Introduction
Sorghum bicolor L. (Moench) is a multi-purpose crop grown for grain, forage and bioenergy (Rooney et al. Reference Rooney, Blumenthal, Bean and Mullet2007; Wang et al. Reference Wang, Xiong and Guo2024). In Australia, around 70–80% of sorghum crops are grown under dryland systems, particularly in the North-Eastern regions (Grains Research and Development Corporation (GRDC) 2025). Sorghum is valued for its resilience to stress, primarily drought and heat stress; however, adequate rainfall and nitrogen nutrition are still required to achieve optimum yields (Worland et al. Reference Worland, Robinson, Jordan, Schmidt and Godwin2017; Ostmeyer et al. Reference Ostmeyer, Bahuguna, Kirkham, Bean and Jagadish2022). In Australia, the soils are typically highly weathered and often contain low nitrogen levels that are limiting for crop production. Improving the nitrogen-use-efficiency, as well as improving stress tolerance, are key aims of breeding programs in these regions.
In northern Australia, the use of forage sorghum is complicated by the production of cyanogenic glucosides (CNglcs): specialized metabolites that release hydrogen cyanide (HCN) upon tissue damage, a process termed cyanogenesis (Gleadow and Møller Reference Gleadow and Møller2014). Sorghum produces the CNglc dhurrin using a simple biosynthesis pathway involving three structural enzymes: CYP79A1, CYP71E1 and UGT85B1 (Halkier and Møller Reference Halkier and Møller1989; Gleadow and Møller Reference Gleadow and Møller2014) (Supplementary Figure S1). Dhurrin provides adaptive benefits, including anti-herbivore defence, abiotic stress tolerance and contributes to nitrogen management (Gleadow and Møller Reference Gleadow and Møller2014; Gruss et al. Reference Gruss, Ghaste, Widhalm and Tuinstra2022). However, HCN is a major concern for livestock producers, as high dhurrin levels can cause illness and death in cattle (Giantin et al. Reference Giantin, Franzin, Brusa, Montemurro, Bozzetta, Caprai, Fedrizzi, Girolami and Nebbia2024). This is of particular concern under the environmental conditions in Australia, where events such as drought and nitrogen application can spike dhurrin concentrations in sorghum tissues.
Efforts to reduce HCN toxicity to date have focussed on eliminating CNglcs from crops, with success in sorghum and cassava (Blomstedt et al. Reference Blomstedt, Gleadow, O’Donnell, Naur, Jensen, Laursen, Olsen, Stuart, Hamill, Møller and Neale2012; Tuinstra et al. Reference Tuinstra, Krothapalli, Dilkes and Buescher2016; Juma et al. Reference Juma, Mukami, Mweu, Ngugi and Mbinda2022; Gomez et al. Reference Gomez, Berkoff, Gill, Iavarone, Lieberman, Ma, Schultink, Karavolias, Wyman, Chauhan, Taylor, Staskawicz, Cho, Rokhsar and Lyons2023). However, removing CNglcs from crops also removes the benefits they contribute, such as resilience to pests, tolerance to abiotic stress and nitrogen management (Gleadow and Møller Reference Gleadow and Møller2014; Gruss et al. Reference Gruss, Ghaste, Widhalm and Tuinstra2022). Rather than eliminating CNglcs from crops, altering patterns of CNglc accumulation could enable retention of these benefits while mitigating the risks associated with cyanogenic plants.
Current knowledge of the molecular regulation of dhurrin accumulation is limited with only one transcription factor identified to date: SbGATA22, a repressor of the dhurrin biosynthesis pathway with maturation (Rosati et al. Reference Rosati, Quinn, Gleadow and Blomstedt2024). While other transcriptional regulators of key CNglcs genes have been characterized in other cyanogenic species, such as PdbHLH2 in Prunus dulcis and LjbHLH7 in Lotus japonicus (Sánchez-Pérez et al. Reference Sánchez-Pérez, Pavan, Mazzeo, Moldovan, Aiese Cigliano, Del Cueto, Ricciardi, Lotti, Ricciardi, Dicenta, López-Marqués and Møller2019; Chen et al. Reference Chen, Liu, Zhang, Niu, Zhao, Liu, Georgiev, Xu, Zhang and Zhou2022), application of these findings to regulation of dhurrin in sorghum is complicated by the evolutionary distances between these species and the independent evolution of cyanogenesis in their lineages (Thodberg et al. Reference Thodberg, Sørensen, Bellucci, Crocoll, Bendtsen, Nelson, Motawia, Møller and Neilson2020; Hansen et al. Reference Hansen, Sørensen, Bellucci, Brandt, Olsen, Goodger, Woodrow, Møller and Neilson2023; Sánchez-Pérez and Neilson Reference Sánchez-Pérez and Neilson2024). Within Sorghum, however, dhurrin biosynthesis appears to be monophyletic and conserved across the 22 undomesticated taxa in the genus (Myrans et al. Reference Myrans, Diaz, Khoury, Carver, Henry and Gleadow2020, Reference Myrans, Vandegeer, Henry and Gleadow2021; Cowan et al. Reference Cowan, Møller, Norton, Knudsen, Crocoll, Furtado, Henry, Blomstedt and Gleadow2022), providing a unique opportunity to study regulatory variation.
Sorghum is classified into five subgenera: Eusorghum (including S. bicolor), Chaetosorghum, Heterosorghum, Parasorghum and Stiposorghum (Garber Reference Garber1950; Dillon et al. Reference Dillon, Lawrence, Henry and Price2007; Ananda et al. Reference Ananda, Norton, Blomstedt, Furtado, Møller, Gleadow and Henry2021). Northern Australia is a hotspot for Sorghum diversity, with 17 taxa either native or endemic to the region (Myrans et al. Reference Myrans, Diaz, Khoury, Carver, Henry and Gleadow2020). These wild relatives represent valuable genetic resources for breeding resilience traits into domesticated sorghum lines, including reduced dhurrin concentration (Ananda et al. Reference Ananda, Myrans, Norton, Gleadow, Furtado and Henry2020; Cowan et al. Reference Cowan, Blomstedt, Norton, Henry, Møller and Gleadow2020). Among these species, Sorghum macrospermum E.D. Garber (subgenus Chaetosorghum) is particularly notable. Endemic to a small area in the Northern Territory, it shares key agronomic traits with S. bicolor, such as annual growth habit, palatable forage, and large, nutritious grain (Lazarides et al. Reference Lazarides, Hacker and Andrew1991). These characteristics, combined with its superior drought tolerance and lower cyanogenic profile, make S. macrospermum a promising candidate for crop improvement (Ananda et al. Reference Ananda, Myrans, Norton, Gleadow, Furtado and Henry2020; Cowan et al. Reference Cowan, Blomstedt, Norton, Henry, Møller and Gleadow2020).
Dhurrin regulation in S. macrospermum differs markedly from that in S. bicolor. Both species accumulate dhurrin during early seedling growth, followed by a decline with development (Cowan et al. Reference Cowan, Blomstedt, Møller, Henry and Gleadow2021). However, S. macrospermum has much lower concentrations overall and shows limited responsiveness to nitrogen supply (Myrans et al. Reference Myrans, Vandegeer, Henry and Gleadow2021). Under drought stress, dhurrin accumulates primarily in roots rather than leaves (Cowan et al. Reference Cowan, Blomstedt, Norton, Henry, Møller and Gleadow2020; Ananda et al. Reference Ananda, Norton, Blomstedt, Furtado, Møller, Gleadow and Henry2022). In contrast, S. bicolor upregulates foliar dhurrin production under both drought stress and high nitrogen conditions (O’Donnell et al. Reference O’Donnell, Møller, Neale, Hamill, Blomstedt and Gleadow2013; Blomstedt et al. Reference Blomstedt, Rosati, Møller and Gleadow2018; Rosati et al. Reference Rosati, Blomstedt, Møller, Garnett and Gleadow2019; Cowan et al. Reference Cowan, Blomstedt, Norton, Henry, Møller and Gleadow2020; Myrans et al. Reference Myrans, Vandegeer, Henry and Gleadow2021). If varieties of sorghum could be developed with the pattern seen in S. macrospermum, the stems and leaves would likely remain safe for grazing under a wide range of conditions.
Because amino acid sequences of the dhurrin biosynthesis genes are highly conserved (Cowan et al. Reference Cowan, Møller, Norton, Knudsen, Crocoll, Furtado, Henry, Blomstedt and Gleadow2022), differences in dhurrin accumulate likely reflect divergence in regulatory mechanisms between S. bicolor and S. macrospermum rather than structural gene variation. Understanding these regulatory factors is critical for developing sorghum suited to dryland and low-input systems, where low toxicity, drought resilience and nitrogen-use efficiency are essential. Furthermore, dhurrin turnover, involving enzymes such as Nitrilase 4B2 (NIT4B2), may contribute to nitrogen recycling (Jenrich et al. Reference Jenrich, Trompetter, Bak, Olsen, Møller and Piotrowski2007; Pičmanovà et al. Reference Pičmanovà, Neilson, Motawia, Olsen, Agerbirk, Gray, Flitsch, Meier, Silvestro, Jørgensen, Sánchez-Pérez, Møller and Bjarnholt2015; Bjarnholt et al. Reference Bjarnholt, Neilson, Crocoll, Jørgensen, Motawia, Olsen, Dixon, Edwards and Møller2018). Targeting dhurrin production and turnover may represent a novel target for improving nitrogen-use efficiency in resource-limited cropping systems.
This study investigated short-term response of nitrogen-limited S. macrospermum to a high nitrogen supply (12 mM). We measured changes in nitrate and dhurrin concentrations and expression of key genes in nitrogen assimilation and dhurrin metabolism over 48 hours. We hypothesized that nitrogen application would induce dhurrin biosynthesis in roots, but not in leaves, consistent with previous observations of higher root dhurrin levels in S. macrospermum. To test this, dhurrin concentrations and expression of the dhurrin biosynthesis gene SmCYP79A1 and the turnover enzyme SbNIT4B2, was quantified in roots and leaves.
Additionally, we hypothesized that if dhurrin metabolism is activated by nitrogen in S. macrospermum, its regulation would involve conserved transcriptional networks shared with S. bicolor. Expression of four candidate transcription factors identified in S. bicolor (SmHRS1, SmTRFL3, SmEIL3 and SmMYB-related) were also assessed. Changes in expression of Glutamine synthetase 2 (GS2), involved in primary nitrogen assimilation, was used as positive control for nitrogen response.
Materials and methods
Plant growth conditions
Seeds of S. macrospermum (AGG 322277 WSOR3 MHA 716; Australian Grains Genebank) were germinated according to Cowan et al. (Reference Cowan, Blomstedt, Norton, Henry, Møller and Gleadow2020). Germinated seeds were transferred to pots containing washed river sand (Bastion River Sand, Dingo Cement Australia) and left to establish in a greenhouse under ambient light and temperatures between 25°C and 35°C. After two weeks, seedlings were transferred to 10-cm diameter pots with sand and irrigated twice daily with an automated drip irrigation system. Nutrients were supplied with a modified Long Ashton Nutrient Solution containing a final concentration of 0.5 mM nitrogen (Miller et al. Reference Miller, Gleadow and Cavagnaro2014; Myrans et al. Reference Myrans, Vandegeer, Henry and Gleadow2021), with 200 mL of nutrient solution delivered three times per week (Supplementary Table S1).
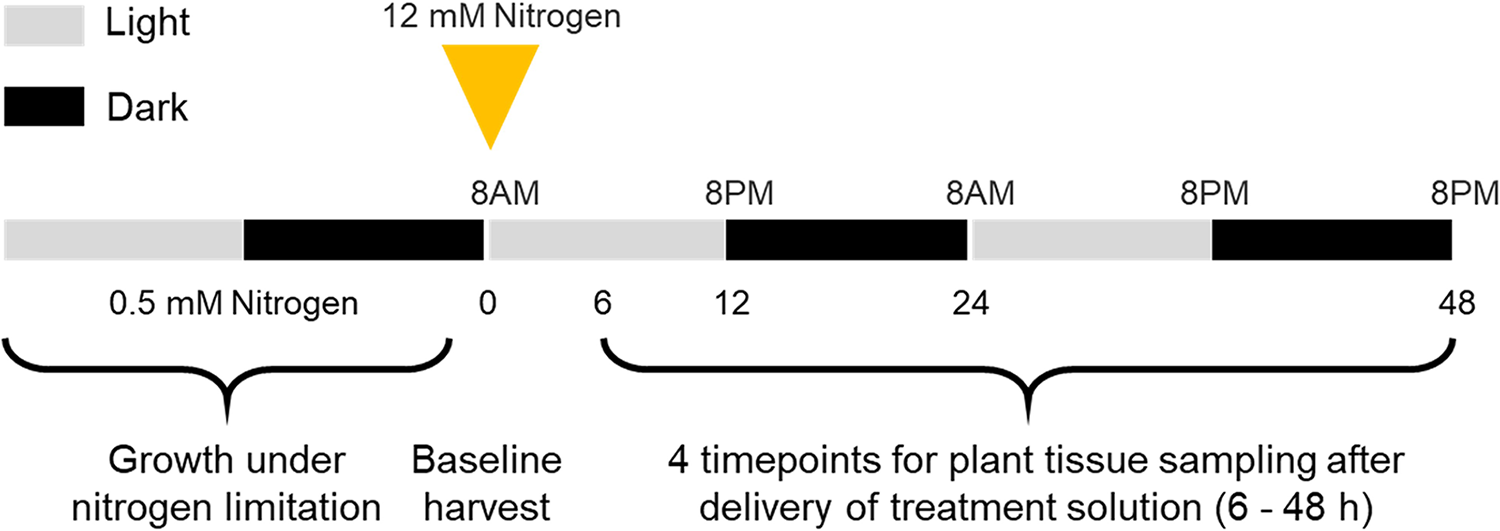
Before application of treatment solutions, plants were randomly assigned to one of five time points: baseline (0-hours), 6-, 12-, 24- or 48-hours. Time points were selected based on data from S. bicolor, where induction of dhurrin biosynthesis genes occurred within 48-hours of nitrogen treatment (Quinn Reference Quinn2025). Due to limited availability of S. macrospermum seeds, each time point included five biological replicates and treatments were compared to the baseline control at the 0-hour time point. A 200 mL of a high-nitrogen treatment solution (12 mM nitrogen) was applied to all plants, except the baseline control (Fig. 1). Measurements were taken for height, leaf number, and chlorophyll content (Apogee Instruments) at each harvest. Tissue was also sampled for chemical and gene expression analysis. Baseline samples for nitrate and biomass measurements were not included.
Overview of experiment set up and time points for sampling. Sorghum macrospermum plants were grown under 0.5 mM nitrogen until they reached the 6–7 leaf stage. Samples were collected from leaf, stem and root tips for RNA at 0-hours (baseline), and remaining plant tissue oven-dried (n = 5). All remaining plants were treated with a 12 mM nitrogen solution, and harvested in the same way at 6-, 12-, 24- and 48-hours after nitrogen treatment (n = 5).

Chemical analyses
Nitrate (NO3−) concentration (mg−1 g−1 DW) was determined spectrophotometrically using 15 mg ± 0.15 mg of finely ground oven-dried plant material following the method of Cataldo et al. (Reference Cataldo, Maroon, Schrader and Youngs1975) as modified by O’Donnell et al. (Reference O’Donnell, Møller, Neale, Hamill, Blomstedt and Gleadow2013).
Cyanogenic glucoside concentrations were determined as hydrogen cyanide potential (HCNp), which is the maximum amount of HCN capable of being released upon hydrolysis of all CNglcs in a sample. HCNp is directly proportional to the dhurrin content in the sample (Gleadow et al. Reference Gleadow, Moldrup, O’Donnell and Stuart2012). HCNp was measured on fresh tissue from leaf, stem, and root tips collected at the time of harvest according to O’Donnell et al. (Reference O’Donnell, Møller, Neale, Hamill, Blomstedt and Gleadow2013). The assay was modified to account for the lower dhurrin concentrations produced in S. macrospermum by using 50 µL NaCN samples undiluted and adjusting concentration of acetic acid to 2 M to neutralize the pH of the sample. Remaining plant tissue was oven dried and accurately weighed for calculation of HCNp as µg−1 g−1 DW.
RNA extraction
Tissues from leaf and root tips were sampled at the time of harvest, frozen immediately in liquid nitrogen and stored at −80°C until RNA extraction. Plant tissue was ground to a fine powder in a bead mill (FastPrep-24, MP Biomedicals) kept cool with dry ice. RNA was extracted using 1 mL Tri Reagent (Sigma, 92389–100 mL), according to the manufacturer’s instructions and resuspended in 100 µL nuclease-free water. RNA integrity was checked with a 2% agarose gel. Quantity and quality were determined using a spectrophotometer (NanoDrop1000, ThermoFisher Scientific). RNA samples where A260/230 < 1.60 were re-precipitated and pellets washed with ethanol to remove additional salt contamination and quality control checks repeated.
Sequence extraction and primer design
No reference sequence for S. macrospermum is currently available. However, a reference sequence for closely related S. bicolor is available (McCormick et al. Reference McCormick, Truong, Sreedasyam, Jenkins, Shu, Sims, Kennedy, Amirebrahimi, Weers, McKinley, Mattison, Morishige, Grimwood, Schmutz and Mullet2018), and short-read sequence data for S. macrospermum is also publicly available (Ananda et al. Reference Ananda, Norton, Blomstedt, Furtado, Møller, Gleadow and Henry2021). For design of primers for target genes in S. macrospermum, an approach similar to Ananda et al. (Reference Ananda, Norton, Blomstedt, Furtado, Møller, Gleadow and Henry2021) was used to align the S. macrospermum short-read sequencing data (AGG accession 302367, retrieved from NBCI, BioSample SAMN17369313; Ananda et al. Reference Ananda, Norton, Blomstedt, Furtado, Møller, Gleadow and Henry2021) to the S. bicolor reference assembly (v3.1.1, (McCormick et al. Reference McCormick, Truong, Sreedasyam, Jenkins, Shu, Sims, Kennedy, Amirebrahimi, Weers, McKinley, Mattison, Morishige, Grimwood, Schmutz and Mullet2018)). Reads for S. macrospermum were analysed with FastQC to check for and remove low-quality reads before aligning to the S. bicolor reference. Reads were aligned using the Map with Bowtie for Illumina tool available in Galaxy Australia, with default parameters for paired-end reads. Alignment was visualized in the Integrative Genomics Viewer (Version 2.17.0) and consensus sequence for target genes in S. macrospermum retrieved from alignment to S. bicolor genes. A similar approach was used by Ananda et al. (Reference Ananda, Norton, Blomstedt, Furtado, Møller, Gleadow and Henry2021) in the construction of the nuclear Sorghum phylogeny, and sequences for genes involved in cyanogenesis in wild sorghum species, including S. macrospermum, have been analysed using this approach (Cowan et al. Reference Cowan, Møller, Norton, Knudsen, Crocoll, Furtado, Henry, Blomstedt and Gleadow2022). Sequences retrieved from the alignment by Ananda et al. (Reference Ananda, Norton, Blomstedt, Furtado, Møller, Gleadow and Henry2021) were used to validate the sequences retrieved in the present study. Consensus sequences for target gene genomic sequences were aligned using MEGA11 (Version 11.0.13) and show similar nucleotide sequences between S. macrospermum and S. bicolor exons (available at https://doi.org/10.26180/31200166). Primers targeting S. macrospermum sequences for use in qPCR were designed using PerlPrimer.(v1.1.21) (Supplementary Table S2).
cDNA synthesis and reverse-transcription qPCR
One microgram of RNA was DNAse-treated (RQ1 DNase, Promega) before cDNA synthesis using First-Strand cDNA synthesis kit (Invitrogen) with oligo(dT) primers. Gene expression was measured using qPCR on a LightCycler 480 instrument (Roche). Reactions were set up in triplicate on 384-well plates using a robotic liquid handling system (Ep-Motion 5070, Eppendorf Research), with each 10 µL reaction containing 5 µL 2 × SensiFAST SYBR qPCR mix (Meridian Bioscience, BIO-98050-2000), 2.5 µL cDNA template (diluted 1:10 with nuclease-free water), and gene-specific forward and reverse qPCR primers to a final concentration of 250 nM each. For each primer pair, the efficiency of the qPCR reaction was determined using a dilution series of a purified PCR product amplified in triplicate alongside the samples. Relative gene expression (mean normalized expression) was calculated using Q-gene software (Simon Reference Simon2003).
Statistical analyses
Statistical analyses were performed using GraphPad Prism (Version 9.3.1, GraphPad Software, LLC). Data for harvest measurements (height, leaf number and chlorophyll concentration) were analysed using a one-way ANOVA. Data for chemistry and gene expression were analysed using two-way ANOVA. Gene expression data was log-transformed to meet assumptions of normality and homogeneity of variance. Post-hoc analyses for significant ANOVA results were conducted with Dunnett’s test for multiple comparisons, where contrasts were made against the baseline time point and subsequent time points.
Results
Harvest measurements
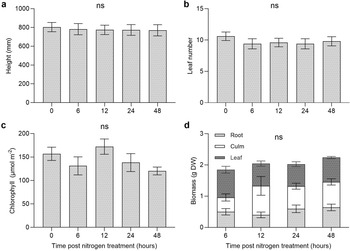
Measurements of growth were taken at the time of harvest for height, leaf number and chlorophyll content (Fig. 2a–c). No significant differences were detected for S. macrospermum at any time point in response to the nitrogen surge application. There was no significant increase in total biomass in response to the nitrogen surge treatment during the sampling timeframe (Fig. 2d).
Measurements for (a) height, (b) leaf number and (c) chlorophyll content for nitrogen-limited S. macrospermum in response to a 12 mM nitrogen treatment at four time points up to 48 hours after treatment (n = 5). Measurements taken at time of harvest. (d) Total biomass for dried tissue (n = 5). Columns represent mean ± SE. No significant differences between time points were detected using a one-way ANOVA.

Nitrate concentration
Nitrate concentration was measured on dried and ground tissue for whole samples of leaf, sheath and roots of S. macrospermum (Fig. 3). No significant differences in nitrate concentration were detected for any tissue between time points; however, differences between tissue type were identified (F(2, 48) = 24.25, P < 0.0001), with leaf tissue containing lower amounts of nitrates than sheath (P < 0.0001) and root (P < 0.0001) (Fig. 3a–c).
Nitrate concentration of (a) leaf, (b) sheath and (c) root tissue for nitrogen-limited S. macrospermum in response to a 12 mM nitrogen treatment (n = 5). Columns represent the mean ± SE. No significant differences between time points were detected using a two-way ANOVA.

Hydrogen cyanide potential
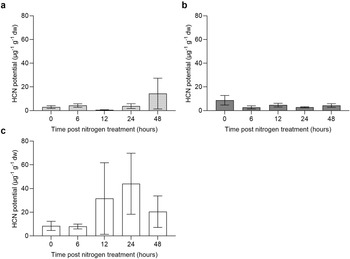
Hydrogen cyanide potential (HCNp) was measured on fresh tissue collected from the last fully emerged leaf, the stem and from root tips of S. macrospermum. HCNp was significantly different between tissue types (F(2,57) = 4.79, P = 0.012), with roots containing higher HCNp than leaf tissue (P = 0.008), which appears to be driven by increased HCNp in a few individuals across the 12-to 48-hour time points in root tissue at the later stages of the sampling period. There was also a slight increase in HCNp in leaf tissue at the 48-hour time point, although no significant effect of nitrogen treatment on the HCNp was detected in any time point in the tissues tested (F(4,57) = 1.02, P = 0.40) (Fig. 4a–c).
Hydrogen cyanide potential (HCNp) of (a) leaf, (b) stem and (c) root tissue for nitrogen-limited S. macrospermum in response to a 12 mM nitrogen treatment (n = 5). Columns represent mean ± SE. No significant differences between time points were detected using a two-way ANOVA.

Expression of key metabolism genes
Glutamine synthetase 2 (GS2) plays a key role in assimilation of inorganic forms of nitrogen in plants. The nitrogen treatment significantly increased the expression of the SmGlutamine Synthetase 2 (SmGS2) in S. macrospermum (F(4, 20) = 9.70, P = 0.0002), with a similar response seen in both root and leaf tissue (Fig. 5a). Expression of SmGS2 in leaf tissue was significantly higher than baseline at 6-, 24- and 48-hours after application of the nitrogen treatment. Expression of SmGS2 in roots also increased from 6-hours after nitrogen treatment, but was only significantly higher than baseline at the 24-hour time point (P = 0.006).
Gene expression normalized to ubiquitin (mean normalized expression) for Sorghum macrospermum leaf and root tissues in response to treatment with a 12 mM nitrogen solution. (a) SmGlutamine Synthetase 2, (b) SmCYP79A1 and (c) SmNitrilase 4B2. Columns represent mean ± SE (n = 3). Stars represent significant differences at P < 0.05 (*), 0.01 (**) and 0.001 (***) using two-way ANOVA and Sidak’s test for multiple comparisons.

Analysis of expression of SmCYP79A1, which encodes the gene for the first step in the dhurrin biosynthesis pathway, showed a significant effect of tissue type (F(1, 16) = 10.43, P = 0.005), but not time point (F(4, 16) = 2.47, P = 0.087) (Fig. 5b). Differences between the two tissues were largely driven by increased expression of SmCYP79A1 in the roots from the 12-hour time point onwards, as there is negligible expression of this gene in either root tissue at 0- and 6-hours after nitrogen treatment or at any time point in leaf. Post-hoc analysis comparing all time points to the baseline within tissue types detected significantly higher expression of SmCYP79A1 from 12-hours after nitrogen application (Fig. 5b).
Like SmCYP79A1, there was no significant difference between time points for expression of SmNitrilase 4B2 (SmNIT4B2), although expression was significantly higher in root that leaf tissue (F(1, 20) = 30.04, P < 0.0001) (Fig. 5c). Multiple comparisons between tissue type at each time point showed that expression of SmNIT4B2 in roots was significantly higher than that in leaf tissue at the 12-hour (P = 0.016) and 24-hour (P = 0.002) time points. This appears to be the result of both a slight decrease in expression of SmNIT4B2 in leaf tissue at these time points and a corresponding increase in expression in root tissue.
Expression of candidate transcription factors
Of the genes encoding putative transcription factors, SmHRS1 was the only candidate with noticeable expression detected in S. macrospermum (Fig. 6). There was essentially no expression of SmEIL3, SmMYB-like or SmTRF-like in either leaf or root at any time point tested (Fig. 6 a–c). Expression of SmHRS1 was detected in both leaf and root tissues, and there was an effect of time point on expression (F(4, 20) = 5.84, P = 0.003), with significantly higher expression of SmHRS1 in leaf tissue at the 12-hour time point compared to baseline expression levels (P = 0.004) (Fig. 6d).
Gene expression normalized to ubiquitin (mean normalized expression) for Sorghum macrospermum leaf and root tissues in response to treatment with a 12 mM nitrogen solution. (a) SmEthylene Insensitive-like 3, (b) SmTRF-like 3, (c) SmMYB-like and (d) SmHRS1. Data show mean ± SE (n = 3). Note differences in ranges for Y-axes between a and c are two orders of magnitude lower than for d. Stars represent significant differences at P < 0.05 (*), 0.01 (**) and 0.001 (***) using two-way ANOVA and Sidak’s test for multiple comparisons.

Discussion
Crop wild relatives offer genetic resources for improving crop resilience and productivity, particularly under dryland conditions where nitrogen is limited and drought is frequent (Ananda et al. Reference Ananda, Myrans, Norton, Gleadow, Furtado and Henry2020). Sorghum macrospermum is a promising candidate for breeding due to its low dhurrin content, high productivity with minimal nitrogen and ability to tolerate drought (Cowan et al. Reference Cowan, Blomstedt, Norton, Henry, Møller and Gleadow2020; Myrans et al. Reference Myrans, Vandegeer, Henry and Gleadow2021). Importantly, S. macrospermum has been successfully crossed with S. bicolor making the prospect of introgression much simpler (Price et al. Reference Price, Dillon, Hodnett, Rooney, Ross and Johnston2005; Kuhlman et al. Reference Kuhlman, Burson, Stelly, Klein, Klein, Price and Rooney2010). Here, we examined dhurrin regulation in nitrogen-limited S. macrospermum by characterizing the pattern of dhurrin biosynthesis in space and time following a single dose of nitrogen. Findings support the hypothesis that dhurrin biosynthesis occurs primarily in the roots of S. macrospermum. We conclude that this trait could be used to develop low-cyanide, stress-resilient sorghum for low-input dryland systems.
Growth rates of S. macrospermum were delayed under nitrogen limitation, with plants taking 12 weeks to produce 12 leaves. These results are similar to previous reports where S. macrospermum is grown in low-nitrogen conditions (Myrans et al. Reference Myrans, Vandegeer, Henry and Gleadow2021). The high-nitrogen treatment did not significantly affect growth or nitrate concentration over 48 hours, contrasting with S. bicolor, which shows rapid nitrate accumulate under similar conditions (Quinn Reference Quinn2025). The lack of increased nitrate concentration suggests S. macrospermum employs a different strategy for nitrogen use and uptake, possibly occurring at a slower rate not detected within the 48-hour sampling timeframe. Alternatively, S. macrospermum may rely more heavily on nitrate stores, possibly an adaptation to the unpredictable monsoonal region where S. macrospermum grows (Lazarides et al. Reference Lazarides, Hacker and Andrew1991).
Despite the absence of higher nitrate concentrations, elevated expression of SmGS2 was observed in both roots and leaves. Increased SmGS2 expression was induced earlier in leaves, likely linked to photosynthesis, while root expression increased gradually, suggesting regulation tied to ongoing nitrogen demand. These findings indicate a transcriptional response to nitrogen application, but further work is needed to clarify additional regulatory mechanisms involved in S. macrospermum nitrogen metabolism, which could provide additional targets for altering NUE in sorghum.
Dhurrin concentrations in S. macrospermum were very low throughout the experiment, both before and after nitrogen application, with no significant changes detected across tissues. These findings align with previous reports showing that Australian sorghum species, including S. macrospermum, typically accumulate less than 50 µg g−1 HCNp even under high-nitrogen conditions (Cowan et al. Reference Cowan, Blomstedt, Norton, Henry, Møller and Gleadow2020, Reference Cowan, Blomstedt, Møller, Henry and Gleadow2021; Myrans et al. Reference Myrans, Vandegeer, Henry and Gleadow2021), compared with > 300 µg g−1 commonly observed in S. bicolor leaves (Blomstedt et al. Reference Blomstedt, Rosati, Møller and Gleadow2018; Myrans et al. Reference Myrans, Vandegeer, Henry and Gleadow2021).
Although overall dhurrin levels remained low, with only a slight increase in root dhurrin after 12 hours, there was a significant upregulation of SmCYP79A1, the gene encoding the first and rate-limiting enzyme in the dhurrin biosynthesis pathway (Busk and Møller Reference Busk and Møller2002) (Supplementary Figure S1). Dhurrin concentration would likely continue to increase, as seen in S. bicolor, where an increase in SmCYP79A1 transcripts has been shown to precede changes in phenotype (Sohail et al. Reference Sohail, O’Donnell, Blomstedt and Gleadow2023). Expression of SmCYP79A1 was restricted to roots, unlike S. bicolor, where nitrogen induces leaf-specific expression of SbCYP79A1 within 24 hours (Quinn Reference Quinn2025). This pattern, together with previous drought studies, suggests that roots are the primary site of dhurrin production in S. macrospermum in response to environmental cues. The functional significance of root-localized dhurrin is not clear, although CNglcs can be effective at managing soil pests, such as nematodes (Abawi and Widmer Reference Abawi and Widmer2000; Smith et al. Reference Smith, Scheikowski and Kafle2024). However, the dhurrin levels in S. macrospermum roots are so low that they are unlikely to be effective in deterring herbivores (Myrans et al. Reference Myrans, Vandegeer, Henry and Gleadow2021).
Alternatively, dhurrin stored in roots may serve as a nitrogen reserve for regrowth after stress, such as grazing or fire, through a recycling pathway that has been identified in S. bicolor and other cyanogenic species (Pičmanovà et al. Reference Pičmanovà, Neilson, Motawia, Olsen, Agerbirk, Gray, Flitsch, Meier, Silvestro, Jørgensen, Sánchez-Pérez, Møller and Bjarnholt2015; Nielsen et al. Reference Nielsen, Stuart, Pičmanovà, Rasmussen, Olsen, Harholt, Møller and Bjarnholt2016; Bjarnholt et al. Reference Bjarnholt, Neilson, Crocoll, Jørgensen, Motawia, Olsen, Dixon, Edwards and Møller2018). SbNIT4B2 has been implicated in this pathway (Nielsen et al. Reference Nielsen, Stuart, Pičmanovà, Rasmussen, Olsen, Harholt, Møller and Bjarnholt2016; Bjarnholt et al. Reference Bjarnholt, Neilson, Crocoll, Jørgensen, Motawia, Olsen, Dixon, Edwards and Møller2018), although it also plays a role in the detoxification of HCN after bioactivation (Jenrich et al. Reference Jenrich, Trompetter, Bak, Olsen, Møller and Piotrowski2007). In S. macrospermum, SmNIT4B2 expression was consistently higher in roots than leaves, though unaffected by nitrogen application, leaving its primary role in either defence or nitrogen management uncertain. Examining expression of other bioactivation and detoxification genes, such as dhurrinase or hydroxynitrile lyase, could help resolve this. Determining whether and how dhurrin contributes to nitrogen recycling in S. macrospermum, and elucidating the underlying mechanisms, will be critical for understanding its multifunctional role and for leveraging this trait to develop sorghum varieties with improved nitrogen-use efficiency and resilience, supporting sustainable dryland cropping systems.
The regulatory mechanisms for the induction of dhurrin biosynthesis appear to be different in S. bicolor and S. macrospermum. To explore whether nitrogen-responsive regulation is conserved, we examined four candidate nitrogen-responsive transcription factors previously identified through a GRN approach in S. bicolor that may control expression of genes in the dhurrin biosynthesis pathway (Quinn Reference Quinn2025). Expression of SmEIL3, SmTRF-like, and SmMYB-like were consistently very low across tissues and time points, indicating they are unlikely to play a role in nitrogen response or dhurrin regulation in S. macrospermum. A fourth candidate, SmHRS1, was expressed in both leaves and roots but showed significant upregulation only in leaves 12 hours after the application of nitrogen. This pattern suggests that it plays a role in the leaf nitrogen response rather than dhurrin regulation per se. HRS1 belongs to the NIGT/HRS1 family of GARP-type transcription factors, widely characterized for nitrogen signalling and stress responses (Kiba et al. Reference Kiba, Inaba, Kudo, Ueda, Konishi, Mitsuda, Takiguchi, Kondou, Yoshizumi, Ohme-Takagi, Matsui, Yano, Yanagisawa and Sakakibara2018; Li et al. Reference Li, Zhou, Li, Zhang and Gao2021). While this group has primarily been characterized in Arabidopsis, roles in regulation of nitrogen responses appear to be conserved in monocots, including rice (OsNIGT1) (Yanagisawa Reference Yanagisawa2013; Medici et al. Reference Medici, Marshall-Colon, Ronzier, Szponarski, Wang, Gojon, Crawford, Ruffel, Coruzzi and Krouk2015), and in networks shared by maize and sorghum (Du et al. Reference Du, Ning, He, Wang, Ge, Xu and Zhao2020). Although the HRS1 orthologue has not been functionally characterized in S. bicolor, its conservation across monocots suggests a similar role in nitrogen signalling within the Sorghum genus, including S. macrospermum.
GARP-type transcription factors could contribute to nitrogen-responsive regulation of dhurrin in S. macrospermum, although SmHRS1 does not appear to control dhurrin biosynthesis in roots. Further interrogation of the differences in regulation of the dhurrin biosynthesis pathway between S. macrospermum and S. bicolor could enable fine-tuning of dhurrin concentrations in sorghum crops, thereby improving safety of forage varieties. Additionally, understanding the regulatory mechanisms that restrict accumulation of dhurrin to roots could enable development of both safer sorghum crops while retaining the benefits of nitrogen storage in the roots. Completion of the S. macrospermum reference genome will allow identification of transcription factor orthologues and CRE differences, providing insight into regulatory evolution and offering genetic resources to improve nitrogen-use efficiency and resilience in sorghum for sustainable dryland cropping systems.
Our study reveals distinct nitrogen-responsive regulation of dhurrin biosynthesis in S. macrospermum, with production restricted to roots rather than leaves as in S. bicolor. Understanding these species-level differences offers valuable genetic resources for breeding sorghum varieties with improved nitrogen-use efficiency and resilience, supporting sustainable dryland cropping systems.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S1479262126100574.
Acknowledgements
This research was supported by the Australian Research Council Discovery Project DP210103010 to R.M.G. AAQ was supported by an Australian Government Research Training Program (RTP) Scholarship, an AW Howard Memorial Trust Research Fellowship, and a Denis and Maisie Carr Award. We thank Dr Sally Norton at the Australian Grains Genebank for provision of the S. macrospermum seed material used in this experiment. AI tools were used in the editing of this manuscript.
Author contributions
The study was conceived by R.M.G., A.A.Q. and C.K.B. A.A.Q. and C.K.B. conducted the greenhouse experiment and harvested plant tissue. A.A.Q. performed experimental analysis, analysed the data and drafted the manuscript. All authors contributed to editing and revising the manuscript.
Conflict of interests
The authors declare no conflict of interest.
Use of artificial intelligence (AI) tools
AI tools were for assistance with editing and restructuring the manuscript to improve clarity, readability and conciseness. All suggestions were reviewed and edited by the authors, who take full responsibility for the accuracy and integrity of the work.