INTRODUCTION
Coccidia in the family Sarcocystidae, the tissue coccidia, are important pathogens of many vertebrates, including humans (Velmurugan and Dubey, Reference Velmurugan and Dubey2008). Many species within the subfamily Toxoplasmatinae (e.g. Toxoplasma gondii, Neospora caninum, Hammondia heydorni and Hammondia hammondi) exhibit facultative or obligatory heteroxenous life cycles. Exogenous sporulation produces disporocystic tetrazoic oocysts without Stieda bodies (Carreno et al. Reference Carreno, Schnitzler, Jeffries, Tenter, Johnson and Barta1998; Mugridge et al. Reference Mugridge, Morrison, Heckeroth, Johnson and Tenter1999). Coprological identification of N. caninum, T. gondii, H. heydorni and H. hammondi is difficult because their unsporulated oocysts are morphologically indistinguishable for all practical purposes (Frenkel and Dubey, Reference Frenkel and Dubey1975; Dubey and Lindsay, Reference Dubey and Lindsay1996; Lindsay et al. Reference Lindsay, Dubey and Blagburn1997). Only some Cystoisospora spp. (e.g. C. canis and C. felis) possess distinctly larger oocysts that can be diagnosed reliably using microscopy. There are only two genera in the Sarcocystinae (Sarcocystis Lankester 1882 and Frenkelia Biocca 1968) distinguished by the location of tissue cysts within intermediate hosts; the monophyly of these genera has been questioned repeatedly based on phylogenetic hypotheses erected using nuclear rDNA sequences (e.g. Mugridge et al. Reference Mugridge, Morrison, Jakel, Heckeroth, Tenter and Johnson2000; LSU rDNA, Morrison et al. Reference Morrison, Bornstein, Thebo, Wernery, Kinne and Mattsson2004). The number of valid genera and species within the coccidian subfamily Toxoplasmatinae has been the subject of ongoing debate (e.g. Dubey, Reference Dubey and Kreier1977; Carreno et al. Reference Carreno, Schnitzler, Jeffries, Tenter, Johnson and Barta1998; Ellis et al. Reference Ellis, Morrison, Liddell, Jeckins, Mohammed, Ryce and Dubey1999; Mehlhorn and Heydorn, Reference Mehlhorn and Heydorn2000; Dubey and Sreekumar, Reference Dubey and Sreekumar2003; Morrison, Reference Morrison2009). Currently, there are seven described genera recognized in the subfamily: Hyaloklossia Labbé 1896; Toxoplasma Nicolle & Manceaux 1908; Besnoitia Henry 1913; Cystoisospora Frenkel 1977; Hammondia Frenkel and Dubey Reference Frenkel and Dubey1975; Neospora Dubey et al. Reference Dubey, Carpenter, Speer, Topper and Uggla1988; and Nephroisospora Wünschmann et al. 2010. Toxoplasma gondii was described from tissue cysts (bradyzoites) observed in rodents in 1908. Dubey et al. (Reference Dubey, Carpenter, Speer, Topper and Uggla1988) proposed the name N. caninum to reflect the distinctiveness of N. caninum from T. gondii and summarized the biological, morphological, molecular and antigenic differences that distinguish these two parasites (Dubey et al. Reference Dubey, Barr, Barta, Bjerkas, Björkman, Blagburn, Bowman, Buxton, Ellis, Gottstein, Hemphill, Hill, Howek, Jenkins, Kobayashi, Koudela, Marsh, Mattsson, McAllister, Modry', Omata, Sibley, Speer, Trees, Uggla, Upton, Williams and Lindsay2002b ). The genus Besnoitia Henry 1913 was named for tissue coccidia that form thick-walled polyzoic cysts within connective tissue of their intermediate hosts (e.g. Besnoitia wallacei Frenkel 1977). Frenkel and Dubey (Reference Frenkel and Dubey1975) erected the genus Hammondia to accommodate a feline coccidium, H. hammondi, with an obligatory two host life cycle. Hammondia heydorni Dubey, Reference Dubey and Kreier1977, a parasite of canids, was characterized by Dubey (Reference Dubey and Kreier1977) and Blagburn et al. (Reference Blagburn, Lindsay, Swango, Pidgeon and Braund1988) from oocysts retrieved from feces of dogs. A second Hammondia sp. of canids, H. triffittae, has been recognized in European foxes by Gjerde and Dahlgren (Reference Gjerde and Dahlgren2011) who distinguished this parasite from other isosporoid coccidia of dogs (i.e. H. heydorni or N. caninum) by subtle differences in oocyst dimensions, genetic differences and the inability of H. triffittae to infect dogs. The separation of these two Hammondia spp. of canids was supported by later molecular comparisons (Schares et al. Reference Schares, Meyer, Bärwald, Conraths, Riebe, Bohne, Rohn and Peters2003; Dahlgren and Gjerde, Reference Dahlgren and Gjerde2010; Gjerde and Dahlgren, Reference Gjerde and Dahlgren2011).
Most molecular phylogenetic studies of apicomplexan parasites in the family Sarcocystidae relied on small subunit (18S) rDNA sequences (e.g. Tenter and Johnson, Reference Tenter and Johnson1997; Mugridge et al. Reference Mugridge, Morrison, Jakel, Heckeroth, Tenter and Johnson2000). The nuclear 18S rDNA locus is a useful genetic target from which to obtain sequence data due to its high copy number and conserved terminal sequences that are convenient for PCR primer design and amplification (Morrison and Ellis, Reference Morrison and Ellis1997; Mugridge et al. Reference Mugridge, Morrison, Jakel, Heckeroth, Tenter and Johnson2000; Ouvrard et al. Reference Ouvrard, Campbell, Bourgoin and Chan2000). These attributes and the relatively conserved nature of the 18S rDNA within the nuclear genome have made 18S rDNA an important and widely exploited genetic target for biodiversity and species identification studies. In most eukaryotes, rDNA copies undergo concerted evolution that homogenizes copy-to-copy variability in the nuclear genome (Hillis and Dixon, Reference Hillis and Dixon1991). However, this is not always true for members of the Apicomplexa. Highly divergent paralogous copies of nuclear rDNA have been demonstrated within haemosporinid parasites (McCutchan et al. Reference McCutchan, Cruz, de la Lal, Gunderson, Elwood and Sogin1988, Reference McCutchan, Li, McConkey, Rogers and Waters1995; Li et al. Reference Li, Gutell, Damberger, Wirtz, Kissinger, Rogers, Sattabongkot and McCutchan1997), cryptosporidia (Le Blancq et al. Reference Le Blancq, Khramstov, Zamani, Upton and Wu1997), piroplasms (Goethert et al. Reference Goethert, Cook, Lance and Telford2006) and, more recently, eimeriid coccidia (Vrba et al. Reference Vrba, Poplstein and Pakandl2011; El-Sherry et al. Reference El-Sherry, Ogedengbe, Hafeez and Barta2013). The possibility of highly divergent rRNA genes (e.g. A, S and O 18S rRNA genes that vary in both primary and secondary structures) could make determining homology at the gene-level problematic. Morrison et al. (Reference Morrison, Bornstein, Thebo, Wernery, Kinne and Mattsson2004) noted that complete 18S rDNA sequences were inadequate to resolve relationships among closely related tissue coccidia. Phylogenetic reconstructions based on 18S rDNA sequences support a monophyletic Sarcocystidae consistently (Ellis and Morrison, Reference Ellis and Morrison1995; Carreno et al. Reference Carreno, Schnitzler, Jeffries, Tenter, Johnson and Barta1998; Mugridge et al. Reference Mugridge, Morrison, Jakel, Heckeroth, Tenter and Johnson2000; Morrison et al. Reference Morrison, Bornstein, Thebo, Wernery, Kinne and Mattsson2004); however, monophyly of individual genera within the Sarcocystidae is not supported in these analyses. Unlike nuclear 18S rDNA sequences, comparatively short sequences obtained from the mitochondrial cytochrome c oxidase subunit I (mt COI) gene provide sufficient sequence divergence to clearly differentiate closely related coccidia. Ogedengbe et al. (Reference Ogedengbe, Hanner and Barta2011) demonstrated that ~500–800 bp partial mt COI sequences could better distinguish closely related coccidia (Eimeriidae) compared with complete or near-complete nuclear 18S rDNA sequences. Until recently (Gjerde, Reference Gjerde2013a , Reference Gjerde b ), lack of primers capable of amplifying the mt COI locus from sarcocystid coccidia restricted the use of mt COI sequences to only eimeriid coccidia (e.g. Eimeriidae) and a few members of the Toxoplasmatinae (see Ogedengbe et al. Reference Ogedengbe, Hanner and Barta2011). Based on the ability to better resolve relationships among closely related coccidia using mt COI sequences (e.g. Ogedengbe et al. Reference Ogedengbe, Hanner and Barta2011; El-Sherry et al. Reference El-Sherry, Ogedengbe, Hafeez and Barta2013), the monophyly of the described genera in the Sarcocystinae might best be tested using this genetic locus.
In this study, we used existing and newly designed PCR primers to amplify and sequence portions of the mt COI gene from a variety of tissue coccidia (i.e. parasites belonging to the genera Toxoplasma, Neospora, Hammondia, Cystoisospora, Nephroisospora and Sarcocystis in the Sarcocystidae). Partial mt COI sequences, nuclear 18S rDNA sequences or a concatenation of sequences from both loci were utilized to infer evolutionary relationships among these tissue coccidia and confirm the monophyly of some named genera in the family Sarcocystidae.
MATERIALS AND METHODS
Parasite sources
Parasites and parasite genomic DNA were obtained from a variety of sources. Fecal specimens containing oocysts of C. felis, C. suis, C. cf. ohioensis, C. canis and H. heydorni were obtained from diagnostic fecal specimens submitted to the Animal Health Laboratory, Laboratory Services Division, University of Guelph (Guelph, ON, Canada). Additional fecal specimens (containing C. felis, C. canis, C. suis, C. cf. ohioensis or C. rivolta) from a variety of hosts were kindly provided by Dr Donald Martin (IDEXX Laboratories, Markham, ON). Some fecal samples containing C. cf. ohioensis were provided by Dr Scott Weese (Department of Pathobiology, University of Guelph, ON). DNA samples of Sarcocystis spp. from experimentally infected hosts (i.e. S. rileyi 908131_Duck2 #4) or from tissue culture (i.e., S. neurona MIH2) were kindly provided by Dr Ben Rosenthal (USDA, Beltsville, MD, USA). Purified parasite genomic DNA was obtained for H. hammondi strain H.H-20 and T. gondii strains GT1, MAS, PTG, TgCat, CalBr64, CalBr5, TgCal and TgToucan prepared from scrapes of tachyzoite-infected cell cultures were provided by Dr Chunlei Su (Department of Microbiology, University of Tennessee, TN); the origins of these lines have been described previously (Su et al. Reference Su, Khan, Zhou, Majumdar, Ajzenberg, Dardé, Zhu, Ajioka, Rosenthal, Dubey and Sibley2012).
Identification of oocysts
Sporulated Cystoisospora oocysts collected and measured for this study were from dogs, cats, and pigs, and were identified as C. canis, C. felis, C. rivolta and C. suis. Small subspherical oocysts shed by dogs were considered part of the C. cf. ohioensis species complex and assigned the name C. cf. ohioensis to reflect the uncertainty in their species identification. In all cases prior to DNA extraction, initial assignment to species was on the basis of oocyst morphometrics. Oocysts images were captured using a Provis AX 70 photomicroscope (Olympus 95 Canada, Richmond Hill, ON) fitted with a digital imaging device (Infinity3–1C, Lumenera Corporation Ottawa, ON) controlled using iSolution Lite image analysis software (Hoskin Scientific, Burlington, ON). Measurements are reported as means ± standard deviation in μm with the range in parenthesis.
Oocyst purification and DNA extraction
Oocysts were isolated from fecal samples using saturated salt (NaCl) flotation followed by a bleach treatment to remove exogenous DNA prior to parasite DNA extraction (Ogedengbe et al. Reference Ogedengbe, Hafeez and Barta2013, Reference Ogedengbe, El-Sherry, Whale and Barta2014). DNA isolation was accomplished using glass bead disruption and DNAzol nucleic acid extraction (Invitrogen, Carlsbad, CA) as previously described (Ogedengbe et al. Reference Ogedengbe, Hafeez and Barta2013). DNA concentration was estimated using a Nanodrop 2000 spectrophotometer (NanoDrop Products, Wilmington, DE) and stored at 4 °C for immediate use or −20 °C for later use.
Primers for PCR amplification of mt COI
Initial PCR reactions attempted with ‘universal’ COI LCO1490 and HCO2198 primers that amplify many metazoan mt COI (Folmer et al. Reference Folmer, Black, Hoeh, Lutz and Vrijenhoek1994) failed to amplify the COI locus from any species within the Toxoplasmatinae. Previously described coccidian-specific primers (Ogedengbe et al. Reference Ogedengbe, Hanner and Barta2011) were able to amplify some, but not all, members of the Sarcocystidae. Consequently, a pair of new Sarcocystidae-specific primers was designed using an alignment of all available COI sequences from tissue coccidia (data not shown). A pair of Sarcocystidae group specific degenerate primers (Sdae–COI 260F, degeneracy 4; Sdae–COI 1147R, degeneracy 2) was designed from two comparatively conserved regions with the aid of Primer3 (Untergrasser et al. Reference Untergrasser, Cutcutache, Koressaar, Ye, Faircloth, Remm and Rozen2012) executed from within the bioinformatics package Geneious (Version 6.1·8; http://www.geneious.com, Kearse et al. Reference Kearse, Moir, Wilson, Stones-Havas, Cheung, Sturrock, Buxton, Cooper, Markowitz, Duran, Thierer, Ashton, Mentjies and Drummond2012). The previously published and newly designed primers were then used in various combinations to amplify ~500–900 bp fragments of the mt COI locus (Table 1). PCR reactions were ran with the anneal Tms determined with the Primer3 program for the primer pairs. Beginning with the lowest possible Tms for a pair of degenerate primers, anneal Tms were move up if multiple product bands were obtained to eliminate amplification of spurious DNA products. Extension times were usually set at the standard 1 min kb−1 expected product length.
Amplification primers for mitochondrial COI and nuclear 18S rDNA loci for various tissue coccidian parasites including anneal temperatures (T a) and expected PCR product sizes.

PCR – partial mt COI and near-complete nuclear 18S rDNA
PCR amplification was carried out in an MJ Mini thermal cycler (Bio Rad, CA) with reactions consisting of 1 × PCR buffer (Invitrogen, Carlsbad, CA) supplemented with 4 mM MgCl2, 200 µ m dNTPs, 0·5 µ m of each of the amplification primers (Table 1), 2·5 U of Platinum® Taq Polymerase (Invitrogen) and 50–100 ng DNA template. Cycling conditions were initial denaturation at 96 °C for 5 min followed by 35 cycles of 94 °C for 30 s, annealing at 50–55 °C (see Table 1 for anneal conditions required for specific primer combinations) for 30 s and extension at 72 °C for 30–90 s (depending on expected product size). The PCR reaction was completed with a final extension step at 72 °C for 7–10 min. PCR reaction products were electrophoresed using a 1·5% submarine agarose gel, stained with ethidium bromide and visualized using UV trans-illumination (Spectronics Corporation, New York, NY). The apparent size of DNA bands were determined by comparison with a 100 bp DNA ladder (Bio Basic Inc., Markham, ON). Bands were excised using a fresh scalpel blade and gel purified using a QIAGEN Gel Extraction Kit (Valencia, CA) according to the manufacturer's instructions. The purified PCR products were cycle sequenced using an ABI Prism 7000 Sequence Detection System (Applied Biosystems Inc., Foster City, CA) by the Molecular Biology Unit of the Laboratory Services Division, University of Guelph (Guelph, ON) using the amplification primers to obtain sequences in both directions. For the longer nuclear 18S rDNA fragments, internal sequencing primers were used so that complete, double-stranded sequencing was obtained.
Sequence assembly
The sequencing reads were assembled using de novo assembly within the Geneious bioinformatics software package. For both loci, the strict consensus sequence, less the amplification primers, was submitted to GenBank to obtain accession numbers reported in Supplementary Table 1.
Added to newly generated sequences, existing publically available COI and corresponding complete or near-complete 18S rDNA sequences generated from the same strain or isolate, when available, were used preferentially for phylogenetic analyses (see Supplementary Table 1 for sequences and strains used in this study). GenBank sequence FJ357797, previously identified as an Isospora sp. (strain Harbin/01/08) retrieved from the Siberian tiger (Panthera tigris altaica), was renamed as Cystoisospora sp. (strain Harbin/01/08) in this study on the basis of sporulated oocysts possessing sporocysts without Stieda bodies and with sporozoites lacking refractile bodies (see Fig. 2A and B of Zhijun et al. Reference Zhijun, Mingwei, Hongliang and Yuping2011); these morphological features are consistent with the genus Cystoisospora rather than Isospora (see Barta et al. Reference Barta, Schrenzel, Carreno and Rideout2005). In addition, the following selected mt COI and 18S rDNA sequences from apicomplexan parasites outside of the Eimeriorina were used as the taxonomic outgroup taxa for rooting the resulting trees: Hepatocystis sp.; Plasmodium juxtanucleare; P. vivax; P. falciparum; P. malariae; Babesia caballi; B. bovis; B. bigemina; B. rodhaini; Theileria annulata and T. parva. Accession numbers for all sequences included in the analyses are indicated on the resultant trees and in Supplementary Table 1.
Where possible, sequences obtained following bacterial cloning were excluded in the analyses to avoid sequence variations resulting from nucleotide misincorporation during PCR amplification (Olivieri et al. Reference Olivieri, Ermini, Rizzi, Corti, Bonnal, Luciani, Marota, De Bellis and Rollo2010; Ogedengbe et al. Reference Ogedengbe, Hafeez and Barta2013). The 18S rDNA sequence from Isospora rivolta (AY618554) was removed from all analyses because a BLAST search determined that this sequence was identified incorrectly; AY618554 is likely derived from a basidiomycete fungus (see Whipps et al. Reference Whipps, Fournie, Morrison, Azevedo, Matos, Thebo and Kent2012). In this study, only the newly generated nuclear 18S rDNA sequences from Cystoisopora canis were obtained from cloned PCR products.
Multiple sequence alignment
Multiple sequence alignments based on the primary structure were generated for the partial mt COI sequences using the ‘Translation Align’ algorithm within Geneious Ver. 6·1·8 bioinformatics software (Cost Matrix = Blosum 62; Gap open penalty = 12; Gap extension penalty = 3) (http://www.geneious.com, Kearse et al. Reference Kearse, Moir, Wilson, Stones-Havas, Cheung, Sturrock, Buxton, Cooper, Markowitz, Duran, Thierer, Ashton, Mentjies and Drummond2012).
Nuclear 18S rDNA sequences were trimmed at both ends to exclude primer regions with sequences starting at the identical homologous nucleotide position 14 and ending at position 1778 if compared with the GenBank sequence DQ060683 of C. belli (Gjerde, Reference Gjerde2013b ). Alignments were performed first with MAFFT (Katoh et al. Reference Katoh, Asimenos and Toh2009; Katoh and Toh, Reference Katoh and Toh2010) and ClustalW (Larkin et al. Reference Larkin, Blackshields, Brown, Chenna, McGettigan, McWilliam, Valentin, Wallace, Wilm, Lopez, Thompson, Gibson and Higgins2007) executed from within Geneious. The occurrence of hypervariable regions in the 18S rRNA gene warranted that the MAFFT alignment was further staggered to address regions for which positional homology was uncertain (Barta, Reference Barta1997). The sequence alignments were analysed as three separate sequence datasets: (1) an ‘18S rDNA dataset’; (2) a ‘COI dataset’; and (3) a ‘concatenated dataset’. The 18S rDNA dataset consisted of 100 aligned nucleotide sequences obtained from 61 spp. There were a total of 3273 character positions in the final 18S rDNA dataset of which 1689 characters were constant, 904 informative characters and 685 variable but uninformative characters. The COI dataset consisted of 91 aligned nucleotide sequences obtained from 59 spp. There were a total of 756 character positions in the final dataset of which 226 characters were constant, 501 informative characters and 29 variable but uninformative characters. The concatenated dataset combined the nuclear 18S rDNA and mt COI datasets into a single dataset without modifying the pre-existing sequence alignments for each genetic locus. The concatenated dataset had 4061 total characters of which 1943 characters were constant, 1408 informative characters and 710 variable but uninformative characters. All positions containing gaps were treated as unknown.
Phylogenetic analyses
Phylogenetic analyses for all datasets (i.e., 18S rDNA, COI or concatenated) were performed using three tree building methods: Bayesian Inference (BI) executed from within MrBayes Version 3.1.2. (Huelsenbeck and Ronquist, Reference Huelsenbeck and Ronquist2001; Ronquist and Huelsenbeck, Reference Ronquist and Huelsenbeck2003); Maximum Likelihood (ML) (Guindon et al. Reference Guindon, Dufayard, Lefort, Anisimova, Hordijk and Gascuel2010) executed from within PAUP version 4.0 Beta (Swofford, Reference Swofford2002); and Maximum Parsimony (MP) (Felsenstein, Reference Felsenstein1985) executed from within PAUP version 4.0 Beta. For MP analyses, all characters were treated as unordered and equal weight with gaps treated as missing.
For the likelihood-based analyses, best fit models and parameters were based on the Akaike Information Criterion (AIC) for 24 models of DNA substitution using the hierarchical likelihood ratio test performed within MrModeltest v2·3 (Nylander J. A. A. 2004. MrModeltest v2 Program; distributed by the author, Evolutionary Biology Centre, Uppsala University). Based on AIC, the Hasegawa, Kishino and Yano (HKY, nst = 2 gamma categories) model with a discrete gamma distribution rate variation among sites (G) and accounting for proportion of invariant sites (I) estimated base frequencies was recommended for the 18S rDNA dataset (Hasegawa et al. Reference Hasegawa, Kishino and Yano1985). The general time reversible model with discrete gamma distribution, including invariable site (GTR + I + G, nst = 6 gamma categories) (Tavaré, Reference Tavaré1986) was recommended for the COI data; a codon nucleotide model (i.e. Nucmodel = Codon) was implemented for all COI data using metazoan mitochondrial translation (i.e. Code = metmt). The concatenated dataset was partitioned in the ML and BI analyses. The COI portion of the concatenated dataset was analysed using a GTR + I + G (nst = 6, nucmodel = codon, code = metmt) substitution model and the 18S rDNA portion of the concatenated dataset was analysed using the HKY + I + G (nst = 2, nucmodel = 4by4) substitution model.
For all BI analyses, 1 000 000 generations of Markov Chain Monte Carlo (heated chains = 4, chain temperature = 0·2, unconstrained branch length with exponential = 10) were executed with a sampling frequency of 1000; burn-in length was set at 10% of the number of generations (i.e. 100 000). For MP and ML analyses, bootstrap consensus trees were constructed from 500 replicates of each analysis. Branches with less than 50% bootstrap support were collapsed. In all analyses, trees were rooted using members of the Haemosporida and Piroplasmida. Only BI trees (Figs 2–5) are shown. Trees were drawn such that horizontal branch lengths are proportional to hypothesized genetic divergence.
RESULTS
Oocyst dimensions
Representative oocysts collected from pigs, cats or dogs are illustrated in Fig. 1. The mean dimensions for oocysts and sporocysts of parasites isolated from fecal material are summarized for these parasites in Table 2.
Sporulated oocysts of some tissue coccidia: Cystoisospora species that infect dogs (A, Cystoisospora canis; B, Cystoisospora ohioensis); Cystoisospora species that infect cats (D, Cystoisospora felis; F, unsporulated Cystoisospora rivolta); Cystoisospora suis, infecting pigs (E); and Hammondia heydorni, a parasite of dogs (C). All photographs are at the same scale (scale bar represents 10 µm).

Oocyst and sporocyst dimensions for some parasites obtained from fecal samples.
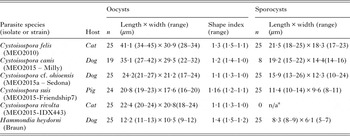
a n/a – sporulated oocysts were not available for this species.
Sequences obtained
Pairs of degenerate coccidia- and Sarcocystidae-specific primers designed to amplify a portion of the mt COI gene (see Table 1) produced PCR products that ranged from 418 to 848 bp in length (excluding primer regions); translations of all sequences indicated that no indels existed within the region amplified by any of these primers from the sarcocystid parasites. New sequences from nuclear 18S rDNA ranged from 573 bp (partial sequence from C. rivolta) to near full-length sequences of ~1800 bp. A total of 31 new COI sequences and 17 new 18S rDNA sequences were generated from 11 different species (see Supplementary Table 1 for the GenBank accession numbers for all sequences used in this study). Among the Sarcocystidae, no shared indels were found within 18S rDNA sequences from parasites in the Toxoplasmatinae; in contrast, Sarcocystis spp. has numerous (at least seven) expanded regions with no homology with members of the Toxoplasmatinae.
Phylogenetic analyses based on COI and 18S rDNA sequences
All phylogenetic analyses using BI, ML or MP methods with COI, 18S rDNA or concatenated sequences produced topologically similar trees that supported the monophyly of the family Sarcocystidae (BI trees illustrated in Figs 2–5).
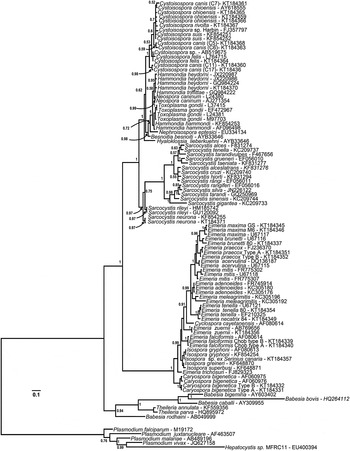
Phylogeny of eimeriid coccidia (Eimeriidae and close relatives) and isosporoid coccidia (Sarcocystidae) based on the 18S rDNA dataset generated by BI using the HKY (nst = 2 gamma categories) model of nucleotide substitution with a discrete gamma distribution rate variation among sites (G) and accounting for proportion of invariant sites (I) estimated base frequencies. The coccidia were rooted using a number of haemosporinid and piroplasmid parasites. The monophyly of the coccidia (eimeriid and isosporoid coccidia) was strongly supported. The 18S rDNA dataset supported monophyly of the eimeriid coccidia as well as the family Sarcocystidae and its subfamilies Toxoplasmatinae (PP = 1·00) and Sarcocystinae (PP = 0·97).
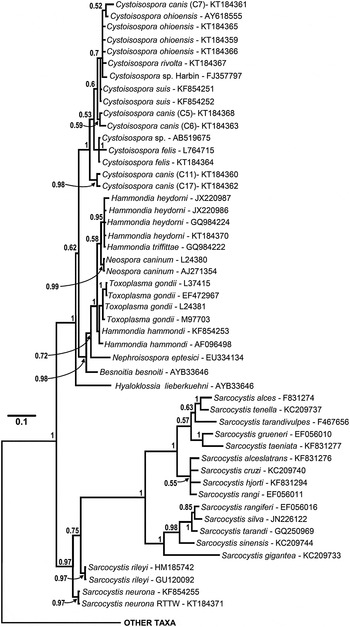
Enlarged portion of the 18S rDNA sequence based tree (see Fig. 2 for complete tree) illustrating relationships among isosporoid coccidia. Hyaloklossia lieberkuehni branched basally to two, well supported clades that contained all remaining members of the Toxoplasmatinae (all Cystoisospora spp. in one; all remaining taxa in the second). Monophyly of individual Cystoisospora species within the first clade was supported rarely; as an example, a single polytomy contained four Cystoisospora spp. from four different host species. In the second clade, Besnoitia besnoiti and Nephroisospora eptesici branched basally to a clade of poorly resolved taxa belonging to the genera Hammondia, Toxoplasma and Neospora. Although monophyly of N. caninum and T. gondii was each supported (PP = 0·99 and 0·71, respectively), sequences from individual Hammondia spp. did not form monophyletic clades and the genus Hammondia itself was paraphyletic as well.
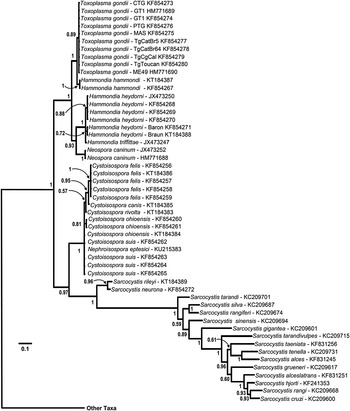
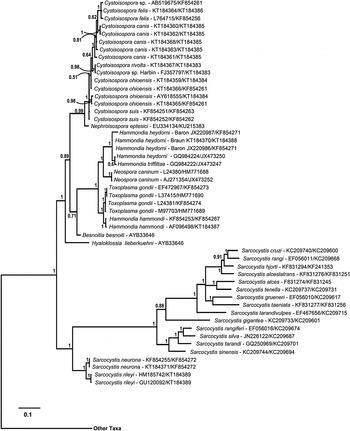
Phylogeny of isosporoid coccidia based on the COI dataset using the same outgroups as in Fig. 2; only the clade containing members of the family Sarcocystidae are illustrated with the branch leading to the other taxa indicated. Trees were generated using BI and the general time reversible model with discrete gamma distribution including invariable site (GTR + I + G, nst = 6 gamma categories) using a codon nucleotide model (i.e. Nucmodel = Codon) implemented using metazoan mitochondrial translation (i.e. Code = metmt). Three major clades were supported within a monophyletic family Sarcocystidae: (1) a clade of Sarcocystis spp.; (2) Cystoisospora spp. plus Nephroisospora eptesici; and (3) a clade of Neospora, Toxoplasma and Hammondia species. Monophyly of the subfamily Toxoplasmatinae was not supported in this tree. Hammondia species do not form a monophyletic group; Hammondia species using canids as their definitive hosts grouped with N. caninum, whereas Hammondia spp. using felids as their definitive hosts grouped with T. gondii.
Phylogeny of isosporoid coccidia based on the concatenated dataset (18S rDNA and COI sequences) using the same outgroups as in Fig. 2. The concatenated dataset was partitioned with the COI portion of the dataset analysed using a GTR + I + G (nst = 6, nucmodel = codon, code = metmt) substitution model and the 18S rDNA portion analysed using the HKY + I + G (nst = 2, nucmodel = 4by4) substitution model. Only the clade containing members of the family Sarcocystidae are illustrated with the branch leading to the other taxa indicated. Monophyly of the Sarcocystidae and its subfamilies Sarcocystinae and Toxoplasmatinae was supported strongly (all PP = 1·00). Besnoitia besnoiti and Hyaloklossia lieberkuehni branched near the base of the Toxoplasmatinae. Nephroisospora eptesici formed a well-supported sister clade to the Cystoisospora spp. Individual species within the clade containing Toxoplasma, Neospora and Hammondia spp. were well supported with the exception of Hammondia spp. infecting canids. As in the tree based on the COI dataset alone, Hammondia species using dogs as their definitive host grouped with N. caninum whereas Hammondia spp. using cats as their definitive host grouped with T. gondii.
The ML analysis on the COI dataset (756 nucleotides) was performed with a gamma distribution shape parameter of 1·2495, a rate category of 4, and proportion of invariable sites (I) of 0·2621; the number of distinct patterns under this model was 561. The MP analysis on the COI dataset generated at tree length of 3641 with a consistency index (CI) of 0·3087 based on 501 parsimony-informative characters.
For the nuclear 18S rDNA dataset (3278 nucleotides), the ML analysis was performed with a gamma shape parameter was 0·2769, and a rate category of 4. The proportion of invariable sites (I) was 0·1791 and the number of distinct data patterns under this model was 1580. The MP analysis on the 18S rDNA dataset generated a tree length of 3100 based on 904 parsimony-informative characters with a CI of 0·7200.
For the concatenated sequence dataset (4061 nucleotides), the ML analysis had 2140 distinct data patterns under the selected nucleotide modes with the proportion of invariable sites (I) equal to 0·2041, the number of rate categories equal to 4 and gamma distribution shape parameter equal to 0·4366. Tree length in the MP analysis based on 1408 parsimony-informative characters was 6766 with CI 0·4957.
In all analyses, three clades were consistently supported within the family Sarcocystidae: (1) a clade containing all Sarcocystis spp. (Sarcocystinae) with PP ≥ 0·97 and bootstrap support of ≥99%; (2) a clade containing all Cystoisospora spp. (including Nephroisospora eptesici in the COI-based tree only) with PP ≥ 0·99 and bootstrap support of ≥99%; and (3) a clade containing all of the included parasites in the genera Hammondia, Neospora and Toxoplasma.
Besnoitia besnoiti and Hyaloklossia lieberkuehni were determined to branch near the base of the Toxoplasmatinae in the 18S rDNA and concatenated dataset trees but there were no COI sequences available for these parasites. The position of N. eptesici varied depending on the genetic loci. In the tree generated from the COI dataset, the N. eptesici clustered within a clade with all Cystoisospora spp. In the tree generated from the 18S rDNA dataset, N. eptesici was found in the same clade with Toxoplasma/Hammondia/Neospora along with B. besnoiti; H. lieberkuehni branched earliest amongst all taxa in the Toxoplasmatinae. However, in the concatenated sequence dataset, N. eptesici formed a well-supported sister clade to the Cystoisospora spp.
In the analyses based on 18S rDNA and concatenated datasets (Figs 3 and 5, respectively) monophyly of both the subfamilies Toxoplasmatinae and Sarcocystidae was well-supported. However, monophyly of the Toxoplasmatinae was not supported in trees generated from COI sequences alone (Fig. 4) because the rooting position within the family Sarcocystidae was between Cystoisospora/Nephroisospora clade and the remaining members of the Toxoplasmatinae rather than between the Sarcocystis spp. and other members of the Sarcocystidae as was found in trees based on the other datasets (Figs 2, 3 and 5).
The relationships among closely related species within the Toxoplasmatinae were not resolved fully, particularly in trees based on the 18S rDNA (Figs 2 and 3) or concatenated datasets (Fig. 5). For example, trees based on the 18S rDNA dataset (Figs 2 and 3) did not support monophyly of Hammondia spp.; 18S rDNA sequences from Hammondia spp. formed a polytomy with those of T. gondii and N. caninum. Similarly, monophyly of individual Cystoisospora spp. was not supported with the exception of the C. felis sequences. The remaining 18S sequences from Cystoisospora spp. formed a poorly resolved, polychotomous multi-species cluster in which sequences from individual species did not form monophyletic clades. Five C. canis cloned 18S rDNA sequences failed to resolve into a monophyletic group in the phylogenetic analyses based on this locus. Based on pairwise sequence comparisons of alignments of these five sequences, the clones clustered into at least two distinct sets of paralogous 18S rDNA sequences. One pair of sequences (‘Type A’ – clones 11 and 17) had pairwise identity of 99·8% and clustered into a well-supported (PP = 0·98) monophyletic clade. The second set of three sequences (‘Type B’ – clones 5–7) had mean pairwise identity of 99·6% but did not resolve into a monophyletic clade in the 18S rDNA analyses.
In the trees based on COI sequences (Fig. 4), monophyletic clades of sequences corresponding to recognized species within the Toxoplasmatinae were supported in most cases. However, sequences from C. suis, C. cf. ohioensis and N. eptesici were found in an unresolved polychotomy basal to the remaining Cystoisospora species. The Cystoisospora spp. that possess large, egg-shaped oocysts (i.e. C. felis and C. canis) formed a well-supported monophyletic clade (PP = 0·95). Similarly, species of typically heteroxenous parasites infecting dogs and cats (species in the genera Toxoplasma, Neospora and Hammondia) were all found in a well-supported monophyletic clade in trees based on COI sequences (Fig. 4). However, species in the genus Hammondia remained paraphyletic. The canid-infecting Hammondia spp. [i.e. H. triffittae, H. heydorni and an unnamed Hammondia sp. from North America (i.e. strains ‘Baron’ and ‘Braun’)] were each monophyletic and formed a sister clade to N. caninum whereas the felid-infecting H. hammondi was sister to T. gondii.
In the concatenated dataset analyses, sequences from canid-infecting Hammondia species were not resolved into monophyletic clades representing named species although they all formed a well-supported monophyletic clade (Fig. 5). Again, as observed in the COI sequence-based analyses (Fig. 4), parasites using canids as definitive hosts (DHs, i.e. H. heydorni, H. triffitae and N. caninum) and parasites using felids as DHs (i.e. H. hammondi and T. gondii) each formed monophyletic clades; but the monophyly of Hammondia spp. was not supported.
DISCUSSION
Monophyly of genera in the Toxoplasmatinae remains uncertain despite numerous phylogenetic analyses based on nuclear rDNA sequences and other loci (e.g. Mugridge et al. Reference Mugridge, Morrison, Jakel, Heckeroth, Tenter and Johnson2000; Morrison et al. Reference Morrison, Bornstein, Thebo, Wernery, Kinne and Mattsson2004; Morrison, Reference Morrison2009). In the present study, Toxoplasmatinae-specific PCR primers were used to obtain partial mt COI sequences with the goal of resolving evolutionary relationships among these tissue coccidia that could not be resolved using nuclear 18S rDNA sequences (e.g. Morrison, Reference Morrison2009). Culture-derived parasites of known identity as well as parasites obtained from naturally infected hosts were used to obtain mt COI and nuclear 18S rDNA sequences (if such sequences were unavailable in public databases).
Nuclear 18S rDNA possesses numerous indels (most commonly within the single-stranded loops between the helices) and some hypervariable regions (e.g. the E21–1, E21–3 and E21–5 helices in the Sarcocystinae) that can make alignments problematic for these parasites (Morrison et al. Reference Morrison, Bornstein, Thebo, Wernery, Kinne and Mattsson2004). Morrison et al. (Reference Morrison, Bornstein, Thebo, Wernery, Kinne and Mattsson2004) concluded that nuclear 18S rRNA gene sequences provide sufficient phylogenetic signal for deep relationships but have insufficient signal to discern species-level relationships within the Eimeriidae and Sarcocystidae. Deleting the ‘problematic’ regions from phylogenetic analysis may compromise the phylogenetic support for the various taxonomic groups (Morrison et al. Reference Morrison, Bornstein, Thebo, Wernery, Kinne and Mattsson2004). Barta (Reference Barta1997) suggested a staggered alignment approach as a solution for retaining hypervariable regions while still maintaining positional homologies among those taxa for which positional homology can be reasonably assumed; this staggered alignment approach was used in the present work to maximize the information content of the 18S rDNA sequences in both the single locus and concatenated datasets. Even when such strategies are employed, 18S rDNA sequences from a single species (e.g. Cystoisospora canis in the present study) can be highly divergent. It is possible that the two paraphyletic clades C. canis identified using 18S rDNA sequences represent paralogous rDNA loci within the nuclear genome of this parasite as demonstrated for eimeriid coccidia previously (e.g. El-Sherry et al. Reference El-Sherry, Ogedengbe, Hafeez and Barta2013).
Hammondia species were paraphyletic in all analyses containing COI sequences and paraphyly was not refuted by the 18S rDNA dataset (see also Morrison et al. Reference Morrison, Bornstein, Thebo, Wernery, Kinne and Mattsson2004). Phylogenetic analyses based on LSU rDNA and ITS1 sequence data (Ellis et al. Reference Ellis, Morrison, Liddell, Jeckins, Mohammed, Ryce and Dubey1999), similarities in G + C content of DNA sequences (Johnson et al. Reference Johnson, Illana, Dubey and Dame1987), as well as oocyst structure and antigenic responses (Dubey et al. Reference Dubey, Barr, Barta, Bjerkas, Björkman, Blagburn, Bowman, Buxton, Ellis, Gottstein, Hemphill, Hill, Howek, Jenkins, Kobayashi, Koudela, Marsh, Mattsson, McAllister, Modry', Omata, Sibley, Speer, Trees, Uggla, Upton, Williams and Lindsay2002b ) have consistently indicated that species in the genus Hammondia do not form a monophyletic group. We confirmed the lack of monophyly of species currently placed in the genus Hammondia as reported previously (Johnson et al. Reference Johnson, Illana, Dubey and Dame1987; Ellis et al. Reference Ellis, Morrison, Liddell, Jeckins, Mohammed, Ryce and Dubey1999; Dubey et al. Reference Dubey, Barr, Barta, Bjerkas, Björkman, Blagburn, Bowman, Buxton, Ellis, Gottstein, Hemphill, Hill, Howek, Jenkins, Kobayashi, Koudela, Marsh, Mattsson, McAllister, Modry', Omata, Sibley, Speer, Trees, Uggla, Upton, Williams and Lindsay2002b ) using a mitochondrial genetic marker (COI) alone or combined with nuclear 18S rDNA sequences.
In the present analyses, Hammondia spp. formed a polytomy with T. gondii and N. caninum based on 18S rDNA sequences. All canid-infecting Hammondia species formed a monophyletic clade that was the sister group to N. caninum as reported by Ellis et al. (Reference Ellis, Morrison, Liddell, Jeckins, Mohammed, Ryce and Dubey1999). Sequences obtained from a canine isolate of a Hammondia spp. from central Canada were nearly identical with those of H. triffittae at both the COI and 18S rDNA loci suggesting that the range of H. triffittae may include the entire temperate north. Although closely related, H. heydorni exhibits genetic and biological differences from H. triffittae (Schares et al. Reference Schares, Heydorn, Cüppers, Mehlhorn, Geue, Peters and Conraths2002, Reference Schares, Meyer, Bärwald, Conraths, Riebe, Bohne, Rohn and Peters2003; Abel et al. Reference Abel, Schares, Orzeszko, Gasser and Ellis2006; Gjerde and Dahlgren, Reference Gjerde and Dahlgren2011).
Hammondia hammondi clustered with T. gondii in all analyses. Partial mt COI sequences from all nine strains of T. gondii were identical. The mt COI sequence from the H.H-20 strain of H. hammondi shared 98·5% identity to the T. gondii sequences. The 1·5% divergence between these two species is identical to the divergence between C. felis and C. rivolta (1·5%) and similar to divergence between two Isospora spp. infecting the same avian host (1·3%, Hafeez et al. Reference Hafeez, Stasiak, Delnatte, El-Sherry, Smith and Barta2014) or between the chicken parasites E. tenella and E. necatrix (1·7%, Ogedengbe et al. Reference Ogedengbe, Hanner and Barta2011). The genetic divergence supports species-level separation of H. hammondi from T. gondii in contrast to Mehlhorn and Heydorn's (Reference Mehlhorn and Heydorn2000) and Heydorn and Mehlhorn's (Reference Heydorn and Mehlhorn2001) arguments against the recognition of H. hammondi as a distinct species. These latter authors had suggested that T. gondii and H. hammondi may be strains of a single species based on oocyst wall structure, tachyzoite ultrastructure and molecular similarities (Mehlhorn and Heydorn, Reference Mehlhorn and Heydorn2000) and cross-immunity (Frenkel and Dubey, Reference Frenkel and Dubey2000). Although partial COI sequences appeared well suited for species delimitation (Ogedengbe et al. Reference Ogedengbe, Hanner and Barta2011) and inferring phylogenetic relationships among the eimeriid coccidia (Ogedengbe et al. Reference Ogedengbe, Hanner and Barta2011 and current study) as well as members of the Hammondia/Neospora/Toxoplasma clade, the COI sequences from Cystoisospora spp. demonstrated remarkably limited sequence variation. For example, the partial COI sequence of C. cf. ohioensis (KT184384) differed from the COI sequences of C. suis (KF854262–KF854265) and C. rivolta (KT184383) by 1 single nucleotide difference (SND), while differing from a second C. cf. ohioensis (KT184365) isolate by 3 SNDs. It is possible that the pairwise differences in the COI sequences of the 2 C. cf. ohioensis isolates is detection of species level differences among parasites in the C. ohioensis spp. complex. Only isolation of a pure line of each parasite and observations on the endogenous development of these parasites would be able to assign these COI sequences unequivocally to C. ohioensis, C. burrowsi or C. neorivolta.
Phylogenetic trees based on COI or concatenated datasets strongly support H. hammondi as a sister species to T gondii; the support for the close relationship of the other Hammondia spp. to N. caninum is equally strong. The COI– and the concatenated sequence-based trees show that relationships among species within the Hammondia/Neospora/Toxoplasma clade reflect the DHs infected by these parasites.
Mehlhorn and Heydorn (Reference Mehlhorn and Heydorn2000) proposed a radically simplified taxonomic scheme for parasites in the Toxoplasmatinae. They suggested that T. gondii and H. hammondi should be synonymized under a single species, T. gondii. The two described Neospora species, N. caninum and N. hughesi, were to be synonymized under the single species H. heydorni, and then H. heydorni was to be transferred to the genus Toxoplasma to give T. heydorni as the name for these parasites of dogs. Synonymizing N. caninum, N. hughesi and H. heydorni under T. heydorni by Mehlhorn and Heydorn (Reference Mehlhorn and Heydorn2000) would appear to be a ‘taxonomic lumping’ too far. Would this taxonomic arrangement reflect both biological features and COI genotypes? Synonymizing T. gondii and H. hammondi (and, presumably, Hammondia pardalis Hendricks, Ernst, Courtney and Speer, 1979 that also infects a felid DH) under T. gondii may reflect their biological similarities but does not reflect the COI genotypes of these distinct species because of the 10 SNDs between the H.H-20 strain of H. hammondi and the various strains of T. gondii over 751 bp of the COI gene (1·3% genetic difference); to put this in perspective, C. felis and C. canis have only 0·9% sequence divergence (6 SNDs) in the same region of the COI gene and they do not share the same DH.
To re-establish monophyly of species currently recognized to belong to the genus Hammondia, taxonomic revision will be necessary. The closely related tissue coccidian currently classified in the genera Toxoplasma, Neospora and Hammondia have biological and genetic affinities that directly conflict (Dubey, Reference Dubey and Kreier1977; Carreno et al. Reference Carreno, Schnitzler, Jeffries, Tenter, Johnson and Barta1998; Ellis et al. Reference Ellis, Morrison, Liddell, Jeckins, Mohammed, Ryce and Dubey1999; Mehlhorn and Heydorn, Reference Mehlhorn and Heydorn2000; Dubey et al. Reference Dubey, Hill, Lindsay, Jenkins, Uggla and Speer2002a ; Dubey and Sreekumar, Reference Dubey and Sreekumar2003; Morrison, Reference Morrison2009). For the tissue coccidia infecting felines as DHs, T. gondii (facultatively heteroxenous exhibiting horizontal transmission among intermediate hosts) is closely related genetically to H. hammondi (and presumably H. pardalis; both obligately heteroxenous, lacking horizontal transmission among intermediate hosts). However, H. hammondi and H. pardalis are more similar biologically to canid-infecting Hammondia species, namely H. heydorni and H. triffittae, both of which are obligately heteroxenous and lack horizontal transmission among intermediate hosts. Genetically, the latter two Hammondia species are related more closely to Neospora species (facultatively heteroxenous, horizontal transmission among intermediate hosts).
Transferring H. hammondi and H. pardalis into the genus Toxoplasma would appear to be a taxonomic change that reflects molecular features of these parasites but would group a facultatively monoxenous parasite (T. gondii) with an obligately heteroxenous parasite (H. hammondi). By the same logic, the Hammondia species infecting canine DHs would then be most suitably placed into a single genus with Neospora species. The only remaining biological distinction between the two genera that remain, Toxoplasma and Hammondia, would be their use of feline vs canine DHs; clearly this is not a distinguishing feature that warrants separation at the level of genus (c.f. Sarcocystis species). So there are only two choices for revising generic assignments of species in the genera Toxoplasma, Hammondia and Neospora: (1) re-assign all of these species to the genus Toxoplasma and redefine a broader definition for this genus; or (2) retain all three genera but, to maintain monophyly of species within each genus, name a new genus for the obligately heteroxenous coccidia closely related genetically to Neospora spp. (i.e. described Hammondia species that utilize canines as DHs).
The lack of comparative molecular data from representatives of all currently recognized genera in the Toxoplasmatinae (i.e. Besnoitia species are not represented) makes the lumping of the parasites examined in this study into single genus premature. To maintain taxonomic stability and to reflect the common usage of many of these genus names in the literature, the most conservative approach is to erect a fourth genus to address the paraphyly of Hammondia species as confirmed in the present study. We therefore propose the erection of a new genus to contain those Hammondia species that utilize canines as DHs.
TAXONOMIC SUMMARY
Apicomplexa Levine, 1980
Conoidasida Levine, 1988
Coccidiasina Leuckart, 1879
Eucoccidiorida Léger, 1911
Eimeriorina Léger & Duboscq, 1911
Sarcocystidae Poche, 1913
Toxoplasmatinae Biocca, 1957
Heydornia n. gen.
DEFINITION: With features of the family Sarcocystidae (heteroxenous or facultatively heteroxenous coccidia; oocysts with two sporocysts lacking Stieda bodies, each with four sporozoites; tissue cysts polyzoic; in vertebrates) and subfamily Toxoplasmatinae (metrocytes not formed in tissue cysts; oocysts sporulate exogenously). Members of the genus Heydornia n. gen. are obligately heteroxenous coccidia of vertebrates using canid DHs (where known); lack horizontal transmission among intermediate hosts; about two named species.
NAME-BEARING TYPE:
Heydornia heydorni nov. comb. (Tadros and Laarman, Reference Tadros and Laarman1976)
Synonym: Isospora bigemina (Stiles, 1891) Luhe, 1906 ‘small form’ of various authors, pro parte
Synonym: Isospora bigemina of Dubey and Fayer, Reference Dubey and Fayer1976
Synonym: Isospora wallacei Dubey, 1976
ETYMOLOGY: The genus is named in honour of Professor Dr Alfred Otto Heydorn (Institute for Parasitology and Tropical Veterinary Medicine, Free University Berlin) who worked extensively on this ‘small form’ of I. bigemina in canids.
TYPE HOST: Canis lupus familiaris
Oocysts: Dubey and Fayer (Reference Dubey and Fayer1976) reported oocysts as spherical, 12 × 11 µm. Nassar et al. (Reference Nassar, Hilali and Rommel1983) gives 11·9 × 11·1 (10·0–14·5 × 9·3–13·1) and Shankar et al. (Reference Shankar, Bhatia and Saleque1991) 11·0–14·0 × 10·5–13·0 µm.
REMARKS: This coccidian is obligatorily heteroxenous and uses a wide range of intermediate hosts, including ruminants, cervids and rodents, in which muscle tissue cysts are formed. Commonly referred to as the ‘small form’ of I. bigemina (Stiles, 1891) Lühe, 1906 in the older literature. Oocysts are passed unsporulated. Members of the genus Heydornia n. gen. can be distinguished from Neospora species because the former lack horizontal transmission among intermediate hosts and their sporulated oocysts are not infective to their DHs.
OTHER NAMED SPECIES:
Heydornia triffittae nov. comb. (Nukerbaeva & Svanbaev, 1973)
Synonyms:
-
Isospora triffitti Nukerbaeva & Svanbaev, 1973
-
Isospora triffittae (Nukerbaeva & Svanbaev, 1973) Levine 1985
-
Hammondia triffittae (Nukerbaeva & Svanbaev, 1973) Gjerde & Dahlgren, 2011
TYPE HOSTS: Red fox (Vulpes vulpes) and arctic fox (Vulpes lagopus).
REMARKS: See Gjerde and Dahlgren (Reference Gjerde and Dahlgren2011) for a comprehensive description of this species and its differentiation from Heydornia heydorni n. comb.
Oocysts: Gjerde and Dahlgren (Reference Gjerde and Dahlgren2011) reported oocysts as 12·5 ± 0·7 × 10·9 ± 0·5 µm (10·3–15·2 × 9·3–12·8 µm, n = 900) with L: W ratio 1·15 ± 0·07 (1·00–1·49, n = 900)
In summary, Heydornia n. gen. is proposed for Hammondia species infecting canid DHs to resolve the well documented paraphyly of coccidia in the genus; the name-bearing type for the genus Hammondia, H. hammondi and H. pardalis both infect felid DHs and remain in the genus. The use of both nuclear and mitochondrial genetic loci was demonstrated to provide good resolution of most species in the Sarcocystidae; inclusion of additional, biologically diverse members of the subfamily Toxoplasmatinae that were not included in the current COI dataset, such as Besnoitia or Hyaloklossia spp., might help stabilise the rooting of the phylogenetic hypothesis for these parasites. Addition of some ‘primitive’ eimeriid coccidia, such as Goussia or Choleoeimeria spp., to the taxonomic outgroup might also promote stability of the rooting point within the Sarcocystidae for this genetic locus. The observations reported herein support the use of combined nu 18S rDNA and mt COI sequences for generating phylogenetic hypotheses that are likely to demonstrate stability of deeper nodes combined with fine resolution of terminal taxa.
SUPPLEMENTARY MATERIAL
To view supplementary material for this article, please visit http://dx.doi.org/10.1017/pao.2015.7
ACKNOWLEDGEMENTS
We thank Dr Donald Martin, Head of Parasitology, IDEXX Laboratories, Markham, Ontario for supplying the IDX Cystoisospora isolates. Dr Ben Rosenthal (USDA, Beltsville, MD, USA) is thanked for supplying Sarcocystis sp. DNA. Dr Scott Weese (Department of Pathobiology, Ontario Veterinary College, University of Guelph, Guelph, ON, Canada) is thanked for providing C. cf. ohioensis isolates. Dr Chunlei Su (Department of Microbiology, College of Arts and Sciences, University of Tennessee, Knoxville, TN, USA) is thanked for providing Toxoplasma gondii strains. This paper was greatly improved by thoughtful comments from anonymous reviewers.
FINANCIAL SUPPORT
M.E.O. was supported by a Ph.D. Scholarship from the Ontario Veterinary College (OVC), University of Guelph. K.E. was supported by a Pfizer Summer Student Research Scholarship held at the OVC. The work reported herein was supported by research grants to J.R.B. from the Natural Sciences and Engineering Research Council of Canada (NSERC Discovery Grant 400566), and the Ontario Ministry of Agriculture, Food and Rural Affairs (OMAFRA No. 200331).
CONFLICT OF INTEREST
None.
ETHICAL STANDARDS
The work is in no violation of ethical standards. Human or laboratory animals were not used in this work.








