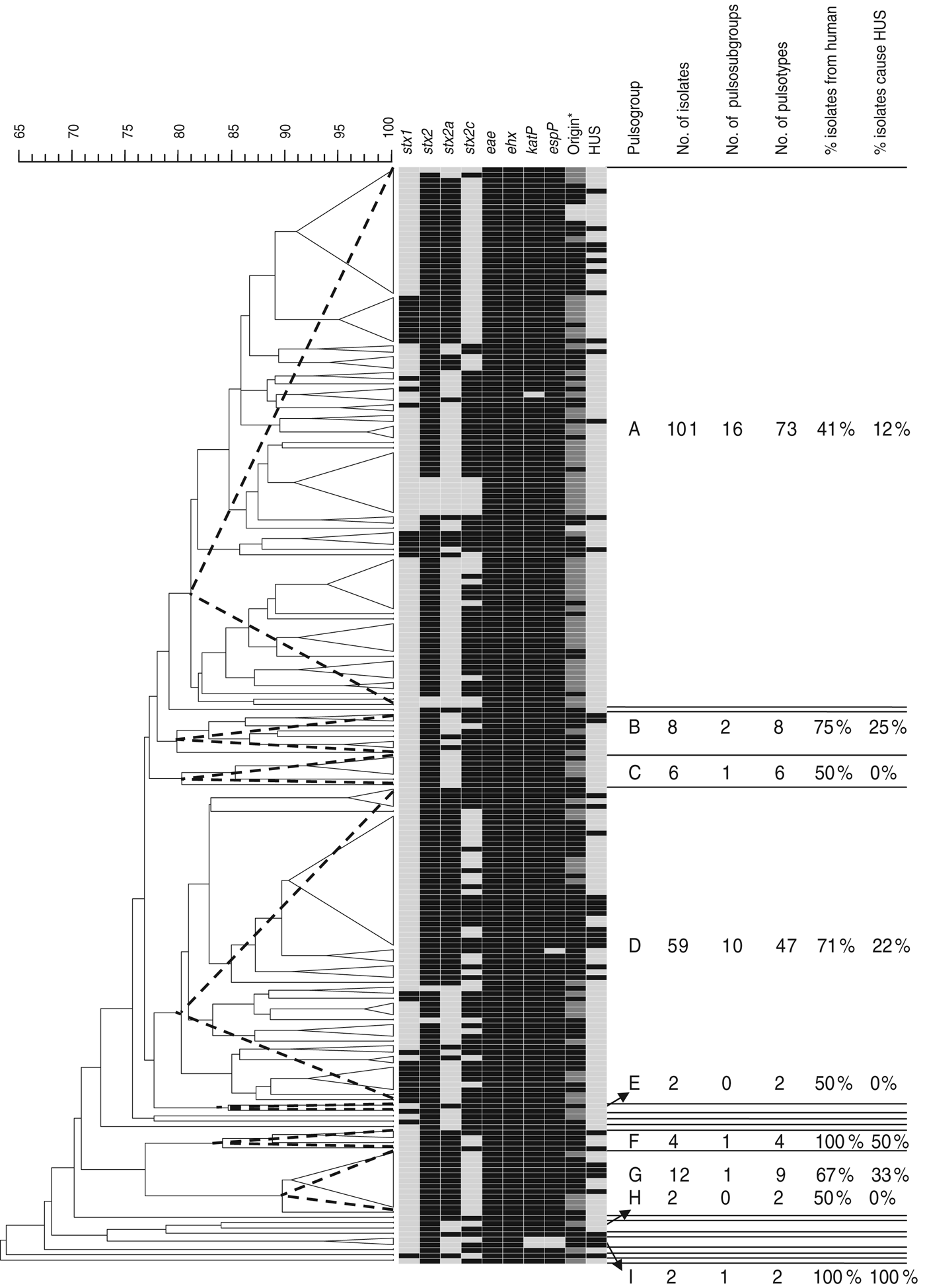
Of the 205 isolates of O157, 160 clustered in two pulsogroups (A, D; Fig. 1). Other pulsogroups contained 2–12 isolates only, and nine single isolates were found. Isolates from food or animal origin did not cluster together, but were distributed in the different pulsosubgroups. Undistinguishable pulsotypes were isolated from human and non-human sources and occasionally they were isolated many years apart. The stx genotypes, and therefore also the virulence profile, were heterogeneously distributed within pulsogroups but clustered together to some extent in pulsosubgroups with some exceptions. This observation is displayed in the Wallace index, which indicated that two strains of the same pulsogroup have only a 5% chance of presenting the same virulence profile, and two strains of the same pulsosubgroup have a 58% chance of presenting the same virulence profile (Table 3). Isolates of the same pulsotype had identical virulence profiles, except for two isolates with an additional stx2c gene compared to the isolate(s) with the same pulsotype: one isolate in a cluster of four pulsotypes from cattle carcasses (pulsogroup A), one isolate in a cluster of five pulsotypes from minced beef and human origin (pulsogroup D), but with no reported clinical manifestations.
Dendrogram, PFGE patterns, and epidemiological data of STEC O157 isolates subjected to PFGE analysis of XbaI-digested genomic DNA and UPGMA similarity analysis using the Dice coefficient and PCR for virulence gene detection. Delineation of pulsogroups (A–I) on the basis of 80% similarity, pulsosubgroups on the basis of 90% similarity and pulsotypes on the basis of one or more bands of difference in the PFGE pattern. Pulsosubgroups are indicated with a dotted-line triangle. Black, positive; light grey, negative. * For origin: black = human; dark grey = food; light grey = animal.

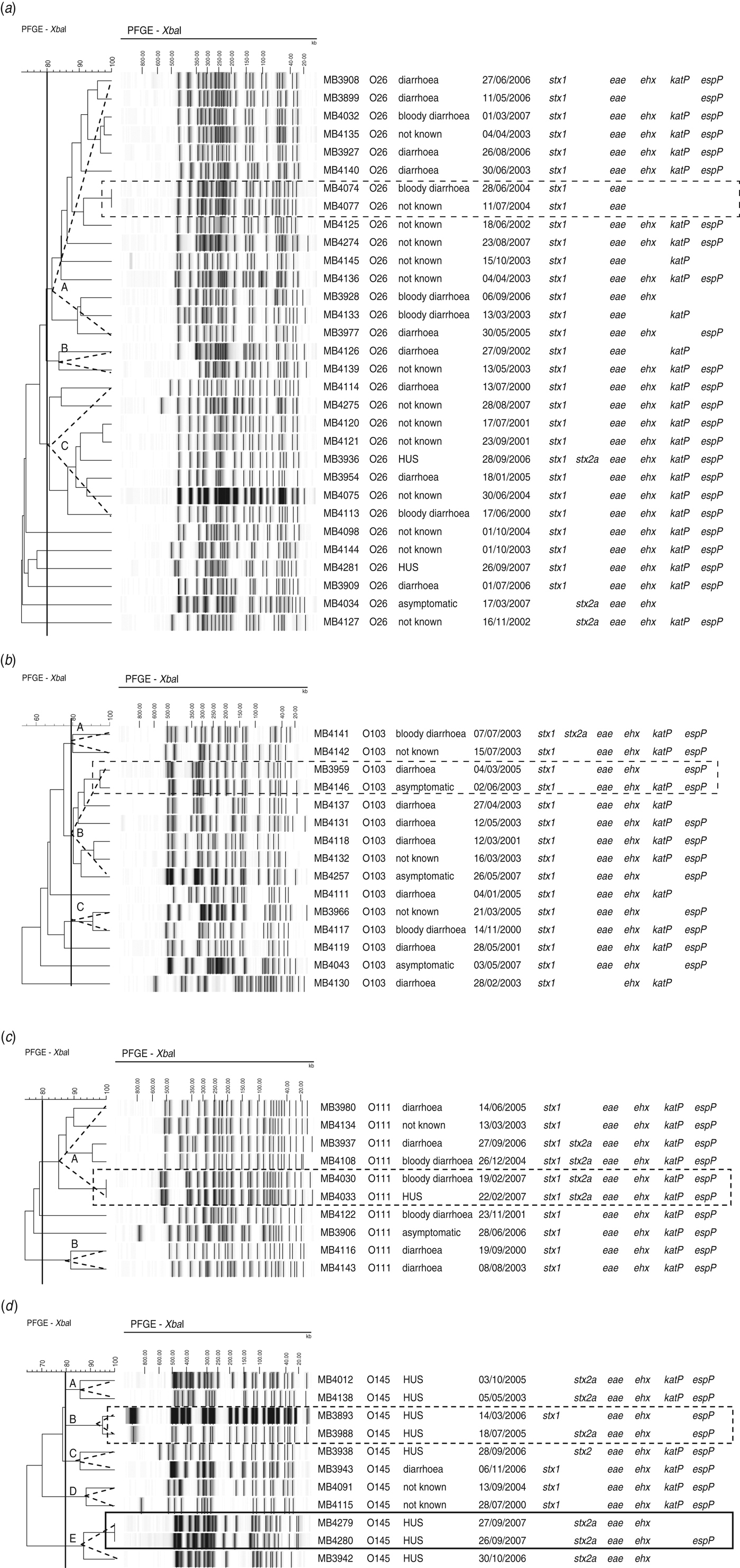
Of the 31 O26 isolates, 25 clustered in three pulsogroups (Fig. 2a, A, B, C), and the other six isolates were single isolates. Virulence profiles were highly heterogeneous within pulsosubgroups (Fig. 2a). Two isolates from humans hospitalized within 13 days of each another were associated to the same pulsotype (pulsogroup A) and had identical virulence profiles (MB4074, MB4077).
Dendrogram, PFGE patterns, epidemiological data, and virulence profile of human (a) STEC O26, (b) O103, (c) O111, and (d) O145 isolates, determined by PFGE analysis of XbaI-digested genomic DNA and UPGMA similarity analysis using the Dice coefficient and PCR for virulence gene detection. Delineation of pulsogroups on the basis of 80% similarity is indicated with a dotted-line triangle. Outbreak isolates are indicated by a solid-line rectangle. Sporadic cases associated with identical pulsotypes or pulsotypes that differ by no more than two bands are indicated by a dotted-line rectangle.

Within two of the three pulsosubgroups of STEC O103, virulence profiles differed in the presence/absence of katP (Fig. 2b). PFGE patterns with only two bands of difference (pulsogroup B) were isolated from sporadic cases that occurred 2 years apart; the virulence profiles differed in the presence/absence of katP.
Of the isolates of serogroup O111, virulence profiles were conserved within the pulsosubgroups (Fig. 2c). This was displayed by a Wallace index of 1 (Table 3). Two sporadic cases that occurred 3 days apart were associated with the same pulsotype (pulsogroup A) with identical virulence profiles.
Within two pulsosubgroups of O145 (in pulsogroups B and E), virulence profiles differed in stx genotype or the absence/presence of espP (Fig. 2d). Two epidemiologically related HUS cases were associated with the same pulsotype (pulsogroup E); the virulence profiles differed in the presence or absence of espP. Two sporadic HUS cases that occurred 6 months apart were associated with PFGE patterns (pulsogroup B) with only one band difference but with a different stx genotype (stx1 or stx2a).
Association between stx genotype, serogroup and age with HUS
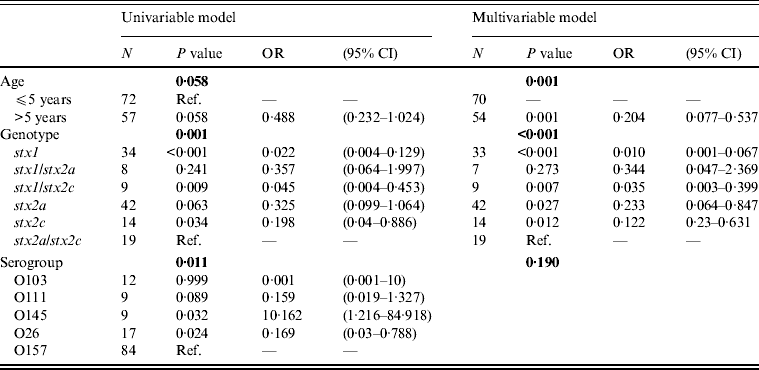
Multivariable logistic regression analysis determined that the odds of having HUS is less in patients with the genotypes stx2a, stx2c, stx1/stx2c, and stx1, compared to patients with the genotype stx2a/stx2c (Table 4). In addition, patients in the >5 years age group have lower odds of developing HUS than patients aged ⩽5 years (Table 4). The development of HUS was not affected by the serogroup, as the effect of the other risk factors, stx genotype and age, predominated (Table 4).
Univariable and multivariable logistic regression model for HUS as outcome variable and age, stx genotype and serogroup as risk factors

HUS, Haemolytic uraemic syndrome; OR, odds ratio; CI, confidence interval.
Standard error of the regression coefficient.
Bold indicates the P value of the risk factor.
O157 cases and outbreaks
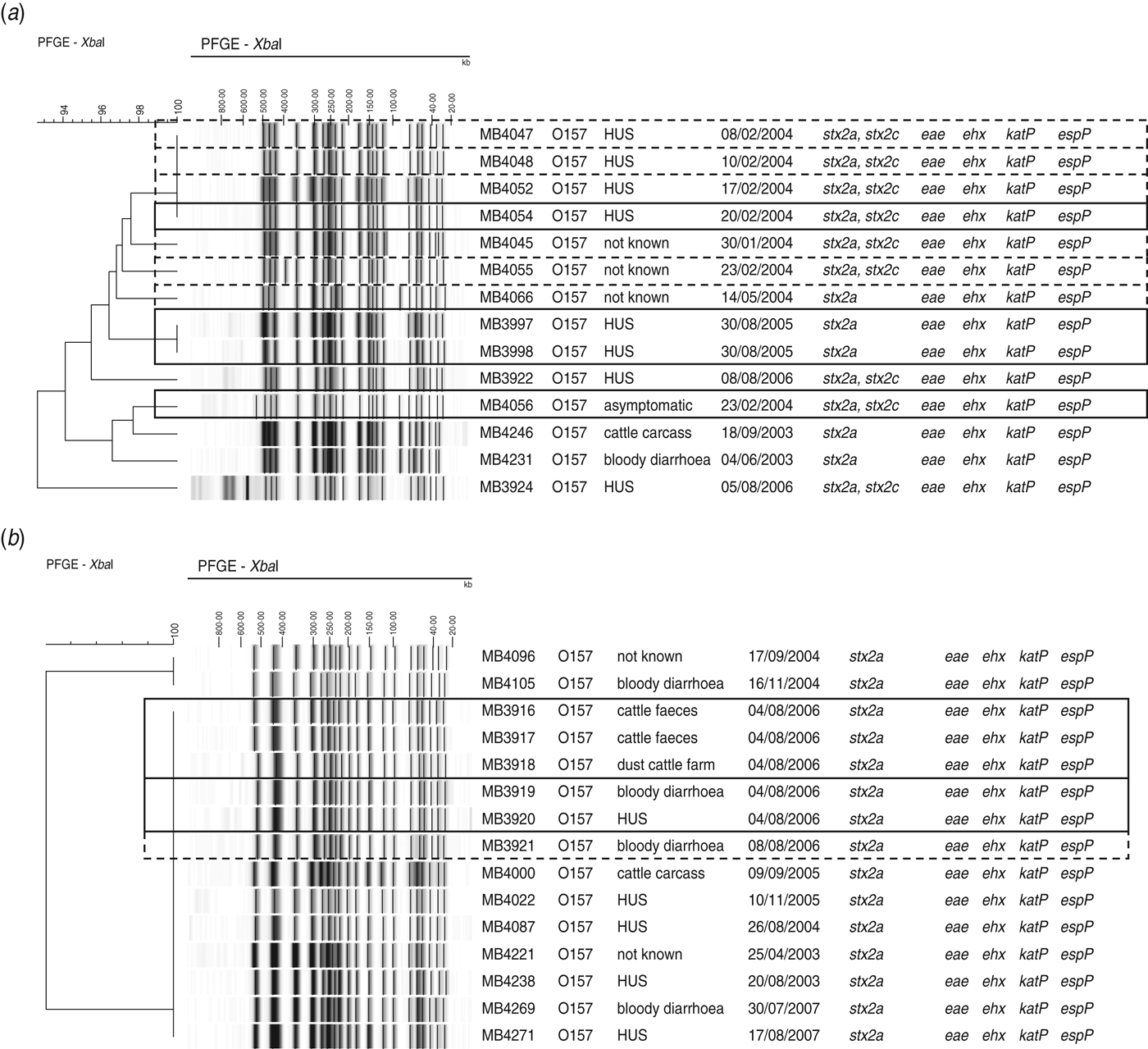
Isolates that could be associated with two outbreaks and sporadic cases were found within the same pulsosubgroup (Fig. 3a). Two of these O157 STEC isolates were associated with an outbreak in a psychiatric institute in Ghent in February 2004. The outbreak involved four HUS cases from which no STEC could be isolated, but two STEC O157 isolates (MB4054, MB4056) could be isolated from contact cases. During the same month as the outbreak, six sporadic cases were reported of which the isolates showed ⩾96% similarity to each other and to the outbreak isolate MB4054. One year later (February 2005), two siblings developed HUS. These isolates (MB3997, MB3998) showed band patterns with 100% PFGE similarity. Isolates of a cattle carcass and sporadic cases that occurred in different years were also found within this pulsosubgroup. Virulence profiles differed only in the presence/absence of stx2c.
Outbreak of STEC O157 in (a) a psychiatric institute in February 2004 and (b) a family outbreak of STEC O157 on a farm in June 2006, found within pulsosubgroups, determined by PFGE analysis of XbaI-digested genomic DNA and UPGMA similarity analysis using the Dice coefficient. Virulence profiles were determined by PCR. Epidemiological data are indicated. Outbreak isolates are indicated by a solid-line rectangle. Sporadic cases associated with identical pulsotypes or pulsotypes that differ by no more than two bands are indicated by a dotted-line rectangle.

Five O157 STEC isolates (MB3916–MB3920) included in this study were associated with a family outbreak in June 2006. The family had spent a weekend at a farm, after which two children contracted bloody diarrhoea and one subsequently developed HUS. STEC O157 could be isolated from the patients’ stools as well as from cattle faeces and dust samples from the stables. The five isolates were of the same pulsotype and were found within a pulsosubgroup with 100% similarity (Fig. 3b), including isolates from cattle carcasses and sporadic cases in different years. Four days after the family outbreak, one sporadic case was reported. Virulence profiles were identical for all isolates in the pulsosubgroup.
DISCUSSION
In Belgium, an average of about 48 cases of STEC infections occur per year. Human STEC isolates collected between 2000 and 2007 were intensively analysed in this study. During the same period, 91 E. coli O157 isolates were recovered from food and animal sources for monitoring and epidemiological studies and included in this study. All isolates were verified by the Belgian national VTEC reference laboratory. In this study, isolates belonging to serogroups O157, O26, O103, O111, and O145 from the current collection were characterized with the objective of determining their virulence potential and genetic relatedness, the association of the stx genotype, age and serotype with HUS, and epidemiological features in Belgium. Most studies include only one or a few serogroups. We have defined several levels of genetic relatedness on the basis of PFGE fingerprinting ranging from pulsogroups (⩾80% similarity) to pulsosubgroups (⩾90% similarity) and pulsotypes (identical fingerprints).
Serogroup O26 was the most common non-O157 serogroup causing human STEC infections in Belgium. This concurs with the incidence of STEC cases in the European Union from 2002 to 2006, which ranks the serogroups in decreasing order as follows: O157, O26, O103, O91, O145, O111 and others [20]. For diagnostic reasons, only STEC isolates of serogroup O157 were recovered from food and animal sources. However, non-O157 serogroups represent a large subset of STEC in cattle [Reference Joris, Pierard and De21] and are also found in food [Reference Madic22]. They were not targeted in this study, therefore the isolates’ genetic relatedness between human and non-human isolates could only be investigated for O157. In addition, the small set of non-human O157 study isolates does not represent well the existing population of O157 isolates in animals and foods. The animal and food isolates did not originate from a substantiated monitoring programme whereas the human isolates did. Due to the difference in completeness between the human and non-human sample set, the diversity within these two sample sets could not be compared.
Isolates of O157 displayed a wide variation of stx genotypes. At the pulsogroup level, isolates of different virulence profiles were heterogeneously distributed. However, at the pulsosubgroup level, concordance was demonstrated using statistical tests, which demonstrated that isolates of the same pulsosubgroup were more likely to have identical virulence profiles. In isolates of O26, O111 and O103, stx genotype stx1 predominated, whereas isolates of O145 displayed a heterogeneous distribution of stx genotypes, with about half of the isolates harbouring genotype stx2a. Similar associations between serogroups and these specific stx genotypes have been described before [Reference Eklund, Scheutz and Siitonen23–Reference Bastos26]. Undefined stx2 subtypes (divergent from stx2a and stx2c) were observed for a number of O103, O145 and O157 isolates. These stx2 genes could either belong to subtypes stx2b, stx2d, stx2e, stx2f or stx2g according to the subtyping nomenclature established at the 7th International Symposium on Shiga Toxin (Verocytotoxin)-Producing Escherichia coli Infections (Buenos Aires, 10–13 May 2009). To specify these stx2 subtypes, specific PCRs [Reference Feng27] or a restriction fragment length polymorphism (RFLP)–PCR assay [Reference Pierard28, Reference Tyler29] can be used.
Isolates of O26, O103 and O145 displayed many different combinations of large plasmid-encoded genes (ehx, espP, katP), whereas these genes were conserved in isolates of O111 and O157. This was in agreement with earlier studies, which reported a great heterogeneity in gene composition of large plasmids observed in non-O157 STEC strains [Reference Brunder13, Reference Zhang30]. The intimin gene (eae) was found in all isolates of this study, with only one exception for one O103 isolate. In human clinical cases, intimin is involved in pathogenesis. In food and animal isolates, however, the presence of eae creates the potential for pathogenicity in humans [Reference Sandhu, Clarke and Gyles31]. Based on the virulence profile, this demonstrated that the O157 isolates originating from food and animal sources are potential human pathogens.
We observed a correlation between the virulence profiles and the clinical manifestations of the human isolates. Isolates with genotypes stx2a, stx2c, stx1/stx2c, and stx1 had lower odds of HUS compared to genotype stx2a/stx2c. This is in agreement with reports of correlation of either stx2a, stx2c, or both with severe human diseases [Reference Persson8, Reference Eklund, Leino and Siitonen32, Reference Friedrich33]. Isolates harbouring stx genotype stx1 were the least likely to cause HUS. This was most prominent in non-O157 isolates which mainly represent stx genotype stx1 (75%), but for which HUS cases were largely associated with stx2 (in 9/11 cases) and stx2a in particular (8/9 cases). The difference in pathogenicity between stx1 and stx2 has been explained by a structural difference and by a difference in biological activity demonstrated in animal models [Reference Paton34]. Isolates harbouring stx1 were homogeneously distributed in human and non-human isolates. Another approach for differentiating the more virulent STEC isolates is single nucleotide polymorphism (SNP) typing [Reference Manning35]. In that study, Manning et al. identified a clade 8 group of STEC O157 strains which was seven times more likely to elicit HUS than the other strains. In our study, however, we were not able to determine the presence of clade 8 isolates in our collection because PFGE cannot predict these hypervirulent variants of STEC O157 and we did not perform SNP typing [Reference Eriksson36].
Despite the correlation between the stx genotype and clinical manifestations, isolates that produced the same clinical manifestation were not seen to be highly genetically related. Furthermore, isolates of the same pulsotype or pulsosubgroup were associated with different clinical manifestations. A possible explanation is that clinical manifestations depend on patient-related factors such as age, gastric acidity, the use of antibiotics, and genetic factors [Reference Karmali, Gannon and Sargeant37].
The serogroups evaluated in this study significantly differed in their association with HUS, but when age and stx genotype were included to the multivariable regression model, the effect of serogroup was ruled out. Patients aged >5 years had lower odds of developing HUS compared to patients aged ⩽5 years. This finding has also been observed in many other studies in the literature [Reference Tarr3].
Epidemiologically related isolates harboured the same pulsotype and virulence profile, except for two isolates of O145 which differed in the presence of espP. Sonntag et al. [Reference Sonntag25] stated that isolates with different virulence profiles cannot be part of a diffuse outbreak. However, our data support the view that genetic mobility may occur during the course of an outbreak, in agreement with Proctor et al. [Reference Proctor38], which may lead to differences in the virulence profile. Epidemiological relationships have been suggested for some sporadic cases based on undistinguishable pulsotypes, identical virulence profiles, a short period between cases, and the restricted area (Belgium). However, indistinguishable PFGE patterns do not equivocally demonstrate an epidemiological connection between cases [Reference Barrett, Gerner-Smidt and Swaminathan39], and although these cases occurred within a short period, infection by means of different routes cannot be excluded [Reference Barrett, Gerner-Smidt and Swaminathan39]. Therefore, epidemiological relationship can only be suggested but not confirmed for sporadic cases. Nevertheless, the same pulsotypes were observed in humans, foods, and animals, which confirms the animal reservoir of STEC and food as a possible vehicle. The epidemiological persistence of isolates was also demonstrated by observing indistinguishable or very similar PFGE patterns during different years. Some virulence profiles were identical, but some showed minor variations due to genetic evolution.
In summary, we have genetically characterized a collection of isolates of STEC O157, O26, O103, O111, and O145 originating from humans, foods and animals in Belgium between 2000 and 2007. This characterization revealed virulence genetic profiles, whole genome genetic variations and relationships between isolates on different levels. Pulsotypes representing pathogenic clones were found in humans, foods and animals over a 7-year period.
ACKNOWLEDGEMENTS
We thank Miriam Levenson for the English-language editing of this manuscript. This research was funded by the Belgian Science Policy grant STECTRACK SD/AF/06A.
DECLARATION OF INTEREST
None.