Introduction
Barley (Hordeum vulgare L.) is a globally important cereal crop with a worldwide annual production of ∼146 million tons, and is the world’s fourth most important cereal crop after maize, wheat and rice (FAO 2024). Barley cultivars can broadly be divided into two main groups, based on their end-use. Two-row barley, in which each rachis node of the inflorescence contains a central fertile floret flanked by two sterile florets, is predominantly used for brewing (Fig. 1). In contrast, six-row barley, in which all three florets at each rachis node are fertile, is mostly used for animal feed (Fig. 1). The wild progenitor of barley (H. vulgare subsp. spontaneum K.Koch) has the two-rowed ear phenotype (Fig. 1), and is native to the Eastern Mediterranean, Central Asia, the Tibetan Plateau and China. Work in cultivated barley has shown that mutation at the homeodomain-leucine zipper I-class homeobox gene Six-rowed spike 1 (VRS1) results in conversion of the wild-type two-row phenotype to the six-row form (Komatsuda et al. Reference Komatsuda, Pourkheirandish, He, Azhaguvel, Kanamori, Perovic, Stein, Graner, Wicker, Tagiri, Lundqvist, Fujimura, Matsuoka, Matsumoto and Yano2007). Three mutant alleles have been described that result in a six-row phenotype (Bregitzer et al. Reference Bregitzer, Lundvist and Carollo Blake2007), each of which results in disruption of the VRS1 predicted protein (Komatsuda et al. Reference Komatsuda, Pourkheirandish, He, Azhaguvel, Kanamori, Perovic, Stein, Graner, Wicker, Tagiri, Lundqvist, Fujimura, Matsuoka, Matsumoto and Yano2007). In the vrs1.a1 allele, a single nucleotide deletion at position 681 (G681/Del) relative to the wild-type allele from cv. ‘Bonus’ leads to a frameshift within the recognition helix and premature termination of the predicted protein. The vrs1.a2 allele carries a 1 bp insertion within exon-2 (T243/Ins) that leads to a frameshift after amino acid 40 and premature truncation of the predicted protein before the start of the homeodomain. Finally, the vrs1.a3 allele has a missense mutation (C349/G) in exon-2 that results in an amino acid substitution at a highly conserved amino acid residue within the homeobox domain (F75/L).
Examples of different barley spikelet morphologies controlled by VRS1. Single spikelets viewed front on are shown in panels (a)–(d) (scale bar = 5 mm), while whole ears are shown in panels (e)–(g) in which spikelets are viewed side on (scale bar = 10 mm). (a) The deficiens type in cv. ‘Craft’ (Vrs1.t1 allele) in which the two sterile florets flanking the central fertile floret are severely reduced in form. (b) The two-row type in barley cv. ‘Maris Otter’ (allele Vrs1.b3). (c) The six-row type in cv. ‘KWS Feeris’ in which all three florets per rachis node are fertile. (d) Wild barley (Hordeum vulgare subsp. Spontaneum) two-row form (HorID accession 10924. Vrs1.b2 allele) in which at each rachis node, the central fertile floret is flanked by two sterile florets. (e–g) Whole ears: (e) the two-row deficiens barley cv. ‘Craft’ (Vrs1.t1), (f) the two-row barley cv. ‘Maris Otter’ (Vrs1.b3) and (g) the six-row barley cv. ‘KWS Feeris’. Fs = fertile spikelet, ss = sterile spikelet, vs = vestigial spikelet.

Barley domestication began in the Fertile Crescent region of the Near East around 10,000 years ago with the shift from hunter-gatherer societies to farming (Hillman Reference Hillman1975; Zohary et al. Reference Zohary, Hopf and Weiss2012; Fuller et al. Reference Fuller, Denham and Allaby2023; Guo et al. Reference Guo, Jayakodi, Himmelbach, Ben-Yosef, Davidovich, David, Hartmann-Shenkman, Kislev, Fahima, Schuenemann and Reiter2025). Post-domestication, human activity led to the spread of barley cultivation from its origins in the Fertile Crescent, first into Europe, Asia and Northern Africa (Zohary et al. Reference Zohary, Hopf and Weiss2012), and subsequently to the Americas and Oceania. The first domesticated barley was two-rowed, first appearing in the archaeobotanical record ∼10,000 years before present at sites in Syria (van Zeist and Bakker-Heeres Reference van Zeist and Bakker-Heeres1985) and Iraq (Braidwood, Reference Braidwood1960; Helbaek Reference Helbaek1959). Around 1,000 years later, six-row types emerged as key Neolithic innovations in the Near East, largely replacing the two-row cultivated form to become the most important crop in the Near East during the Neolithic (Helbaek Reference Helbaek1959; Harlan Reference Harlan1968; Pourkheirandish and Komatsuda Reference Pourkheirandish and Komatsuda2007; Zohary et al. Reference Zohary, Hopf and Weiss2012). Prior to the use of crop cultivars with the advent of industrial plant breeding approaches in the late 19th century, the post-domestication spread of barley was via the use and transport of germplasm stocks now termed ‘landraces’ (germplasm grown and selected by growers for use in their local agro-ecological regions of cultivation and for specific end-uses) (Jones et al. Reference Jones, Civan, Cockram, Leigh, Smith, Jones, Charles, Molina-Cano, Powell, Jones and Brown2011). Analysis of genetic markers and VRS1 haplotypes associated with the different Vrs1 alleles in barley cultivars, landraces and wild barley has found that the six-row alleles vrs1.a2 and vrs1.a3 likely arose post-domestication in cultivated barley via single mutations in their respective wild-type alleles (Tanno et al. Reference Tanno, Taketa, Takeda and Komatsuda2002; Komatsuda et al. Reference Komatsuda, Pourkheirandish, He, Azhaguvel, Kanamori, Perovic, Stein, Graner, Wicker, Tagiri, Lundqvist, Fujimura, Matsuoka, Matsumoto and Yano2007; Saisho et al. Reference Saisho, Pourkheirandish, Kanamori, Matsumoto and Komatsuda2009). Specifically, vrs1.a2 is thought to have directly arisen from Vrs1.b2 in barley cultivated in the Western Mediterranean, and vrs1.a3 from Vrs1.b3 in East Asia. However, while vrs1.a1 is the most commonly used six-row allele in modern barley cultivars (Komatsuda et al. Reference Komatsuda, Pourkheirandish, He, Azhaguvel, Kanamori, Perovic, Stein, Graner, Wicker, Tagiri, Lundqvist, Fujimura, Matsuoka, Matsumoto and Yano2007; Saisho et al. Reference Saisho, Pourkheirandish, Kanamori, Matsumoto and Komatsuda2009; Casas et al. Reference Casas, Contreras-Moreira, Cantalapiedra, Sakuma, Gracia, Moralejo, Molina-Cano, Komatsuda and Igartua2018), the hypothetical progenitor Vrs1.b1 allele (i.e. an allele represented by a haplotype that is identical to that carried by the vrs1.a1 allele, save for the single causative 1 bp deletion that results in truncation of the vrs1.a1 predicted protein) has not been identified in cultivated barley.
Allelic variation at VRS1 also affects additional barley traits. For example, the extreme suppression of the infertile lateral spikelets is observed in the naturally occurring deficiens allele Vrs1.t1 (Fig. 1c), which carries three amino acid substitutions in its predicted protein compared to the wild-type allele from cv. ‘Bonus’ (D8/G, D26/E and S184/G) has been shown via comparison to the induced deficiens mutants Def2 (Vrs1.t2, S184/G) and Def3 (Vrs1.t3, S186/P) to be due to the S184/G amino acid substitution at a conserved residue towards the C-terminus of the VRS1 protein (Sakuma et al. Reference Sakuma, Lundqvist, Kakei, Thirulogachandar, Suzuki, Hori, Jianzhong, Tagiri, Rutten, Koppolu, Shimada, Houston, Thomas, Waugh, Schnurbusch and Komatsuda2017). The six-row vrs1.a1 allele also confers increased leaf size, predominantly due to an increase in leaf width, as well as increased leaf vein number and percentage nitrogen content (Thirulogachandar et al. Reference Thirulogachandar, Alqudah, Koppolu, Rutten, Graner, Hensel, Kumlehn, Bräutigam, Sreenivasulu, Schnurbusch and Kuhlmann2017). VRS1 is expressed during leaf primordia initiation and development, where it represses the proliferation of leaf and lateral spikelet primordia (Sakuma et al. Reference Sakuma, Lundqvist, Kakei, Thirulogachandar, Suzuki, Hori, Jianzhong, Tagiri, Rutten, Koppolu, Shimada, Houston, Thomas, Waugh, Schnurbusch and Komatsuda2017; Thirulogachandar et al. Reference Thirulogachandar, Alqudah, Koppolu, Rutten, Graner, Hensel, Kumlehn, Bräutigam, Sreenivasulu, Schnurbusch and Kuhlmann2017). The six-row allele vrs1.a1, as well as the intermedium alleles vrs1(int-d.11) and vrs1(int-d.22), are also known to confer reduced tiller number at maturity (Liller et al. Reference Liller, Neuhaus, von Korff, Koornneef and van Esse2015; Thirulogachandar et al. Reference Thirulogachandar, Alqudah, Koppolu, Rutten, Graner, Hensel, Kumlehn, Bräutigam, Sreenivasulu, Schnurbusch and Kuhlmann2017). This reduction in tiller number is not observed at earlier stages of development, indicating VRS1 does not affect the initiation and initial growth of tillers. The increased grain number per spike and reduced absolute spikelet abortion observed in six-row types, despite the presence of three times the number of fertile spikelets compared to two-row barley, is hypothesized to be due, at least in part, to the increased leaf area found in six-row types (Alqudah and Schnurbusch Reference Alqudah and Schnurbusch2015; Thirulogachandar et al. Reference Thirulogachandar, Alqudah, Koppolu, Rutten, Graner, Hensel, Kumlehn, Bräutigam, Sreenivasulu, Schnurbusch and Kuhlmann2017). Given the numerous positive effects of six-row vrs1 alleles on ear and leaf traits, the reduction of tiller number at later stages in the plant lifecycle is thought to be due to negative trade-off interactions that occur after tiller initiation (Thirulogachandar et al. Reference Thirulogachandar, Alqudah, Koppolu, Rutten, Graner, Hensel, Kumlehn, Bräutigam, Sreenivasulu, Schnurbusch and Kuhlmann2017).
Given the key role of VRS1 in the barley domestication and post-domestication process, and the emerging findings that VRS1 has pleiotropic effects on other traits of potential agronomic interest in barley, here we screen a panel of wild, landrace and cultivated barley accessions previously phenotyped for ear-row type with the aim of (i) Identifying sources of VRS1 missense variation in the wider barley genepool, and (ii) resolving the evolutionary origin of the six-row vrs1.a1 allele.
Materials and methods
Barley germplasm and DNA extraction
Barley's scientific name nomenclature follows that outlined by POWO (2025). Seed for 69 wild barley and 170 barley landrace accessions was obtained from the genebanks listed in Supplementary Table S1, as previously described by Jones et al. (Reference Jones, Leigh, Mackay, Bower, Smith, Charles, Jones, Jones, Brown and Powell2008, Reference Jones, Civan, Cockram, Leigh, Smith, Jones, Charles, Molina-Cano, Powell, Jones and Brown2011). Additionally, seed for 98 of the barley cultivars described by Rostoks et al. (Reference Rostoks, Ramsay, MacKenzie, Cardle, Bhat, Roose, Svensson, Stein, Varshney, Marshall, Graner, Close and Waugh2006) was sourced from the James Hutton Institute, Dundee, UK (Supplementary Table S1). Genomic DNA was extracted from seedling leaf material collected from a single plant per accession using a DNeasy Kit (Qiagen), following the manufacturer’s instructions. DNA concentration was estimated using a NanoDrop spectrophotometer (Thermo Fisher Scientific) and diluted to 10 ng/μl using molecular biology-grade water (Thermo Fisher Scientific).
VRS1 polymerase chain reaction amplification and sequencing
VRS1 was amplified via polymerase chain reaction (PCR) using the primers OSU-VRS1-F1 (5’-CCGATCACCTTCACATCTCC-3’) and OSU-VRS1-R1 (5’-GGTTTCTGCCGATCTTGAAGC-3’). PCR amplification was undertaken in 10 µl reactions using the FastStart Taq DNA polymerase kit (Roche) and a Veriti 96-well Thermo Cycler (Applied Biosystems) using the following PCR cycle: 6 min at 96°C, followed by 35 cycles of 50 sec at 96°C, 50 sec at 60°C, 90 sec at 72°C, and a final extension of 7 min at 72°C. PCR products were subsampled to check for adequate amplification and expected product size via electrophoresis across ethidium bromide stained 1% agarose gels alongside a 1 kb DNA ladder. Amplification products were ExoSap treated in reactions containing 1.5 U of Exonuclease I (New England Biolabs) and 1 U of Shrimp Alkaline Phosphatase (Promega) and incubated at 37°C for 60 min, followed by 15 min at 96°C using a Veriti 96-well Thermo Cycler. Sanger sequencing was then undertaken using the ExoSap-treated PCR products using BigDye Kit v3.1 (Applied Biosystems) in 10 μl reactions following the manufacturer’s instructions, and the resulting fluorescently end-labelled single-stranded extension products were separated and visualized using a 3730 DNA Analyzer (Applied Biosystems). Five primers were used for Sanger sequencing: OSU-VRS1-F1, OSU-VRS1-R1, HvVRS1-F1 (5’- CAGAACAACCTACCGTGTCT-3’), HvVRS1-F2 (5’- AGATGGACGGAGGAGGGGAC-3’) and HvVRS1-R1 (5’- TGTCATCAGCTTAGCCAGCA-3’). DNA sequence traces were manually quality controlled and contigged using Contig Express (Thermo Fisher Scientific). VRS1 DNA sequence polymorphisms, including single-nucleotide polymorphisms (SNPs) and 1-2 bp insertions/deletions (InDels), were positioned with reference to the genomic sequence of the two-row Vrs1.b3 allele from barley cultivar ‘Bonus’ (GenBank accession AB489121), with the A of the ATG start codon representing position 1 bp.
Bioinformatic, haplotype and geographical analyses
VRS1 DNA sequences, and their corresponding predicted protein sequences determined using EMBOSS Transeq, were aligned using Clustal Omega (Madeira et al. Reference Madeira, Madhusoodanan, Lee, Eusebi, Niewielska, Tivey, Lopez and Butcher2024). Genomic VRS1 sequences were used for haplotype analysis using the software PopArt, which implements the minimum spanning tree method (Leigh and Bryant Reference Leigh and Bryant2015). Using the collection location latitude and longitude information Supplementary Table S1), the VRS1 haplotypes for the wild and landrace barley accessions were plotted on geographic maps using the packages ggplot2 (Wickham Reference Wickham2016) and rnaturalearth (Massicotte and South Reference Massicotte and South2025) in R version 4.4.0 (R Core Team 2024). Visualization of VRS1 haplotypes across germplasm groups was conducted using the R packages ggplot2 and ggalluvial (Brunson and Quentin Reference Brunson and Quentin2023). Two-row wild-type (Vrs1.b2, Vrs1.b3), six-row mutant (vrs1.a1, vrs1.b1, vrs1.c1) and two-row deficiens (Vrs1.t1) alleles are designated as previously described (e.g. Komatsuda et al. Reference Komatsuda, Pourkheirandish, He, Azhaguvel, Kanamori, Perovic, Stein, Graner, Wicker, Tagiri, Lundqvist, Fujimura, Matsuoka, Matsumoto and Yano2007); VRS1 haplotypes are designated here based on the allele nomenclature in non-italic, along with the inclusion of a numerical suffix (.X, where X = a number). Within each major haplotype (e.g. Vrs1.a1), sub-haplotype numbering is sequential based on the position of the first DNA variant in the sequenced region that differs from the reference allele used here for variant calling (e.g. Vrs1.a1.1, for which the first DNA variant that distinguishes it from the canonical Vrs1.a1 haplotype is A-189/G, located in the VRS1 promoter region). Protein 3D structure was investigated using the AlphaFold protein models for the VRS1 gene model HORVU.GOLDEN_PROMISE.PROJ.2HG00164530.1 from two-row barley cv. ‘Golden Promise’ (genome assembly: Hvulgare_cv_Golden_Promise_BPGv2, INSDC Assembly, European Nucleotide Archive assembly GCA_949783185.1), accessed using Ensembl Plants (Yates et al. Reference Yates, Allen, Amode, Azov, Barba, Becerra, Bhai, Campbell, Carbajo, Chakiachvili and Chougule2022).
Results
VRS1 haplotype analysis in cultivated, landrace and wild barley
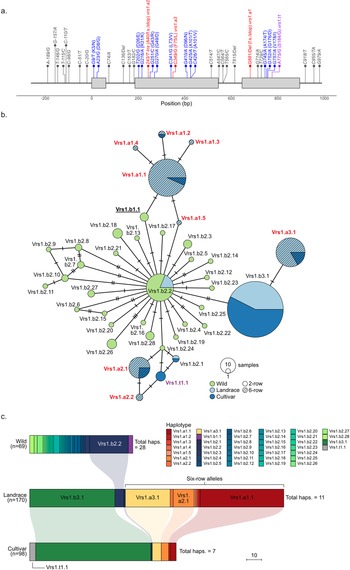
To provide baseline data from the modern cultivated barley genepool, sequencing VRS1 in the 98 predominantly European cultivars from the ‘pre-AGOUEB’ barley collection was undertaken, with haplotype analysis on the resulting nine DNA variants identified (Fig. 2a), and found all to possess one of six common VRS1 alleles (Fig. 2b and c; Supplementary Tables S1–S2). Amongst the 82 two-row cultivars analyzed, four were phenotypically deficiens and carried a VRS1 haplotype consistent with the Vrs1.t1 deficiens allele. Of the remaining two-row cultivars, one carried the Vrs1.b2 haplotype (cv. ‘Camargue’, a German spring barley variety released in 1986), while the remaining 78 cultivars all carried the Vrs1.b3 allele. For the 16 six-row cultivars, VRS1 haplotype analysis found roughly equal numbers of cultivars to carry the six-row alleles vrs1.a1 (five cultivars), vrs1.a2 (five cultivars) and vrs1.a3 (five cultivars and 1 morphological stock).
VRS1 DNA variants and haplotypes identified in wild, landrace and cultivated barley accessions. (a) Positions of VRS1 DNA variants, relative to the Vrs1.b3 two-row allele from cv. ‘Bonus’ (GenBank accession AB489121). Variants leading to an amino acid substitution are indicated in blue, those resulting in the six-row phenotype in red, and the mutation resulting in the Vrs1.t1 deficiens allele highlighted in purple. Variants that do not lead to a change in the predicted protein are shown in grey. (b) VRS1 haplotype network indicating the relationships between the 39 haplotypes identified across the three barley genepools. The size of each haplotype is proportional to the number of accessions, with haplotypes representing accessions with the two- or six-row phenotype indicated (see figure key). The number of DNA variants distinguishing linked haplotypes is indicated using the short bisecting lines. (c) VRS1 haplotype proportions across the three barley genepools. Vrs1.a mutant haplotypes associated with six-row alleles are indicated in the brown-to-Orange range, while the Vrs1.t1.1 haplotype corresponding to the two-row deficiens allele is shown in grey. All remaining Vrs1.b haplotypes correspond to two-row alleles. Haplotypes present in more than one genepool are linked via correspondingly coloured ribbons. The number (n) of accessions within each genepool is indicated, and the scale bar represents a group size of 10 accessions.

Next, VRS1 sequencing in a collection of 170 European barley landraces (identifying 11 DNA variants) and 69 wild barley accessions from across its natural range (33 DNA variants; Fig. 2a) allowed haplotypes and corresponding alleles to be determined across all 239 accessions (Fig. 2b; Supplementary Tables S1–S2). Similar to the finding in modern varieties, the most common two-row landrace allele, as determined by VRS1 haplotype, was Vrs1.b3 (57 accessions, representing 90% of the two-row barley landraces), with the remaining carrying either vrs1.b2 (HorID 1393 from France) or a sub-haplotype of the vrs1.b2 allele, which lacks the intron-2 C564/T SNP (six accessions, from Syria, Afghanistan, the Czech Republic and Bolivia) and termed here haplotype Vrs1.b2.2 (Fig. 2b and c). Of the 107 six-row landraces, 53% carried haplotypes consistent with a vrs1.a1 allele (57 accessions), while the remaining possessed one of the following four sub-haplotypes: Vrs1.a1.2 (HorID 2672, carrying an A-189/G SNP in the upstream region), Vrs1.a1.3 (HorID 47, carrying a C-20/G SNP in the upstream region), Vrs1.a1.4 (HorID 1173 from Ukraine, the only landrace carrying a missense mutation: SNP C426/T, amino acid substitution A101/V), and Vrs1.a1.5 (HorID 1214, which lacked the 3’ UTR C918/T variant found in all other Vrs1.a1 haplotypes) (Fig. 2b). In the landraces, haplotype analysis identified just one accession predicted to carry the two-row Vrs1.b2 allele (HorID 1393. Collection location: France), while analysis of haplotype genealogy found no landraces carried a VRS1 haplotype directly ancestral to the six-row vrs1.a1 allele (Fig. 2b).
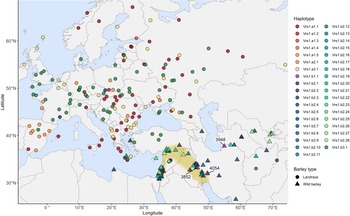
VRS1 haplotype analysis found 66 of the 69 two-row wild barley accessions to possess haplotypes indicative of the wild-type Vrs1.b2 allele (Fig. 2b; Supplementary Tables S1–-S2). A total of 26 Vrs1.b2 sub-haplotypes were identified, which possessed between 1 and 3 DNA variants relative to the Vrs1.b2.1 haplotype found in cultivated barley. Twenty-seven of the 28 wild barley Vrs1.b2 sub-haplotypes lacked the intron-2 C564/T SNP found in cultivated barley. The exception was a wild barley accession from Cyprus (HorID 4042), which, although it possessed the intron-2 C564/T SNP, also contained a 3’ UTR SNP (G997/T) that differentiated it from the canonical Vrs1.b2.1 haplotype found in modern barley. The most common wild barley haplotype was Vrs1.b1.2, which was present in 30% of the accessions screened, with the remaining Vrs1.b1 sub-haplotypes representing rare occurrences, each present in between 1 and 4 accessions. Notably, analysis of VRS1 haplotype genealogy identified three wild barley accessions as carrying a haplotype consistent with the hypothesized Vrs1.b1 allele directly ancestral to the six-row allele vrs1.a1 (the most common mutant six-row allele found in cultivated barley) (Fig. 2b). All three of the wild barley accessions carry the Vrs1.a1 haplotype originated within the Fertile Crescent: two from Iran and one from Turkmenistan (HorIDs 3852, 4054 and 3948, respectively) (Fig. 3).
Geographic locations of the landrace (n = 161) and wild barley (n = 69) accessions with collection location information, indicating their VRS1 haplotypes. Haplotypes corresponding to the wild-type Vrs1.b1, Vrs1.b2 and Vrs1.b3 two-row alleles are indicated using the blue-to-green colour scale. Haplotypes corresponding to the mutant vrs1.a1, vrs1.a2 and vrs1.a3 six-row alleles are indicated using the red-to-yellow colour scale. The three wild barley accessions (HorID 3852, 3948 and 4054) were found to carry the Vrs1.b1.1 haplotype ancestral to the mutant vrs1.a1 six-row allele is indicated. The region representing the Fertile Crescent is shaded in brown.

Landrace and wild barley VRS1 missense mutations
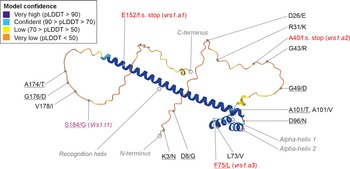
Overall, 42 SNPs or InDels across the 1,251 bp sequenced region of VRS1 were identified in the landrace and wild barley accessions. VRS1 haplotype diversity decreased between the wild barley and landrace genepools (27 and 11 haplotypes, respectively), with a further contraction in diversity found across the transition from landrace to modern cultivars (7 haplotypes) (Fig. 2c). While much novel variation was identified at the DNA level in landrace and wild barley accessions, only a subset of variants resulted in novel changes to the predicted VRS1 protein. While these haplotypes conferred wild-type two-row alleles, the increasing evidence of pleiotropic effects of VRS1 on traits other than row number means that such mutations may be of wider interest. In wild barley, all wild-type two-row VRS1 haplotypes contained the two missense mutations that lead to amino acid substitutions D8/G and D26/E, which in modern two-row barley cultivars are only found in the relatively rare Vrs1.b2 allele (Fig. 2b and c, Supplementary Table S1). In addition to these, 10 two-row Vrs1.b2 sub-haplotypes contained missense mutations that resulted in a change in amino acid residue elsewhere within the predicted protein. Of these, six mutations were located either within the N-terminus (K3/N, R31/K, G43/R) or C-terminus (A174/T, G176/D, V178/I) regions. While both have low model confidence scores in the VRS1 AlphaFold 3D protein model (pLDDT ≤52), the acid substitutions in the C-terminus region are located at residues with good levels of amino acid conservation across plant VRS1 homologues (Fig. 4). In contrast to the unstructured N- and C-terminus regions, the predicted homeobox-leucine zipper domain (Panther ID PTHR24326) has high AlphaFold 3D model confidence (pLDDT typically >90 across almost all but the first 15 and last 6 amino acid residues of the domain). This domain is predicted to consist of three principal structures: the first and second alpha-helixes, which serve as primary structural elements, and the recognition helix, which determines DNA binding and specificity. Four VRS1 haplotypes were found to contain amino acid changes within the homeobox-leucine zipper domain at amino acid residue with high conservation in plant VRS1 homologues: missense mutation L73/V (haplotype Vrs1.b2.15, 68% conservation) located towards the end of the first alpha-helix, D96/N (haplotype Vrs1.b2.20, 100% conservation) located in the connecting loop between the second alpha-helix and the recognition helix, and A101/T (haplotype Vrs1.b2.21, 100% conservation) close to the start of the recognition helix itself (Fig. 4; Supplementary Figure S1).
VRS1 AlphaFold 3D protein model for two-row barley cultivar ‘Golden Promise’ (UniProt accession D2KV17, encoded by the Vrs1.b3 allele) overlaid with the positions of the amino acid substitutions coded for by DNA variants identified in the wild, landrace and cultivar barley genepools investigated. 3D protein model confidence is colour-coded as indicated in the key. The homoeobox-leucine zipper domain (Panther ID PTHR24326) overlaps with the ‘very high’ protein model confidence, and consists of alpha-helix 1, alpha-helix 2 and the recognition helix. Non-conservative substitutions are underlined. Mutations causative for the three known six-row alleles are indicated in red. The mutations underlying the two-row deficiens allele are indicated in purple. F.S. = frame shift. Stop = premature stop codon.

Discussion
Domestication of the six-row phenotype preserved VRS1 haplotype diversity in cultivated barley
All wild barley accessions analyzed here were phenotypically two-row and carried two-row VRS1 haplotypes, indicating no instances of feral six-row individuals in our panel. Haplotype analysis of landraces and cultivars identified all three previously described six-row alleles, vrs1.a1, vrs1.a2 and vrs1.a3, consistent with previous studies of cultivated barley germplasm from this and other regions (Komatsuda et al. Reference Komatsuda, Pourkheirandish, He, Azhaguvel, Kanamori, Perovic, Stein, Graner, Wicker, Tagiri, Lundqvist, Fujimura, Matsuoka, Matsumoto and Yano2007; Saisho et al. Reference Saisho, Pourkheirandish, Kanamori, Matsumoto and Komatsuda2009; Cuesta-Marcos et al. Reference Cuesta-Marcos, Szűcs, Close, Filichkin, Muehlbauer, Smith and Hayes2010; Sakuma et al. Reference Sakuma, Lundqvist, Kakei, Thirulogachandar, Suzuki, Hori, Jianzhong, Tagiri, Rutten, Koppolu, Shimada, Houston, Thomas, Waugh, Schnurbusch and Komatsuda2017). In our analysis of 337 barley accessions, we found VRS1 haplotype diversity declined sharply from wild barley to landraces (28 → 11), but only modestly from landrace to cultivars (11 → 7, with six-row haplotypes predominating in both of these genepools). This suggests that six-row mutations that arose post-domestication were opportunistically selected due to their visually striking mutations, thus increasing haplotype diversity at the locus. Indeed, previous studies based on 32 cultivars and three wild barley accessions suggested that the six-row alleles vrs1.a2 and vrs1.a3 originated via single point mutations from the corresponding wild-type Vrs1.b2 and Vrs1.b3 alleles in landraces from East Asia and the Western Mediterranean, respectively (Komatsuda et al. Reference Komatsuda, Pourkheirandish, He, Azhaguvel, Kanamori, Perovic, Stein, Graner, Wicker, Tagiri, Lundqvist, Fujimura, Matsuoka, Matsumoto and Yano2007). Our data broadly support these hypotheses: landraces predicted to carry the vrs1.a2 allele originated from Spain, while the haplotypes are predictive of vrs1.a3 occurred primarily in Russia and Eastern Europe (Fig. 3). In summary, we conclude that the use of six-row alleles ameliorates to some extent the loss in haplotype diversity that would otherwise be expected across the transition from wild barley to modern cultivars, and that the domestication origin of these six-row alleles is still evident in their distribution within landraces.
Evidence that the Vrs1.b1 allele arose in the wild barley genepool
The geographic origin of the vrs1.a1 six-row allele has remained unclear (Komatsuda et al. Reference Komatsuda, Pourkheirandish, He, Azhaguvel, Kanamori, Perovic, Stein, Graner, Wicker, Tagiri, Lundqvist, Fujimura, Matsuoka, Matsumoto and Yano2007) due to the absence of a directly ancestral wild-type two-row haplotype representing the hypothetical Vrs1.b1 allele. This is somewhat unexpected, given that vrs1.a1 has been reported as the most common six-row allele in landraces (Saisho et al. Reference Saisho, Pourkheirandish, Kanamori, Matsumoto and Komatsuda2009) and cultivars (Komatsuda et al. Reference Komatsuda, Pourkheirandish, He, Azhaguvel, Kanamori, Perovic, Stein, Graner, Wicker, Tagiri, Lundqvist, Fujimura, Matsuoka, Matsumoto and Yano2007). In the 170 predominantly European barley landraces investigated, we also found vrs1.a1 to be the most common landrace six-row allele, widely distributed across most of the geographic regions assessed here, suggesting it was the first to be domesticated. This is consistent with a recent genome-wide sequence-based estimate of barley haplotype origin around the VRS1 locus, which found the haplotype spanning vrs1.a1 to be the most ancient of the three six-row alleles (Guo et al. Reference Guo, Jayakodi, Himmelbach, Ben-Yosef, Davidovich, David, Hartmann-Shenkman, Kislev, Fahima, Schuenemann and Reiter2025). Indeed, while only two six-row landraces within the Fertile Crescent were investigated here, both were found to contain haplotypes consistent with a vrs1.a1 allele (Fig. 3). However, we identified three wild barley accessions carrying a Vrs1.b1.1 haplotype, differing from the vrs1.a1 allele by just the causative 1 bp exon-3 deletion that leads to a truncated VRS1 protein. Two of these Vrs1.a1.1 accessions were located in western Iran (Ilam and Khuzestan provinces) in the foothills of the Zagros mountains, north-east of the Tigris river within the Fertile Crescent, while the third was from Turkmenistan near the Atrak river that forms the border between Turkmenistan and Iran, and situated to the south-east of the Caspian Sea. This Caspian Sea region also played a notable role in the domestication of the related temperate cereal crop, bread wheat (Triticum aestivum), which arose in Neolithic farmers’ fields via rare natural hybridization events between cultivated tetraploid wheat and the wild diploid wheat Aegilops tauschii (Wang et al. Reference Wang, Luo, Chen, You, Wei, Zheng and Dvorak2013; Gaurav et al. Reference Gaurav, Arora, Silva, Sánchez-Martín, Horsnell, Gao, GS, Widrig, John, Singh and Wu2022). Vrs1.b1.1 haplotypes were recently identified in two wild barley accessions via analysis of genomic sequence data generated using Illumina HiSeq 200 sequencing: FT897 and FT879 (ENA records ERX692648 and ERX692649, respectively). The collection sites for both were from the Karmanshah province of north-western Iran, within the foothills of the Zagros mountains. Collectively, these findings suggest that Vrs1.b1 alleles were present at low frequency in stands of wild barley growing in regions of the Fertile Crescent where barley domestication occurred, as well as in the regions flanking the southern shores of the Caspian Sea where major Neolithic cereal domestication events occurred.
Reduction and shift in two-row VRS1 haplotype diversity during domestication
In contrast to the emergence and proliferation of six-row alleles in the landraces, we found two-row VRS1 haplotypes to have undergone a sharp reduction in diversity from wild barley to landraces (29 → 4), an expected general outcome of domestication (e.g. Saisho et al. Reference Saisho, Pourkheirandish, Kanamori, Matsumoto and Komatsuda2009; Milner et al. Reference Milner, Jost, Taketa, Mazón, Himmelbach, Oppermann, Weise, Knüpffer, Basterrechea, König and Schüler2019; Cheng et al. Reference Cheng, Feng, Wingen, Cheng, Riche, Jiang, Leverington-Waite, Huang, Collier, Orford and Wang2024; Civáň et al. Reference Civáň, Fricano, Russell, Pont, Özkan, Kilian and Brown2024). Two features stand out: Firstly, the predominant haplotype in wild barley (Vrs1.b2.2) sharply declined in frequency amongst landraces, and was completely absent in cultivars (although the Vrs1.b2 allele was transmitted between the landrace and cultivar genepools via haplotype Vrs1.b2.1, albeit at extremely low frequency in both germplasm groups). Secondly, the two-row haplotype Vrs1.b3.1, which is absent from wild barley, predominates in both landrace (89%) and cultivar (94%) genepools. These patterns indicate that two-row haplotypes experienced a strong genetic bottleneck, likely driven by a founder effect followed by subsequent expansion of Vrs1.b3.1, potentially reinforced by human selection. Notably, haplotypes consistent with the Vrs1.b2 allele carry two missense mutations (D8/G and D26/E) in the VRS1 N-terminus region also found in the rare Vrs1.b1 allele, the deficiens Vrs1.t1 allele, and the six-row vrs1.a1 and vrs1.a3 alleles that are characterized by truncated VRS1 proteins. Although these amino acid substitutions occur at relatively un-conserved residues within an unstructured region of the protein, they may be associated with pleiotropic effects. For example, the six-row mutant allele vrs1.a1 (which shares both substitutions in the region upstream of the premature stop codon) is associated with reduced tillering, while the induced six-row mutants vrs1(hex-v.3) (a null VRS1 deletion) and vrs1(hex-v.6) (derived from cv. ‘Foma’, Vrs1.b3), and so do not carry the D8/G and D26/E substitutions, show no significant tiller reduction (Liller et al. Reference Liller, Neuhaus, von Korff, Koornneef and van Esse2015).
VRS1 variation affecting protein sequence in wild barley
Ten wild barley haplotypes contained variants that altered the VRS1 protein sequence. Allelic variation at VRS1 is known to affect additional barley traits, making novel variants that affect the VRS1 protein sequence of potential modern-day utility. Some of these pleiotropic traits are directly linked to the spikelets themselves, such as the extreme suppression of the infertile lateral spikelets observed in the deficiens allele Vrs1.t1 (conferred by the S184/G amino acid substitution at a conserved residue towards the C-terminus of the VRS1 protein) (Sakuma et al. Reference Sakuma, Lundqvist, Kakei, Thirulogachandar, Suzuki, Hori, Jianzhong, Tagiri, Rutten, Koppolu, Shimada, Houston, Thomas, Waugh, Schnurbusch and Komatsuda2017) and reduced lemma extension (Saisho et al. Reference Saisho, Pourkheirandish, Kanamori, Matsumoto and Komatsuda2009). However, VRS1 has also been shown to affect tiller number (Liller et al. Reference Liller, Neuhaus, von Korff, Koornneef and van Esse2015; Alqudah et al. Reference Alqudah, Koppolu, Wolde, Graner and Schnurbusch2016; Thirulogachandar et al. Reference Thirulogachandar, Alqudah, Koppolu, Rutten, Graner, Hensel, Kumlehn, Bräutigam, Sreenivasulu, Schnurbusch and Kuhlmann2017), leaf size (Thirulogachandar et al. Reference Thirulogachandar, Alqudah, Koppolu, Rutten, Graner, Hensel, Kumlehn, Bräutigam, Sreenivasulu, Schnurbusch and Kuhlmann2017), and plant height (Alqudah et al. Reference Alqudah, Koppolu, Wolde, Graner and Schnurbusch2016). Accordingly, VRS1 negatively controls lateral floret development and leaf primordia initiation and development, where it is hypothesized to suppress the cell proliferation of leaf founder cells that form the leaf primordia. Four VRS1 haplotypes carried missense mutations within the highly conserved homeobox-leucine zipper (HD-ZIP) domain, of which two (L73/V in Vrs1.b2.15 and D96/N in Vrs1.b2.20) fall in or near the first two alpha helices known to play an essential role in protein positioning at the target DNA site in other HD-ZIP proteins. The proximity of the conservative L73/V substitution to the causal F75/L substitution underlying the six-row vrs1.a3 allele, and three independent six-row VRS1 mutants (Komatsuda et al. Reference Komatsuda, Pourkheirandish, He, Azhaguvel, Kanamori, Perovic, Stein, Graner, Wicker, Tagiri, Lundqvist, Fujimura, Matsuoka, Matsumoto and Yano2007), underscores the domain’s functional importance. Additional wild barley mutations within the connecting loop between the second alpha helix and the recognition helix (non-conservative D96/N; haplotype Vrs1.b2.20) and close to the start of the recognition helix (conservative A101/T; haplotype Vrs1.b2.21) lie near residues where ≥16 induced missense mutants cause six-row or intermedium phenotype (see Supplementary Text 1) (Komatsuda et al. Reference Komatsuda, Pourkheirandish, He, Azhaguvel, Kanamori, Perovic, Stein, Graner, Wicker, Tagiri, Lundqvist, Fujimura, Matsuoka, Matsumoto and Yano2007; Gottwald et al. Reference Gottwald, Bauer, Komatsuda, Lundqvist and Stein2009). Substitutions at residue 107 exemplify this effect: non-conservative (R107/L from hex-v.42 and hex-v.43) mutations confer six-row alleles, while conservative substitutions (R107/H from cv. ‘Barke’ TILLING line 11657-1) result in intermedium phenotype.
While the three haplotypes leading to amino acid changes in the N-terminus regions were not located within named protein domains, two are found within a region predicted to form an alpha fold structure at amino acid residues with high conservation across homologous plant proteins (conservative R31/K and non-conservative G43/R substitutions, originating from haplotypes Vrs1.b2.17 and Vrs1.b2.18, respectively). Finally, while the C-terminus region in which the three wild barley mutations affecting protein sequence (conservative A174/T, non-conservative G176/D and conservative V178/I substitutions) does not contain notable 3D structure, much of this region is relatively highly conserved across VRS1 homologues. The importance of this region is highlighted by the presence of the non-conservative D184/G amino acid mutation underlying the deficiens allele Vrs1.t1 (Gottwald et al. Reference Gottwald, Bauer, Komatsuda, Lundqvist and Stein2009; Hansson et al. Reference Hansson, Youssef, Zakhrabekova, Stuart, Svensson, Dockter, Stein, Waugh, Lundqvist and Franckowiak2024), as well as the intermedium mutation C194/S in vrs1(Int-d.11) (Komatsuda et al. Reference Komatsuda, Pourkheirandish, He, Azhaguvel, Kanamori, Perovic, Stein, Graner, Wicker, Tagiri, Lundqvist, Fujimura, Matsuoka, Matsumoto and Yano2007). Indeed, removal of the end of the recognition alpha-helix and downstream C-terminus region prevents interaction with the floral development gene HvMADS13 and the inhibition of HvMADS13 expression associated with lateral floret abortion (Shen et al. Reference Shen, Sun, Liu, Shen, Wang, Ye, Wang and Guo2025). In summary, the naturally occurring VRS1 variants identified here in wild barley often cluster in conserved, functionally important regions and may influence key developmental traits, warranting further study of their phenotypic effects.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S1479262126100665.
Acknowledgements
We thank the genebanks and institutes that provided barley germplasm: INRA (National Institute of Agricultural Research Biological Resource Centres, France), IPK (Leibniz Institute of Plant Genetics and Crop Plant Research genebank, Germany), IRTA (Institute of Agrifood Research and Technology, Spain), JHI (James Hutton Institute, UK), JIC GRU (John Innes Centre Germplasm Resource Unit, UK, a National Bioscience Research Infrastructure supported by the UKRI-BBSRC, grant number BBS/E/JI/23NB0001 for conserving and supplying germplasm through www.seedstor.ac.uk), NordGen (Nordic Genetic Resources Center, Sweden), RAC (Station de Recherche Agroscope Changins, Switzerland), USDA NSGC (U.S. Department of Agriculture National Small Grains Collection, USA), VIR (N.I. Vavilov Research Institute of Plant Industry genebank, Russia).
Author contributions
J.A.B. undertook haplotype analyses and geographic plotting. H.J. provided project resources. J.C. undertook sequence and bioinformatic analysis. The AEGIS Consortium facilitated discussions and provided the wider scientific context in which the work was analyzed and interpreted. J.C. wrote the manuscript. J.A.B., H.J. and J.C. edited the manuscript.
Funding statement
The work was supported via the Natural Environment Research Council via the project ‘The Domestication of Europe’ and by ‘AEGIS: Ancient Environmental Genomics Initiative for Sustainability’ (Novo Nordisk Foundation grant code NNF24SA0092560; Wellcome Trust grant code 313162/Z/24/Z).
Competing interests
J.C. is an editor at the Plant Genetic Resources journal. The other authors declare no other competing interests.
Data availability statement
Genomic DNA sequences for the cultivars, as well as one exemplar from each VRS1 haplotype identified in landrace and wild barley accessions, have been deposited in GenBank using the accession numbers listed in Supplementary Table S1. Germplasm is available via the relevant genebanks and institutes, as listed in Supplementary Table S1.