Introduction
Pigs are an important source of human dietary protein worldwide. Intensive pig farming systems have gained popularity due to their ability to achieve higher productivity and streamlined management. However, intensive farming systems may cause stress in animals (Barnett et al. Reference Barnett, Hemsworth and Cronin2000), and behavioral signs of stress may include increased defecation, aggression, sow infanticide, lethargy, decreased appetite, and postpartum depression and can have adverse effects on productivity of pigs (Jarvis et al. Reference Jarvis, Moinard and Robson2006). For example, when subjected to overcrowding and mixing, pigs experience decreased growth rates of 15.7% and 7.1%, respectively, while the feed-to-meat conversion ratio is estimated to decline by 15% and 10%, respectively (Hyun et al. Reference Hyun, Ellis and Johnson1998). Additionally, the reproductive performance of sows in close proximity to aggressive sows is compromised as a result of stress (Stookey and Gonyou Reference Stookey and Gonyou1994), which has a significant effect on the economic efficiency of pig farming. Aggression is recognized as a paramount concern in modern pig production systems because it poses significant challenges to the health, welfare and economic aspects of the pigs (Kongsted Reference Kongsted2004).
Selection signatures may be effective approaches for elucidating genetic mechanisms of porcine mental health (PMH), even in the absence of detailed phenotypic data (Burgos-Paz et al. Reference Burgos-Paz, Souza and Megens2013). Intensively farmed pig breeds, such as Duroc, Landrace, Yorkshire, Hampshire, and Pietrain pigs, have gone through long-term intensive production management (Petrelli et al. Reference Petrelli, Buglione and Maselli2022), in which breeding pigs prone to mental health problems will be eliminated because of their reduced production performance. Under intense systemic selection pressure, pigs experiencing mental health issues are typically culled due to decreased production performance; meanwhile, genes related to mental health will also experience selection. In contrast, Chinese indigenous pig breeds such as Jinhua pigs and Meishan pigs experience less systemic selection pressure due to their lower level of intensification in farming. Therefore, by comparing the genomes of intensively and nonintensively farmed pig breeds using selection signature detection methods, candidate genes associated with PMH can be detected. Specifically, it was possible to construct intrapopulation selection signatures within intensively farmed pig breeds and interpopulation selection signatures between intensive and nonintensively farmed pig breeds. By employing various selection signature methods, it was feasible to detect candidate PMH genes.
Pigs are an excellent model animal species for gaining insights into the human brain and neurodevelopment (Jelsing et al. Reference Jelsing, Nielsen and Olsen2006). The pig genome exhibits high sequence similarity and chromosomal synteny with the human genome, which is useful for genomic applications (Chen et al. Reference Chen, Baxter and Muir2007). Additionally, Chen et al. (Reference Chen, Guo and Yang2009) reported the identification of chromosomal regions associated with infanticidal behavior in pigs that contain genes comparable to those of humans and rodents that are involved in anxiety, bipolar affective disorder, or coping behaviors. Pigs exhibit anatomical similarities to humans, with the porcine brain having similarities to the human brain, exhibiting a gyrencephalic cranial structure (Jelsing et al. Reference Jelsing, Nielsen and Olsen2006). In particular, the hippocampus and sensory cortex arrangement in pigs are more similar to those of humans than are those in mice (Zhang et al. Reference Zhang, Hu and Xu2014). In terms of practical considerations, pigs exhibit several advantages over primates and other livestock models. These features include a shorter generation time, bigger litter size, and suitability for straightforward genetic editing (Lunney et al. Reference Lunney, Van Goor and Walker2021). Since 2007, numerous robust and replicable genetic findings have been reported for psychiatric disorders. These findings have mostly been reported via genome-wide association (GWAS) and structural variation studies (Sullivan et al. Reference Sullivan, Daly and O’Donovan2012) and were systematically collected in disease gene databases (such as MalaCards (Rappaport et al. Reference Rappaport, Twik and Plaschkes2017) and DISEASE and DisGeNET (Piñero et al. Reference Piñero, Queralt-Rosinach and Bravo2015)). Previous studies have suggested that human brain regions such as the amygdala, striatum, and hippocampus are especially relevant to mental disorders (Matthies et al. Reference Matthies, Rüsch and Weber2012). For example, the hippocampus, which is part of the limbic system, is involved in emotion, memory formation, stress responses, emotional regulation, and memory processes, and damage to this region can lead to memory and learning impairments, as well as behavioral disorders such as anxiety, depression, and schizophrenia (Rastegar-Moghaddam et al. Reference Rastegar-Moghaddam, Mohammadipour and Hosseini2019). Since there are fewer studies on pig mental health trait, we consulted databases related to human mental disorders. By annotating genes from databases associated with human mental illnesses, we constructed a dataset for the PMH gene, thereby narrowing down the scope of candidate PMH genes.
In this study, we aimed to first compile a list of high-confidence psychiatric disease risk genes identified from human psychiatric disease databases and then further screen for selection signatures that reflect positive selection under an intensive pig farming system. Finally, we systematically revealed the characteristics of candidate PMH genes, including their expression levels and changes in different types of tissues and cells, and their potential effects on pig production traits. Our findings provide novel insight into elucidating the genetic basis of PMH traits.
Materials and methods
Datasets and their sources
Human disease–gene data sets
From three databases, DISEASE, DisGeNET (Piñero et al. Reference Piñero, Queralt-Rosinach and Bravo2015), and MalaCards & GeneCards (Rappaport et al. Reference Rappaport, Twik and Plaschkes2017), we collected a total of 1642 gene–disease records with gene–disease association scores (Table S1).
Genotyping dataset
We collected the genotypes of 348 intensively farmed pigs (including pigs of the Duroc, Landrace, Large White, Bershire, Hampshire, and Pietrain breeds) and 446 nonintensively farmed pigs (including 39 Chinese indigenous pig breeds, Table S2) from the PHARP v2 database (Wang et al. Reference Wang, Zhang and Chen2022), a pig genotyping source containing a total of 2,048 individuals of intensively farmed breeds, Chinese indigenous pig breeds, and wild boars. The genotypes contained approximately 23.1 million SNPs (Single Nucleotide Polymorphism) after removing those with minor allele frequencies of less than 0.05. All the statistical analyses were performed using the above genotype data unless otherwise specified.
Refinement of human psychiatric candidate genes
Given the challenges in the past in ascertaining the phenotypes of mental health traits in pigs, studies on the genetic basis of these traits have been limited. Nevertheless, genes associated with human psychiatric conditions have been well identified and compiled in the aforementioned human disease–gene databases. Through homologous gene analysis between pigs and humans, we were able to identify potential mental health genes in pigs by refining human psychiatric genes. Here, we used three human disease–gene databases to assemble a list of human psychiatric genes. Briefly, we first classified the disease names in each database into the DSM-5 and ICD-10-CM mental disorders classification code (criteria for the diagnosis of various mental disorders) utilizing UMLS Metathesaurus (Schuyler et al. Reference Schuyler, Hole and Tuttle1993) and then manually reviewed uncategorized disease names in five databases – DSM-5-TR, OMIM (Amberger et al. Reference Amberger, Bocchini and Schiettecatte2015), MalaCards (Rappaport et al. Reference Rappaport, Twik and Plaschkes2017), Disease Ontology (Schriml et al. Reference Schriml, Arze and Nadendla2012), and UMLS Metathesaurus Browser (Bodenreider Reference Bodenreider2004) – to determine whether they were associated with mental disorders and were classified into the DSM5-TR code (Tables S3–S5). According to the DSM5-TR and ICD10-CM codes, there are a total of 22 distinct categories for classified diseases. The above steps were used to classify the different diseases and determine whether a disease was related to a human mental disorder. We next filtered disease–gene items with disease–gene association scores below the top 2.5% to retain the high confidence disease–gene items for each database. Genes supported by at least two of these three disease–gene databases were considered relevant to human mental disorders. The Biomart tool (Durinck et al. Reference Durinck, Moreau and Kasprzyk2005) was used to analyze homologous genes between humans and pigs, thereby establishing a dataset of genes associated with PMH.
Population structure analysis
We applied three methods to analyze the population genetic structure. First, we performed principal component analysis (PCA) using PLINK v1 90 (Purcell et al. Reference Purcell, Neale and Todd-Brown2007) software to cluster individuals. Next, we used the ADMIXTURE program (Alexander and Lange Reference Alexander and Lange2011) to estimate the proportion of common ancestors among individuals. Finally, we built the NJ-tree with MEGA11 (Miura et al. Reference Miura, Tamura and Tao2020) using the genetic distance among individuals calculated by PLINK v1.90 and refined it with iTOL v6 (Letunic and Bork Reference Letunic and Bork2021).
Selection signature detection
Here, we employed five approaches, including three interpopulation methods (F ST (Weir and Cockerham Reference Weir and Cockerham1984), cross-population compound likelihood ratio [XP-CLR] (Chen et al. Reference Chen, Patterson and Reich2010), and cross-population extended haplotype homozygosity [XP-EHH] (Sabeti et al. Reference Sabeti, Varilly and Fry2007)) and two intrapopulation methods (composite likelihood ratio [CLR] (Nielsen et al. Reference Nielsen, Williamson and Kim2005) and runs of homozygosity [ROH] (Gibson et al. Reference Gibson, Morton and Collins2006)) to identify putative genomic regions undergoing positive selection. Considering that genes associated with pig mental health have undergone soft positive selection, we integrated selection signatures identified by five complementary approaches to identify potential PMH candidate genes.
FST test
F ST is a method for detecting selection signatures based on population differentiation (Weir and Cockerham Reference Weir and Cockerham1984). We screened the mean F ST value for each genomic region with a window size of 50 kb and a step size of 10 kb between intensively and nonintensively farmed pig populations by VCFtools v0.1.16 software (Danecek et al. Reference Danecek, Auton and Abecasis2011) and considered the genomic regions with the top 1% mean F ST values as putative positive selection signatures.
XP-CLR test
The XP-CLR test is a method used to estimate the CLR of a genomic region to detect genome-wide selection signatures (Chen et al. Reference Chen, Patterson and Reich2010). We estimated the XP-CLR across each chromosome by using xpclr v1 1 2 (Chen et al. Reference Chen, Patterson and Reich2010) with the parameters ld 0.95 – maxsnps 200 – size 50,000 – step 10,000 and defined the genomic regions with XP-CLRs in the top 1% as putative positive selection signatures.
XP-EHH test
XP-EHH is a haplotype-based method for detecting selection signatures (Sabeti et al. Reference Sabeti, Varilly and Fry2007). We used the intensively farmed pig population as the observed population and the nonintensively farmed population as the reference population using Selscanv1 3 0 (Szpiech and Hernandez Reference Szpiech and Hernandez2014) to calculate XP-EHH values and considered genomic regions with positive XP-EHH values in the top 2.5% as putative positive selection signatures.
CLR test
The CLR test is a method for detecting selection signatures based on the distribution of allele frequencies in a population. We calculated the CLR value for every 50 kb genomic region across the genome using SweeD software (Pavlidis et al. Reference Pavlidis, Živković and Stamatakis2013) and then ranked each genomic segment based on the CLR value. The genomic regions with 5% right-tail empirical p values were regarded as potential positive selection regions.
ROH detection
ROH is an approach for detecting selection signatures by identifying genomic regions with reduced variation relative to the genome average. We estimated the ROH for each individual using genotypes after linkage disequilibrium pruning (with the parameter – indep-pairwise 50 10 0.2) by “-homozyg” in PLINK v1.90 software (Purcell et al. Reference Purcell, Neale and Todd-Brown2007) with the parameters – homozyg-snp 100, – homozyg-window-snp 50, and – homozyg-window-missing 5. The criteria applied for ROH identification were as follows (Fang et al. Reference Fang, Hao and Xu2021): (1) 50 SNPs were contained in each sliding window; (2) an ROH consisted of no less than 100 consecutive SNPs; and (3) the density was higher than one SNP per 50 kb (Fang et al. Reference Fang, Hao and Xu2021). We calculated the frequency of occurrence for each SNP within ROH segments across all individuals and defined SNPs with frequencies in the top 1% as significant regions.
Putative candidate genes under selection
We considered genes within the 50 kb region upstream and downstream of identified selection signatures as potentially selected candidate genes and utilized the GALLO package (Fonseca et al. Reference Fonseca, Suárez-Vega and Marras2020) with the pig reference genome Sus11 1 (Warr et al. Reference Warr, Affara and Aken2020) to annotate selection signatures in the pursuit of potential selected candidate genes.
Characterization of candidate PMH genes
Functional enrichment analysis
We used the clusterProfiler package (Wu et al. Reference Wu, Hu and Xu2021) to identify Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways and Gene Ontology (GO) terms enriched with candidate PMH genes. P values were adjusted for multiple tests using the BH (Benjamini-Hochberg) method, and the adjusted P value was set to 0.05 as the significance threshold. Furthermore, we used the pig quantitative trait loci (QTL) database (https://www.animal-genome.org/cgi-bin/QTLdb/SS/index) (Hu et al. Reference Hu, Park and Reecy2022) to annotate economic traits associated with the candidate genes.
Tissue gene expression analysis
To investigate gene-tissue expression specificity, we examined the expression profiles of candidate PMH genes across different tissues from Pig-GTEx (Teng et al. Reference Teng, Gao and Yin2022) (Table S6). Using the Euclidean distance and Ward’s clustering methods, we grouped the candidate PMH genes into different clusters according to their different expression profiles across tissues. A heatmap of gene expression was generated using the R “gplots” package with the “heatmap.2” function.
Weighted expression cell-type enrichment analysis
Expression-weighted cell-type enrichment (EWCE) (Skene and Grant Reference Skene and Grant2016) was used for PMH candidate gene enrichment analysis with default parameters. This analysis was performed separately for each brain region dataset (Darmanis et al. Reference Darmanis, Sloan and Zhang2015) and for the hippocampus of pig single-cell data derived from our unpublished data. Then, the specificity matrix for each dataset was calculated by using the EWCE package. All the detected genes in each brain region dataset were used as background, and bootstrap resampling was performed 10,000 times. The adjusted P value was corrected using the BH method, and 0.05 was used as the significance threshold.
Differentially expressed genes (DEGs)
We used the “scanpy.tl.rank_genes_groups” function in Scanpy (v1.9.1) (Wolf et al. Reference Wolf, Angerer and Theis2018) with the Wilcoxon rank-sum test to identify the DEGs among candidate PMH genes in the inhibitory neurons of the hippocampus in both Jinhua and Duroc pigs. The parameters were adjusted P value < 0.01 and |logfold change| > 1.
Protein–protein interaction (PPI) network
We constructed a network using the GeneMANIA database (Warde-Farley et al. 2010) online tool (https://genemania.org/) to explore the connections between genes.
GWAS signal enrichment analysis
To explore the potential effects of candidate PMH genes on economic traits, we performed GWAS signal enrichment analysis using our unpublished GWAS summary data (the effects considered were sex, parity, year-season, and three principal components) of six traits, including corrected backfat thickness of 115 kg (BFT115), corrected day of age to reach 115 kg (DAY115), feed conversion rate, feed intake, loin muscle depth, offset intramuscular fat (OffsetIMF), and represent feed intake. We implemented a summation-based marker set significance test method (Fang et al. Reference Fang, Cai and Liu2020) using the QGG software package (Rohde et al. Reference Rohde, Fourie Sørensen and Sørensen2020) for the above six traits. We considered false discovery rate-corrected P values < 0.05 as the threshold for identifying significantly correlated traits.
Results
Pig mental health candidate genes
From the DisGeNET database, we assembled a list of 1,269 human mental disorder candidate genes related to 122 diseases and 2,786 gene–disease items (Fig. 1A and Fig. S1A). Using the disease database, we assembled a list of 2,467 human mental disorder candidate genes related to 108 mental diseases and 6,378 gene–disease items (Fig. 1B and Fig. S1B). Similarly, from the MalaCards database, we assembled a list of 2,661 human mental disorder candidate genes related to 377 diseases and 42,384 gene–disease items (Fig. 1C and Fig. S1C). Finally, we narrowed down a total of 1,642 genes identified as candidates for human mental disorders that appeared in at least two out of three databases (Fig. 1D and Table S7). Subsequently, we conducted a human-to-pig homologous gene search using Biomart for these candidate genes, resulting in 1,549 pig genes (Table S8), which included 1,439 genes located on autosomes (Fig. S2 and Table S9).
Summary of human disease gene databases.

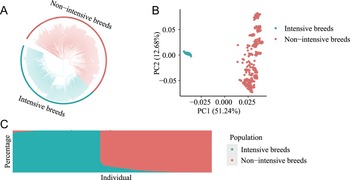
Population genetic structure analysis of the NJ-tree, PCA, and ADMIXTURE showed that individuals were well clustered into two groups (intensive and nonintensively farmed pig breeds, Fig. 2A–C). The PCA plot showed that PC1 could clearly separate individuals into intensively farmed pig breeds and nonintensively farmed pig breeds. PC1 and PC2 accounted for 51.24% and 12.68% of the total variance, respectively (Fig. 2B). Similarly, the intensively farmed pigs were clearly distinguished from the nonintensively farmed pigs when K = 2 in the admixture analysis (Fig. 2C). Next, we conducted selection signatures analysis on these intensive and nonintensively farmed pig breed. According to the selection criteria for each method (see details in Materials and Methods), the F ST, XP-EHH and XP-CLR, CLR, and ROH methods revealed selection signatures associated with 537, 922, 1,013, 912, and 622 genes, respectively (Fig. 3A-E and Tables S10-S14). A total of 2,844 unique positively selected genes (PSGs) were detected by these five selection signature detection methods (Fig. 3F).
Population structure.

Selection signatures across the autosomes of pigs.

We then compared PSGs with human mental disorders genes, resulting in 254 genes identified as candidate PMH genes (Fig. 4A, Figs. S3-S7, and Table S15). Functional enrichment analysis of the 254 candidate PMH genes revealed that they were overrepresented in 83 KEGG pathways and in 275 GO terms (Fig. 4B and C). Many of these pathways or GO terms are highly relevant to signal transduction. The cellular components in which the candidate PMH genes were enriched were those such as dendrites, synapses, neuronal projections, and postsynapses, indicating their involvement in signal transmission (Table S16). The GO terms in which these genes were also enriched were related to the regulation of signal transmission (Table S17). KEGG pathway enrichment analysis revealed that the dopaminergic synapse, cAMP signaling pathway, neuroactive ligand‒receptor interaction, calcium signaling pathway, glutamatergic synapse, GABAergic synapse, and serotonergic synapse genes were enriched (Table S18). The QTL enrichment analysis indicated that the loci encompassed by these genes were linked to meat quality traits such as meat color b*, intermuscular fat content, and drip loss (Fig. 4D). GWAS signal enrichment analysis revealed that PMH candidate genes were significantly (p < 0.05) associated with corrected 115 kg backfat thickness, corrected age in days to reach 115 kg body weight, and offset intramuscular fat, suggesting that PMH candidate genes may influence routine production performance (Fig. 4E).
Functional annotations of candidate PMH genes.

Spatial and temporal specificity of PMH candidate gene expression
Based on the expression profiles of PMH candidate genes in various brain tissues, we observed high expression of these genes in the frontal cortex, hippocampus, and hypothalamus tissues (Fig. 4F and Fig. S8). Genes, such as DRD1, CACNA1I, CACNA1E, GABRB2, and SCN2A, related to calcium channels, potassium channels, and GABA transporters exhibited high expression in the hippocampus. The DRD1 gene is a dopamine receptor encoding a protein that initiates G-protein-coupled receptor activity and dopamine neurotransmitter receptor activity (Li et al. Reference Li, Liangsheng and Yang2024). The CACNA1I and CACNA1E genes encode calcium channel proteins that mediate the entry of calcium ions into excitable cells and participate in various calcium-dependent processes (El Ghaleb et al. Reference El Ghaleb, Schneeberger and Fernández-Quintero2021; Helbig et al. Reference Helbig, Lauerer and Bahr2018). The GABRB2 gene encodes the β-2 subunit, influencing the major inhibitory system in the brain – the GABA system (Barki and Xue Reference Barki and Xue2022). The SCN2A gene encodes a sodium channel protein, and variations can lead to various neurological disorders, including benign epilepsy, epileptic encephalopathy, and autism spectrum disorders (Sanders et al. Reference Sanders, Campbell and Cottrell2018).
To investigate the candidate PMH genes spatiotemporal specificity expression pattern in pigs of different ages, we subsequently compared the transcriptomic expression data between days 38 and 56 in the prefrontal cortex region of the porcine brain. During this period, pigs have just experienced weaning and regrouping, which may induce some stress responses (Fig. 4G). We detected an increase in the expression of GABRA4 at the 56-day time point. These two genes encode the β subunits of GABA-A receptors, which are inhibitory neurotransmitter receptors. The expression of the SLC17A7 gene was also significantly higher on day 56 than on day 38. The SLC17A7 gene is primarily expressed in neurons and plays a crucial role in the packaging and transport of the neurotransmitter glutamate into synaptic vesicles. The SLC1A1 gene encodes a protein that removes excess glutamate from the synaptic gap to maintain neurotransmitter homeostasis. The DRD1 gene functions as a type of dopamine receptor. The SCN2A and SCN1A genes also encode sodium channel proteins, while the KCNQ2 and KCNC2 genes are related to potassium channels. The DTNBP1 gene encodes a membrane-associated protein. Notably, the expression of these genes was higher at 56 days than at 38 days. Calcium ion-related genes, CACNA1I, CACNA1A, and CACNA1E, did not change significantly between these two periods. Compared with the expression of the HSPA5 gene on day 38, that on day 56 significantly decreased.
High expression of candidate PMH genes in inhibitory neurons in the hippocampus
Intriguingly, we observed that the expression of PMH genes was significantly enriched in inhibitory neurons in the hippocampus of JH pigs (a nonintensively farmed pig breed) (Fig. 5A), whereas the enrichment was not significant in Duroc pigs (an intensively farmed pig breed) (Fig. 5B). Therefore, we compared the expression of candidate PMH genes in inhibitory neurons in the hippocampus between the two breeds (Fig. 5C and Table S19). In Duroc pigs, there were 11 genes whose expression was greater than that in JH pigs and eight genes whose expression was lower than that in JH pigs. Moreover, PPI network analysis of these eight downregulated genes revealed that the HSPA5 and HSPA8 genes were hub genes (Fig. 5D). The differences in hippocampal gene expression patterns between the two breeds led us to focus on specific cell types for studying PMH traits in the future. Moreover, we conducted enrichment analyses of candidate PMH genes using human brain data, and the candidate PMH genes were also enriched in neuron cell types in the human brain (Fig. 5E and F). This finding suggested a significant association between PMH genes and neurons in the hippocampus of pigs.
The expression pattern of pig mental health candidate genes in single cells.

Flowchart of PMH candidate gene construction and functional annotation.

Discussion
With the intensification of pig farming in China, large-scale and intensive production management methods have imposed significant stress on pigs, leading to decreased production performance and substantial economic losses in the swine industry. However, there has been limited research focusing on PMH, as these traits are challenging to measure and identify. Currently, there is a scarcity of studies and a lack of phenotype data in this area. Given the high homology between humans and pigs, we aim to leverage information on human diseases to identify candidate genes related to PMH through comparative genomics. Our findings are beneficial for conducting molecular design breeding and genetic improvement to adapt pigs to intensive environments, reduce stress, and enhance economic trait performance.
In our study, we identified 254 candidate PMH genes that underwent positive selection during the transition from nonintensive to intensive farming. The KEGG pathways and GO terms in which these genes were enriched were related to neurotransmission, such as the dopaminergic synapse, neuroactive ligand‒receptor interaction, calcium signaling pathway, glutamatergic synapse, and GABAergic synapse. Dysregulation in the dopamine system is one of the factors contributing to the development of schizophrenia, and current antipsychotic drugs mainly act by antagonizing dopamine receptors (Barki and Xue Reference Barki and Xue2022). A previous study also showed that imbalances in glutamate and GABA (the primary inhibitory neurotransmitter) can directly disrupt endocrine activity, leading to behavioral abnormalities (Sanders et al. Reference Sanders, Campbell and Cottrell2018).
Furthermore, PMH candidate genes are significantly expressed in brain regions involved in mood disorders, such as the prefrontal cortex and hippocampus. Dysfunction in the prefrontal cortex and its connectivity with other brain regions, particularly the limbic system, has been observed in patients with schizophrenia (Amato et al. Reference Amato, Vernon and Papaleo2018). The hippocampus is another important brain structure for cognition and emotion (Nian et al. Reference Nian, Ding and Feng2023). These findings highlighted that the prefrontal cortex and hippocampus might be important brain regions associated with PMH.
Interestingly, the candidate genes are significantly expressed in inhibitory neurons in the hippocampus, suggesting a potential role in regulating neural activity and maintaining the balance between excitatory and inhibitory signals. Imbalances in excitatory and inhibitory neurotransmission have been implicated in mental disorders, including major depressive disorder (Duman et al. Reference Duman, Sanacora and Krystal2019). Stress during development has been shown to affect neuronal proliferation and differentiation in the mammalian (e.g., rats and monkeys) hippocampus (Lemaire et al. Reference Lemaire, Lamarque and Le Moal2006).
In addition, we found that the expression levels of certain candidate genes were lower in individuals from the intensively farmed pig breed (Duroc) than in those from the nonintensively farmed pig breed (Jinhua). These genes, including HSPA5, HSPA8, KCNC2, etc., have been implicated in genetic risk factors for bipolar disorder and schizophrenia (Wedenoja et al. Reference Wedenoja, Tuulio-Henriksson and Suvisaari2010). The downregulation of these genes may contribute to mental health in individuals exposed to intensive farming conditions.
Our findings highlighted that PMH might affect economic traits. First, we observed that PMH candidate genes were significantly enriched in meat quality and carcass traits in pigs. Additionally, through differential gene expression and protein interaction network analysis, HSPA5 and HSPA8 were identified as hub genes, indicating their close association with heat stress, which can effect meat quality and the immune response (Bejaoui et al. Reference Bejaoui, Sdiri and Ben Souf2023). Furthermore, the GWAS signal enrichment analysis of PMH candidate genes (Fig. 4D) revealed a significant association between these candidate genes and important production traits for meat quality, such as corrected 115 kg backfat thickness, corrected day of age to reach 115 kg, and offset intramuscular fat, both of which are important production traits for meat quality. Among these traits, backfat thickness is recognized as an essential indicator of fattening performance in commercial pigs. Notably, we identified seven genes (ASS1, MC4R, C3, INSR, LMNB2, TUBB4A, and SCARB2) with a total of 64 specific loci that were significantly associated with backfat thickness. These findings provide novel valuable insights into the relationship between PMH genes and meat quality traits, specifically focusing on backfat thickness and off set intramuscular fat.
Conclusion
Overall, in this study, we systematically identified a candidate gene list related to PMH at the genome level for the first time. Moreover, we provided comprehensive functional annotations of these putative genes. Our results will aid in the genomic selection of mental health traits in pigs, improve animal welfare, and facilitate the use of pigs as models of human psychiatric disorders.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/anr.2024.20.
Data availability statement
The genotype datasets generated and/or analyzed during the current study are available at PHARP (http://alphaindex.zju.edu.cn/PHARP/index.php) and at the SRA repository, https://www.ncbi.nlm.nih.gov/sra. See the “Materials and Methods” section above for their availability. Computer code for data processing is available from the authors upon request.
Acknowledgements
We thank C. Cao and Q. Xie for help in collecting the brain tissues of pigs. We thank Z. Wang and Z. Zhang et al. for providing the whole-genome data for pigs.
Author contributions
Z. Wang designed the research framework of the paper and directed the writing of the manuscript. X. Wei wrote the manuscript. X. Wei, L. Xu, and J. Jiang performed the bioinformatics analyses. Z.Z., Q.S.W., and Y.C.P. provided useful suggestions for the analysis. Z. Chen, F. Wu, and J. Miao conducted a rigorous review of the manuscript.
Funding statement
This work was supported by the Natural Science Foundation of China (32172691) and the National Key Research and Development Program of China (2022YFF1000500 and 2021YFD1200802).
Conflicts of interest
The authors declare that they have no competing interests.