Introduction
Cereal aphids are important insect pests of many grasses and cereals, including wheat and barley (Leybourne et al., Reference Leybourne, Storer, Marshall, Musa, Telling, Abel, White, Ellis, Yang and Berry2024b; Van Emden and Harrington, Reference Van Emden and Harrington2007). Cereal aphids are widely distributed across Europe and cause significant damage to cereal crops through direct feeding (Dedryver et al., Reference Dedryver, Le Ralec and Fabre2010) and the transmission of plant viruses, including barley yellow dwarf virus and cereal yellow dwarf virus (Perry et al., Reference Perry, Kolb, Sammons, Lawson, Cisar and Ohm2000). In Europe, the two cereal aphid species of greatest concern are the grain aphid (Sitobion avenae Fabricius) and the bird cherry-oat aphid (Rhopalosiphum padi Linnaeus). During the main period of crop growth (spring–winter), aphid populations reproduce through asexual parthenogenesis, leading to the establishment of clonal populations. This method of reproduction facilitates rapid growth of aphid populations and heightens the likelihood that aphids will migrate throughout the crop and between fields, increasing the risk of virus infection and spread (Fabre et al., Reference Fabre, Dedryver, Leterrier and Plantegenest2003). When present in high numbers, direct feeding damage can reduce cereal yields by 5% (Dedryver et al., Reference Dedryver, Le Ralec and Fabre2010) and severe levels of virus infection can decrease yields by 20–80% (Perry et al., Reference Perry, Kolb, Sammons, Lawson, Cisar and Ohm2000; Zhao and Zhou, Reference Zhao and Zhou2025). Often, cereal fields are dominated by a small number of prolific clonal populations and understanding the processes behind the dominance of these populations can help with pest and disease management.
Several studies have described variable phenotypes between different aphid clonal lineages (Figueroa et al., Reference Figueroa, Simon, Le Gallic, Prunier-Leterme, Briones, Dedryver and Niemeyer2004; Gao and Liu, Reference Gao and Liu2013; Guo et al., Reference Guo, Moreau and Lapierre1996; Simon et al., Reference Simon, Dedryver, Pierre, Tanguy and Wegorek1991; Wang et al., Reference Wang, Zhao, Bai, Shang, Zhang, Hou, Chen and Han2021), including phenotypic traits of agricultural importance (e.g. virus transmission capacity, development time, reproduction, insecticide resistance, response to environmental stress). These distinct phenotypes can influence the agricultural risk presented by the aphid population. For example, aphid populations with faster development times will have shorter periods where they are susceptible to parasitoid biocontrol, populations with a higher reproductive rate carry a greater risk of causing significant feeding damage, and populations that produce more alate (winged) aphids have a greater virus dispersal risk. These life-history and fitness traits can be associated with heritable factors within the aphid population: genetic diversity in the aphid population and the presence of endosymbionts (Figueroa et al., Reference Figueroa, Simon, Le Gallic, Prunier-Leterme, Briones, Dedryver and Niemeyer2004; Gonzalez-Gonzalez et al., Reference Gonzalez-Gonzalez, Cabrera, Rubio-Meléndez, Sepúlveda, Ceballos, Fernández, Francis, Figueroa and Ramirez2024). Being able to associate these heritable factors with key phenotypes can generate biological insights into the processes underpinning the dominance of specific aphid clones and can be used to develop pest and disease management strategies that are based on the phenotypic risk of the aphid populations present.
Genetic variation is a key driver of phenotype in many aphid species (Figueroa et al., Reference Figueroa, Simon, Le Gallic, Prunier-Leterme, Briones, Dedryver and Niemeyer2004; Leybourne et al., Reference Leybourne, Bos, Valentine and Karley2020a; Martinez-Chavez et al., Reference Martinez-Chavez, Roberts, Karley, Shaw and Pope2024). In Europe, cereal aphid populations are genetically diverse, can be grouped into genetically similar populations, and genotypic variation can influence aphid phenotype (Leybourne et al., Reference Leybourne, Bos, Valentine and Karley2020a; Łukasik et al., Reference Łukasik, Dawid, Ferrari and Godfray2013). For example, resistance to pyrethroids in S. avenae has been linked to mutations in the kdr gene and is associated with S. avenae genotype/clone sa3 (Foster et al., Reference Foster, Paul, Slater, Warren, Denholm, Field and Williamson2014). When compared with other S. avenae clones, the sa3 clone has increased vulnerability to parasitism (Jackson et al., Reference Jackson, Malloch, McNamara and Little2020). For R. padi, aphid genotype can influence mass gain in nymphs, adult reproduction (Leybourne et al., Reference Leybourne, Bos, Valentine and Karley2020a), and the transmission of barley yellow dwarf virus (Leybourne et al., Reference Leybourne, Whitehead and Will2024a).
Association with endosymbiotic bacteria is a second heritable trait that can influence aphid phenotype. The majority of aphid species form an essential relationship with the endosymbiont Buchnera aphidicola, with B. aphidicola providing nutritional supplementation to the aphid diet (Douglas and Prosser, Reference Douglas and Prosser1992). Aphids also form non-essential, or facultative, relationships with a range of endosymbionts (Guo et al., Reference Guo, Hatt, He, Chen, Francis and Wang2017; Zytynska and Weisser, Reference Zytynska and Weisser2016). Facultative endosymbiotic relationships occur naturally in cereal aphid populations, with endosymbiont prevalence varying for different aphid-endosymbiont combinations (Guo et al., Reference Guo, Liu, Poncelet, He, Francis and Wang2019; Leybourne et al., Reference Leybourne, Bos, Valentine and Karley2020a, Reference Leybourne, Melloh and Martin2023; Łukasik et al., Reference Łukasik, Dawid, Ferrari and Godfray2013; Zytynska et al., Reference Zytynska, Sturm, Hawes, Weisser and Karley2023). When present, these facultative endosymbionts substantially alter cereal aphid phenotype: One trait often conferred across a range of aphid-endosymbiont combinations is resistance against parasitoid wasps and fungal pathogens (Leybourne et al., Reference Leybourne, Bos, Valentine and Karley2020a; Luo et al., Reference Luo, Monticelli, Li, Ahmed, Pandharikar, Zhao, Desneux and Hu2020; Zytynska et al., Reference Zytynska, Tighiouart and Frago2021). In cereal aphids, the majority of studies characterising endosymbiont-associated phenotypes have focussed on S. avenae, with few studies describing endosymbiont-derived phenotypes for R. padi. For S. avenae, endosymbiont-associated phenotypes include: altered fecundity (Hamiltonella defensa (Łukasik et al., Reference Łukasik, Dawid, Ferrari and Godfray2013); Regiella insecticola (Ramírez-Cáceres et al., Reference Ramírez-Cáceres, Moya-Hernández, Quilodrán, Nespolo, Ceballos, Villagra and Ramírez2019)), phenotypic plasticity (R. insecticola (Wang et al., Reference Wang, Xiaoqin, Peng, Deguang, Xinjia, Zheming, Zhaohong and Xiuxiang2016)), greater population growth (Rickettsia spp. (Zhu et al., Reference Zhu, Li, Desneux, Gatti, Hu and Luo2024)), and a reduction in winged morph production (R. insecticola (Liu et al., Reference Liu, Lei and Chen2019)). However, these endosymbiont-conferred traits are often plastic and can be influenced in both direction and strength by additional external factors, including host plant suitability and quality (Gonzalez-Gonzalez et al., Reference Gonzalez-Gonzalez, Cabrera, Rubio-Meléndez, Sepúlveda, Ceballos, Fernández, Francis, Figueroa and Ramirez2024; Leybourne et al., Reference Leybourne, Valentine, Bos and Karley2020b; Ramírez-Cáceres et al., Reference Ramírez-Cáceres, Moya-Hernández, Quilodrán, Nespolo, Ceballos, Villagra and Ramírez2019).
Together, these heritable factors (genotype, endosymbiont association) influence a diverse range of cereal aphid phenotypes. However, there are significant gaps in our understanding of some of the processes that contribute towards the success of aphids in agriculturally important systems, including how intra-species variation in aphid populations underpins phenotypic traits of agricultural importance, this is particularly apparent for the understudied aphid species, R. padi.
Here, multiple populations for the two main cereal aphid species in Europe are used to determine how heritable factors influence aphid phenotype. Several agriculturally important phenotypes are examined (development time, reproductive output, winged morph production) and both endosymbiont- and genotype-derived phenotypes are identified. These results provide insight into the biological drivers behind phenotypic diversity in two agriculturally important aphid species.
Materials and methods
Aphid lines: characterisation of intra-specific factors
Aphid populations comprised 25 S. avenae populations and 8 R. padi populations. Aphids were collected from agricultural fields in Lower Saxony, Germany during summer and autumn 2021 (table 1). All aphid populations were maintained under controlled environment conditions (18 ± 2°C; 16:8 h L:D cycle) in a plant growth room. Populations were recently sequenced and assigned genotypes based on the clustering of populations into genetically similar clades (Leybourne et al., Reference Leybourne, Whitehead and Will2024a).
Intra-species characterisation for each aphid population
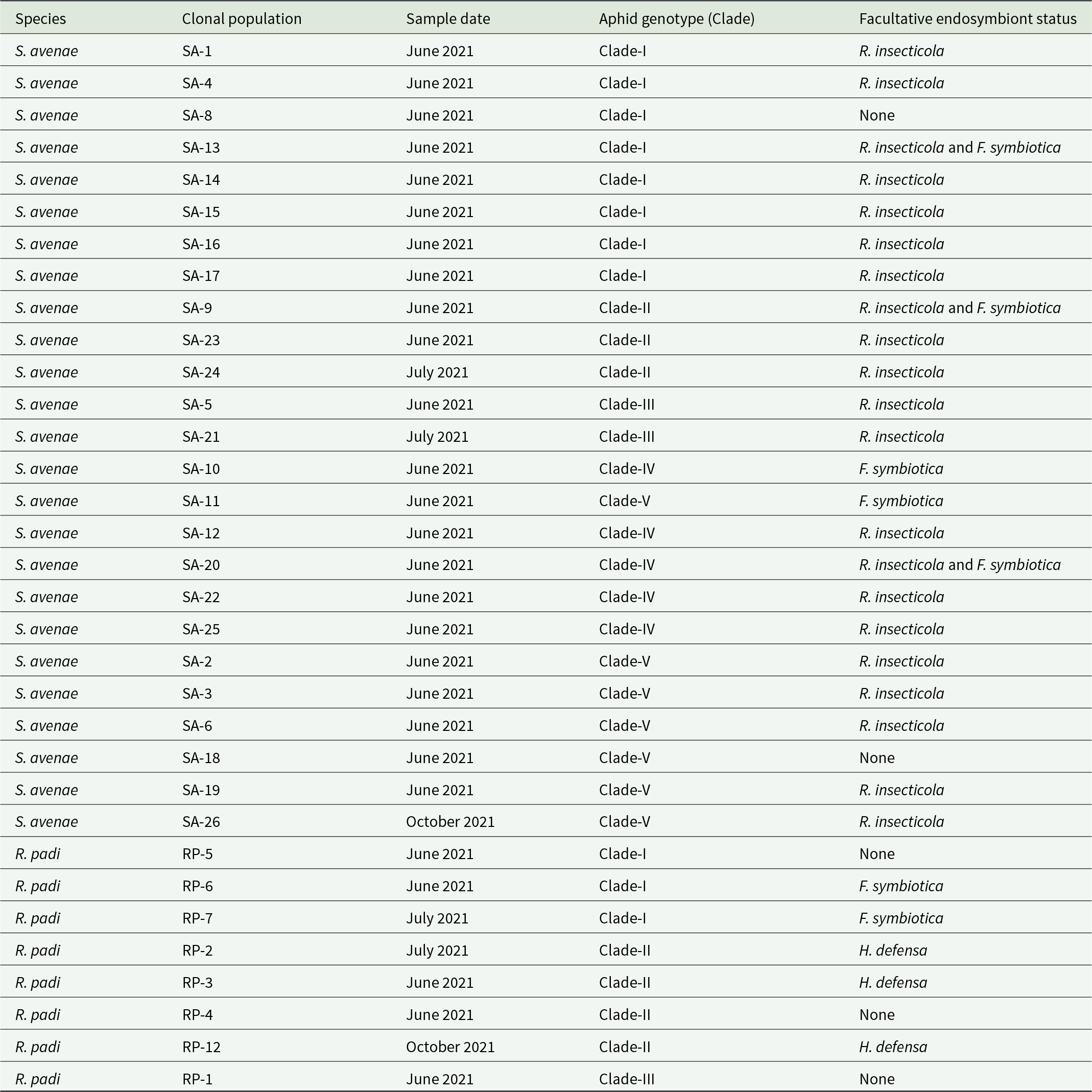
Endosymbiont communities were previously described for most aphid populations used in this study (Leybourne et al., Reference Leybourne, Melloh and Martin2023). For populations where facultative endosymbionts were not previously characterised, namely RP-12, the facultative endosymbiont profile was determined using the following approach: A sample of five aphids (mixture of adults and nymphs) were collected and DNA was extracted using the Norgen® Plant and Fungi DNA extraction kit (Norgen Biotek, Germany) following manufacturer’s instructions. Successful DNA extraction was confirmed using the Polymerase Chain Reaction (PCR) marker for the primary aphid symbiont Buchnera aphidocola. The presence of facultative endosymbionts was determined using a three-step multiplex diagnostic PCR assay (Beekman et al., Reference Beekman, Donner, Litjens, Dicke, Zwaan, Verhulst and Pannebakker2022; Manentzos et al., Reference Manentzos, Pahl, Melloh, Martin and Leybourne2024). Multiplex assays were used to detect the presence of the main secondary endosymbionts: Spiroplasma spp., Regiella insecticola, Hamiltonella defensa, Rickettsiella spp., Fukatsuia symbiotica, Seratia symbiotica, Rickettsia spp., and Arsenophonus spp. All PCR primer details are described in table S1. PCR assays were completed in a final reaction volume of 12 µL consisting of 2 µL DNA and 6 µL 2X Kappa2G Fast PCR Ready Mix (Merck, Germany). Primer concentrations and volumes differed between the multiplex assays and are detailed in table S1. The final reaction mixture was made to 12 µL using nuclease-free diethyl pyrocarbonate -treated water (CarlRoth, Germany). PCR conditions followed Beekman et al. (Reference Beekman, Donner, Litjens, Dicke, Zwaan, Verhulst and Pannebakker2022), i.e. denaturation at 94°C for 3 min followed by 35 cycles of 94°C for 30 s, 58°C for 30 s, and 72°C for 60 s with a final extension step at 72°C for 10 min. Positive DNA was included as a control, an extraction blank was used as an extraction negative control, and DNA-free PCR mastermix was included as PCR negative control. Endosymbiont presence was detected by separation of PCR products on a 1% agarose gel stained with GelRed® (Biotium, Germany), and reactions were visualised under UV light; a 100 bp DNA ladder (ThermoFischer, Germany) was used to estimate band size. Positive identification of the presence of endosymbionts in the multiplex assay was confirmed in additional singleplex assays. All PCR assays were completed in a Biometra TRIO 48, Thermocycler (Analytik Jena, Germany).
Age-synchronisation of test aphids
A cohort of age-synchronised aphids were produced for each population by placing three adult aphids onto the base of a young wheat (Triticum aestivum) plant, covering the plant with a microperforated bag to contain the aphids, and allowing the adult aphids to reproduce overnight. The following day all adults were removed, with all nymphs (<12 h old) left on the plant. Nymphs were retained on the plant and allowed to develop for 5 days, this was done to reduce mortality caused by handling recently larviposited aphids. Age-synchronisation was completed under controlled environment conditions (18 ± 2°C; 16:8 h L:D cycle) in a plant growth room. Three simultaneous age-synchronised cohorts were produced for each population.
Aphid life-history and population growth experiments
To investigate how intra-species diversity affects aphid phenotype a series of bioassays were completed. To manage the experiments, a split-block approach was followed. There were four experimental rounds for S. avenae, each containing up to four replicates per aphid population and two experimental rounds for R. padi, each containing up to ten replicates per aphid population; this split-block design was followed due to space limitations. However, due to aphid mortality, the final replication levels ranged from 7–15 (S. avenae) and 17–20 (R. padi). Each experimental run was arranged in a complete randomised block design. All experiments were completed under controlled environment conditions (18 ± 2°C; 16:8 h L:D cycle) in a plant growth room. New cohorts of age-synchronised aphids were produced for each experimental round and all experiments were complete within 9–15 months of the aphids being collected from the field. Aphid endosymbiont status was confirmed at the start of the life-history experiments.
For the experiments, winter wheat was sown into compost in individual plastic pots and left to grow to the seedling stage (GS10–11). Following this, a single age-synchronised aphid nymph (the ‘focal aphid’) was placed at the base of the plant stem and contained using microperforated bags. Focal aphids were monitored daily, and the date of reproductive maturity (observation of new nymphs) was noted. Once reproductive maturity was reached, the morph of the focal aphid was noted, these data were used to model aphid morph (binary variable) and to calculate the proportion of adult aphids comprising apterous or alate morphs for each population. Reproductive output was monitored across a time-period equivalent to the pre-reproductive period and this information was used to calculate the intrinsic rate of population increase (rm), following the equation of Wyatt and White (Reference Wyatt and White1977).
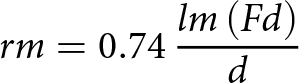 \begin{equation*}rm = 0.74\,\frac{{lm\left( {Fd} \right)}}{d}\end{equation*}
\begin{equation*}rm = 0.74\,\frac{{lm\left( {Fd} \right)}}{d}\end{equation*}Equation 1: The intrinsic rate of population increase (rm) equation of Wyatt and White (Reference Wyatt and White1977): d is the time period between aphid birth and production of first progeny, and Fd is the total progeny over a time period equal to d.
Statistical analysis
All statistical analysis was carried out using R (v.4.1.2) and R Studio (v.1.3.1093). The following additional packages were used to support data analysis and data visualisation: car v.3.0-11, ggplot2 v.3.3.5, ggpubr v.0.4.0, ggsci v.2.9.
In all models, explanatory variables included aphid genotype and facultative endosymbiont status (a factor comprising either ‘none’ or the species present), with facultative endosymbiont status tested as a nested variable within aphid genotype. A general linear model with binomial distribution and experimental block included as a fixed factor was used to test whether development into an apterous or alate aphid was influenced by aphid genotype or facultative endosymbiont status, with Type-II Χ 2 tests used to identify significant explanatory variables. Pre-reproductive period and rm were modelled as individual response variables in linear models, aphid morph (apterous or alate) and block were included as additional fixed factors. In these models, significant explanatory variables were identified using a Type II ANOVA. Tukey tests with honest significant differences were used to identify pairwise differences between aphid genotypes.
Results
Association with the facultative endosymbiont Regiella insecticola promotes alate production in S. avenae
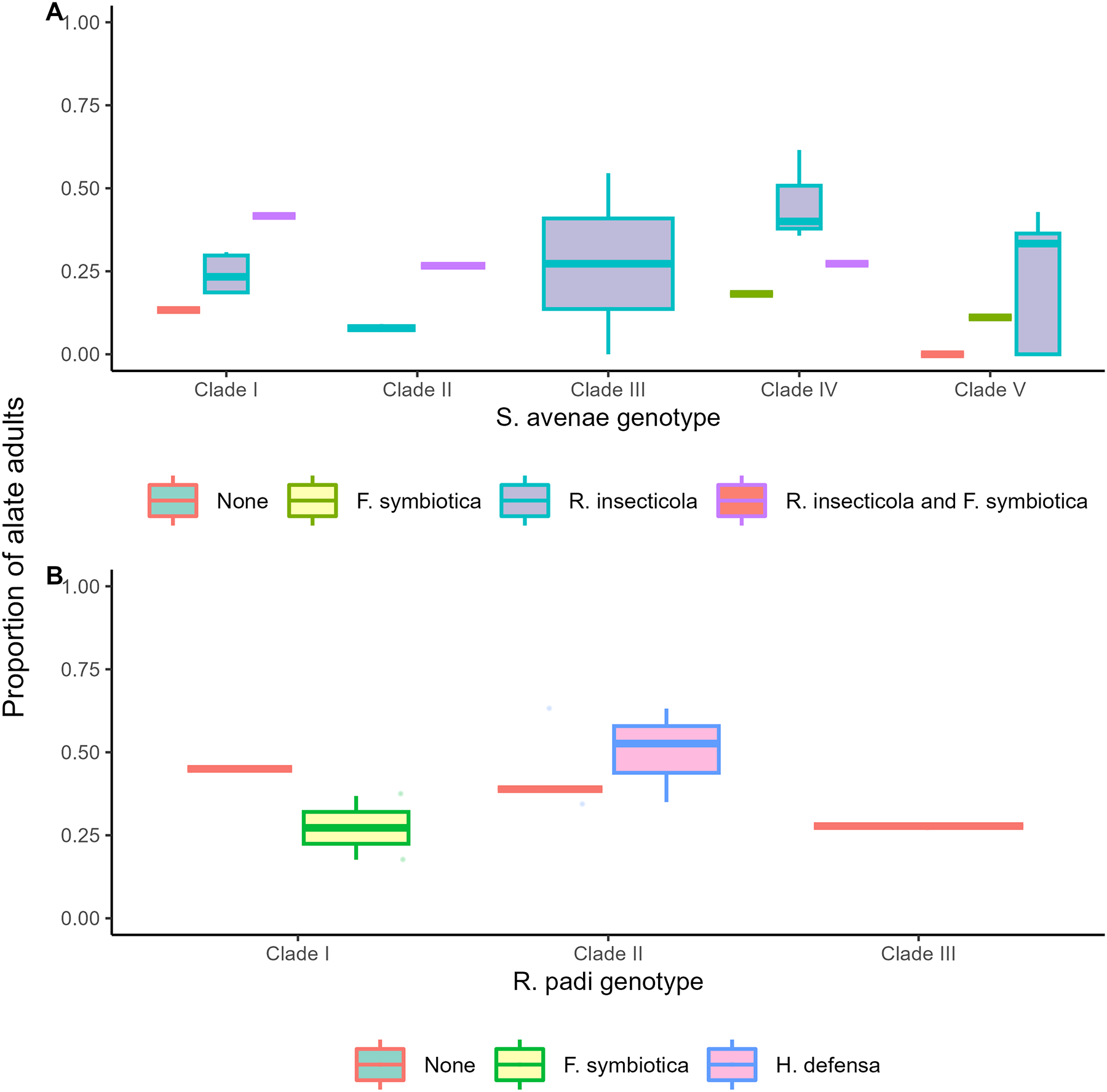
Facultative endosymbiont status (Χ 27 = 15.12; p = 0.034) and genotype significantly influenced adult morph in S. avenae (Χ 24 = 10.99 p = 0.024). Generally, S. avenae populations harbouring the facultative endosymbiont R. insecticola, either through single infections or co-infections with F. symbiotica, produced a higher proportion of alate adult morphs compared with populations harbouring no facultative endosymbionts or F. symbiotica in single infections (Figure 1A). For R. padi, no significant influence of facultative endosymbiont (Χ 2 = 2.02, p = 0.365) or genotype (Χ 22 = 2.54, p = 0.280) was observed (Figure 1B).
Proportion of S. avenae (A) and R. padi (B) individuals that develop into an alate (winged) adult morph. Data are pooled across all clonal populations into genotypes and facultative endosymbiont status; plot colour indicates facultative endosymbiont status.
Development time to reproductive maturity is driven by aphid morph and genotype
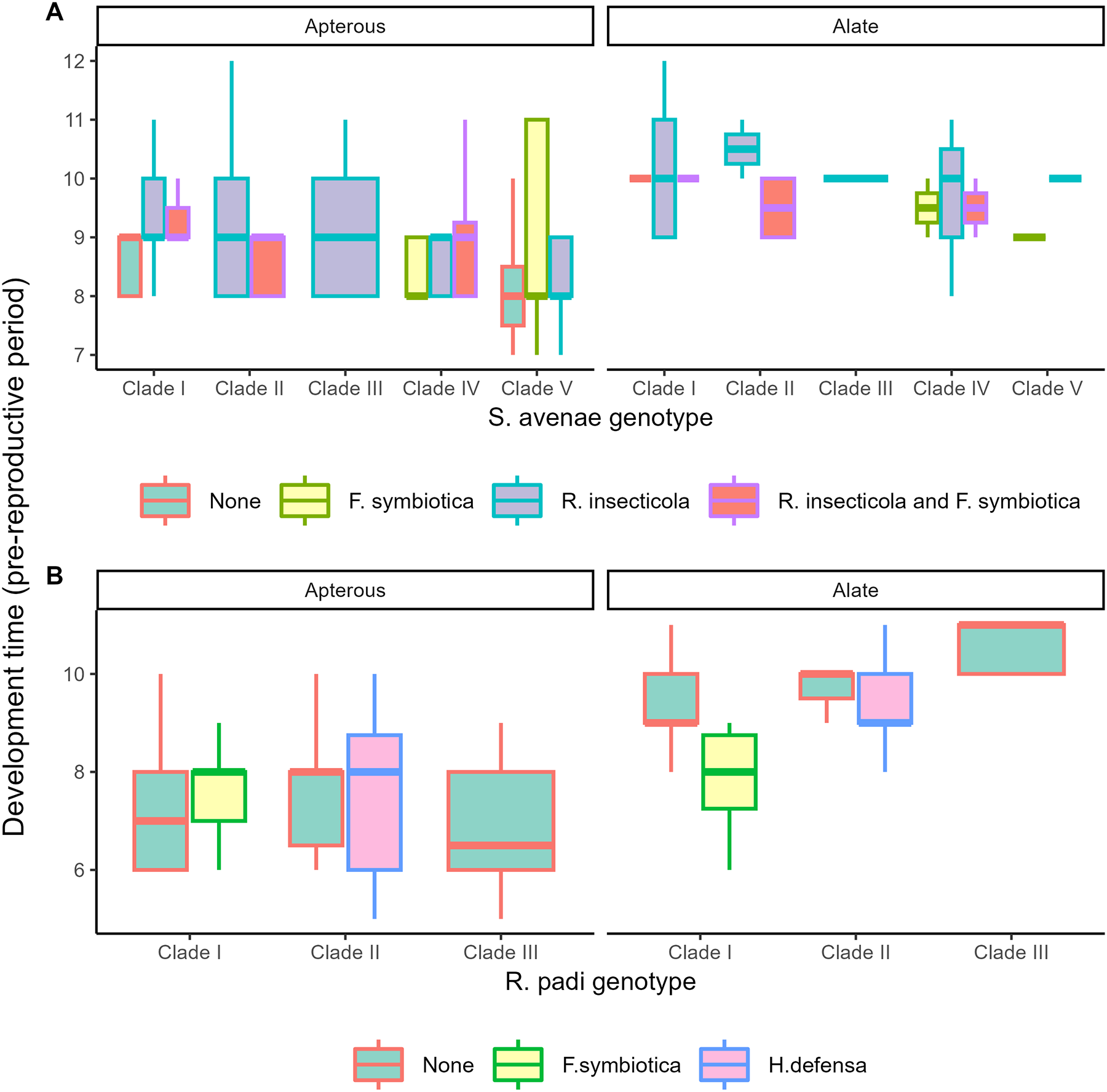
In both species, aphids that developed into alate morphs took longer to reach reproductive maturity than aphids developing into apterous morphs (S. avenae: F 1,277 = 57.22; p = <0.001; R. padi: F 1,121 = 142.99; p = <0.001). For S. avenae (Figure 2A), pre-reproductive development time was also influenced by aphid genotype (F 4,277 = 4.91; p = <0.001); post hoc analysis identified differences between Clade V with Clades III (p = 0.041) and Clade I (p = <0.001). For R. padi (Figure 2B), pre-reproductive period was primarily influenced by the interaction between genotype and aphid morph (F 2,121 = 6.31; p = 0.027), with longer development times for alate aphids across all genotypes. Endosymbiont had no significant influence on development time for either species (S. avenae: F 7,277 = 1.19; p = 0.303; R. padi: F2,121 = 0.98; p = 0.375).
Development time to reproductive maturity for S. avenae (A) and R. padi (B). Data are pooled across all clonal populations into genotype, facultative endosymbiont status, and adult morph (apterous or alate); plot colour indicates facultative endosymbiont status and individual plots are shown for the two adult morphs.
Reproductive output in S. avenae is influenced by genotype and the facultative endosymbiont Regiella insecticola
Aphid morph was a key factor determining reproductive output in both species, with apterous morphs having a greater reproductive output than alate morphs (S. avenae: F 1,277 = 24.82; p = <0.001; R. padi: Χ 21 = 51.88; p = <0.001; Figure 3). For S. avenae (Figure 3A), the intrinsic rate of population increase was also influenced by genotype (F 4,277 = 11.94; p = <0.001), endosymbiont association (F 1,277 = 3.71; p = <0.001), and the interaction between aphid genotype and morph (F 1,277 = 4.47; p = 0.002). On average, apterous aphids belonging to Clade III were the least fecund with post hoc testing identifying differences when compared with Clade I (p = <0.001), Clade II (p = <0.001), Clade IV (p = <0.001), and Clade V (p = <0.001). Apterous aphids from Clade V were the most fecund, producing more offspring when compared with Clade I (p = <0.001) and Clade II (p = 0.019). No differences were observed between genotypes for reproduction of alate morphs. For endosymbionts, pairwise analysis within genotypes found that for aphids belonging to Clade II, aphids containing a co-infection of R. insecticola and F. symbiotica had a higher fecundity when compared with individuals containing only single R. insecticola infections (p = <0.001).
Reproductive output (the intrinsic rate of fitness, rm) for S. avenae (A) and R. padi (B). Data are pooled across all clonal populations into genotype, facultative endosymbiont status, and adult morph (apterous or alate); plot colour indicates facultative endosymbiont status and individual plots are shown for the two adult morphs.
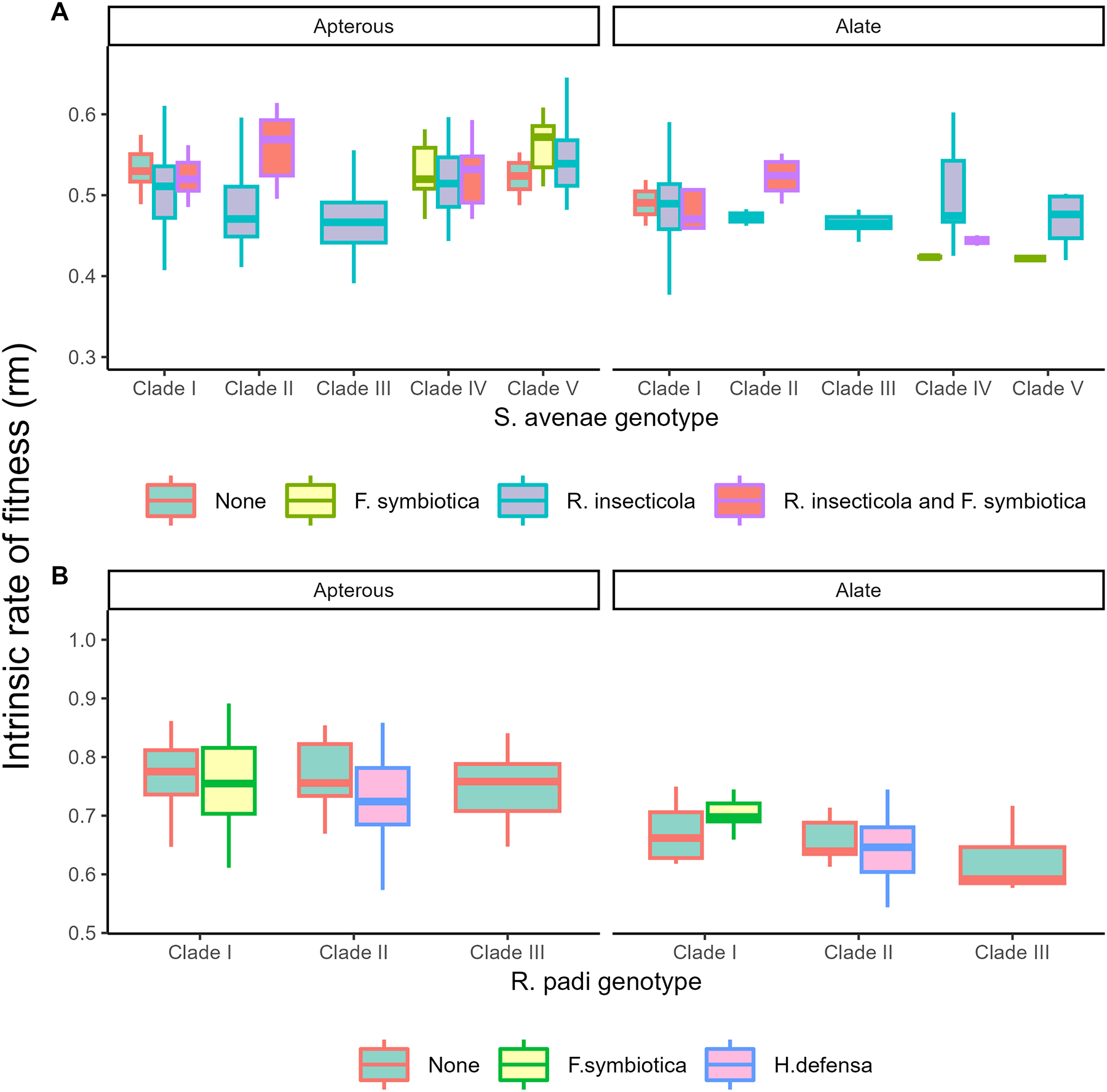
For R. padi (fig. 4B), there was no significant influence of aphid genotype (F 2,121 = 2.78; p = 0.066), endosymbiont (F 2,121 = 1.02; p = 0.363), or genotype × morph interaction (F 2,121 = 1.81; p = 0.168), with aphid morph the key driver of reproduction (F 1,121 = 51.88; p = <0.001).
Discussion
This study links phenotype with heritable factors in agriculturally important cereal aphid species. Two heritable factors were characterised, genetic diversity (aphid genotype) and association with facultative endosymbionts. Both genotypic and endosymbiotic drivers of cereal aphid phenotype were identified.
Genotype is a key heritable factor determining aphid phenotype (life-history and fitness), with genotypic variation underpinning phenotypic variation for numerous aphid species (Figueroa et al., Reference Figueroa, Simon, Le Gallic, Prunier-Leterme, Briones, Dedryver and Niemeyer2004; Khan and Port, Reference Khan and Port2008; Leybourne et al., Reference Leybourne, Bos, Valentine and Karley2020a; Martinez-Chavez et al., Reference Martinez-Chavez, Roberts, Karley, Shaw and Pope2024). Here, genotype was found to influence several life-history traits in S. avenae, with aphids belonging to Clade V developing faster and apterous aphids belonging to Clade III being the least fecund. Previous studies have made similar findings, with aphid genotype influencing mean relative growth rate (Jackson et al., Reference Jackson, Malloch, McNamara and Little2020), reproductive output (Figueroa et al., Reference Figueroa, Simon, Le Gallic, Prunier-Leterme, Briones, Dedryver and Niemeyer2004), and development time (Khan and Port, Reference Khan and Port2008). These phenotypic traits are also plastic and can vary in response to environmental factors such as host plant species (Figueroa et al., Reference Figueroa, Simon, Le Gallic, Prunier-Leterme, Briones, Dedryver and Niemeyer2004) and cultivar (Caillaud et al., Reference Caillaud, Dedryver, Di Pietro, Simon, Fima and Chaubet1995). In contrast, for R. padi, there was limited influence of aphid genotype on the phenotypic traits examined (pre-reproductive development time, reproductive output, alate production), with aphid morph (alate or apterous) the main factor influencing development time and reproductive output in R. padi. Increased development time and reduced fecundity for alate vs. apterous morphs has been commonly observed across different R. padi (Khan and Port, Reference Khan and Port2008; Yu et al., Reference Yu, Yang, Gill, Chirgwin, Gu, Joglekar, Umina and Hoffmann2024) and S. avenae (Khan and Port, Reference Khan and Port2008; Wu et al., Reference Wu, Hu, Wu, Yao and Xu2023) populations, and although a direct effect of R. padi genotype on aphid fitness was not detected a combined effect of aphid morph and aphid genotype was observed. This interaction influenced development time to reproductive maturity, with limited variation in aptera development time across the three genotypes assessed.
For R. padi, endosymbiont association did not influence any of the phenotypic traits measured. Still, other studies have identified endosymbiont-derived fitness traits in R. padi, including increased longevity in R. padi transfected with the endosymbiont R. insecticola (Yu et al., Reference Yu, Yang, Gill, Chirgwin, Gu, Joglekar, Umina and Hoffmann2024) and a fitness cost of reduced nymph mass gain associated with H. defensa presence (Leybourne et al., Reference Leybourne, Bos, Valentine and Karley2020a, Reference Leybourne, Valentine, Bos and Karley2020b). Variability in observed endosymbiont-mediated phenotypes could be driven by specific aphid-symbiont combinations, or costly phenotypes might only be observed under less favourable conditions. For example, the H. defensa-mediated phenotype of reduced nymph mass gain was observed when aphids are feeding on less favourable (partially-resistant) host plants (Leybourne et al., Reference Leybourne, Valentine, Bos and Karley2020b), indicating that endosymbiont-mediated effects are potentially influenced by more complex aphid-environment interactions.
For the S. avenae populations, some facultative endosymbiont phenotypes were observed. This included higher production of alate morphs for S. avenae that associated with the endosymbiont R. insecticola (either alone or through co-infections with F. symbiotica) when compared with populations that harboured no endosymbionts or only hosted F. symbiotica. Facultative endosymbionts also influenced reproductive output, with this phenotype potentially linked to aphid genotype. In Clade II, aphid harbouring a co-infection of R. insecticola and F. symbiotica had a greater reproductive output compared with populations harbouring only R. insecticola in a single infection, however, these endosymbiont-derived effects were less apparent in other genotypes. Similar observations of increased alate production and higher reproduction have been described for S. avenae that associate with R. insecticola (Mahieu et al., Reference Mahieu, González-González, Rubio-Meléndez, Moya-Hernández, Francis and Ramírez2024). However, studies also describe contrasting results: reduced alate production in S. avenae that associate with R. insecticola (Liu et al., Reference Liu, Lei and Chen2019; Ramírez-Cáceres et al., Reference Ramírez-Cáceres, Moya-Hernández, Quilodrán, Nespolo, Ceballos, Villagra and Ramírez2019) and negative effects of R. insecticola association on reproductive output in S. avenae (Liu et al., Reference Liu, Lei and Chen2019; Luo et al., Reference Luo, Luo, Meng, Wan, Zhao and Hu2017, Reference Luo, Monticelli, Li, Ahmed, Pandharikar, Zhao, Desneux and Hu2020; Ramírez-Cáceres et al., Reference Ramírez-Cáceres, Moya-Hernández, Quilodrán, Nespolo, Ceballos, Villagra and Ramírez2019). This variability in endosymbiont-derived phenotypes indicates that the observed phenotypes are often underpinned by host aphid genotype, potentially acting through more complex genotype × endosymbiont interactions. However, this requires further examination.
Fukatsuia symbiotica (previously pea aphid x-type symbiont) was found to associate with both S. avenae (two populations in single infections, three populations in co-infection with R. insecticola) and R. padi (two populations in single infections). Little is known about the influence F. symbiotica infection has on cereal aphid phenotype. However, in other aphid species, F. symbiotica is often found in co-infections with H. defensa and Spiroplasma spp. (Zytynska et al., Reference Zytynska, Tighiouart and Frago2021). These F. symbiotica co-infections can influence aphid interactions with natural enemies and reduce reproductive output (Zytynska et al., Reference Zytynska, Tighiouart and Frago2021). Here, no direct effect of F. symbiotica was observed in R. padi. However, for S. avenae, co-infections of F. symbiotica with R. insecticola enhance aphid reproductive output in apterous aphids belonging to Clade II. This finding provides further insight into the role of F. symbiotica and suggests that symbiont-mediated phenotypes potentially follow a genotype x endosymbiont interaction.
Conclusion
Cereal crops are often dominated by a small number of prolific clonal aphid populations. The prevalence of a given clonal population over other populations suggests that the dominant population has a phenotypic advantage. Characterising phenotypic diversity between distinct clonal aphid populations can help to identify which phenotypic traits help a given clonal population proliferate. Many aphid phenotypes are inherited through two heritable factors – genetic diversity (genotype) and association with facultative (vertically transmitted) endosymbiotic bacteria. By characterising these heritable factors and associating these with agriculturally important phenotypes (such as development time, reproductive output, winged morph production), it is possible to generate biological insights into the processes underpinning the dominance of specific aphid clones. Future research could use these insights to develop pest and disease management strategies based around the phenotypic risk of the aphid populations present within the locality, potentially contributing towards the development of sustainable pest management programmes.
For example, recent findings using the same aphid populations show that genotypic diversity in R. padi is a key driver behind efficient transmission of the aphid-vectored virus barley yellow dwarf virus PAV (Leybourne et al., Reference Leybourne, Whitehead and Will2024a), and similar studies have indicated that R. insecticola can reduce the ability of R. padi to transmit barley yellow dwarf virus PAV (Yu et al., Reference Yu, Yang, Gill, Chirgwin, Gu, Joglekar, Umina and Hoffmann2024). By combining information on aphid life-history (this study) with an estimation of virus transmission risk (Leybourne et al., Reference Leybourne, Whitehead and Will2024a; Yu et al., Reference Yu, Yang, Gill, Chirgwin, Gu, Joglekar, Umina and Hoffmann2024) and associating these with heritable factors (genotype, endosymbionts), we can work towards the development of models and decision support systems that combine these insights to estimate aphid dispersal and virus spread for distinctive aphid genotypes or endosymbiont combinations.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0007485325000124.
Acknowledgements
The author would like to thank Petra Melloh for technical and experimental support.
Funding
The Royal Commission for the Exhibition of 1851 through a Research Fellowship (RF-2022-100004) and The Alexander von Humboldt Foundation through a Postdoctoral Research Fellowship (ALAN).
Competing interests
There is no conflict of interest associated with this research.






