Introduction
Over the past decades, our knowledge of living cells’ inner molecular and biochemical workings has reached a formerly unimaginable depth. Nevertheless, despite all the novel techniques, scientific discoveries, and advances, the living cells remain a mystery, particularly its living aspect (Schrödinger Reference Schrödinger1944). Thus, one of the significant scientific challenges of this century is to uncover what creates a living artificial cell (Ehmoser-Sinner and Tan Reference Ehmoser-Sinner, Tan, Ehmoser-Sinner and Tan2018). The current strategies toward this goal lie on the spectrum between two drastically different approaches: top-down and bottom-up.
In the top-down approach, a living cell is stripped of all unnecessary, that is, non-vital parts. A prime example of this approach was the mutagenesis study of the Mycoplasma mycoides genome, where only 473 of the 901 genes were kept to produce a minimal synthetic JCVI syn3.0 cell (Hutchison et al. Reference Hutchison, Chuang, Noskov and Venter2016). However, the fact that 79 out of those 473 genes remain unassigned highlights the fundamental property of living systems: the whole is more than the sum of its parts. The bottom-up approach in synthetic biology tackles this by taking an opposite rationalistic and somewhat mechanical approach to reconstituting cellular functions, properties, and structures. Building blocks, natural or synthetic cellular components, are assembled and combined to create a system that could be considered living. As scientists focus on particular cellular systems and develop new biophysical techniques to interact with them, they also gain a profound understanding of how these systems work and converge with other building blocks.
One of the essential systems required to achieve a synthetic cell is the processing of the genetic material, DNA, and protein expression. The latter is necessary to guarantee the function of maintaining itself by recreating its own parts, which is typical for living systems (Maturana and Varela Reference Maturana and Varela1972). Without the transcription and translation systems, the synthetic cell cannot produce necessary metabolites, divide, and grow, that is, fulfill all fundamental criteria of the minimal viable organism.
Another requirement imposed for a prospective synthetic cell to function like its natural counterparts is the boundary, separating it from the environment. The selective permeability of the cellular membrane allows for extensive regulation and control of the interior processes. For this reason, attempts to produce synthetic cells and study cellular systems in vitro commonly involve encapsulation (Noireaux and Libchaber Reference Noireaux and Libchaber2004) in giant unilamellar vesicles (GUVs) (Van de Cauter et al. Reference Van de Cauter, van Buren, Koenderink and Ganzinger2023). GUVs allow the reproduction of spatial separation conditions and can be produced in various ways, which will be discussed in this review, along with the implications of transcription–translation encapsulation, its products, and the means to image those processes, as well as the challenges that arise to do so.
Finally, when it comes to storing, processing, and inheriting genetic information by the proto-synthetic cells, one way is to encapsulate multiple copies of the same chromosome or a set of plasmids (Nordström and Austin Reference Nordström and Austin1989). In this case, random portioning will create offspring cells with some copies of the necessary genetic information. However, over the generations, this will lead to an increasing heterogeneity in the number of gene copies (Huh and Paulsson Reference Huh and Paulsson2011). Additionally, these chromosomes, containing 100s of genes, would take increasingly more space and energy. An alternative nature-inspired approach is to encapsulate a single chromosome and design the system to yield single-chromosome offspring cells, for example, through active (Lemon and Grossman Reference Lemon and Grossman2001) or entropy-driven separation of chromosomes (Jun and Mulder Reference Jun and Mulder2006) before the division (Olivi et al. Reference Olivi, Berger, Creyghton and van der Oost2021). Favoring the second approach, we highlight techniques that allow single-molecule transcription tracking at low DNA copy numbers in GUVs.
Achieving transcription–translation in vitro
Two major yet conceptually diametrical approaches for in vitro transcription–translation are using cell lysate (Didovyk et al. Reference Didovyk, Tonooka, Tsimring and Hasty2017) and the Protein synthesis Using Recombinant Element (PURE) system (Shimizu et al. Reference Shimizu, Inoue, Tomari and Ueda2001, Reference Shimizu, Kanamori and Ueda2005). Cell lysate can be viewed as the top-down solution, capturing all the necessary components to achieve transcription–translation in vitro as they are present in live bacteria, commonly Escherichia coli lysate is used for this purpose (Didovyk et al. Reference Didovyk, Tonooka, Tsimring and Hasty2017), coming with innately coupled transcription and translation systems, thus ensuring high yields (Hansen et al. Reference Hansen, Ventosa Rosquelles, Yelleswarapu and Huck2016b). Furthermore, crowding in the solution emulates a native intracellular environment and can improve yields (Vibhute et al. Reference Vibhute, Schaap, Maas and Huck2020). Finally, the use of different lysate dilutions allows researchers to regulate the kinetic rates of mRNA production and degradation. The use of lysate does come at a cost: RNases and proteases are present and active in the lysate solution. They are actively working against the transcription and translation systems, digesting synthesized RNA and protein molecules. It is because of the previously mentioned coupling of the transcription–translation system that this potential issue is mitigated. Once the systems are decoupled, the yields decrease significantly (Hansen et al. Reference Hansen, Ventosa Rosquelles, Yelleswarapu and Huck2016b). There is virtually no control over the composition of the lysate, making it essentially a black box where some of the cellular pathways remain active while others cease. This lack of information about components was partially circumvented by using mass-spectrometry to reveal the detailed proteome of the lysate (Foshag et al. Reference Foshag, Henrich, Hiller and Bernhard2018). Additionally, theoretical models for lysate are being developed, such as a system of ordinary differential equations that was used to describe the expression of the reporter gene. Experimental data supplemented this system and showed the regulatory effects of various promoters and ribosome-binding sites (Marshall and Noireaux Reference Marshall and Noireaux2019). Nevertheless, more studies of the cell lysate composition and fine-tuned pathway models are needed to disentangle the mesh of metabolic interaction inside it. This data could then be applied to artificially enhance specific pathways by supplementing additional metabolites depending on the needs (Miguez et al. Reference Miguez, Zhang, Piorino and Styczynski2021).
PURE system was initially engineered bottom-up through the combination of T7 bacteriophage (Figure 1) RNA polymerase responsible for transcription, the E. coli translation machinery, and necessary metabolites to provide building blocks for RNA and protein (Shimizu et al. Reference Shimizu, Inoue, Tomari and Ueda2001, Reference Shimizu, Kanamori and Ueda2005). All its components, including the choice of salts, and their concentrations are listed in the Supplementary section. The composition of PURE was highly optimized, making it possible to create a kinetic description of the PURE system (Doerr et al. Reference Doerr, De Reus, Van Nies and Danelon2019). Though it is possible to produce all PURE system components and solutions in the lab, it is far more complex than lysate production since ribosome isolation and over 36 protein purifications are required (Grasemann et al. Reference Grasemann, Lavickova, Elizondo-Cantú and Maerkl2021). Therefore, several commercial PURE system kits are currently available, optimized for different applications, and sold by GeneFrontier (Japan), New England Biolabs (USA), and Creative Biolabs (USA). The precise control over the content minimizes any RNase and protease activity, which the manufacturers claim to reduce ‘greatly’. However, the combination of transcription and translation complexes from two different, though compatible, organisms makes the PURE system underperform by at least an order of magnitude compared with cell lysate when it comes to protein expression (Hillebrecht and Chong Reference Hillebrecht and Chong2008). Ultimately, the choice of PURE or lysate comes down to the particular application, protein of interest, and technical constraints. We provide their overview comparison in Table 1.
Comparison of cell lysate and PURE system for in vitro transcription–translation in the context of bottom-up synthetic cells
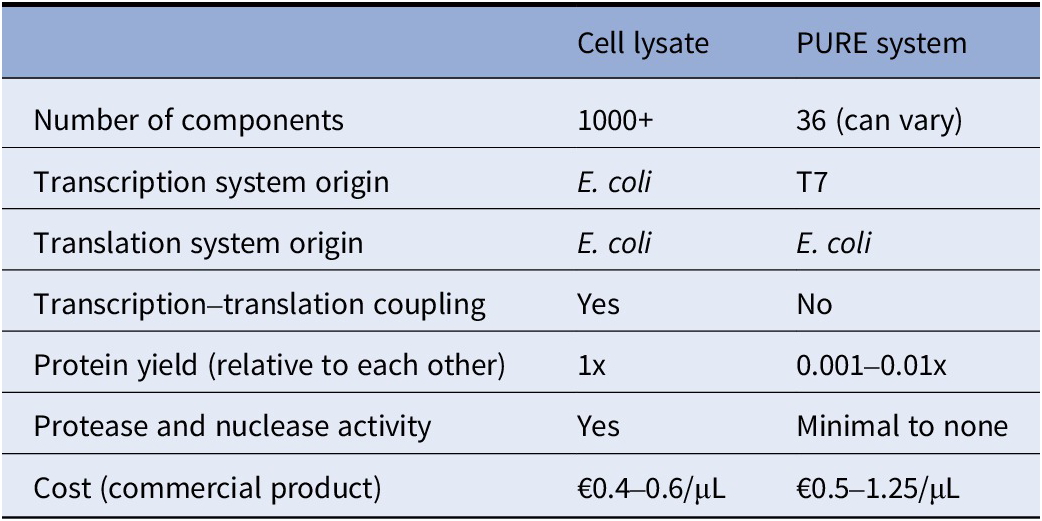
Note: Cell lysate costs were calculated using Biotechrabbit (Germany) and Cube Biotech (Germany) catalogs; (Nirenberg and Matthaei Reference Nirenberg and Matthaei1961; Hansen et al. Reference Hansen, Ventosa Rosquelles, Yelleswarapu and Huck2016b; Didovyk et al. Reference Didovyk, Tonooka, Tsimring and Hasty2017; Foshag et al. Reference Foshag, Henrich, Hiller and Bernhard2018; Marshall and Noireaux Reference Marshall and Noireaux2019). PURE system costs were calculated using GeneFrontier (Japan) and New England Biolabs (USA) catalogs and their distributors in the Netherlands; (Shimizu et al. Reference Shimizu, Inoue, Tomari and Ueda2001, Reference Shimizu, Kanamori and Ueda2005; Doerr et al. Reference Doerr, De Reus, Van Nies and Danelon2019; Cauter et al. Reference Cauter, Fanalista, Buren and Ganzinger2021; Grasemann et al., Reference Grasemann, Lavickova, Elizondo-Cantú and Maerkl2021), reviews (Li et al. Reference Li, Zhang, Huang and Church2017; Laohakunakorn et al. Reference Laohakunakorn, Grasemann, Lavickova and Maerkl2020; Garenne et al. Reference Garenne, Haines, Romantseva, Freemont, Strychalski and Noireaux2021a; Cui et al. Reference Cui, Chen, Wang and Lu2022).
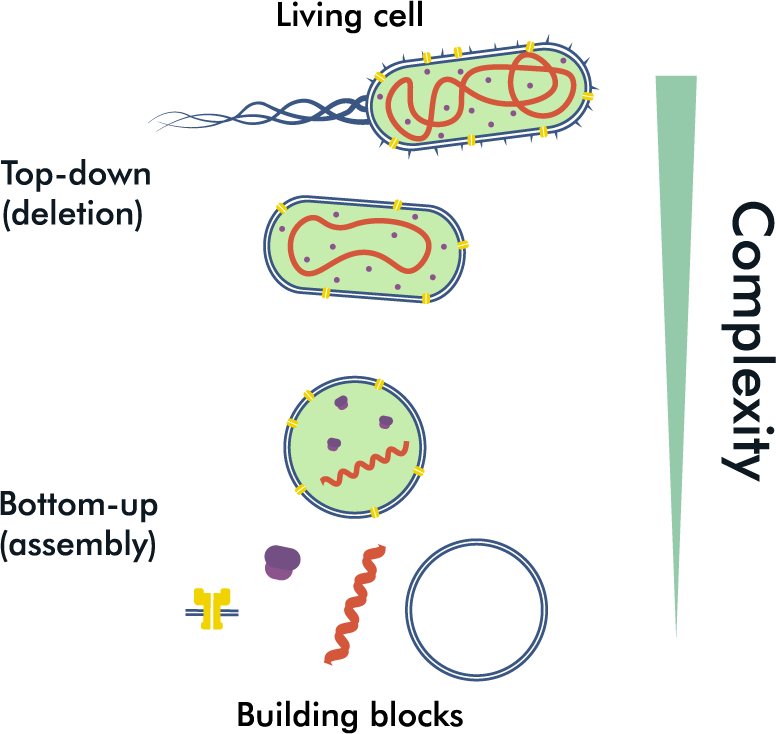
Two diametrically opposite approaches for synthetic cell production. The top-down approach aims to decrease the complexity of the system by discarding unnecessary elements, while the bottom-up approach combines building blocks to increase the complexity of the system.
In the context of synthetic cells, independent of which approach is used for in vitro transcription and translation, the end goal is not just to maximize protein yields but to reconstitute cellular systems, gene interactions, and metabolic pathways in a coordinated manner (Noireaux et al. Reference Noireaux, Bar-Ziv and Libchaber2003; Shin and Noireaux Reference Shin and Noireaux2012). This is somewhat problematic, considering that fluorescent proteins, most commonly used to study in vitro expression, are great for reporting but play no functional role. An alternative to that is the use of transcription–translation systems to produce proteins involved in cellular functions such as lipid synthesis (Scott et al. Reference Scott, Noga, de Graaf, Westerlaken, Yildirim and Danelon2016), cytoskeleton formation (Cauter et al. Reference Cauter, Fanalista, Buren and Ganzinger2021; Litschel et al. Reference Litschel, Kelley, Holz and Schwille2021), or genome replication (Van Nies et al. Reference Van Nies, Westerlaken, Blanken, Salas, Mencía and Danelon2018). Another possibility is to design systems with genetic circuits, such as in the case of the TX-TL Toolbox (myTXTL is the commercial name), which is based on E. coli lysate (Shin and Noireaux Reference Shin and Noireaux2012; Garamella et al. Reference Garamella, Marshall, Rustad and Noireaux2016; Garenne et al. Reference Garenne, Thompson, Brisson, Khakimzhan and Noireaux2021b). This system was numerically described (Marshall and Noireaux Reference Marshall and Noireaux2019), characterized by mass spectrometry (Garenne et al. Reference Garenne, Beisel and Noireaux2019), and used to create biocircuits made of sigma factor-regulated plasmids (Garamella et al. Reference Garamella, Marshall, Rustad and Noireaux2016; Agrawal et al. Reference Agrawal, Marshall, Noireaux and Sontag2019). Studying how these synthetic cell-tailored transcription and translation systems are reconstituted inside GUVs is particularly relevant, which is why compatible GUV production methods will be the focus of the next section.
Encapsulating transcription–translation in GUVs
The methods for GUV production are generally divided into two categories: swelling-based and emulsion-based approaches. The swelling-based approaches involve the rehydration of lipids on, among other things, gel (Weinberger et al. Reference Weinberger, Tsai, Koenderink and Marques2013), electrode (Angelova and Dimitrov Reference Angelova and Dimitrov1986), and glass beads (Nourian et al. Reference Nourian, Roelofsen and Danelon2012; Tanasescu et al. Reference Tanasescu, Mettal, Colom, Roux and Zumbuehl2018). These methods offer limited encapsulation efficiency, and only the lipid-coated glass beads method has been repeatedly and successfully used for in vitro transcription and translation. Hence, we only consider this approach in our table GUV production methods (Table 2). It does not offer size control nor produce a monodisperse GUV population, but the simplicity and cost-effectiveness of the method make it very easy to reproduce in the lab. The second large category, the emulsion-based methods, includes diverse methods that produce a lipid bilayer at the most fundamental level by fusing two lipid monolayers. The first one surrounds the water-in-oil droplet with the encapsulated solution, and the second one is on the oil–water interface, through which that droplet passes. In practice, this can be achieved by various approaches of increasing complexity: from the centrifugation in the inverted emulsion method (Nishimura et al. Reference Nishimura, Suzuki, Toyota and Yomo2012; Moga et al. Reference Moga, Yandrapalli, Dimova and Robinson2019) and the use of 3D printed spinning chamber with a water–oil interface, into which the encapsulated solution is pumped through a capillary in cDICE (Abkarian et al. Reference Abkarian, Loiseau and Massiera2011; Cauter et al. Reference Cauter, Fanalista, Buren and Ganzinger2021) to the clean-room manufacturing of intricate PDMS chips in microfluidic methods, such as OLA (Deshpande et al. Reference Deshpande, Caspi, Meijering and Dekker2016), double-emulsion dewetting (Yan et al. Reference Yan, Bauer, Fischlechner, Hollfelder, Kaminski and Huck2013; Kang et al. Reference Kang, Zhu, Kong and Wang2016; Deng et al. Reference Deng, Vibhute, Zheng, Zhao, Yelleswarapu and Huck2018) and surfactant-free microfluidic approach (Yandrapalli et al. Reference Yandrapalli, Petit, Bäumchen and Robinson2021). These microfluidic methods could be viewed as automating a more manual inverted emulsion and making use of surface tension differences to shed the oil phase from the water-in-oil droplet and produce GUVs.
Comparison of GUV production methods for in vitro transcription and translation
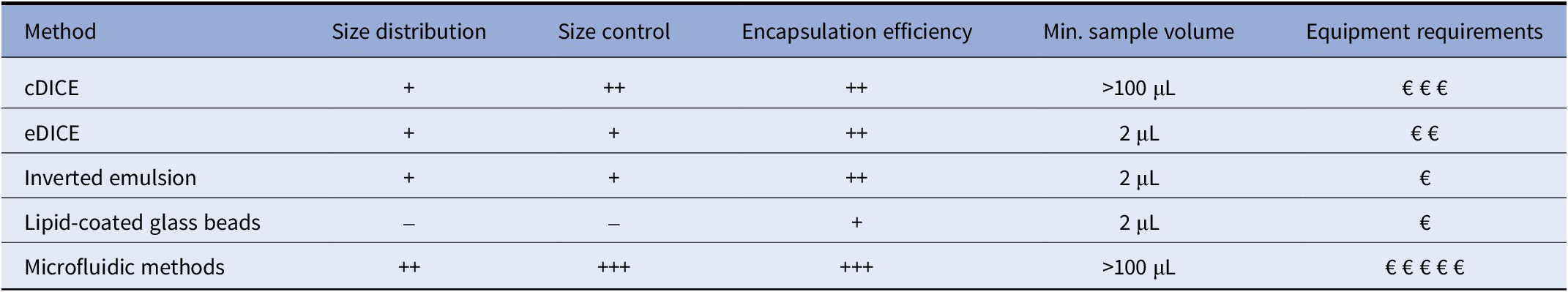
Note: Size distribution: – polydisperse, + broad monodisperse, ++ narrow monodisperse; Size control: – no size control; + limited empirical control; ++ limited mechanical control; +++ precise control; Encapsulation efficiency: + low, ++ moderate, +++ high; Equipment requirements: cDICE – syringe pump, specially-designed capillary stand, specially-designed rotating stage with chamber holder, 3D printed chamber, glove box for improved yields (Abkarian et al. Reference Abkarian, Loiseau and Massiera2011; Blosser et al. Reference Blosser, Horst and Keller2016; Bashirzadeh et al. Reference Bashirzadeh, Wushet, Litschel, Schwille and Liu2021b, Reference Bashirzadeh, Redford, Lorpaiboon and Liu2021a; Cauter et al. Reference Cauter, Fanalista, Buren and Ganzinger2021, Reference Van de Cauter, Jawale, Tam and Ganzinger2024; Litschel et al. Reference Litschel, Kelley, Holz and Schwille2021); eDICE – specially-designed rotating stage with chamber holder, 3D printed chamber, glove box for improved yields (Baldauf et al. Reference Baldauf, Frey, Arribas Perez, Idema and Koenderink2023a, Reference Baldauf, Frey, Perez and Koenderink2023b; Wubshet et al. Reference Wubshet, Wu, Veerapaneni and Liu2023); Inverted emulsion – centrifuge (Nishimura et al. Reference Nishimura, Suzuki, Toyota and Yomo2012; Soga et al. Reference Soga, Fujii, Yomo, Kato, Watanabe and Matsuura2014; Litschel et al. Reference Litschel, Ramm, Maas, Heymann and Schwille2018; Berhanu et al. Reference Berhanu, Ueda and Kuruma2019; Moga et al. Reference Moga, Yandrapalli, Dimova and Robinson2019; Yoshida et al. Reference Yoshida, Kohyama, Fujiwara, Nishikawa and Doi2019; Garenne and Noireaux Reference Garenne and Noireaux2020; Zhang et al. Reference Zhang, Obuchi and Toyota2023); Lipid-coated glass beads – none (Nourian et al. Reference Nourian, Roelofsen and Danelon2012; Tanasescu et al. Reference Tanasescu, Mettal, Colom, Roux and Zumbuehl2018; Blanken et al. Reference Blanken, Foschepoth, Serrao and Danelon2020; Kattan et al. Reference Kattan, Doerr, Dogterom and Danelon2021; Gonzales et al. 2022); Microfluidic methods – microfluidic pressure-driven flow controller, microscope to observe production, clean room as the manufacturing of narrow-channeled PDMS chips is highly complex (Teh et al. Reference Teh, Khnouf, Fan and Lee2011; Yan et al. Reference Yan, Bauer, Fischlechner, Hollfelder, Kaminski and Huck2013; Deshpande et al. Reference Deshpande, Caspi, Meijering and Dekker2016; Kang et al. Reference Kang, Zhu, Kong and Wang2016; Deng et al. Reference Deng, Vibhute, Zheng, Zhao, Yelleswarapu and Huck2018; Vibhute et al. Reference Vibhute, Schaap, Maas and Huck2020; Yandrapalli et al. Reference Yandrapalli, Petit, Bäumchen and Robinson2021; Gonzales et al. 2022).
Continuing with the analogy, cDICE, with its microfluidic capillary and a macrochamber, lies somewhere in between. Another similar method, eDICE, combines manual mechanical agitation to produce water-in-oil droplets of the inverted emulsion and the spinning chamber with a water–oil interface of cDICE. Notably, at this time, no peer-reviewed papers have demonstrated in vitro transcription and translation using eDICE. However, the successful use of both inverted emulsion and cDICE, moderate encapsulation efficiency, and low costs of necessary components make eDICE a promising approach for future research. Additionally, emulsion-based methods that employ centrifugation of a tube with a specially designed microfluidic insert, such as droplet-shooting and size-filtration (DSSF) (Morita et al. Reference Morita, Onoe, Yanagisawa and Takinoue2015) and modification (Venero et al. Reference Venero, Sato, Heili, Deich, Adamala, Karim and Jewett2022; Deich et al. Reference Deich, Gaut, Sato, Engelhart and Adamala2023), have not been included due to the limited publications on them.
While all emulsion-based approaches generate monodisperse GUV populations (Figure 2), only the microfluidic approaches ensure highly monodisperse and continuous GUV production. The same could be said about the size control, except that in cDICE, different diameter capillaries offer limited GUV size control. In contrast, the sizes resulting from inverted emulsion/eDICE could only be adjusted to a limited degree empirically by changing the number of times the tube is dragged along a rack to agitate the solutions and mix the encapsulated water-based solution into the oil. The encapsulated volume for the inverted emulsion, eDICE, and lipid-coated glass beads could be as small as a few micrometers, while cDICE and microfluidic methods require >100 μL samples to be used each time because of the inner volume of the tubing. This volume loss could be problematic when using costly solutions such as commercial PURE systems (1€/μL). Costs are the downside of the microfluidic methods, which generally require more expertise as well as special facilities and equipment to establish them in the lab. While they allow precision manufacturing of GUVs, other less complex methods we listed could be adopted in a shorter time by more different kinds of labs. Furthermore, the comparison table contains several non-numerical but comparative columns, such as Encapsulation efficiency and size control. The challenges of GUV synthesis are not only in choosing the best method and protocol for one’s needs but also in ensuring it works optimally in the conditions of a specific lab, as even environmental factors such as air humidity can affect the resulting yields (Cauter et al. Reference Cauter, Fanalista, Buren and Ganzinger2021). For an extensive overview of GUV requirements and production methods for synthetic cell applications beyond in vitro transcription and translation, consider the review (Van de Cauter et al. Reference Van de Cauter, van Buren, Koenderink and Ganzinger2023).
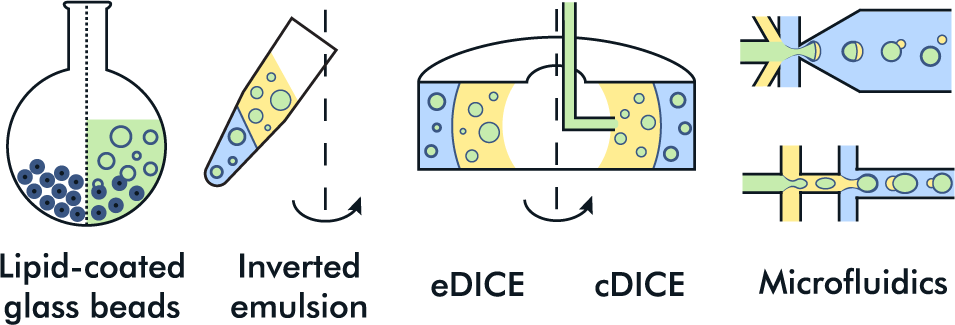
Examples of GUV production methods used to encapsulate transcription-translation reaction. The solutions used in each of them are color-coded: encapsulated solution (green), lipids diluted in oil phase (yellow), and exterior aqueous phase (blue).
Imaging transcription in vitro
After discussing the methods to encapsulate the in vitro transcription–translation mixture in GUVs, we next examine the ways to follow those two processes in bulk and inside GUVs. In particular, we highlight approaches with single-molecule resolution, which are or could be of use in synthetic cell studies. Among the detection approaches, reverse-transcriptase quantitative PCR (RT qPCR) variations are the go-to method for mRNA studies (Park and Magan Reference Park and Magan2011; Green and Sambrook Reference Green and Sambrook2018; Sato et al. Reference Sato, Sharon, Deich and Adamala2022). Meanwhile, mass spectrometry offers unparalleled precision when it comes to protein quantification and sensitivity sufficient even to resolve posttranslational modifications (Gerber et al. Reference Gerber, Rush, Stemman, Kirschner and Gygi2003). Both, however, are the end-of-reaction methods used to calculate the quantity of mRNA or protein after the process is finished unless aliquoting is used. For real-time approaches, it is common to employ fluorescent probes that activate upon binding to the mRNA.
One such probe is called a molecular beacon (Tyagi and Kramer Reference Tyagi and Kramer1996; Giesendorf et al. Reference Giesendorf, Vet, Tyagi, Mensink, Trijbels and Blom1998; Tsourkas et al. Reference Tsourkas, Behlke, Rose and Bao2003a; Goel et al. Reference Goel, Kumar, Puniya, Chen and Singh2005; Bratu et al. Reference Bratu, Catrina, Marras and Gerst2011). It is typically a 20+ oligonucleotide loop, functionalized with a fluorophore and a quencher at opposing ends. Molecular beacons are designed to stay closed in the solution because of the interaction of typically 4–6 oligos-long double-stranded stem segments and open up when its target-binding loop oligos complementary sequence on the mRNA. The third state of the molecular beacon is the open-in-solution state, which produces an intense background signal of unquenched dye and is the major downside of this probe. Another downside is that molecular beacons, being single-stranded ribonucleotides, are prone to degradation by nucleases unless particular chemical strategies, such as the use of 2′-O-methyl ribonucleotides, are used (Majlessi et al. Reference Majlessi, Nelson and Becker1998). The exact structure, stem length, and fluorophore of the MB have to be optimized for a particular application. For example, having a longer G-C-rich stem improves the selectivity of the probe and results in a lower background signal at the cost of hybridization rate and vice versa (Tsourkas et al. Reference Tsourkas, Behlke, Rose and Bao2003a) unless some of the neck segment is also involved in target-binding (Tsourkas et al. Reference Tsourkas, Behlke and Bao2002).
Molecular beacons offer the benefits of bright and stable commercial dyes, which allow the development of various enhanced imaging strategies, particularly for in vivo applications (Mao et al. Reference Mao, Ying, Wu and Chen2020), but potentially useful in vitro as well. These include using donor and acceptor molecular beacons with a FRET dye pair for the minimization of background noise (Bratu et al. Reference Bratu, Cha, Mhlanga, Kramer and Tyagi2003; Tsourkas et al. Reference Tsourkas, Behlke, Xu and Bao2003b). Other approaches focus on using multiple molecular beacon repeats to achieve better signal-to-noise ratio and even single-molecule resolution for the localization of mRNA molecules. In one study, the localization precision was comparable to that of smFISH, starting with 8x molecular beacon repeats, which came at the cost of significant elongation of the construct. Notably, the molecular beacon-tagged RNA molecules, in this case, were sufficiently fluorescent for imaging by conventional widefield fluorescence microscopy (Chen et al. Reference Chen, Ma, Wu and Chen2017). Meanwhile, another in vitro study utilized confocal microscopy and 32x molecular beacon repeats to showcase how crowding in picoliter droplets leads to mRNA concentration in individually resolved spots instead of homogeneous distribution observed in the absence of a crowding agent (Hansen et al. Reference Hansen, Meijer, Spruijt and Huck2016a).
Aptamers (Ellington and Szostak Reference Ellington and Szostak1990) are oligonucleotides or polypeptides developed to bind a specific target ligand. One particular type of aptamers that is of interest for the scope of this review is called fluorescent light-up aptamers (Bouhedda et al. Reference Bouhedda, Autour and Ryckelynck2017). These on-when-bound oligonucleotide aptamers are developed using the Systematic Evolution of Ligands by EXponential enrichment (SELEX) process (Tuerk and Gold Reference Tuerk and Gold1990), which enables scanning through large libraries of sequences for the one with the highest affinity and specificity for the target ligand. Among them, Spinach, a 98-nucleotide RNA aptamer, was developed as a mimic of Green Fluorescent Protein (GFP), where the interaction of oligos forms a G-quadraplex with affinity for synthetic dye 3,5-difluoro-4-hydroxybenzylidene imidazolinone (DFHBI). This interaction creates a very similar atomical motive to that of amino acids in GFP (Pothoulakis et al. Reference Pothoulakis, Ceroni, Reeve and Ellis2014), enabling widefield fluorescence imaging of the produced RNA molecules in solution. The stabilization of the fluorophore by the tertiary aptamer structure increases its fluorescence by preventing premature decay from the excited state. To improve the thermal properties and stability of Spinach aptamer, various modifications (Strack et al. Reference Strack, Disney and Jaffrey2013; Warner et al. Reference Warner, Chen, Song and Ferré-D’Amaré2014; Zhang et al. Reference Zhang, Fei, Leslie, Han, Kuhlman and Ha2015; Autour et al. Reference Autour, Westhof and Ryckelynck2016) were developed along with Broccoli (Filonov et al. Reference Filonov, Moon, Svensen and Jaffrey2014; Okuda et al. Reference Okuda, Fourmy and Yoshizawa2017; Kartje et al. Reference Kartje, Janis, Mukhopadhyay and Gagnon2021).
Meanwhile, other wavelengths and fluorophores were used to produce aptamers (Figure 3), such as a single-molecule oriented Mangos (Dolgosheina et al. Reference Dolgosheina, Jeng, Panchapakesan and Unrau2014; Autour et al. Reference Autour, Jeng, Cawte and Unrau2018; Cawte et al. Reference Cawte, Unrau and Rueda2020), photostable Corn (Song et al. Reference Song, Filonov, Kim and Jaffrey2017), and multicolor Pepper (Chen et al. Reference Chen, Zhang, Su and Yang2019; Tang et al. Reference Tang, Afasizheva, Cano, Plath, Black and Franco2024) that, depending on the bound dye can emit cyan-to-red photons. Importantly, Pepper aptamer was imaged not only with conventional wide-field microscopy but also using structured illumination microscopy and two-photon confocal microscopy (Chen et al. Reference Chen, Zhang, Su and Yang2019). The range of emission wavelength spectra of aptamers enables the imaging of multiple species of RNA being synthesized in parallel, for example, using a multimode microplate reader (Yan et al. Reference Yan, Eshed, Tang and Green2024). Or aptamers can be designed in a way that they can bind the ligand and fluoresce only when another activator molecule is present, allowing for even more metabolic studies (Wang and Simmel Reference Wang and Simmel2023). The downsides of aptamer transcription imaging are related to their degradation by nucleases, limited dye diversity, low thermal stability, and background noise (Odeh et al. Reference Odeh, Nsairat, Alshaer and Ismail2019). Nevertheless, aptamers have been extensively used to quantify transcription yields in cell-free expression systems. In particular, for PURE studies, a single Spinach aptamer sequence separated by a sufficiently long linker of 36 bases not to impede ribosomal activity or aptamer folding (Van Nies et al. Reference Van Nies, Canton, Nourian, Danelon and Burke-Aguero2015, p. 201) has been shown to be an effective transcription reporter (Van Nies et al. Reference Van Nies, Canton, Nourian, Danelon and Burke-Aguero2015; Doerr et al. Reference Doerr, De Reus, Van Nies and Danelon2019). Aptamers were also used to produce numerical models of transcription dynamics in vitro (Doerr et al. Reference Doerr, De Reus, Van Nies and Danelon2019; Zhao and Wang Reference Zhao and Wang2024). Moreover, applications of multiple repeats or tandem aptamers showcase a promising strategy to enhance the sensitivity and brightness of aptamers (Chinnappan et al. Reference Chinnappan, Dubé, Lemay and Lafontaine2013; Zhang et al. Reference Zhang, Fei, Leslie, Han, Kuhlman and Ha2015) for future synthetic cell studies.
(A) Schematic depiction of transcription in the PURE system. (B) Imaging of transcription can be achieved using molecular beacons and aptamers. (C) Examples of the ways to monitor translation using fluorescent reporters. All elements are not to scale.
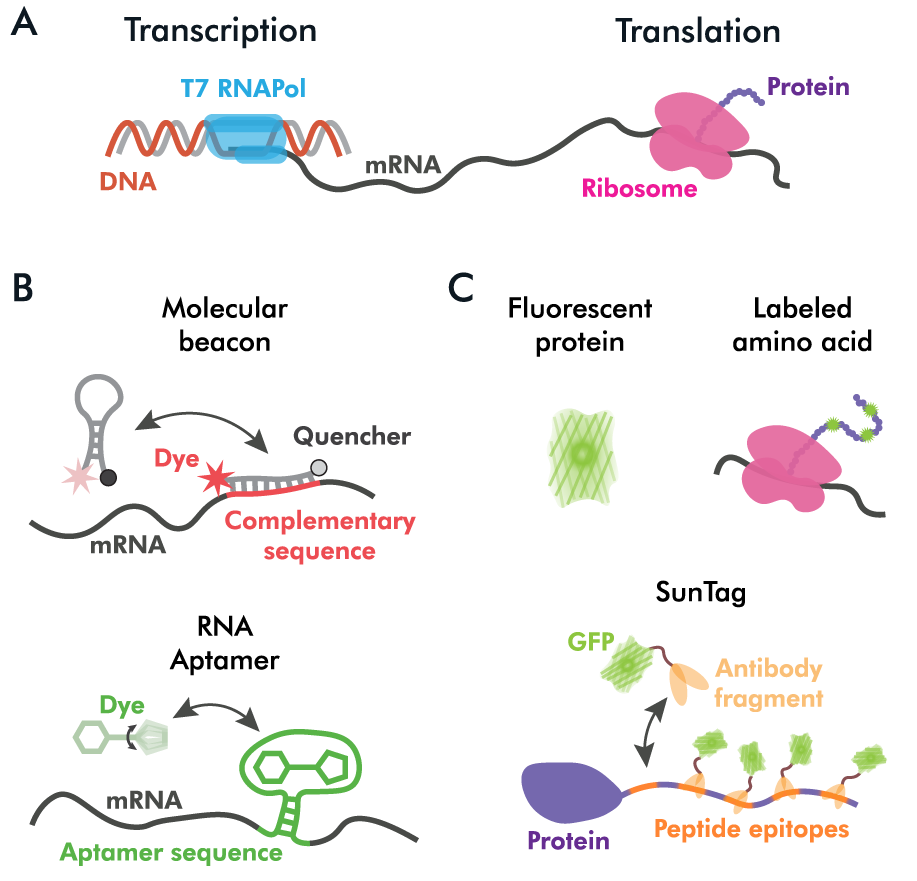
Imaging translation in vitro
The most commonly used approach for in vitro translation imaging is direct reporting with fluorescent proteins. This can be achieved using green fluorescent protein (GFP) (Prendergast and Mann Reference Prendergast and Mann1978; Prasher et al. Reference Prasher, Eckenrode, Ward, Prendergast and Cormier1992; Chalfie et al. Reference Chalfie, Tu, Euskirchen, Ward and Prasher1994) and its variants, such as enhanced GFP (eGFP) with superior fluorescent properties (McRae et al. Reference McRae, Brown and Bushell2005), superfolder GFP with better folding and higher stability (Pédelacq et al. Reference Pédelacq, Cabantous, Tran, Terwilliger and Waldo2006), and split GFP, split into two non-fluorescent fragments, thus making it apt for colocalization studies (Cabantous et al. Reference Cabantous, Terwilliger and Waldo2005; Cabantous and Waldo Reference Cabantous and Waldo2006), or with other colorful fluorescent proteins (Rodriguez et al. Reference Rodriguez, Campbell, Lin and Tsien2017). The advances in this field over the past 30 years have made it possible to monitor the expression of several proteins and to track simultaneous transcription–translation in vivo and in vitro. Examples of the in vitro applications include using Spinach aptamer in combination with yellow fluorescent protein for quantitative description of the PURE system in GUVs (Van Nies et al. Reference Van Nies, Canton, Nourian, Danelon and Burke-Aguero2015; Doerr et al. Reference Doerr, De Reus, Van Nies and Danelon2019), transcription–translation coupling studies in lysate with AlexaFluor 488 molecular beacon and eGFP (Hansen et al. Reference Hansen, Ventosa Rosquelles, Yelleswarapu and Huck2016b) and characterization of riboswitch functions in cell extract employing Mango-(IV) aptamer together with shifted GFP (Bains et al. Reference Bains, Qureshi, Ceylan, Wacker and Schwalbe2023). Going forward with transcription–translation studies, careful consideration has to be given to the spectral separation of the two probes, DNA sequence design that does not impede ribosomal activity (Lentini et al. Reference Lentini, Forlin, Martini and Mansy2013), and fluorescent protein folding times, which cause maturation delay in translation reporting.
In some cases, expressed proteins can have a substantial effect on in vitro system metabolism and the properties of the GUV. Thus, the transcription could be tracked indirectly, such as in the case of the replication machinery production inside of the GUVs (Van Nies et al. Reference Van Nies, Westerlaken, Blanken, Salas, Mencía and Danelon2018) and lipid synthesis (Scott et al. Reference Scott, Noga, de Graaf, Westerlaken, Yildirim and Danelon2016) imaged using scanning confocal microscopy. Alternatively, reporters could be incorporated into the produced protein, as in the case of FluoroTect™ GreenLys (Promega), which uses lysine-charged tRNA labeled with BODIPY®-FL to incorporate fluorescent amino acid into the protein sequence. There is limited literature on its use outside of imaging in vivo-produced proteins on a gel. Although the use of GreenLys can affect the functionality of protein when incorporated into functional domains and produce high background noise from the unincorporated amino acids, a poly-lysine appendix to the protein sequence in live bulk measurements could be a promising direction of research.
Finally, there is an option to deploy fluorescent labels that are associated with the expressed protein. SunTag is an approach that requires multiple peptide epitopes to be added to the protein sequence. Once the protein is expressed, these epitopes bind tags, comprised of single-chain variable fragment antibody, GCN4 peptide, and fluorescent protein, typically GFP. The multiplexing of 10–24 GFP copies on the SunTag scaffold enables long-term single-molecule imaging at the cost of substantial molecular weight (Tanenbaum et al. Reference Tanenbaum, Gilbert, Qi, Weissman, Vale and Edu2014). It can also carry a functional protein, for example, a DNA-interacting protein, instead of a fluorescent one and perform regulatory functions (Shakirova et al. Reference Shakirova, Ovchinnikova and Dashinimaev2020). This customizability makes SunTag another viable candidate for imaging transcription in vitro and reconstituting complex genetic networks in synthetic cells.
Discussion and outlook
Having discussed the means of reconstituting and observing transcription–translation in GUVs, we now assess the main challenges ahead. The first one is the lack of standards when it comes to developing cell-free expression systems, which are developed with the maximization of the yield in mind. After all, the motivation behind the currently available commercial cell-free expression kits, such as PURE variants, was to produce high quantities of protein of interest from a single gene plasmid in vitro (Shimizu et al. Reference Shimizu, Inoue, Tomari and Ueda2001). This is a straightforward metric that is easy to conceptualize and optimize. However, the bottom-up synthetic cell would require a different type of transcription–translation system, optimized instead for a chromosome containing a complex gene network with embedded regulatory interactions necessary for ensuring metabolic activity and reconstituting the cell cycle (Olivi et al. Reference Olivi, Berger, Creyghton and van der Oost2021). It is a whole separate problem, identifying how such a cell-free expression system would differ in design and content, yet there have already been promising steps toward this, like in the case of lysate-based and commercially available myTxTl kit, adaptable to program gene circuits with sigma factors (Shin and Noireaux Reference Shin and Noireaux2012; Garamella et al. Reference Garamella, Marshall, Rustad and Noireaux2016; Garenne et al. Reference Garenne, Thompson, Brisson, Khakimzhan and Noireaux2021b, Reference Garenne, Beisel and Noireaux2019).
On top of that, the bottom-up approach entails the convergence of various building blocks, which are often at odds with each other. As more effort is put into synthetic biology, there will be both the demand for and the spark of innovation in biophysical techniques. One such innovative field could come out of disentangling transcription–translation decoupling, which is currently focused on combining the T7 transcription and E. coli translation machinery in a bottom-up PURE system. Some optimization has already been done to address the underlying issues of inefficient ribosomal usage (Doerr et al. Reference Doerr, De Reus, Van Nies and Danelon2019, Reference Doerr, Foschepoth, Forster and Danelon2021) by increasing ribosome recycling efficiency, reducing ribosome stalling, and raising the fraction of functional full-length protein through the addition of associated factors (Li et al. Reference Li, Gu, Aach and Church2014, Reference Li, Zhang, Huang and Church2017). These attempts highlight the rational way forward. Cell lysate is a mixture of components just like the PURE system, yet the latter is 1–2 orders of magnitude less efficient, even though the transcription in it is more rapid (Gregorio et al. Reference Gregorio, Levine and Oza2019). This means that through the rational optimization of the solution composition, it should be possible to enhance the bottom-up system by an order of magnitude.
In practice, this would entail a crude, labor- and time-intensive exploration of a multiparameter space to optimize reaction performance. Among the parameters to explore are the components already included in the PURE system (Shimizu et al. Reference Shimizu, Kanamori and Ueda2005) and the additional elements taken from the list of lysate components (Foshag et al. Reference Foshag, Henrich, Hiller and Bernhard2018), but also the buffer condition components, such as crowding agents, that affect transcription–translation by changing the environment where the reaction is taking place (Deng et al. Reference Deng, Vibhute, Zheng, Zhao, Yelleswarapu and Huck2018). The number of parameters is limited but sizeable, and some of them are interconnected, such as the nucleotide and MgCl2 concentrations (Kartje et al. Reference Kartje, Janis, Mukhopadhyay and Gagnon2021), while others could be assigned a fixed value, like using a single DNA per synthetic cell (Olivi et al. Reference Olivi, Berger, Creyghton and van der Oost2021). Together, the resulting list of parameters would include several hundreds of salts, proteins, and nutrients. The exploration of this vast set of conditions using manual labor could be a waste of funding and time for highly qualified scientific staff. Instead, we envision the use of automated pipetting robots, similar to how automatization is now employed in other industries involving precise manipulation, be it pharmaceutical or machinery manufacturing. Furthermore, the navigation through this parameter space would require an algorithmic approach, where the conditions of all the past experiments and their results are considered. Bayesian optimization, commonly used for global optimization without assuming any functioning form, has found an array of applications in the time of machine learning advances (Garnett Reference Garnett2023). This approach’s focus on both random exploration of the uncharted areas of the parameter space and the narrowing down on the prospective regions to find the optimal conditions will make it useful for this task (Wakabayashi et al. Reference Wakabayashi, Otsuka, Krockenberger, Sawada, Taniyasu and Yamamoto2022). Even then, the convergence of components, the scale of the challenge, and the potential benefits warrant this to be a very costly, large-scale, long-term project similar to the previous megaprojects of the past years.
For example, the Human Genome Project (‘Human Genome Project Fact Sheet’ n.d.) took from 1990 to 2003, with some work not being finished till 2022. As a result, genomes were sequenced for species of increasing complexity, starting with Haemophilus influenzae bacteria and culminating in the entire human genome. The project, with a total cost exceeding $3 billion, was a collaboration between 20 universities, laboratories, and companies from six countries. While the financial costs may seem immense, they were covered by the economic effects of the project, which led to the explosive growth of the bioinformatics field (Venter et al. Reference Venter, Adams, Sutton, Kerlavage, Smith and Hunkapiller1998), produced unprecedented insights into our genetics used in therapeutics nowadays, and sparked the development of technologies.
Similarly, a synthetic cell project would be very costly, yet it would bring innovation to the fields of therapeutics, agriculture, and biomaterials (Khalil and Collins Reference Khalil and Collins2010). Moving toward completion would require international collaboration and long-term funding. For that reason, it is promising to see the increasing focus on synthetic biology in the last decades. The examples include institutes like JCVI (USA), collaborative networks like Build-A-Cell (USA), fabriCELL (UK), and SynCellEU (EU), and funded projects like BaSyC & EVOLF (NL) and MaxSynBio (DE). With the growing scale of these networks and projects, we could see significant progress toward understanding life as the convergence of building blocks, the development of new biophysical technologies, and the rapid growth of synthetic biology applications.
Open peer review
To view the open peer review materials for this article, please visit http://doi.org/10.1017/qrd.2024.27.
Supplementary material
The supplementary material for this article can be found at http://doi.org/10.1017/qrd.2024.27.
Acknowledgements
We express our gratitude to David Dulin for his extensive contribution to discussions on transcription–translation studies; to Kristina Ganzinger and Lori van Cauter for their collaborative spirit and knowledge-sharing on GUV production methods; to Christophe Danelone, Marileen Dogterom, Gijsje Koenderink, their lab members and other people involved in BaSyC for the workshops on the PURE system and insightful scientific meetings.
Authors contribution
VGB: writing and visualization, GJLW: writing and supervision.
Financial support
NWO funded the work as a part of Building a Synthetic Cell (BaSyC) project.
Competing interest
GJLW is a co-founder of LUMICK B.V., producing setups for single-molecule fluorescence studies of DNA–protein interactions.







Comments
Dear Editorial Board,
We hereby submit the manuscript “Challenges in transcription-translation for bottom-up synthetic biology”, which we kindly request to be considered for publication in the QRB Discovery. The manuscript presents a perspective/review of the different aspects of in vitro transcription and translation systems. This work includes different approaches taken in synthetic biology field towards making a viable artificial organism, the overview of commonly used in vitro transcription and translation systems, the comparison of diverse methods used for their encapsulation in GUVs and the imaging approaches to resolve the produced mRNA and proteins in bulk as well as with single-molecule resolution. The perspective/review aims to provide a concise overview of the developments in the bottom-up synthetic cell development, related to the transcription and translation modules, essential for a viable artificial cell. The perspective/review was written in a manner that makes it approachable for readers not familiar with the topic and insightful for people who already have knowledge of the topic.
Thank you for your consideration of our manuscript for publication in the QRB Discovery. We look forward to your reply.