Highlights
What is already known?
-
• The scientific literature is growing at increasing rates, with meta-analyses and systematic reviews (MA/SRs) seen as a gold-standard of evidence synthesis.
-
• Large language models (LLMs) can assist with paper screening in MA/SRs.
-
• LLM task performance depends strongly on the instructions given to the LLM, known as the prompt.
-
• Single LLM prompt templates have been retrospectively developed for optimal paper selection versus comparison MA/SRs on specific datasets.
What is new?
-
• Five hundred fifteen screening-related prompts were prospectively tested with open-access LLMs against MA/SRs from four different medical fields, revealing prompt engineering principles applicable across the included MA/SRs.
-
• Prompt performance heterogeneity across MA/SRs was quantified and demonstrated, and workload/cost reduction of top-performing prompts calculated.
Potential impact for RSM readers
-
• Findings highlight a previously overlooked recall/sensitivity–precision/specificity trade-off in LLM prompt design for MA/SRs and contribute an evidence-based framework for prompt engineering in LLM-based paper screening.
-
• The Python-based LLM screening pipeline and list of prompts used here are publicly available and allow for flexible testing of more MA/SRs and open-source or commercial LLMs, supporting further development of LLM-based evidence synthesis methods.
1 Introduction
The scientific literature is growing at increasing rates.Reference Chen, Hu and Peng 1 , Reference González-Márquez, Schmidt, Schmidt, Berens and Kobak 2 By 2024, PubMed alone contained over 36 million articles,Reference Chen, Hu and Peng 1 , Reference Sayers, Bolton and Brister 3 and over 1 million new articles in biomedicine are published every year.Reference González-Márquez, Schmidt, Schmidt, Berens and Kobak 2 With this expansion comes a greater need to efficiently synthesize information,Reference Wang, Cao, Danek, Jin, Lu and Sun 4 with meta-analyses and systematic reviews (MA/SRs) widely regarded as a gold-standard of scientific evidence synthesis.Reference Basile, Villaschi and Maggioni 5 , Reference Wang, Nayfeh, Tetzlaff, O’Blenis and Murad 6 However, it is becoming exceedingly difficult to publish high-quality MA/SRs at a rate required for the rapidly growing scientific literature base, given time and economic demands.Reference Borah, Brown, Capers and Kaiser 7 – Reference Syriani, David and Kumar 9 For example, across 195 MA/SRs, the average time from initiation to publication was 67.3 weeks,Reference Borah, Brown, Capers and Kaiser 7 and another analysis estimated the total labor cost for a single MA/SR at $141,194.80.Reference Michelson and Reuter 8 Up to 20% of the time required for MA/SRs is spent on screening alone,Reference Haddaway and Westgate 10 which involves sorting search results into studies to include or exclude.
The rapid growth of published research and the time-intensive nature of MA/SRs have increased interest in artificial intelligence (AI) tools.Reference Wang, Cao, Danek, Jin, Lu and Sun 4 , Reference Cao, Sang and Arora 11 – Reference Kim, Chua, Li, Rickard and Lorenzo 13 In a 2018 survey of 168 reviewers, most of whom worked in biomedicine, 32% reported using automation tools for at least some aspects of the review process,Reference Van Altena, Spijker and Olabarriaga 12 a portion that has likely increased since the advent of large language models (LLMs).Reference Wang, Cao, Danek, Jin, Lu and Sun 4 , Reference Cao, Sang and Arora 11 , Reference Kim, Chua, Li, Rickard and Lorenzo 13 – Reference Thirunavukarasu, Ting, Elangovan, Gutierrez, Tan and Ting 16 For the screening process in particular, LLMs offer the opportunity to efficiently analyze large amounts of text to identify relevant studies for MA/SRs.Reference Wang, Cao, Danek, Jin, Lu and Sun 4 , Reference Syriani, David and Kumar 9 , Reference Cao, Sang and Arora 11 , Reference Delgado-Chaves, Jennings and Atalaia 14 , Reference Huotala, Kuutila, Ralph and Mäntylä 17 , Reference Strachan 18 LLMs, such as those powering applications like ChatGPT,Reference Cao, Sang and Arora 11 , Reference Kim, Chua, Li, Rickard and Lorenzo 13 , 19 process and generate text based on large training datasets and are increasingly used in diverse fields, including medicine.Reference Liu, Zhou and Gu 15 , Reference Thirunavukarasu, Ting, Elangovan, Gutierrez, Tan and Ting 16 However, LLM performance depends largely on the text instruction given to the LLM, referred to as the prompt.Reference Strachan 18 , Reference Lautrup, Hyrup, Schneider-Kamp, Dahl, Lindholt and Schneider-Kamp 20 , Reference Nguyen, MacKenzie and Kim 21 For example, Nguyen et al.Reference Nguyen, MacKenzie and Kim 21 showed that differences in prompting significantly influenced ChatGPT’s performance on the 2022 American College of Radiology In Training Test Questions for Diagnostic Radiology Residents. Another study analyzed 123 prompts on advice given by LLMs in response to medical questions and found that while LLMs generally provided concise and correct advice, they could also give erroneous and even medically dangerous answers depending on the prompt.Reference Lautrup, Hyrup, Schneider-Kamp, Dahl, Lindholt and Schneider-Kamp 20
Prompt engineering is also recognized as important in the screening of papers for MA/SRs. However, prior research on the use of LLMs to automate the screening process has primarily focused on either developing single prompts or prompt templatesReference Syriani, David and Kumar 9 , Reference Cao, Sang and Arora 11 , Reference Strachan 18 or has employed prompt engineering strategies not tailored to the task of MA/SR paper screening specifically.Reference Cao, Sang and Arora 11 , Reference Delgado-Chaves, Jennings and Atalaia 14 , Reference Huotala, Kuutila, Ralph and Mäntylä 17 For example, a 2025 study of 48,425 citations from 10 MA/SRs optimized individual prompt templates with the LLM GPT4-0125-preview in a train-test-split design and achieved a mean abstract-screening sensitivity of 97.7% and specificity of 85.2%.Reference Cao, Sang and Arora 11 Another 2024 study of LLM title and abstract screening against three MA/SRs found comparable performance between commercial models (e.g., GPT) and open-source alternatives (llama3-8b-8192, mixtral:8x22b), without consistent advantage for larger models.Reference Delgado-Chaves, Jennings and Atalaia 14 Agreement was low to moderate, with Matthews Correlation Coefficients of 0.24–0.35 and kappa values of 0.48–0.62, the highest of which were observed with Mistral:v0.2. Two further 2024 studies benchmarked GPT-based screening against gold-standard MA/SRs. One reported sensitivities of 95.8% and 100% against a single review,Reference Strachan 18 and the other reported an accuracy of 82% across five reviews.Reference Syriani, David and Kumar 9 However, both studies developed only a single optimized prompt template for maximizing performance.Reference Syriani, David and Kumar 9 , Reference Strachan 18 Therefore, these important prior studiesReference Syriani, David and Kumar 9 , Reference Cao, Sang and Arora 11 , Reference Delgado-Chaves, Jennings and Atalaia 14 , Reference Strachan 18 provided optimized prompts and prompt templates but did not identify specific, generalizable prompt characteristics that can guide prompt design for new MA/SRs.
To the best of our knowledge, to date, no study has prospectively examined the impact of specific prompt characteristics on paper selection by LLMs compared to medical gold-standard MA/SRs, that is, published and peer-reviewed MA/SRs serving as the reference. In this study, we developed the open-access Python-based pipeline MA-LLM to compare LLM-based paper selection against MA/SRs from four different medical fields published after the LLMs’ training cut-offs. We tested the performance of 515 unique prompts with systematically varied screening-related characteristics across three LLMs and four MA/SRs, yielding a total of 12,360 runs of the MA-LLM pipeline.
2 Methods
2.1 Gold-standard meta-analysis and systematic review selection
This study employed two search strategies to source high-quality MA/SRs. First, articles were identified through direct access to the websites of Q1-registered journals in the Web of Science Journal Info database. 22 Second, a PubMed search was performed with the terms (“Systematic Review” [Publication Type]) OR (“Meta-Analysis” [Publication Type]), and a publication date filter starting from July 1, 2024. The date filter was applied to select only MA/SRs published after the LLMs’ training cut-offs, freely available on the LLM’s websites, to avoid the MA/SRs being part of the LLMs’ training data. The search was purposive rather than exhaustive and aimed to identify MA/SRs from different medical fields that met predefined methodological criteria.
Potential MA/SRs were cross-referenced against the inclusion criteria, with only those meeting all criteria selected: adhered to PRISMA 2020 statement for MA/SRs reporting;Reference Page, McKenzie and Bossuyt 23 employed dual-reviewer screening;Reference Waffenschmidt, Knelangen, Sieben, Bühn and Pieper 24 indexed on PubMed; published in a Q1-ranked journal; 22 published after July 1, 2024; written in English with full-text availability; included any type of study design (e.g., prospective, retrospective, cohort, RCTs, etc.); any type of MA/SRs including network MAs, to reflect applicability to a diverse range of study designs and research aims; studies included in the MA/SR were published in English; included at least 10 studies but no more than 50 studies (to keep the workload required to analyze the characteristics of each study at a manageable level for this study); provided fully accessible data on search queries, included studies, cohorts, and estimates; PubMed was used as one of the search engines, and all PubMed search terms were explicitly provided in the MA/SR. From eligible MA/SRs, four were selected to represent different medical fields (psychiatry, endocrinology, pediatric pulmonology, orthodontics).
2.2 Prompt design
Prompts were developed following prior prompt engineering research.Reference Wang, Cao, Danek, Jin, Lu and Sun 4 , Reference Wang, Chen and Deng 25 , Reference Chen, Zhang, Langrené and Zhu 26 The aim of the present study was to evaluate the impact of predefined prompt characteristics on LLM screening performance compared with gold-standard manual meta-analysis/systematic review (MA/SR) screening. Prompts were developed along predefined screening dimensions, with each dimension included a null variant as the reference category for statistical comparison. First, prompts differed by screening focus: (1) no focus (reference category), (2) topics/research questions, (3) methods, or (4) results/findings. Second, prior research suggests that additional context and task instructions may improve performance.Reference Chen, Zhang, Langrené and Zhu 26 Hence, prompts differed by the amount and type of MA/SR information provided: (1) no information (reference category), (2) the information that the papers would be screened for a MA/SR, (3) the title of the MA/SR, and (4) the inclusion and exclusion criteria of the MA/SR. Third, prompts varied by length to evaluate the independent effect of word count on agreement with MA/SR selection. Prompts were developed across all combinatorial combinations, with ≥10 prompts per combination.
While the present study aimed to evaluate these screening-specific prompt characteristics, other prompt engineering principles may affect performance.Reference Wang, Cao, Danek, Jin, Lu and Sun 4 , Reference Wang, Chen and Deng 25 . Reference Chen, Zhang, Langrené and Zhu 26 Prior research distinguishes zero-shot, one-shot, and few-shot prompting (varying amount of examples and specifications per prompt), and performance may also be affected by role-based prompting, which specifies the role/personality of the model.Reference Chen, Zhang, Langrené and Zhu 26 Other research suggests that repeated specification may improve performance.Reference Wang, Chen and Deng 25 We incorporated these principles in prompt design (e.g., “You are a world-class clinical researcher […]”) to evaluate screening-related characteristics across different prompt contexts. Furthermore, LLMs naturally exhibit probabilistic behavior, where same or similar prompts can produce different outputs.Reference Wang, Cao, Danek, Jin, Lu and Sun 4 , Reference Wang, Chen and Deng 25 , Reference Chen, Zhang, Langrené and Zhu 26 To address this issue, Markov chains were applied using the markovify package in Python to increase prompt diversity while preserving predefined characteristics.Reference Millard 27 For each combinatorial group, additional prompts were generated from word sequences of the manual set, removing duplicates and syntactically or semantically incorrect prompts.
Overall, this process resulted in 515 prompts. Across three LLMs, four MA/SRs, and title vs. abstract screening, this procedure yielded a total of 3 × 4 × 2 × 515 = 12,360 runs of the MA-LLM pipeline.
2.3 MA-LLM pipeline and large language models
PMIDs of screened articles were extracted from the National Institutes of Health (NIH) database, discarding duplicate citations. Titles and abstracts of screened papers were extracted automatically as part of the MA-LLM pipelineReference Van Rossum and Drake 28 using the BioC application programming interface (API), and the extracted information was stored to an SQLite database. Unlike manual screening, which typically proceeds sequentially from title screening to abstract screening of retained records, the MA-LLM pipeline iterated over the list of prewritten prompts, using each prompt in turn to screen the full list of titles or abstracts. With each iteration, the order of titles or abstracts was randomized to avoid systematic effects of the order of screened papers. The LLM response format was specified, and parsing was applied as part of the MA-LLM pipeline, with manual runs ensuring complete correctness of the response format and parsing. The full MA-LLM pipeline, a graphic usage tutorial, and all prompts are available at: https://zenodo.org/records/19097181 (see also Supplementary Figure S1). This pipeline allows for the integration of any open-source or commercial LLM, run via API or locally, and flexible testing against any SR/MA with automatic paper information extraction from the NIH database, as well as a separate setting for paper screening independent of MA/SR which was not used in the present study. Only publicly available models were used in the present study. Models of different sizes were chosen, varying by their number of parameters: llama3-8b-8192 (8 billion parameters, cut-off: March, 2023), 29 llama3-70b-8192 (70 billion parameters, cut-off: December, 2023), 30 and Mistral-Large-Instruct-2407-gptq-4bit (123 billion parameters, cut-off: June, 2024). 31
2.4 Selection performance evaluation
LLM selection quality was evaluated using both traditional metrics from signal detection theory commonly used in other fields of medicine and related works (sensitivity, specificity, accuracy),Reference Syriani, David and Kumar 9 , Reference Cao, Sang and Arora 11 , Reference Strachan 18 , Reference Leeflang, Deeks, Gatsonis and Bossuyt 32 – Reference Zou, O’Malley and Mauri 34 as well as more recent information retrieval performance metrics from machine learning (recall, precision, F1-score (F1)).Reference Delgado-Chaves, Jennings and Atalaia 14 , Reference Liu, Zhou and Gu 15 , Reference Powers 35 Papers selected by both the MA/SR and the LLM were counted as true positives, and papers discarded by both as true negatives. Papers discarded by the MA/SR but selected by the LLM were counted as false positives, whereas papers selected by the MA/SR but discarded by the LLM were counted as false negatives. Hence, the selection by the MA/SR served as the gold-standard comparison for the LLM-based selection. Comparison metrics were calculated with the following formulas:
2.5 Outcome metrics and statistical analysis
The primary outcome of this study was F1 as calculated above, including both recall/sensitivity and precision. The secondary outcome was recall/sensitivity in isolation, to evaluate conditions under which LLMs are more likely to identify all relevant articles selected by the MA/SR, which is particularly relevant in early-stage title and abstract screening. Both outcomes were analyzed using univariate linear mixed-effect models.Reference Brauer and Curtin 36 All statistical analyses were conducted in R (Version 4.4.3) 37 using the lme4 package (version 1.1–37)Reference Bates, Mächler, Bolker and Walker 38 for the linear mixed-effect models and the lmerTest package (version 3.1–3)Reference Kuznetsova, Brockhoff and Christensen 39 to calculate p-values via the Satterthwaite approximation. The fixed factors were (1) the specific LLM used, (2) whether titles or abstracts were screened, and (3) the prompt characteristics specified above (section reference, MA/SR information, length). The MA/SRs were included as the grouping variable with random intercepts to ensure result applicability across MA/SRs. Dummy coding was applied for the factors.
Analysis of variance (ANOVA) was used to compare differences in performance between individual prompts, with each prompt assigned a unique ID across LLMs, MA/SRs, and title versus abstract screening. ANOVA was also used to compare selection performance across MA/SRs. ANOVAs were performed using the aov() function in base R. 37
2.6 Workload and cost reduction analysis
In workload/cost reduction analyses, LLM use was modeled as a first-pass preselection. Any record the LLM rejected was considered permanently excluded without later manual review: That is., if the LLM mistakenly excluded a relevant paper (a false negative), it was counted as a missed inclusion in the workload and cost reduction analysis. P was defined as the portion of included articles in the average MA/SR among all screened papers. The expected proportion of records removed before human screening R was calculated as follows:
R reflected two scenarios: (i) a paper was ineligible (1 − P) and correctly removed by the LLM (specificity) or (ii) a paper was eligible (P) but erroneously removed by the LLM (1 − sensitivity).
Screening was assumed to constitute a fixed fraction fs
of the total workload and cost. Therefore, the portion of total workload saved was
![]() ${f}_s\times R$
. We defined
${f}_s\times R$
. We defined
![]() ${T}_{\mathrm{total}}$
as the total project duration in weeks and
${T}_{\mathrm{total}}$
as the total project duration in weeks and
![]() ${C}_{\mathrm{total}}$
as the total cost in USD for an average manual MA/SR. The absolute savings from LLM prescreening were consequently given by the following:
${C}_{\mathrm{total}}$
as the total cost in USD for an average manual MA/SR. The absolute savings from LLM prescreening were consequently given by the following:
To perform workload and cost reduction analyses, findings from prior studies on the average research extend and timelines of MA/SRs were used.
![]() ${f}_s$
was based on analyses estimating screening to account for 20% (7%–45%) of total MA/SR effort, the majority of which is spent on early-stage screening, particularly of the initial list of titles.Reference Haddaway and Westgate
10
${f}_s$
was based on analyses estimating screening to account for 20% (7%–45%) of total MA/SR effort, the majority of which is spent on early-stage screening, particularly of the initial list of titles.Reference Haddaway and Westgate
10
![]() ${C}_{\mathrm{total}}$
was based on an economic model that combined information of labor time and average salary to calculate the point estimate of $141,194.80 per MA/SR.Reference Michelson and Reuter
8
${C}_{\mathrm{total}}$
was based on an economic model that combined information of labor time and average salary to calculate the point estimate of $141,194.80 per MA/SR.Reference Michelson and Reuter
8
![]() ${T}_{\mathrm{total}}$
was based on a study of 195 prospectively registered and completed medical MA/SRs, reporting a mean time to publication of 67.3 weeks (SD=31 weeks; IQR=42 weeks).Reference Borah, Brown, Capers and Kaiser
7
Finally, to acknowledge heterogeneity between MA/SRs in our analyses, we calculated workload and cost reduction for a proportion of eligible studies P ranging from 1% to 10%, consistent with prior studies,Reference Borah, Brown, Capers and Kaiser
7
and matching portions found in the MA/SRs used here.Reference Geta, Terefa and Hailu
40
–
Reference Yang, Fang and Lyu
43
Using the top-performing prompts in the present study for F1-score and recall/sensitivity, we calculated total workload reduction (
${T}_{\mathrm{total}}$
was based on a study of 195 prospectively registered and completed medical MA/SRs, reporting a mean time to publication of 67.3 weeks (SD=31 weeks; IQR=42 weeks).Reference Borah, Brown, Capers and Kaiser
7
Finally, to acknowledge heterogeneity between MA/SRs in our analyses, we calculated workload and cost reduction for a proportion of eligible studies P ranging from 1% to 10%, consistent with prior studies,Reference Borah, Brown, Capers and Kaiser
7
and matching portions found in the MA/SRs used here.Reference Geta, Terefa and Hailu
40
–
Reference Yang, Fang and Lyu
43
Using the top-performing prompts in the present study for F1-score and recall/sensitivity, we calculated total workload reduction (
![]() ${f}_s\times R$
), weeks saved (
${f}_s\times R$
), weeks saved (
![]() ${T}_{\mathrm{reduction}}$
), and cost saved (
${T}_{\mathrm{reduction}}$
), and cost saved (
![]() ${C}_{\mathrm{reduction}}$
), under the assumptions of linear proportionality of time/cost with records screened, unchanged manual extraction/synthesis effort, and unchanged portion of eligible papers.
${C}_{\mathrm{reduction}}$
), under the assumptions of linear proportionality of time/cost with records screened, unchanged manual extraction/synthesis effort, and unchanged portion of eligible papers.
3 Results
3.1 Overall performance by screening characteristics
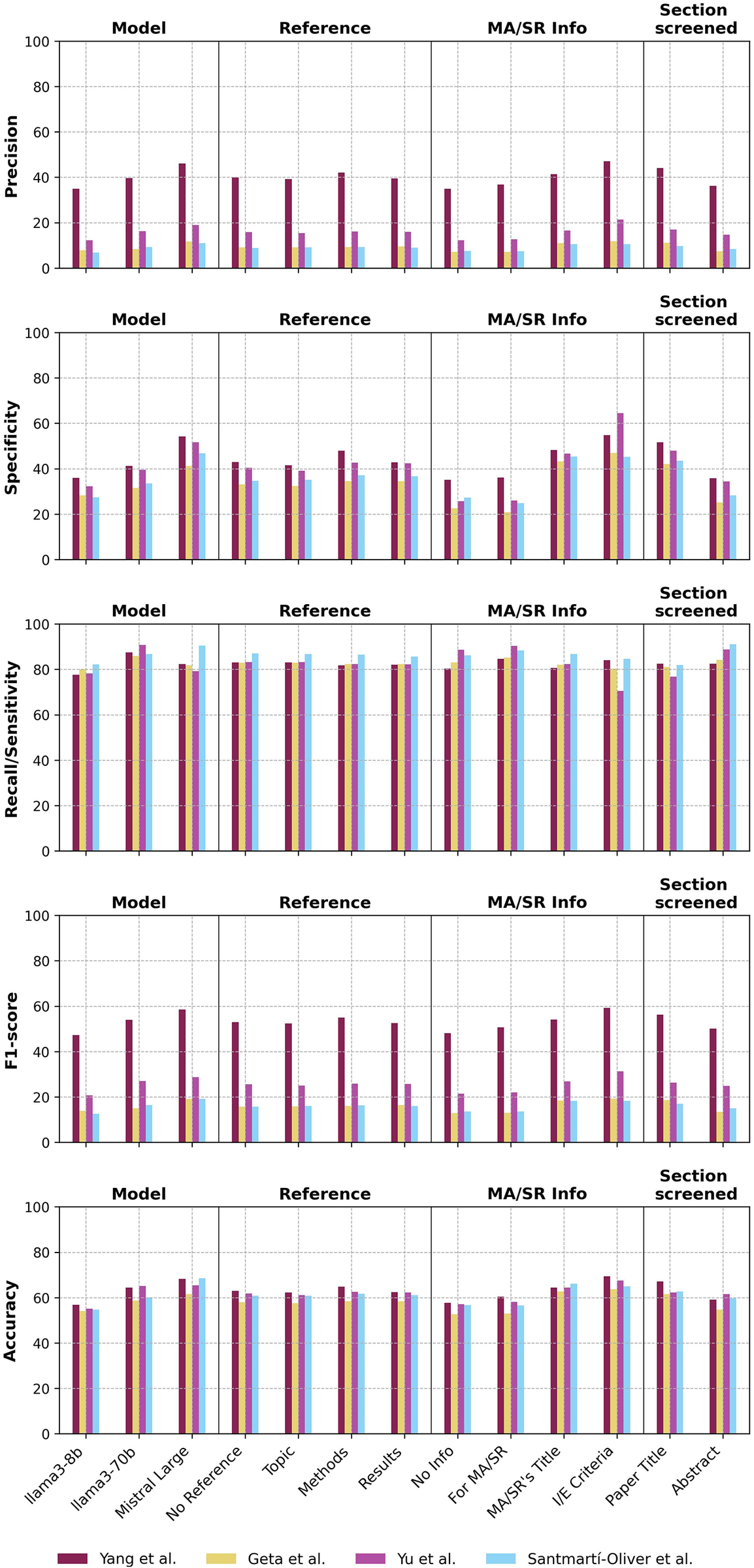
Over 12,360 pipeline runs across four MA/SRs and three LLMs, the 515 prompts achieved averages of recall/sensitivity = 83.6 ± 17.0%, precision = 18.5 ± 15.6%, F1 = 27.6 ± 17.2%, and accuracy = 61.1 ± 11.0% (Table 1, Figure 1) . Descriptively, the highest average recall/sensitivity scores reached by category were 87.7 ± 13.6% with the model llama3-70b, 87.1 ± 14.7% when prompting the LLM that selection would be for a MA/SR, 86.6 ± 16.0% when screening abstracts overall, and 86.4±15.6% for the MA/SR by Santmartí-Oliver et al.Reference Santmartí-Oliver, Jorba-García, Moya-Martínez, de-la-Rosa-Gay and Camps-Font 41 The highest average F1 reached by category were 53.2 ± 9.9% for the MA/SR by Yang et al.,Reference Yang, Fang and Lyu 43 32.0 ± 19.1% when including the MA/SR’s inclusion and exclusion criteria in the prompt, and 31.3 ± 18.5% with the model Mistral Large 2407.
Table 1 Overall mean and standard deviation of selection performance metrics by category, minimum and maximum by prompt average

Abbreviations: SD, standard deviation; min, minimum; max, maximum.

Figure 1 Selection performance across metrics and prompt characteristic categories.
Abbreviations: MA/SR, meta-analysis or systematic review; I/E, inclusion/exclusion.
3.2 Impact of specific prompt characteristics on screening performance
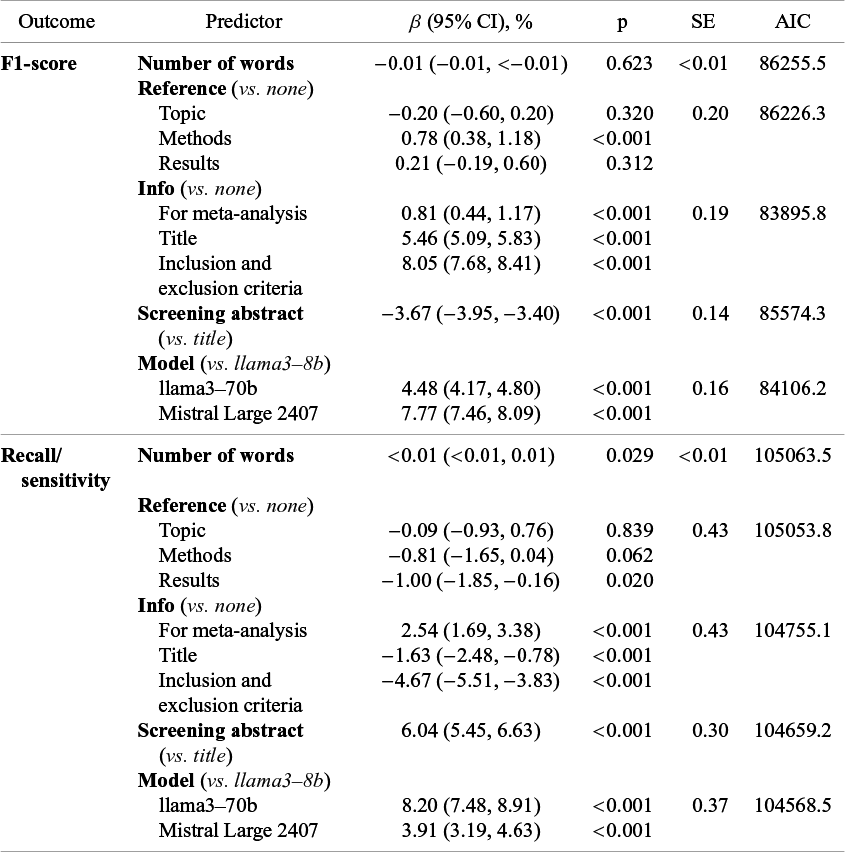
In linear mixed-effects models, F1 increased with reference to the methods of the screened papers in the prompt (β = 0.78%; 95% confidence interval (CI) = 0.38%–1.18%; p < 0.001) but not with reference to topics or results (Table 2). F1 also increased with each additional level of information about the MA/SR provided, with minor improvement when explicitly instructing the LLM to screen papers for MA/SR in general (β = 0.81%; CI = 0.44%–1.17%; p < 0.001), further improvement when including the MA/SR’s title (β = 5.46%; CI = 5.09%–5.83%; p < 0.001), and most improvement when including the MA/SR’s inclusion and exclusion criteria (β = 8.05%; CI = 7.68%–8.41%; p < 0.001). Conversely, F1 was significantly lower when screening abstracts as opposed to titles (β = −3.67%; CI = −3.95 to −3.40%; p < 0.001). Lastly, F1 improved with model size as compared to llama3-8b (8 billion), with higher accuracy with llama3-70b (70 billion, β = 4.48%; CI = 4.17%–4.80%; p < 0.001) and highest accuracy with Mistral Large 2407 (123 billion, β = 7.77%; CI = 7.46%–8.09%; p < 0.001).
Table 2 Effects of prompt and screening characteristics on paper selection performance

β-coefficients indicate change in percent in the outcome metric for that predictor category compared to the reference category. Abbreviations: CI, confidence interval; p, p-value; SE, standard error; AIC, Akaike information criterion.
Linear mixed-effects models with recall/sensitivity as the dependent variable showed partially contrasting effects compared to F1. Recall/sensitivity significantly increased with more words in the prompt (β < 0.01%; 95% CI < 0.01%; p = 0.029) (Table 2). Recall/sensitivity significantly decreased when including explicit reference to results of the screened papers in the prompt (β = −1.00%; CI = −1.85 to −0.16%; p = 0.020), whereas including reference to topics or methods did not significantly change recall/sensitivity. As for F1, recall/sensitivity was improved by explicitly prompting the LLM that papers should be screened for a MA/SR (β = 2.54%; CI = 1.69%–3.38%; p < 0.001) but decreased by including the MA/SR’s title (β = −1.63%; CI = −2.48 to −0.78%; p < 0.001) or selection criteria (β = −4.67%; CI = −5.51 to −3.83%; p<0.001). In contrast to F1, recall/sensitivity was significantly greater when screening abstracts as opposed to titles (β = 6.04%; CI = 5.45%–6.63%; p < 0.001). As with F1, recall/sensitivity was improved with model size compared to llama3-8b (8 billion), but the greatest improvements were observed with llama3-70b (70 billion, β = 8.20%; CI = 7.48%–8.91%; p < 0.001), followed by Mistral Large 2407 (123 billion, β = 3.91%; CI = 3.19%–4.63%; p < 0.001).
3.3 Differences between individual prompts
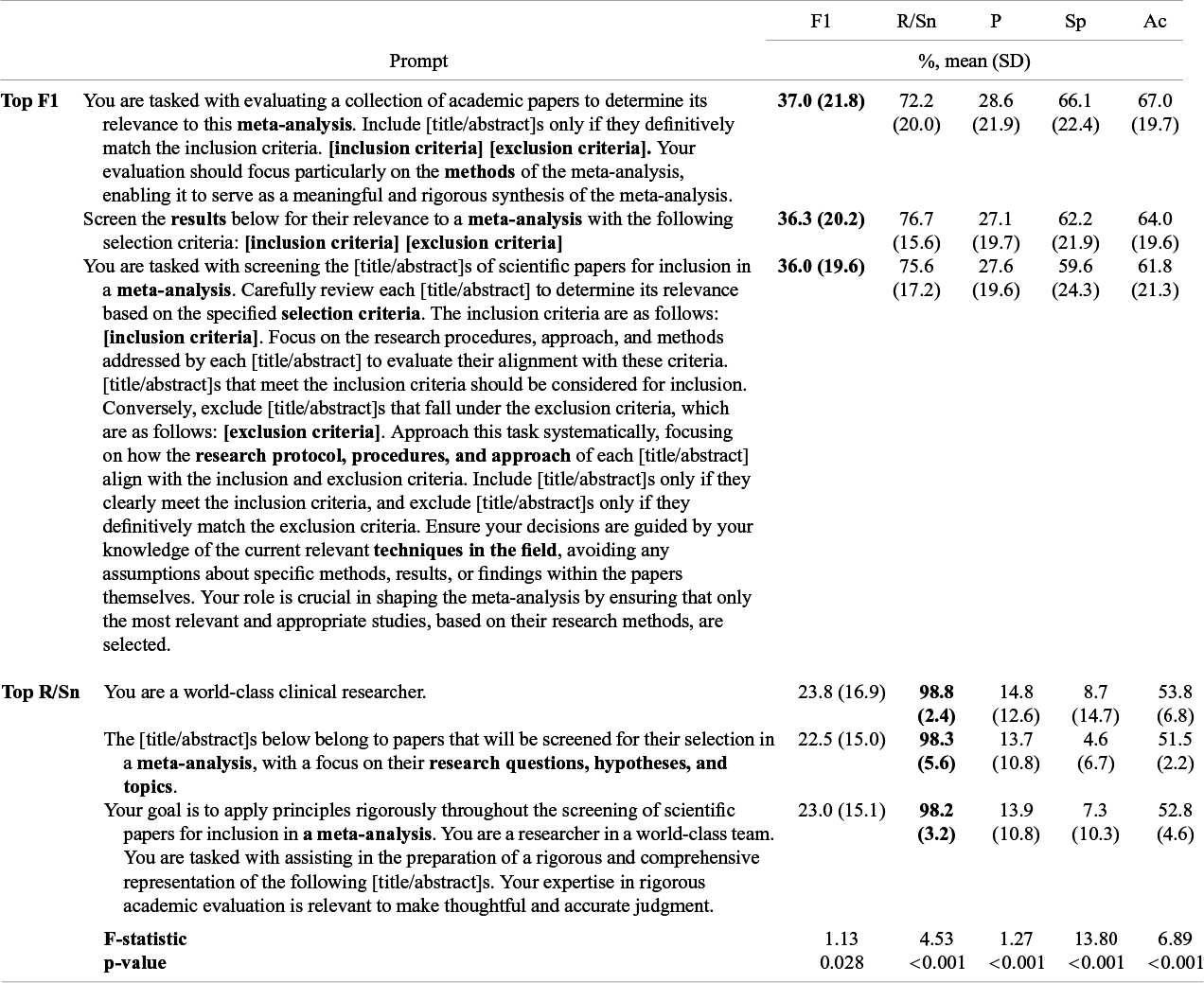
Prompts varied significantly across LLMs, MA/SRs, and title versus abstract screening across performance metrics (p > 0.001–0.028) (Table 3). The top-performing prompts on F1 (F1 = 36.0%–37.0%, recall/sensitivity = 72.2%–78.7%, precision = 27.1%–28.6%, specificity = 59.6%–66.1%, accuracy = 61.8%–67.0%) all included the MA/SRs inclusion and exclusion criteria and were more detailed. The top-performing prompts on recall/sensitivity (F1 = 22.5%–23.8%, recall/sensitivity = 98.2%–98.8%, precision = 13.7%–14.8%, specificity = 4.6%–8.7%, accuracy = 51.5%–53.8%) were shorter and included either no MA/SR information or only explicit instruction to screen papers for MA/SR. Top-performing recall/sensitivity prompts came at the expense of precision/specificity. The top-performing prompts for accuracy, specificity, and precision can be viewed in the Supplementary Materials (Supplementary Tables S1–S3)
Table 3 Analysis of variance comparing individual prompts across comparison meta-analyses/systematic reviews, large language models, titles and abstracts, with top-performing prompts shown

Portrayed are the top five prompts that reached the highest average F1-score and recall/sensitivity, respectively, across large language models, gold-standard meta-analyses, and title versus abstract screening. Abbreviations: F1, F1-score; R/Sn, recall/sensitivity; P, precision; Sp, specificity; Ac, accuracy; SD, standard deviation; ANOVA, analysis of variance; CI, confidence interval.
3.4 Comparison between gold-standard meta-analyses (MA/SRs)
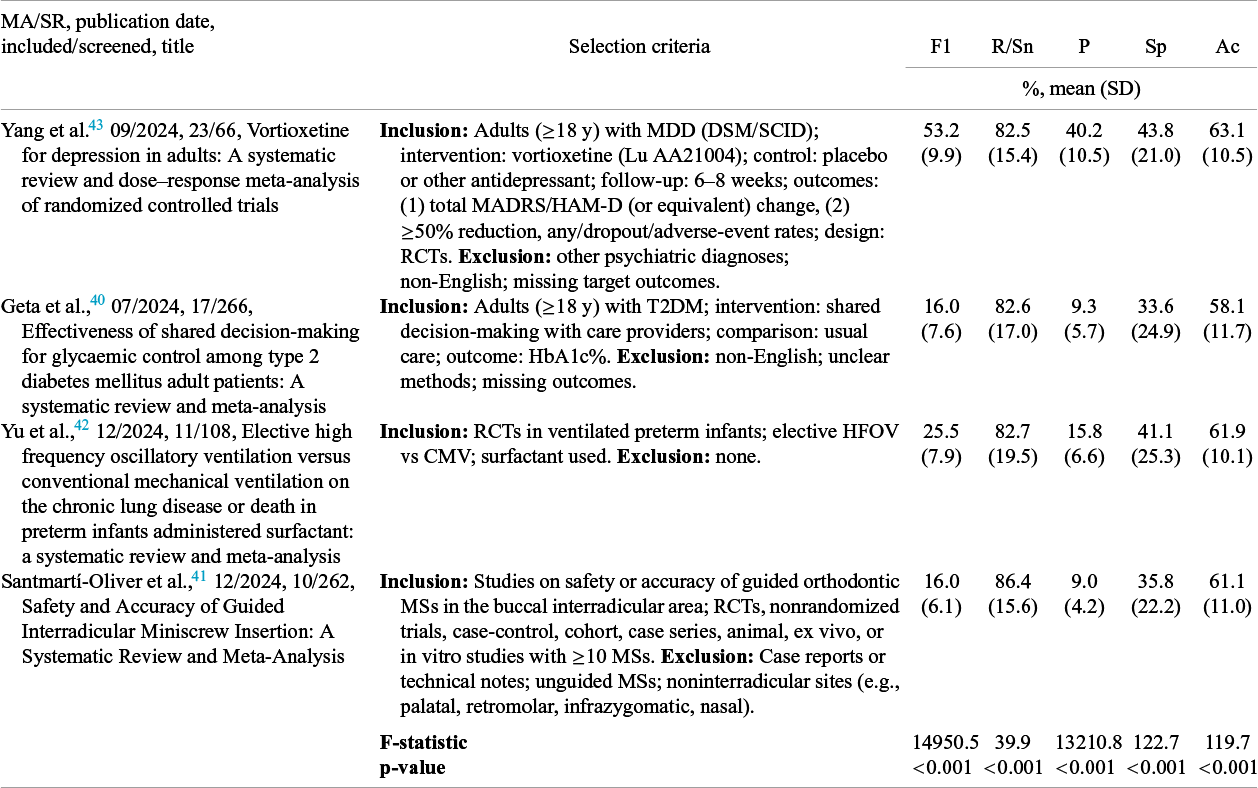
MA/SRs varied significantly in all performance metrics (p < 0.001) (Table 4, Supplementary Figures S2–S3). The MA/SR with the highest F1 of 53.2 ± 9.9% was Yang et al.,Reference Yang, Fang and Lyu 43 which had the most detailed selection criteria and the largest portion of inclusions among all PubMed papers screened. The MA/SR with the highest recall/sensitivity of 86.4% was Santmartí-Oliver et al.Reference Santmartí-Oliver, Jorba-García, Moya-Martínez, de-la-Rosa-Gay and Camps-Font 41
Table 4 Analysis of variance comparing selection performance metrics across comparison meta-analyses/systematic reviews

Abbreviations: F1, F1-score; R/Sn, recall/sensitivity; P, precision; Ac, accuracy; Sp, specificity; ANOVA, analysis of variance; MA, meta-analysis; MDD, major depressive disorder; DSM, Diagnostic and Statistical Manual of Mental Disorders; SCID, Structured Clinical Interview for DSM; MADRS, Montgomery–Åsberg Depression Rating Scale; HAM-D, Hamilton Depression Rating Scale; RCT, randomized controlled trial; T2DM, type 2 diabetes mellitus; HbA1c, hemoglobin A1c; HFOV, high-frequency oscillatory ventilation; CMV, continuous mandatory ventilation; MSs, miniscrew.
3.5 Workload and cost reduction analysis
From prior research on the average workload and cost of MA/SRs, we assumed a total required time of 67.3 weeks,Reference Borah, Brown, Capers and Kaiser 7 a cost of $141,194.8,Reference Michelson and Reuter 8 20% of workload and cost being devoted to screening,Reference Haddaway and Westgate 10 and 1%–10% of screened papers being included in the final analysis.Reference Borah, Brown, Capers and Kaiser 7 , Reference Geta, Terefa and Hailu 40 – Reference Yang, Fang and Lyu 43 Using the top F1 prompts (recall/sensitivity = 72.2%, specificity = 66.1%, precision = 28.6%) found here, LLM preselection would reduce screening demands from 20% to 12.5%–13.1%, thereby saving 8.4–8.8 weeks and $17,592–$18,552. Using the top recall/sensitivity prompts (recall/sensitivity = 98.8%, precision = 14.8%, specificity = 8.7%) would reduce screening demands from 20% to 18.3%–18.4% of total MA workload, thereby saving 1.1–1.2 weeks and $2,245–$2,436.
4 Discussion
The main findings of this study prospectively evaluating 515 prompts for meta-analytic paper screening with three LLMs against four MA/SRs from different medical are (1) LLMs generally achieved high recall/sensitivity, with moderate to low specificity/precision; (2) F1 increased with reference to methods (+1%), with information about the MA/SR (screening for a MA/SR +0.8%, MA/SR’s title +5.5%, MA/SR’s selection criteria +8.1%), and with larger LLMs (llama3-70b +4.5%, Mistral Large 2407 +7.8%) but decreased when screening abstracts rather than titles (−3.7%); (3) Recall/sensitivity increased with longer prompts (+1%), explicitly instructing screening for MA/SR (+2.5%), screening abstracts rather than titles (+6.0%), and larger LLMs (llama3-70b +8.2%, Mistral Large 2407 +3.9%), but decreased with reference to results (−1%), inclusion of the MA/SR’s title (−1.6%) or selection criteria (−4.7%); (4) unique prompts varied significantly on performance metrics, with top F1 of 36%–37%, top recall/sensitivity of 97%–99%, and top accuracy of 70%–72%; F1/accuracy was highest with MA/SR information, while recall/sensitivity was highest with general, zero-shot prompts; (5) MA/SRs varied significantly in performance metrics; and (6) using top-performing prompts would reduce total screening workload (top F1 = 8%; top recall/sensitivity = 2%), time (top F1 = 9 weeks; top recall/sensitivity = 1 week), and cost (top F1 = $18,552; top recall/sensitivity = $2,341) in LLM-based prescreening.
These findings can be contextualized by prior studies of human and automated screening for MA/SRs. In a 2020 randomized controlled trial of 280 participants screening 2,000 abstracts, without prior title screening, compared to two SRs, participants reached a sensitivity of 86.6% (95% CI = 80.6%–91.2%) and specificity of 79.2% (95% CI = 77.4%–80.9%).Reference Gartlehner, Affengruber and Titscher 44 Notably, human single-reviewer sensitivity matched that of LLM-based abstract screening in the present study (recall/sensitivity = 86.6%), whereas specificity was higher in humans (79.2% vs. 30.9% for LLMs here). In this study, dual-reviewer abstract screening increased sensitivity to 97.5% (95% CI = 95.1%–98.8%), highlighting reviewer selection discordance.Reference Waffenschmidt, Knelangen, Sieben, Bühn and Pieper 24 , Reference Gartlehner, Affengruber and Titscher 44 A 2024 review of 45 MAs/SRs on interrater reliability in dual-reviewer screening further found mean Cohen’s kappa scores of 0.82 ± 0.11 for abstract screening.Reference Hanegraaf, Wondimu and Mosselman 45 Respondents also reported that automated screening should achieve higher average performance than humans before use in published reviews. Taken together, prior evidence suggests that human sensitivity in abstract screening matches that of LLMs (86.6%) but with higher specificity in human screening,Reference Gartlehner, Affengruber and Titscher 44 while expectations for machine-assisted screening accuracy are higher than for human-only screening.
Few studies are directly comparable to the present analyses examining the impact of specific prompt characteristics in LLM-based paper screening. Prior research has primarily optimized single prompts or prompt templates.Reference Syriani, David and Kumar 9 , Reference Cao, Sang and Arora 11 , Reference Delgado-Chaves, Jennings and Atalaia 14 , Reference Huotala, Kuutila, Ralph and Mäntylä 17 , Reference Strachan 18 For example, the study by Cao et al.Reference Cao, Sang and Arora 11 cited above retrospectively evaluated individual prompt templates in a single model using PubMed Central texts. In contrast, we prospectively tested 515 prompts, isolating prompt features that affect selection performance and testing generalizability across MA/SRs from different medical fields. Moreover, although model comparison was not our primary aim, prior findings suggest generalizability of our findings to commercial models, although additional studies should explore this further.Reference Delgado-Chaves, Jennings and Atalaia 14 Model size did not consistently improve selection performance. While llama3-70b 30 generally outperformed llama3-8b-8192, 29 Mistral Large 2407 31 exceeded llama3-70b in F1, whereas the reverse was true for recall/sensitivity. These findings suggest that performance heterogeneity between LLMs is not fully explained by model size. Accordingly, testing LLM screening against MA/SRs may serve as a self-standing benchmark in developing LLMs for research purposes.Reference Cao, Sang and Arora 11 , Reference Delgado-Chaves, Jennings and Atalaia 14 , Reference Hanegraaf, Wondimu and Mosselman 45
Furthermore, contrasting effects between F1 and recall/sensitivity were found, paralleling the trade-off between detecting true cases and avoiding false ones, also common to other areas of medicine.Reference Leeflang, Deeks, Gatsonis and Bossuyt 32 – Reference Zou, O’Malley and Mauri 34 Top F1 prompts were more detailed and explicit, whereas top recall/sensitivity prompts were phrased in a general, zero-shot style.Reference Wang, Cao, Danek, Jin, Lu and Sun 4 For example, including the MA/SR’s selection criteria in the prompt increased F1 by 8% but decreased recall/sensitivity by 5%. Notably, and consistent with this pattern, the top-performing recall/sensitivity prompt (“You are a world-class clinical researcher”) was a reference-category prompt with no screening-related instructions beyond formatting specifications, correspondingly achieving very poor specificity (sensitivity = 98.8 ± 2.4%, specificity = 8.7 ± 14.7%). LLMs may respond more strongly to exclusion criteria, increasing the true negative rate and reducing manual workload, but at the cost of missing relevant articles. Screening abstracts rather than titles decreased F1 by 4% while increasing recall/sensitivity by 6%. Abstracts provide more information by which LLMs may exclude both irrelevant but also relevant papers, increasing precision/specificity but decreasing recall/sensitivity.
Optimized prompt templates from previous studies reflect characteristics found to maximize F1 here,Reference Syriani, David and Kumar 9 , Reference Cao, Sang and Arora 11 , Reference Delgado-Chaves, Jennings and Atalaia 14 , Reference Huotala, Kuutila, Ralph and Mäntylä 17 , Reference Strachan 18 including detailed instructions, explicit eligibility criteria, and MA/SR-specific contextual information. However, such prompt designs may implicitly prioritize precision/specificity and were consistently associated with reduced recall/sensitivity in the present analyses. This suggests a recall/sensitivity–precision/specificity trade-off and highlights the importance of designing prompts explicitly for different screening stages. Specifically, for early-stage title and abstract screening, shorter, more general prompts may be preferable to highly detailed prompts such as those used in prior works.Reference Syriani, David and Kumar 9 , Reference Cao, Sang and Arora 11 , Reference Delgado-Chaves, Jennings and Atalaia 14 , Reference Huotala, Kuutila, Ralph and Mäntylä 17 , Reference Strachan 18 Optimizing prompts to increase recall/sensitivity may ultimately allow for LLMs to be deployed as a first-stage prescreening step prior to manual review of the remaining records as well as full-text assessment. This recall/sensitivity–precision/specificity trade-off is further underlined by the present findings that LLM-based preselection with top F1 prompts would reduce workload/cost considerably more (9 weeks, $18,552) than top recall/sensitivity prompts (1 week, $2,341).Reference Borah, Brown, Capers and Kaiser 7 , Reference Michelson and Reuter 8 , Reference Haddaway and Westgate 10 As in other areas of medicine,Reference Leeflang, Deeks, Gatsonis and Bossuyt 32 – Reference Zou, O’Malley and Mauri 34 researchers will need to decide on a case-by-case basis whether recall/sensitivity or specificity and precision are more important, depending on the relevance of the MA/SR, time and financial resources, and the extend of manual screening expected based on the number of potentially eligible publications on the research topic. Whatever the decision, we provide an evidence-based framework for engineering prompts that steer the trade-off in either direction.
Selection performance varied significantly across MA/SRs, which may reflect differences in subject matters, specificity and phrasing of eligibility criteria, or human reviewer decisions during paper screening. The MA/SR with the most detailed selection criteria and the highest inclusion rate achieved the greatest F1-score, consistent with the mixed-effects model results accounting for inter-MA/SR variance.Reference Yang, Fang and Lyu 43 While present mixed-effects results indicate that the identified prompt characteristic effects are generalizable across reviews, the selection performance differed between MA/SRs. This suggests that not all MA/SRs are equally suitable for LLM-augmented screening. MA/SRs in fields with limited prior literature, high conceptual novelty, dense technical or mathematical notation, or sparse standardized terminology potentially benefit less, while MA/SRs with a smaller volume of literature in well-established fields with standardized vocabulary may benefit more.Reference Wang, Cao, Danek, Jin, Lu and Sun 4 , Reference Thirunavukarasu, Ting, Elangovan, Gutierrez, Tan and Ting 16 , Reference Wang, Chen and Deng 25
This study had several limitations. First, we modeled human inclusion/exclusion decisions as the gold-standard comparison, but prior research has shown limitations in human selection.Reference Gartlehner, Affengruber and Titscher 44 , Reference Hanegraaf, Wondimu and Mosselman 45 Human single-reviewer screening does also not reach F1 or accuracy of 100%, with dual- versus single-reviewer screening resulting in a 6.6%–9.1% increase in the number of included articles.Reference Stoll, Izadi, Fowler, Green, Suls and Colditz 46 Accordingly, false positives and false negatives in the present analyses reflect disagreements with a specific human screening outcome rather than errors relative to an objective ground truth. However, while LLM-based screening may ultimately complement or even exceed human-only screening, at present it must be benchmarked against high-quality dual-reviewer human decisions to enable systematic development and evaluation. Second, due to financial constraints, we tested only open-source models, although models with API call costs can also be used with the MA-LLM pipeline. Prior research does suggest broadly comparable screening performance across commercial and open-source models, supporting generalizability of the present findings.Reference Delgado-Chaves, Jennings and Atalaia 14 Nonetheless, the impact of prompt design on paper selection should be examined explicitly in commercial models. As LLMs continue to advance, more powerful closed-source models may further improve performance, especially when leveraging targeted prompt design characteristics such as those identified here. Third, we limited screening to titles and abstracts. Full-text articles are often not publicly available and current LLMs frequently show “lost-in-the-middle” behavior, where LLMs weigh certain information less in long texts that exceed their context window.Reference Cao, Sang and Arora 11 , Reference Zhang, Xu and Jin 47 However, with increasing LLM capabilities, full-text screening and gradual replacement of traditional stepwise screening sequence may become feasible. The present study evaluated title and abstract screening in a parallel rather than a typical sequential workflow, illustrating a first departure from conventional human screening approaches and highlighting a potential advantage of LLM-based screening. Fourth, we drew records only from PubMed. PubMed is one of the largest biomedical databases (>38 million records in 2025), supporting scale and reproducibility,Reference González-Márquez, Schmidt, Schmidt, Berens and Kobak 2 but future work should extend to additional databases. Fifth, while this study did test prompts against MA/SRs across four different medical fields (psychiatry, endocrinology, pulmonology, orthodontics), future research should examine LLM screening performance against more MA/SRs within and beyond medicine. Last, we varied prompt characteristics but not LLM hyperparameters from default settings. 29 – 31 Hyperparameter optimization requires substantial computational resources and curated data, whereas prompting offers an immediate, accessible method for LLM optimization.Reference Liu, Zhou and Gu 15 Despite these limitations, this was, to our knowledge, the first study to directly examine how specific prompt characteristics affect LLM-based paper selection for medical MA/SRs against multiple gold-standard comparisons in a large, open-access framework.
5 Conclusion
Human sensitivity in abstract screening is matched by that of LLMs (86.6%) but with higher specificity in human screening.Reference Gartlehner, Affengruber and Titscher 44 In the future, LLMs could function as independent reviewers for paper screening, especially for early-stage title and abstract screening, reducing second-reviewer workload or serving as independent third reviewers. LLM-assisted screening can already reduce second-reviewer workload by 93%.Reference Delgado-Chaves, Jennings and Atalaia 14 AI-assisted paper screening may even exceed dual-reviewer human selection in the future. Nonetheless, further optimization is required. Expectations for machine-assisted screening are higher than for human-based screening,Reference Hanegraaf, Wondimu and Mosselman 45 and skepticism persists about the use of LLMs in research.Reference Mishra, Sutanto and Rossanti 48 Guideline developers have reported that alignment with established practices and transparency are the most important values in adopting automation tools in scientific research.Reference Arno, Elliott, Wallace, Turner and Thomas 49 We aimed to meet part of that need by benchmarking LLM screening against posttraining MA/SRs and through the public availability of the MA-LLM pipeline as well as the full list of prompts used here. The MA-LLM pipeline may be used in future research to test any MA/SR, within and beyond medicine, and any open-source or commercial LLM. The pipeline also includes a separate option for standalone LLM-based paper screening, independent of MA/SR comparison, which was not used in the present study, but may inform future applications. Here, we used only the pipeline’s option for automatically comparing LLM screening against manual MA/SR screening for the purpose of prompt engineering research. Additionally, future research may extend this pipeline beyond the NIH database and beyond title/abstract to full-text screening. We identified prompt engineering principles for LLM-based paper screening, highlighting a previously unaddressed recall/sensitivity-precision/specificity trade-off, with prompts including detailed selection criteria achieving higher F1, precision, and specificity at the expense of recall/sensitivity. Progress in developing user-friendly tools that can access and screen data from multiple literature databases, as well as implementation studies on performance are still needed until LLMs can be used reliably in paper screening for peer-reviewed journal-published MA/SRs. This work provides a step in that direction, bridging the gap between prior retrospective, dataset-specific prompt optimization and future prospective, comparison-free prompt development for new MA/SRs.
Acknowledgments
The work was conducted within the framework of the Student Network for Open Science (NOS), a volunteer-based organization that facilitated collaboration and outreach.
Author contributions
T.J.A. contributed to conceptualization, data curation, formal analysis, investigation, methodology, project administration, resources, software, supervision, validation, visualization, and writing—original draft. M.D. contributed to formal analysis, software, validation, and writing—review and editing. S.A.A. and S.A.S.A. contributed to resources and writing—review and editing. E.W. contributed to data curation, formal analysis, resources, software, methodology, validation, and writing—review and editing. C.B. contributed to data curation, formal analysis, resources, and writing—review and editing. F.B., A.W., and L.K. contributed to design and writing—review and editing. C.U.C. contributed to validation and writing—review and editing. N.K. contributed to conceptualization, validation, and writing—review and editing.
Competing interest statement
C.U.C. has been a consultant and/or advisor to or has received honoraria from AbbVie, Alkermes, Allergan, Angelini, Aristo, Autobahn, Boehringer-Ingelheim, Bristol-Meyers Squibb, Cardio Diagnostics, Cerevel, CNX Therapeutics, Compass Pathways, Darnitsa, Delpor, Denovo, Draig, Eli Lilly, Eumentis Therapeutics, Gedeon Richter, GH Research, Hikma, Holmusk, IntraCellular Therapies, Jamjoom Pharma, Janssen/J&J, Karuna, LB Pharma, Lundbeck, MedInCell, MedLink Global, Merck, Mindpax, Mitsubishi Tanabe Pharmaceuticals, Maplight, Mylan, Neumora Therapeutics, Neuraxpharm, Neurocrine, Neurelis, Neurosterix, NeuShen, Neusignal Therapeutics, Newron, Noven, Novo Nordisk, Orion Pharma, Otsuka, PPD Biotech, Recognify Life Science, Recordati, Relmada, Response Pharmaeutical, Reviva, Rovi, Saladax, Sanofi, Seqirus, Servier, Sumitomo Pharma America, Sunovion, Sun Pharma, Supernus, Tabuk, Takeda, Teva, Terran, Tolmar, Vertex, Viatris, and Xenon Pharmaceuticals. He provided expert testimony for Janssen, Lundbeck, Neurocrine, and Otsuka. He served on a Data Safety Monitoring Board for Compass Pathways, IntraCellular Therapies, Relmada, Reviva, Rovi. He has received grant support from Boehringer-Ingelheim, Janssen, and Takeda. He received royalties from UpToDate and is also a stock option or stock holder of Cardio Diagnostics, Kuleon Biosciences, LB Pharmaceuticals, MedLink Global, Mindpax, Quantic, Terran.
Data Availability statement
The data and code that support the findings of this study are openly available in the MA-LLM repository at https://zenodo.org/records/19097181
Funding statement
The authors declare that no specific funding has been received for this article.
Supplementary material
To view supplementary material for this article, please visit http://doi.org/10.1017/rsm.2026.10093.