Introduction
From conception to birth, the developing fetus relies on the maternal environment to provide the required nutrients for growth and development. During development, the fetal genome is modulated by intrauterine environmental factors (Wu et al. Reference Wu, Bazer and Cudd2004). Among them, the adequate availability of macro- and micronutrients, such as vitamins and minerals, is essential for the enzymatic and metabolic pathways underlying fetal growth and organ development (Hostetler et al. Reference Hostetler, Kincaid and Mirando2003). These nutrients can modulate gene expression by direct gene regulation or epigenetic changes such as histone modification, DNA methylation, and noncoding RNAs (Beckett et al. Reference Beckett, Yates and Veysey2014; Wessels Reference Wessels, Patel and Preedy2017). Through these mechanisms, maternal nutrient status drives adaptive or responsive fetal programming events in anticipation of postnatal conditions (Diniz et al. Reference Diniz, Reynolds and Ward2024; Maloney and Rees Reference Maloney and Rees2005). Evidence from human studies has shown that nutrient deficiency, even during fertilization, can lead to developmental and metabolic diseases (Christian and Stewart Reference Christian and Stewart2010; Khayat Reference Khayat2017).
Fetal organogenesis starts during early pregnancy. Although the nutrient demand may seem to be negligible during this period, the required amounts for specific nutrients are most likely transitory (Caton et al. Reference Caton, Crouse and McLean2020) depending upon the timing of specific developmental events. Thus, seasonal changes in forage availability and quality may limit the availability of essential nutrients during the prenatal period (NASEM, 2016), with adverse effects on uteroplacental blood flow and nutrient transport (Reynolds et al. Reference Reynolds, Dahlen and Ward2023; Vonnahme et al. Reference Vonnahme, Arndt and Johnson2008). Vitamin and mineral supplementation combined with a moderate rate of body weight (BW) gain during early gestation increased the concentrations of amino acids in maternal serum and allantoic fluid of beef heifers at day 83 when compared with no vitamin and a low rate of BW gain (Menezes et al. Reference Menezes, McCarthy and Kassetas2021). Transcriptome analyses of fetal and maternal placental tissues from the same study reported changes in the abundance of genes involved with energy metabolism and nutrient transport (Diniz et al. Reference Diniz, Reynolds and Borowicz2021b). Although no differences were observed in the expression of angiogenic factors or placental vascular area density at day 83, vitamin and mineral supplementation through gestation increased placental vascularity at parturition (Dávila Ruiz et al. Reference Dávila Ruiz, Dahlen and McCarthy2024). Placental adaptation in response to maternal nutrient stores can influence the immediate availability of substrates to fetal tissues, while programming fetal development and physiology (Reynolds et al. Reference Reynolds, Dahlen and Ward2023).
Nutrient partitioning is tissue-dependent, with the fetal brain receiving high priority, followed by the fetal liver and muscle (Du et al. Reference Du, Tong and Zhao2009; Sookoian et al. Reference Sookoian, Gianotti and Burgueño2013). Periconceptual vitamin and mineral supplementation combined with different rates of BW gain (71 days pre-breeding to day 83 of gestation) affected the expression of energy- and lipid-related genes in the fetal liver as well as their metabolome and lipidome (Crouse et al. Reference Crouse, McCarthy and Menezes2022; Menezes et al. Reference Menezes, Dahlen and McCarthy2023). In the same experimental cohort, McCarthy et al. (Reference McCarthy, B Menezes and Kassetas2022) reported increased concentrations of trace minerals in the liver and muscle of fetuses from supplemented dams. Likewise, vitamin supplementation was associated with heavier fetal livers (Menezes et al. Reference Menezes, McCarthy and Kassetas2022). When analyzed as a percentage of fetal BW, fetal liver weight was greater in fetuses from LG heifers than in those from the MG group, again suggesting its priority for nutrient partitioning. We have demonstrated that global nutrient restriction in heifers during the first 50 days of pregnancy results in differential expression of key metabolic and epigenetic genes in the cerebrum, liver, and muscle tissues (Crouse et al. Reference Crouse, Caton and Cushman2019). Interestingly, maternal nutrition influences the reprogramming of master regulators involved in the mTOR and phosphatidylinositol 3-kinase–protein kinase B (PI3K/Akt) signaling pathways (Diniz et al. Reference Diniz, Crouse and Cushman2021a), which regulate neuronal fate through protein synthesis and energy homeostasis (Takei and Nawa Reference Takei and Nawa2014).
The hypothalamus coordinates feeding, energy expenditure, and nutrient partitioning (Manceau et al. Reference Manceau, Majeur and Alquier2020). Although this relationship remains largely unexplored in livestock, evidence from rodents suggests that poor maternal nutrition may program feeding behaviors and temperament through changes in cerebral function (Georgieff Reference Georgieff2007). Hurlbert et al. (Reference Hurlbert, Baumgaertner and Menezes2024) reported that F1 generation heifers born to dams supplemented with vitamins during gestation exhibited different feeding behaviors and activity levels compared with those from non-supplemented dams. Moderate rate of BW gain (0.79 kg d−1 gain – MG, vs. 0.28 kg d−1 gain – LG) from breeding to day 84 of pregnancy resulted in changes that extend to F2 fetuses (Baumgaertner et al. Reference Baumgaertner, Darío Ramírez-Zamudio and Clara2024a, Reference Baumgaertner, Menezes and Diniz2024b). Baumgaertner et al. (Reference Baumgaertner, Menezes and Diniz2024b) reported that calves born to MG dams were 2.14 kg heavier at birth compared with the LG group. Furthermore, during postnatal development, heifers from MG dams continued to have greater BW gain and feed efficiency (Baumgaertner et al. Reference Baumgaertner, Menezes and Diniz2024b). Metabolic differences were also observed in F1 heifers, with greater concentrations of glucose observed in heifers from MG dams (Baumgaertner et al. Reference Baumgaertner, Darío Ramírez-Zamudio and Clara2024a). Although no effects of F0 dam nutrition on F2 fetal morphometric characteristics were observed, the weight of the mammary gland relative to fetal BW was greater in F2 fetuses born to LG dams (Baumgaertner et al. Reference Baumgaertner, Darío Ramírez-Zamudio and Clara2024a).
From our previous research, we hypothesized that maternal vitamin and mineral supplementation from pre-breeding to day 83 of pregnancy and moderate rates of BW gain would be associated with altered fetal brain and muscle transcriptomes. To test this hypothesis, we carried out a genome-wide transcriptomic approach to determine the effects of maternal vitamin and mineral supplementation and the rate of maternal BW gain during early gestation on fetal brain and muscle gene expression. Furthermore, we retrieved the biological processes (BPs) and pathways underlying tissue development and function. Our results show that maternal diet was associated with molecular changes in the fetal brain and muscle, manifested in the transcriptomic profiles of key metabolic and developmental genes.
Materials and methods
The experimental design and animal procedures were approved by the North Dakota State University Institutional Animal Care and Use Committee (IACUC #A19012).
Animal management and experimental design
Thirty-five crossbred Angus heifers were used in this study, as described previously (McCarthy et al. Reference McCarthy, B Menezes and Kassetas2022; Menezes et al. Reference Menezes, McCarthy and Kassetas2022). Briefly, heifers were randomly assigned to a 2 × 2 factorial arrangement, with the main factors of vitamin and mineral supplementation (VTM vs. NoVTM) and the rate of BW gain (low gain [LG, avg. daily gain of 0.28 kg d−1] or moderate gain [MG, avg. daily gain of 0.79 kg d−1]).
The VTM treatment was initiated at least 71 days before artificial insemination (AI) by supplementing heifers with 113 g heifer–1 d−1 of a pelleted mineral premix (Purina Wind & Rain Storm All-Season 7.5 Complete). At breeding and within VTM groups, heifers were assigned to one of two rates of gain (LG or MG). Thus, the treatments were as follows: NoVTM_LG (no vitamin and mineral supplementation under low gain, n = 9); VTM_LG (vitamin and mineral supplementation under LG, n = 9); NoVTM_MG (no vitamin and mineral supplementation under MG, n = 9); and VTM_MG (vitamin and mineral supplementation and MG, n = 8). The diet and mineral premix compositions were previously reported (Menezes et al. Reference Menezes, McCarthy and Kassetas2022). Diets were delivered once daily via a total mixed ration (TMR) in an electronic head-gate facility (American Calan; Northwood, NH, USA), which allowed individual heifer access to each head-gate. While heifers on the LG treatment were maintained on the basal TMR, heifers under MG were supplemented with a blend of ground corn, dried distiller’s grains plus solubles, wheat midds, fish oil, urea, and ethoxyquin top-dressed on the TMR at 0.58% of BW as-fed daily. Heifers in the NoVTM treatment received a pelleted carrier product fed at 0.45 kg−1 heifer−1 day−1 with no added vitamins or minerals. The diet and premix compositions are described in Supplementary Table S1.
Heifers were bred by AI using female-sexed semen from a single sire after a 7-d CO-Synch + CIDR estrus synchronization protocol (Lamb et al. Reference Lamb, Dahlen and Larson2010). Pregnancy was diagnosed 35 days after AI, and fetal sex was determined on day 65, both via transrectal ultrasonography. The experiment finished on day 83 ± 0.27 after breeding, when the fetus was removed through ovariohysterectomy as described by McLean et al. (Reference McLean, Dahlen and Borowicz2016). This time point was defined based on the fact that fetal organogenesis has already finished and secondary myogenesis is beginning (Du et al. Reference Du, Tong and Zhao2009).
Sample collection and RNA isolation
After ovariohysterectomy, fetal muscle (Longissimus dorsi) and brain tissues were collected as described elsewhere (Crouse et al. Reference Crouse, Caton and Cushman2019). The muscle tissue was snap-frozen on dry ice while the brain was stored in Allprotect Tissue Reagent (Qiagen, Germantown, MA, USA), and both were then stored at −80°C until RNA isolation.
Total RNA from the brain and muscle tissues (30 mg per sample) was isolated following the RNeasy Plus Universal Mini Kit (Qiagen, Germantown, MA, USA) protocol. Sample integrity and purity analyses were determined through the Agilent 2100 Bioanalyzer and agarose gel electrophoresis. Samples meeting the established quality thresholds according to Novogene (RNA Integrity Number - RIN value ≥ 5.0, flat baseline, and no detectable degradation or contamination) were sequenced. The samples had average RINs of 8.3 for muscle and 8.8 for brain. Considering that, 31 samples (n = 8 per group, except VTM_MG, n = 7) proceeded to library preparation. Strand-specific RNA libraries were prepared using the NEBNext Ultra II Directional RNA Library Prep Kit for Illumina (New England BioLabs, Ipswich, MA, USA). Libraries were sequenced on the Illumina NovaSeq 6000 platform, according to Novogene Co. (Nanjing, China) recommendations. A minimum depth of 20 million stranded, paired-end reads per sample was generated.
RNA-Seq quality control, mapping, and quantification
Raw read quality control (QC) and sequencing statistics were assessed using FastQC v0.11.8, and the reports were aggregated by MultiQC v1.9 (Ewels et al. Reference Ewels, Magnusson and Lundin2016). After the QC, reads were mapped to the Bos taurus reference genome (ARS-UCD 1.2) (Rosen et al. Reference Rosen, Bickhart and Schnabel2020) using the STAR aligner v. 2.7.3a (Dobin et al. Reference Dobin, Davis and Schlesinger2013) and the gene annotation file (release 100) retrieved from the Ensembl database. The –quantMode GeneCounts flag from STAR was used to retrieve the raw counts per gene. MultiQC, NOISeq v.2.26.0 (Tarazona et al. Reference Tarazona, Furió-Tarí and Turrà2015), and edgeR v.3.24.3 (Robinson et al. Reference Robinson, McCarthy and Smyth2010) were used for post-mapping QC.
Genes not expressed or lowly expressed (10 counts per million in 70% of samples) were filtered out using the edgeR filterByExpr function (Chen et al. Reference Chen, Lun and Smyth2016). We applied a generalized linear model based on the negative binomial distribution from edgeR to perform the differential gene expression analysis (Robinson et al. Reference Robinson, McCarthy and Smyth2010). The RNA isolation batch (n = 2) was included as a covariate in the model. Pairwise comparisons among the four treatment groups were used to test (1) the main effects of gain (MG vs LG; VTM_MG vs. VTM_LG; and NoVTM_MG vs. NoVTM_LG; (2) the main effects of VTM (VTM vs. NoVTM; VTM_LG vs. NoVTM_LG; and VTM_MG vs. NoVTM_MG); and (3) the crossed comparisons between groups differing in both VTM supplementation and maternal gain (VTM_LG vs. NoVTM_MG and VTM_MG vs. NoVTM_LG). Differentially expressed genes (DEGs) were identified after multiple testing correction of the p-values based on the Benjamini–Hochberg methodology (false discovery rate [FDR] ≤ 0.1).
Overrepresented BPs and KEGG pathways underlying the DEGs were identified using the WebGestalt 2024 web tool based on individual and meta-list overrepresentation analyses (Elizarraras et al. Reference Elizarraras, Liao and Shi2024). Overrepresented terms were retrieved after p-value multiple testing correction (FDR ≤ 0.05).
Results
We applied a differential gene expression approach to characterize the transcriptomic responses of fetal brain and muscle at day 83 of gestation to maternal vitamin and mineral supplementation, combined or not with the two rates of BW gain. We have previously reported the main effects of maternal diet on dam and fetal development (McCarthy et al. Reference McCarthy, B Menezes and Kassetas2022; Menezes et al. Reference Menezes, McCarthy and Kassetas2022). By design, the moderate rate of gain (MG vs. LG) led to greater average daily gain and dam BW at day 83 of gestation (Menezes et al. Reference Menezes, McCarthy and Kassetas2022). However, no significant effects were observed on dam performance or dry matter intake due to the VTM treatment (Menezes et al. Reference Menezes, McCarthy and Kassetas2022). Fetal BW and most fetal organ measurements were not affected by the VTM supplementation, rate of gain, or their interaction. However, VTM supplementation led to greater fetal liver mass, whereas fetal intestinal weight was affected by a VTM × GAIN interaction (Menezes et al. Reference Menezes, McCarthy and Kassetas2022).
Greater concentrations of Se, Cu, and Co were observed in the liver of fetuses from VTM-supplemented dams, whereas Mo and Co were greater in fetuses from LG compared with MG dams (McCarthy et al. Reference McCarthy, B Menezes and Kassetas2022). In fetal muscle, Se and Zn concentrations were greater in the VTM group, and in addition, maternal rate gain influenced Zn concentration, with LG fetuses showing greater Zn than MG fetuses (McCarthy et al. Reference McCarthy, B Menezes and Kassetas2022).
Fetal brain and muscle gene expression were affected by maternal diet
Herein, the transcriptomic data from 31 female fetuses at day 83 of gestation for brain and muscle tissues were analyzed. Paired-end sequencing resulted in, on average, 21.07 and 20.6 million cleaned reads for the brain and muscle, respectively. An average mapping rate of 96.7% and 97% was obtained from the brain and muscle reads, respectively. The number of reads and mapping statistics per sample are shown in the Supplementary Table S2. After post-mapping QC (see methods), out of 27,607 genes, 14,788 genes from the brain and 14,765 genes from the muscle were kept for DEG analysis.
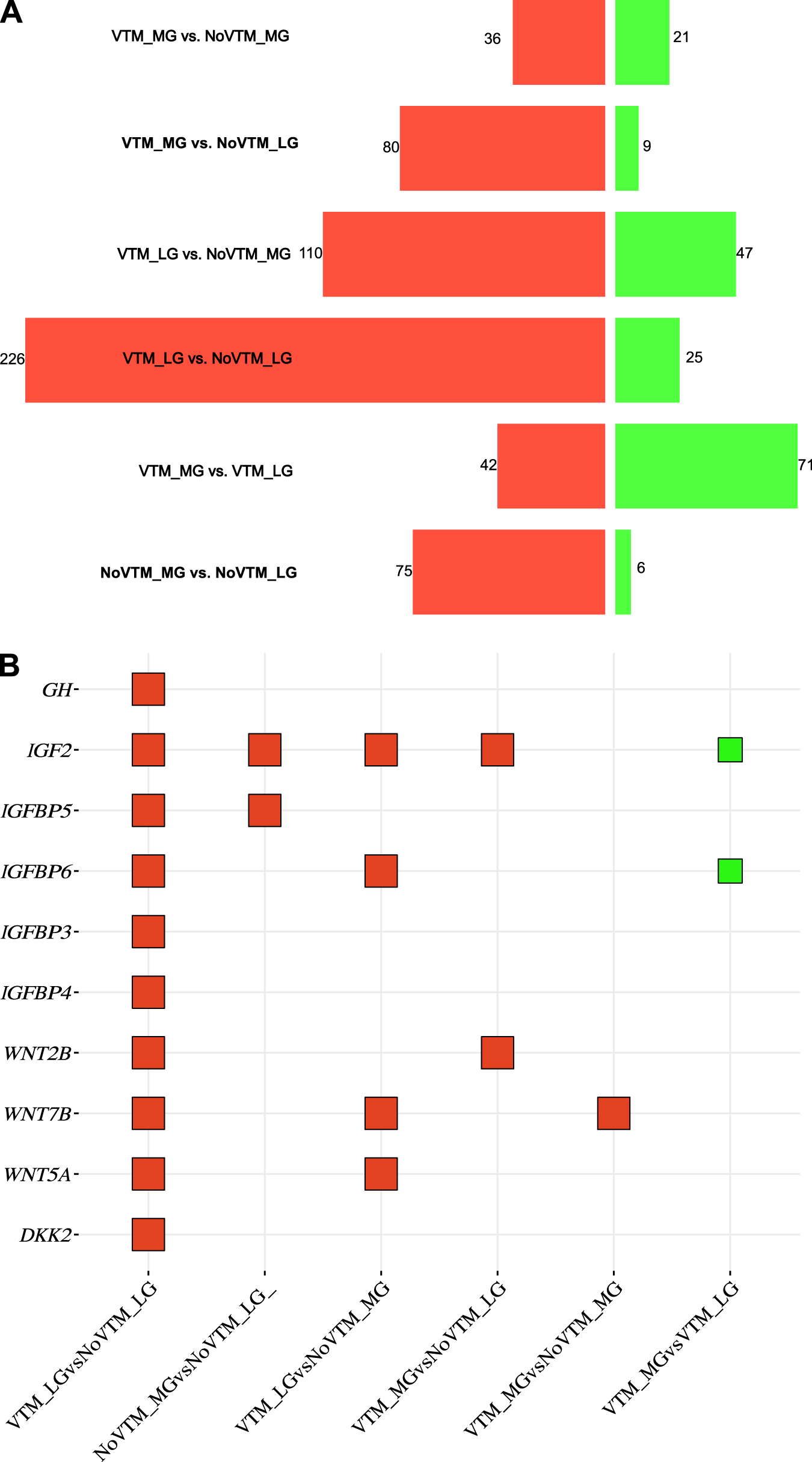
For the fetal brain, we identified 748 DEGs (corresponding to 399 unique genes) across all six contrasts (FDR ≤ 0.1) (Fig. 1A; Supplementary Table S3). Among them, 569 were upregulated and 179 were downregulated. Among these genes, we can highlight those involved with nutrient transport (AQP8, SLC13A4, SLC6A13, and SLC22A6) and the transforming growth factor-beta (TGF-β) superfamily (BMP5 and TGFBR3). Upregulated genes affected by the VTM supplementation under LG (VTM_LG vs. NoVTM_LG) included structural genes from the collagen family (COL1A2, COL8A1, COL18A1, COL6A1, and COL6A2) and those involved in neuronal development, such as WNT2B, WNT5A, WNT7B, and DKK2 (Fig. 1B). For the main effect of gain (VTM_MG vs. VTM_LG), most genes were downregulated in the VTM_MG group. Interestingly, 10 genes (Supplementary Table S3) from the collagen family were downregulated in the VTM_MG group; conversely, mitochondrial genes, such as CYTB, COX1, COX2, and COX3, were upregulated. Contrasts with major changes included VTM_LG vs. NoVTM_LG (main effect of VTM; n = 251) and VTM_LG vs. NoVTM_MG (n = 157). Most of the shared genes (n = 49) were observed among NoVTM_MG vs. NoVTM_LG, VTM_LG vs. NoVTM_LG, and VTM_MG vs. NoVTM_LG (Fig. 1C).
Transcriptomic profile of the brain tissue of female fetuses at day 83 of gestation from heifers receiving or not receiving vitamin–mineral supplementation (VTM or NoVTM) and fed to achieve different rates of gain (low gain [LG] or moderate gain [MG]). (A) Number of differentially expressed genes per contrast. Up- and downregulated genes are represented by red and green bar colors, respectively. Significant DEGs were defined based on FDR ≤ 0.1. (B) Square bubble plot showing the expression profile of selected genes across contrasts involved with the nutrient-sensing and neurogenesis pathways. Up- and downregulated genes are represented by red and green colors, respectively. (C) Number of shared DEGs across contrasts. The UpSet plot provides the overlap between the sets of DEGs from different comparisons. The number of overlapping genes is represented in the vertical bars. The overlapping groups are represented in the dot plots, and the horizontal bar graph reports the total of DEGs. The treatments were arranged as follows: NoVTM_LG – no vitamin and mineral supplementation and LG; VTM_LG – vitamin and mineral supplementation and LG; NoVTM_MG – no vitamin and mineral supplementation and MG; and VTM_MG – vitamin and mineral supplementation and moderate gain. The butterfly bar (A) and square bubble plots (B) were created on the SRplot platform (Tang et al. Reference Tarazona, Furió-Tarí and Turrà2023).


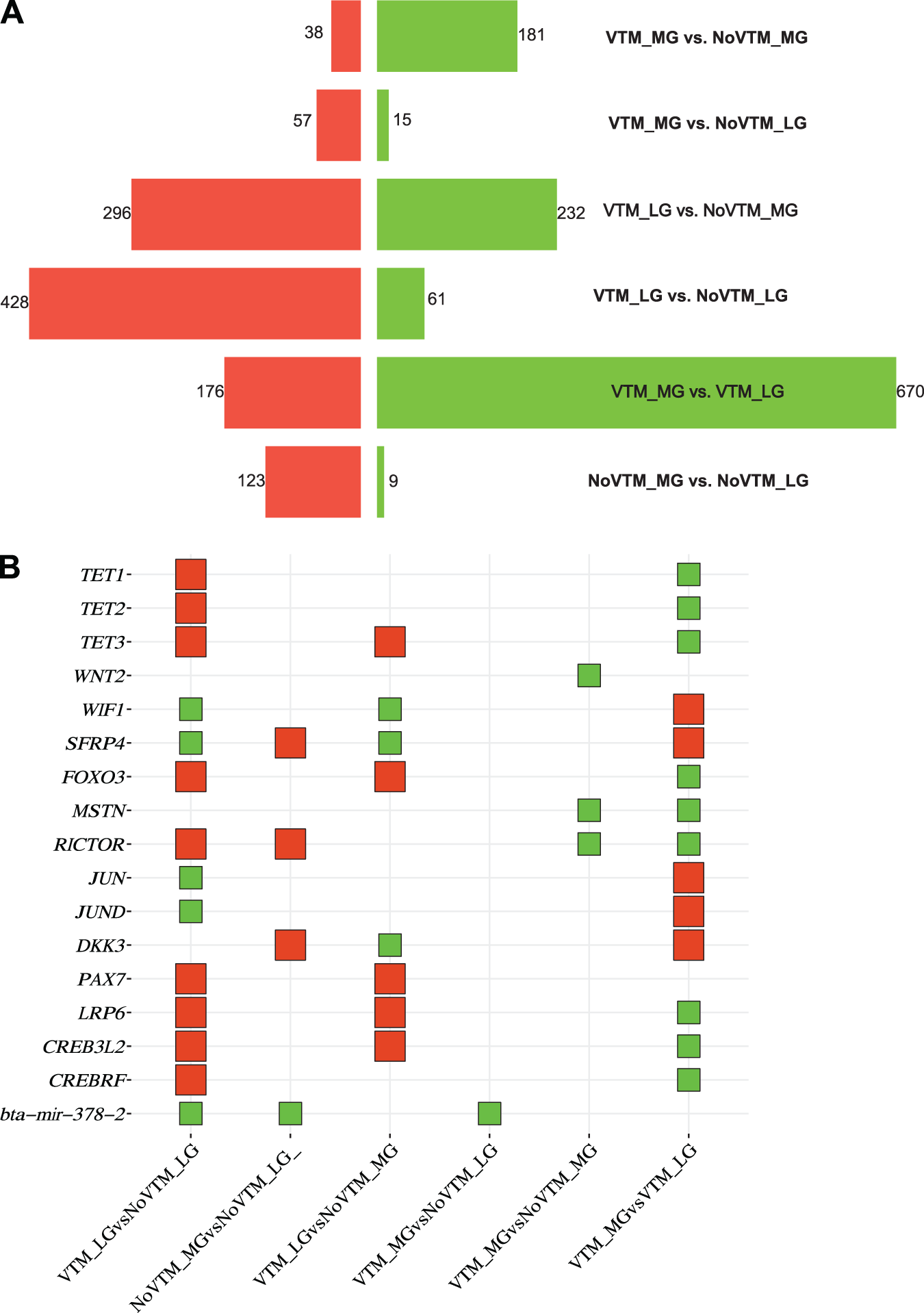
For the fetal muscle, we identified 2,286 DEGs across all six comparisons corresponding to 1,273 unique genes (FDR ≤ 0.1) (Fig. 2A, Supplementary Table S4). The main effect of gain (VTM_MG vs. VTM_LG) resulted in the greatest number of DEGs. The DEGs were mostly downregulated in the MG group, which included TET1, TET2, and TET3. However, VTM supplementation under LG increased gene upregulation (n = 428). Upregulated genes in the VTM-supplemented group coding for DNA demethylation enzymes included the TET1, TET2, and TET3 genes. Genes involved with muscle development and nutrient sensing, such as WIF1, FoxO3, and RICTOR, were also upregulated in the VTM_LG vs. NoVTM_LG comparison (Fig. 2B). Under MG, VTM supplementation led to a downregulation of 181 genes in the VTM_MG group, including MSTN and RICOR. We found the greatest number of shared genes (n = 372) between the contrasts VTM_LG vs. NoVTM_LG and VTM_MG vs. VTM_LG (Fig. 2C). Among the genes were IREB2, TFRC, TET1, TET2, and TET3, which were downregulated on the first contrast but upregulated on the second. The overlap between the DEG lists from brain and muscle tissues identified 95 shared genes, underlying developmental, organismal, and homeostatic processes. Shared genes between tissues included GNAS, LAMA2, MYH8, and those from the collagen family (COL1A1, COL1A2, COL11A2, and COL9A2).
Transcriptomic profile of the muscle tissue of female fetuses at day 83 of gestation from heifers receiving or not receiving vitamin–mineral supplementation (VTM or NoVTM) and fed to achieve different rates of gain (low gain [LG] or moderate gain [MG]). (A) Number of differentially expressed genes per contrast. Up- and downregulated genes are represented by red and green bar colors, respectively. Significant DEGs were defined based on FDR ≤ 0.1. (B) Square bubble plot showing the expression profile of selected genes across contrasts involved with muscle development and epigenetic modification. Up- and downregulated genes are represented by red and green colors, respectively; (C) Number of shared DEGs across contrasts. The UpSet plot provides the overlap between the sets of DEGs from different comparisons. The number of overlapping genes is represented in the vertical bars. The overlapping groups are represented in the dot plots, and the horizontal bar graph reports the total of DEGs. The treatments were arranged as follows: NoVTM_LG – no vitamin and mineral supplementation and low gain; VTM_LG – vitamin and mineral supplementation and LG; NoVTM_MG – no vitamin and mineral supplementation and MG; and VTM_MG – vitamin and mineral supplementation and MG. The butterfly bar (A) and square bubble plots (B) were created on the SRplot platform (Tang et al. Reference Tarazona, Furió-Tarí and Turrà2023).


Functional overrepresentation analyses
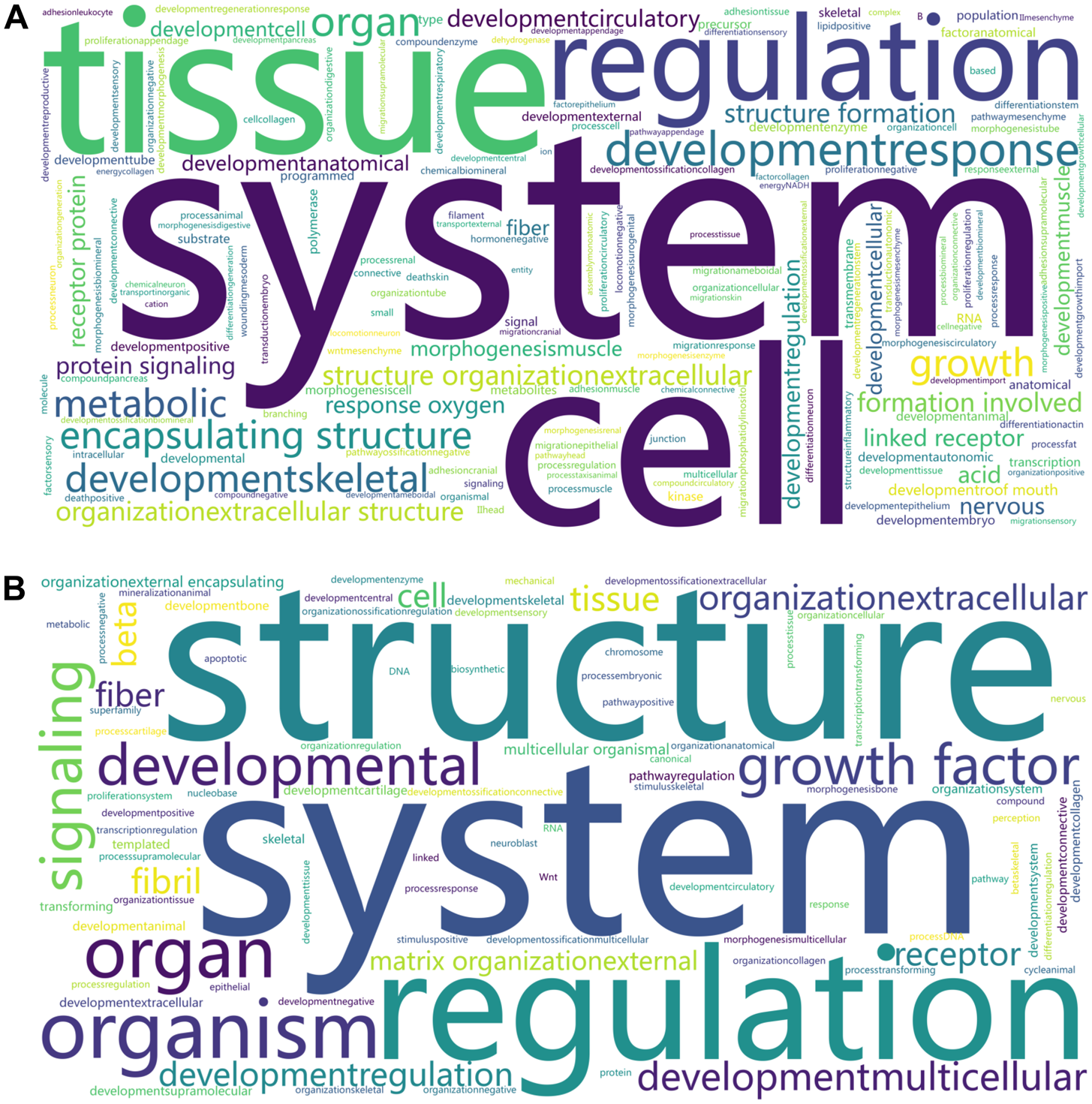
Overrepresented KEGG pathways and BPs underlying the DEGs were retrieved for each comparison within each tissue. From the brain, a multi-list analysis based on all six contrasts resulted in 52 overrepresented BPs and 17 KEGG pathways (meta-FDR ≤ 0.05). Anatomical structure formation involved in morphogenesis, tube development, animal organ morphogenesis, and neuron migration were among the significant BP terms. We identified protein digestion and absorption, ECM–receptor interaction, focal adhesion, and PI3K–Akt signaling pathway among the KEGG pathways. We performed a word cloud analysis of each overrepresented gene list (Fig. 3A). The main words were system, cell, tissue, regulation, and developmental response. A complete list of overrepresented BPs and pathways is reported in Supplementary Tables S5 and S6.
Word cloud of the overrepresented biological processes terms associated with the differential expression of brain (A) and muscle (B) genes of female fetuses at day 83 of gestation from heifers receiving or not receiving vitamin–mineral supplementation (VTM or NoVTM) and fed to achieve different rates of gain (low gain [LG] or moderate gain [MG]). The larger the font size, the more frequently the word is in the overrepresented terms. Significant BP terms (FDR ≤ 0.05) from all the comparison lists (Supplementary Tables S5 and S6) within each tissue were used to draw the cloud using the SRplot platform (Tang et al. Reference Tang, Chen and Huang2023).

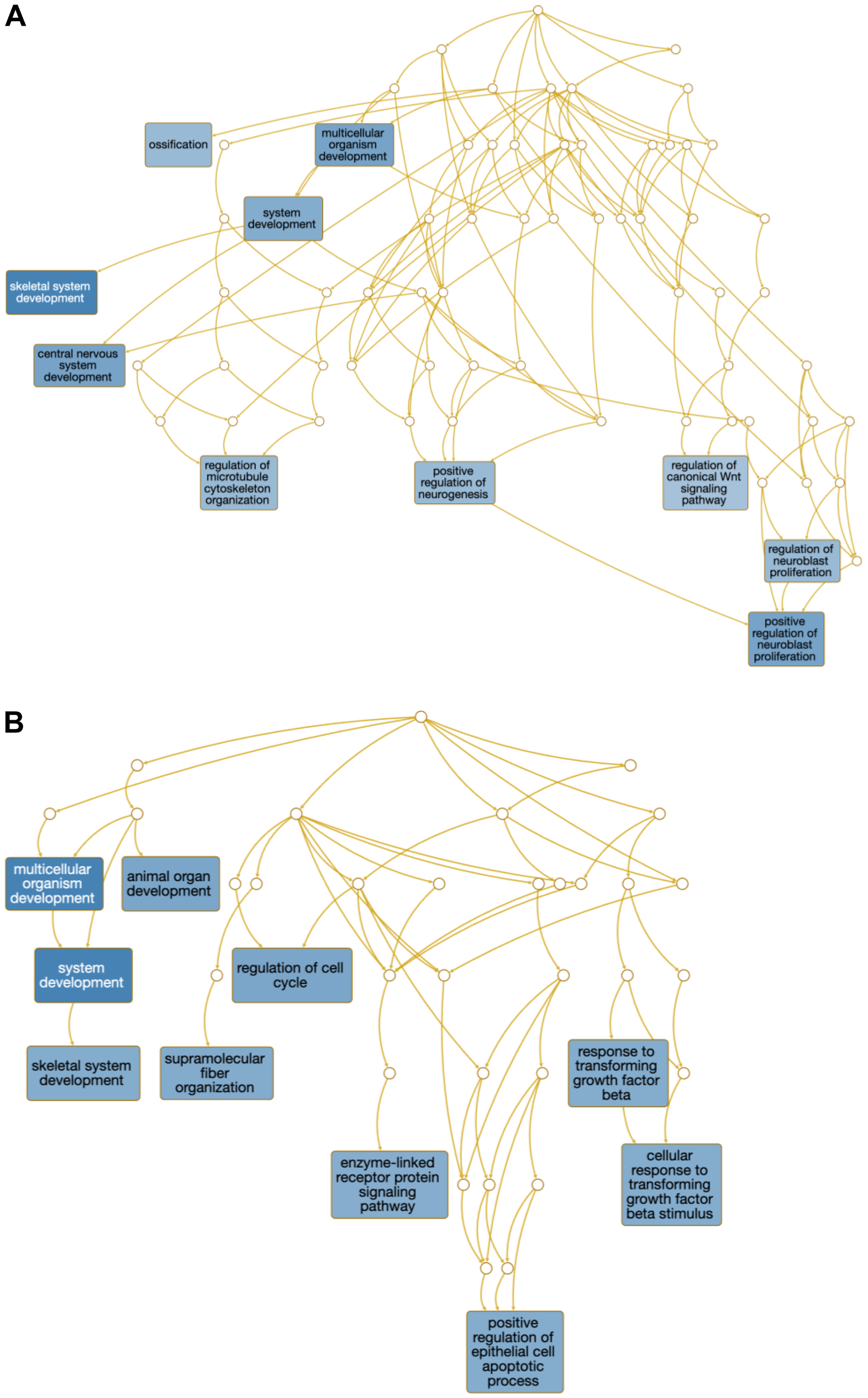
Regarding the DEG lists from the muscle, we identified 78 BP-related terms across all comparisons (FDR ≤ 0.05, Supplementary Table S6). No significant GO terms were identified for the VTM_MG vs. NoVTM_MG. The number of BP terms per contrast ranged from 5 to 20. To summarize the processes across lists, we generated a word cloud analysis from each overrepresented gene list (Fig. 3B). The main words were structure, system, and regulation. Additionally, skeletal system development, tissue development, and animal organ development were identified as overrepresented through the multi-list analysis on WebGestalt (Meta-FDR ≤ 0.05). We also mined the overrepresented BPs underlying the main effects of vitamin supplementation under LG (VTM_LG vs. NoVTM_LG) and MG (VTM_MG vs. VTM_LG). Overrepresented terms included skeletal muscle development and regulation of the canonical Wnt signaling pathway (Fig. 4). The multi-list analysis retrieved eight significant KEGG pathways, including PI3K–Akt, TGF-beta, and Wnt signaling. A complete list of overrepresented BPs and pathways is reported in Supplementary Tables S7 and S8.
Overrepresented biological processes of differentially expressed genes from the muscle of female fetuses at day 83 of gestation from heifers receiving or not receiving vitamin–mineral supplementation (VTM or NoVTM) and fed to achieve different rates of gain (low gain [LG] or moderate gain [MG]) during early gestation. Overrepresented biological processes from the (A) VTM_LG vs. NoVTM_LG and (B) VTM_MG vs. VTM_LG comparisons. Only the top 10 significant terms containing five or more genes are shown. The darker the blue background color of the squares, the more significant is the term (FDR ≤ 0.05).

Discussion
We investigated the transcriptome changes in fetal brain and muscle to determine the effects of maternal vitamin and mineral supplementation and the rate of maternal BW gain from pre-breeding to day 83 of gestation. We previously observed changes in the intrauterine environment that affected the nutrients available for fetal development, although no differences were observed in fetal BW at day 83 (McCarthy et al. Reference McCarthy, B Menezes and Kassetas2022; Menezes et al. Reference Menezes, McCarthy and Kassetas2021, Reference Menezes, McCarthy and Kassetas2022). Maternal diet was associated with molecular changes in the fetal brain and muscle, as evidenced by the transcriptomic profiles of key metabolic and developmental genes. Previously, we reported changes in the placental transcriptome and fetal liver (Diniz et al. Reference Diniz, Reynolds and Borowicz2021b, Reference Diniz, Ward and McCarthy2023). Additionally, micronutrients, such as Se and Zn, had greater concentrations in the muscle of fetuses from VTM dams. Likewise, fetuses from the LG treatment had a greater concentration of Zn than those from the MG treatment. These minerals are essential for synthesizing anabolic hormones and their receptors, tissue development, metabolism, and cognitive function (Adebayo et al. Reference Adebayo, Adenuga and Sandhir2016; Rahman et al. Reference Rahman, Rao and Aldughpassi2025; Sahin et al. Reference Sahin, Orhan and Tuzcu2019). Although the fetus may be buffered by the dam against nutrient imbalances, we observed that maternal liver concentrations of Se and Zn decreased from the beginning to the end of the experiment (day 83), even in the VTM-supplemented group (McCarthy et al. Reference McCarthy, B Menezes and Kassetas2022). This change may reflect the metabolic status of nutrients in fetal tissues, as Hurlbert et al. (Reference Hurlbert, Baumgaertner and Menezes2024) reported decreased hepatic Se concentrations in VTM-supplemented dams from day 84 post-breeding through calving.
Shared DEGs between the brain and muscle were overrepresented in GO terms related to tissue development
While the findings suggest that molecular changes were tissue-specific, the 95 overlapping DEGs across tissues were involved with tissue structure and development. Overrepresented GO terms included tissue development, extracellular matrix (ECM) organization, and animal organ development. Shared genes included LAMA2, MYH8, and those from the collagen family (COL1A1, COL1A2, COL11A2, and COL9A2). However, we found no differences in the fetal brain and hindlimb mass in response to maternal diet (Menezes et al. Reference Menezes, McCarthy and Kassetas2022). Likewise, Peñagaricano et al. (Reference Peñagaricano, Wang and Rosa2014) reported enrichment of similar BPs in ewe fetuses at day 130 of gestation, showing that maternal feed sources during mid-to-late gestation influenced genes involved in tissue and organ development. Pillai et al. (Reference Pillai, Sereda and Hoffman2016) reported that maternal nutrient restriction or overnutrition during gestation reduced offspring mesenchymal stem cell proliferation by 50%. In the current study, morphological and histological parameters of the brain and muscle were not evaluated, warranting further investigation to determine whether the observed molecular changes are reflected in tissue structure and function.
Interestingly, the muscle was more responsive to maternal diet than the brain, as more genes were differentially expressed. Similarly, Crouse et al. (Reference Crouse, Caton and Cushman2019) reported more DEGs in the fetal liver and muscle tissues compared with the cerebrum of nutrient-restricted fetuses on day 50 of gestation. The brain has a high priority in nutrient partitioning (Du et al. Reference Du, Tong and Zhao2009) and controls this process through hormonal and neural signals (Manceau et al. Reference Manceau, Majeur and Alquier2020). Thus, the increased transcriptional changes observed in the muscle likely enable muscle cells to adapt to the maternal nutritional environment (Sartori et al. Reference Sartori, Romanello and Sandri2021). Differences in maternal gain (MG vs. LG) also led to greater transcriptional changes, with most of the genes downregulated in the VTM_MG vs. VTM_LG. These findings are likely a potential compensatory mechanism, as the genes were upregulated in fetuses from LG dams. Although VTM supplementation under MG conditions led to differences in gene expression in both tissues, supplementation under LG triggered extensive transcriptional reprogramming, particularly in fetal muscle (428 up- and 61 downregulated genes), indicating that the effects of VTM are amplified under nutrient-restricted conditions.
Nutrient-sensing pathways were overrepresented in the brain in response to maternal nutrition
Nutrient-sensing pathways are responsive to fluctuations in nutrient availability. We identified DEGs in the brain, including GHR and IGF2, associated with the PI3K–Akt and Wnt signaling pathways. The growth hormone–insulin-like growth factor (GH–IGF) axis plays a central role in mammalian growth and development and is sensitive to maternal nutrition (Gluckman and Pinal Reference Gluckman and Pinal2003; Kaur et al. Reference Kaur, Muhlhausler and Roberts2021). For the samples used in the current study, we have reported a GAIN × day interaction for IGF-1, with greater concentrations for MG on day 83 (Menezes et al. Reference Menezes, McCarthy and Kassetas2022). The GH receptor (GHR) and IGF2 genes were upregulated in the VTM_LG vs. NoVTM_LG comparison, suggesting that VTM supplementation may enhance fetal GH sensitivity, potentiating the GH–IGF axis and nutrient-driven growth signaling (Gluckman and Pinal Reference Gluckman and Pinal2003). The binding of GH to its receptor (GHR) modulates somatic growth and development, carbohydrate and lipid metabolism, and positively modulates IGF2 (Kaur et al. Reference Kaur, Muhlhausler and Roberts2021). In turn, IGF1 and IGF2 stimulate growth and development via metabolic and mitogenic effects (Gluckman and Pinal Reference Gluckman and Pinal2003). The activity of IGFs is also modulated by insulin-like growth factor-binding proteins (IGFBPs), of which we identified IGFBP3, IGFBP4, IGFBP5, and IGFBP6 upregulated in the VTM_LG vs. NoVTM_LG contrast. Moreover, Se and Zn were reported as determinants of IGF-1 bioactivity (Maggio et al. Reference Maggio, De Vita and Lauretani2013). Together, these components of the IGF axis modulate the PI3K–Akt pathway to adjust cellular growth rates (Hietakangas and Cohen Reference Hietakangas and Cohen2009). This pathway participates in important neural development processes, including having neuroprotective effects (Dong et al. Reference Dong, Rovnaghi and Anand2014). In the present study, we identified the PI3K–Akt pathway overrepresented in all contrasts, except for the VTM_MG vs. NoVTM_MG.
Wnt signaling genes were upregulated in the fetal brain of VTM-supplemented groups. Members of the Wnt family, including WNT2B, WNT5A, and WNT7B, play essential roles in neurogenesis and synaptic plasticity (Cho et al. Reference Cho, Jung and Kim2018). We also observed the DKK2 gene upregulated in the VTM_LG vs. NoVTM_LG comparison. DKK2 has a context-dependent regulatory role, functioning as an inhibitor or activator of the Wnt signaling pathway (Mao et al. Reference Mao, Wu and Li2001). For example, DKK2 plays an activator role during neural crest specification in Xenopus embryos (Devotta et al. Reference Devotta, Hong and Saint-Jeannet2018). A review by Cortés-Albornoz et al. (Reference Cortés-Albornoz, García-Guáqueta and Velez-van-meerbeke2021) reported that inadequate maternal intake of vitamins, minerals, and macronutrients was associated with brain defects and neuropsychiatric disorders in human offspring. Furthermore, among the pathways, we identified neurodegeneration and Alzheimer’s disease overrepresented by the main effect of VTM supplementation under MG (VTM_MG vs. NoVTM_MG). We previously reported the effects of maternal vitamin and mineral supplementation during pregnancy on the feeding behavior of the F1 offspring (Hurlbert et al. Reference Hurlbert, Baumgaertner and Menezes2024). Although no significant differences in birth weight were observed, offspring born to dams supplemented throughout gestation consumed more feed and had greater intake at each eating event, compared with those born to dams not supplemented (Hurlbert et al. Reference Hurlbert, Baumgaertner and Menezes2024). The coordinated upregulation of Wnt genes, alongside GH–IGF axis components and PI3K–Akt signaling, suggests an integrated molecular network through which maternal vitamin and mineral supplementation programs the fetal brain.
Epigenetic modifiers, signaling molecules, and transcriptional regulators were affected by maternal nutrition
The foundation for future muscle growth and function is established during early gestation (Du et al. Reference Du, Tong and Zhao2009). Maternal nutrition alters fetal skeletal muscle development by regulating myogenesis, adipogenesis, and fibrogenesis (Du et al. Reference Du, Tong and Zhao2009). Additionally, fetal skeletal muscle has a low priority in nutrient partitioning, making it susceptible to nutrient imbalances (Sookoian et al. Reference Sookoian, Gianotti and Burgueño2013). Our results show that both maternal rate of gain and vitamin and mineral supplementation influence signaling pathways in fetal muscle through the regulation of epigenetic modifiers (TET1, TET2, and TET3), signaling molecules (WNT2, WIF1, SFRP4, DKK3, LRP6, MSTN, and RICTOR), and transcriptional regulators (FOXO3, JUN, JUND, PAX7, CREB3L2, and CREBRF). These genes were differentially expressed and associated with the Wnt, PI3K–Akt, and TGF-β signaling pathways, all of which play a role in myogenesis (Cossu Reference Cossu1999; Jin et al. Reference Jin, Peng and Jiang2016). Furthermore, these pathways were overrepresented in our meta-pathway analysis using the multi-list of DEGs across contrasts. It is also important to note the interplay between the brain and muscle tissue through these pathways. Although the overlap of DEGs between tissues was limited, the enrichment of shared pathways suggests a coordinated systemic response, whereby tissues activate complementary molecular mechanisms to support fetal development (Espina et al. Reference Espina, Cordeiro and Barriga2023).
When heifers were managed to achieve low BW gain, VTM supplementation at LG seems to reprogram myogenesis-related genes epigenetically. Genes involved in epigenetic regulation (TET1, TET2, and TET3), progenitor maintenance (PAX7), and signaling (LRP6, FOXO3, and RICTOR) were upregulated in the VTM_LG compared with the NoVTM_LG group, although they were not consistently differentially expressed across all comparisons. The ten-eleven translocation (TET) family members play a role in DNA demethylation, allowing the expression of target genes (Zhang et al. Reference Zhang, Zhang and Wang2023). In vitro studies using mouse myoblast cells indicate that DNA demethylation would stimulate muscle hypertrophy by activating intracellular signaling pathways (Senesi et al. Reference Senesi, Luzi and Montesano2014). However, DNA methylation was not performed in the current study, and the observed transcriptomics changes in epigenetic-related genes require further validation. FOXO3 was upregulated under VTM_LG, whereas JUN and JUND were downregulated. FOXO3 encodes a transcription factor that acts as a positive regulator of muscle atrophy through protein degradation pathways (Schiaffino et al. Reference Schiaffino, Dyar and Ciciliot2013). Likewise, downregulation of JUNB causes muscle atrophy via FOXO3 signaling (Raffaello et al. Reference Raffaello, Milan and Masiero2010). Consistent with these findings, protein digestion and absorption were overrepresented in our meta-pathway analysis, which indicates molecular adjustments in fetal muscle tissue in response to maternal nutritional status.
On the other hand, genes involved with active myogenesis were upregulated by maternal VTM supplementation and rate of gain (VTM_LG vs. NoVTM_LG, NoVTM_MG vs. NoVTM_LG, or VTM_LG vs. NoVTM_MG). PAX7 encodes a transcription factor that stimulates the development of myogenic progenitor cells in prenatal development and satellite cells during postnatal growth (Gauvin et al. Reference Gauvin, Pillai and Reed2020). This progenitor cell population serves as an important source of myogenic cells for skeletal muscle formation (Relaix et al. Reference Relaix, Rocancourt and Mansouri2005). PAX7-positive cells were decreased in the fetal semitendinosus and triceps brachii muscles of lamb fetuses at day 90 of gestation when dams were under- or overnourished from day 30 of gestation, compared to control (Gauvin et al. Reference Gauvin, Pillai and Reed2020). In contrast, antagonistic genes of the Wnt signaling pathway (Mashhadikhan et al. Reference Mashhadikhan, Kheiri and Dehghanifard2020), such as WIF1 and SFRP4, were downregulated in our study. Mashhadikhan et al. (2020) reported that the downregulation of Wnt modulators is necessary to promote osteoblastic differentiation. Furthermore, Wnt family signaling molecules are required to activate myogenesis. The LRP6 gene encodes a co-receptor needed to activate the Wnt/β-catenin signaling pathway (Alrefaei and Abu-Elmagd Reference Alrefaei and Abu-Elmagd2022). According to Liu et al. (Reference Liu, Amorín and Moriel2020), maternal methionine supplementation from pre-breeding to day 90 of gestation affected the expression of muscle genes involved with myogenesis, adipogenesis, fibrogenesis, and the canonical Wnt/β-catenin pathway in calves at 1 month of age.
Results from a bovine model with rates of BW gain from breeding to day 83 of gestation, similar to the current study, demonstrated phenotypic effects both at birth and throughout the postnatal development of F1 heifers (Baumgaertner et al. Reference Baumgaertner, Menezes and Diniz2024b). Muscle transcriptome analyses at birth identified 275 DEGs, with 177 upregulated in F1 offspring born to MG dams (Ramírez-Zamudio et al. Reference Ramírez-Zamudio, Diniz and Baumgaertner2026). The overlap between DEGs from our study and those reported by Ramirez-Zamudio et al. (Reference Ramírez-Zamudio, Diniz and Baumgaertner2026) revealed 24 shared genes, including those involved in organ and tissue development (LRP6, ATF3, USP47, and EGR1) and response to starvation (ATF3) (Galves et al. Reference Galves, Sperber and Amer-Sarsour2023). These shared genes, together with the BPs identified, suggest that some molecular effects may persist beyond the first trimester of gestation and influence postnatal development, even though some genes are no longer differentially expressed. Our findings provide new insights into the effects of maternal vitamin and mineral supplementation and rates of BW gain on fetal tissue programming.
This study also has limitations. It was not possible to untangle the specific contributions of individual nutrients to the regulation of specific genes nor to fully capture the complexity of their roles in tissue development. In addition, analyses were conducted at a single time point, which limits the ability to track the dynamics of adaptive mechanisms across gestation. Furthermore, this study focused only on female fetuses, which limits the generalization of the findings to male offspring. Given known sex-specific differences in developmental and transcriptomic responses to maternal nutrition, caution should be exercised when extrapolating these findings to male offspring. Future studies directly comparing both sexes are needed to better understand sex-dependent effects of maternal nutritional programming. Finally, while our results suggest potential involvement of epigenetic regulation (e.g., differential expression of DNA demethylation enzymes), direct epigenetic measurements were not performed. Therefore, additional studies integrating epigenomic approaches are required to confirm the role of epigenetic mechanisms in mediating the observed transcriptomic changes. Future longitudinal and integrative studies are needed to provide a more comprehensive understanding of how maternal nutrient availability influences fetal tissue programming, metabolic pathways, developmental trajectories, and long-term performance.
From a beef producer’s standpoint, these findings highlight the potential for targeted supplementation strategies to improve muscle development. From a research perspective, further studies are needed to functionally validate the genes and pathways identified and clarify their causal roles in muscle and brain development. Integrating epigenetic analyses will also be key to determining how maternal nutrition leaves lasting molecular marks that influence gene expression postnatally, including transgenerationally. Together, such efforts will advance our mechanistic understanding of developmental programming and help translate molecular insights into practical nutritional strategies for livestock production.
Conclusions
Maternal vitamin and mineral supplementation, in combination with moderate or low rates of BW gain during early gestation, alters the molecular profile of the brain and muscle tissues of female fetuses at day 83 (approximately the end of the first trimester of gestation). Although gene expression differences were tissue-specific, the overrepresentation of shared nutrient-sensing pathways, such as Wnt and PI3K–Akt signaling, suggests a coordinated systemic response to support fetal development. Notably, changes in the expression of DNA demethylation-associated TET genes in muscle highlight the potential role of epigenetic regulation in modulating myogenesis and long-term muscle development. These findings highlight the importance of maternal nutrition in establishing the developmental trajectory of key fetal tissues. Further investigation is warranted to determine potential epigenetic mechanisms programming fetal tissue development, as well as how early nutritional interventions influence lifelong productivity and efficiency in beef production systems.
Supplementary Material
The supplementary material for this article can be found at https://doi.org/10.1017/anr.2026.10042
Data Availability Statement
All relevant data are included in the paper and its Supplementary Information files. All sequencing data is publicly available on NCBI’s Gene Expression Omnibus database under GEO accession number GSE307490.
Acknowledgements
The authors would like to thank Purina Animal Nutrition LLC (Land O’Lakes, Inc., Arden Hills, MN, USA) for providing financial support for this research. The authors would also like to thank the North Dakota State Board of Agricultural Research and Education, Graduate Research Assistantship Program, and the North Dakota Agricultural Experiment Station for their support of this effort and additional product support from Zoetis Animal Health (Parsippany, NJ, USA) and ST Genetics (Navasota, TX, USA). Appreciation is extended to the personnel at the Central Grasslands Research Extension Center and the Animal Nutrition and Physiology Center for their assistance with animal handling and feeding, as well as to the NDSU Animal Science Nutrition Laboratory. This work used resources of the Center for Computationally Assisted Science and Technology (CCAST) at North Dakota State University and the Auburn University Easley Cluster.
Author Contributions
Conceptualization: C.R.D., K.L.M., J.S.C., J.C.F. R.R.S., and W.J.S.D.; animal management: K.L.M., C.J.K., F.B., J.D.K. and K.K.S.; sample collection: C.R.D., K.L.M., J.S.C., C.J.K., F.B., L.P.R., P.P.B., A.K.W., T.L.N. and S.T.D.; Methodology: W.J.S.D.; formal analysis: W.J.S.D; writing – original draft preparation: W.J.S.D.; writing – review and editing: all authors; supervision: A.K.W. and C.R.D.; funding acquisition: C.R.D., K.L.M. and J.S.C. All authors have read and agreed to the published version of the manuscript.
Funding Statement
This research was funded by the North Dakota Agricultural Experiment Station (NDAES), the North Dakota State Board of Agricultural Research and Education (SBARE) – grant number 19-23-0155, and Purina Animal Nutrition LLC, Gray Summit, MO, USA. The first author (W.J.S.D.) was financially supported by the Agricultural Research Service, U.S. Department of Agriculture, under Agreement No. 58-6010-1-005 and by the Alabama Agricultural Experiment Station – Hatch program of the National Institute of Food and Agriculture, U.S. Department of Agriculture.
Conflicts of Interest
J. C. Forcherio and R. Scott are employees of Purina Animal Nutrition LLC (Land O’Lakes, Inc., Arden Hills, MN, USA), which sponsored the sample analysis for this experiment. Purina Animal Nutrition LLC manufactured the Purina Wind & Rain Storm AllSeason 7.5 Complete mineral, the VTM and NoVTM pellets, and the protein/energy supplement used in this study. The funders had no role in the collection, analyses, or interpretation of the data; in the writing of the manuscript, or in the decision to publish the results, which were done entirely independently by personnel from North Dakota State University and Auburn University. The other authors declare no conflict of interest.
Institutional Review Board Statement
All experiments and methods were performed in accordance with the relevant guidelines and regulations. The experimental design, animal management, and tissue collection were approved by the North Dakota State University Institutional Animal Care and Use Committee (IACUC A19012).