Introduction
During weaning, various physiological and environmental stressors negatively affect the nutrient utilization and intestinal resilience of piglets, resulting in stunted development, intestinal injury, diarrhea, and increased mortality, ultimately causing economic losses in the swine industry (Zhou et al., Reference Zhou, Liang and Xiong2024). Alleviating weaning stress in piglets is crucial for enhancing both the productivity and sustainability of swine production systems. Antibiotics have been used to prevent piglet diarrhea; however, the overuse has raised concerns about antimicrobial resistance and potential effects on public well-being and environment (Ben Lagha et al., Reference Ben Lagha, Haas and Gottschalk2017). Consequently, developing potent treatment alternatives to antibiotics has become a critical priority (Allen et al., Reference Allen, Trachsel and Looft2014).
Antimicrobial peptides synthesized by probiotics are known as bacteriocins and viable alternatives to antibiotics (Silveira et al., Reference Silveira, Roque-Borda and Vicente2021; Sugrue et al., Reference Sugrue, Ross and Hill2024). Compared to live probiotics, antimicrobial peptides exhibit distinct advantages: (i) ability to rapidly pass through the gut mucosa, (ii) lack of infection risk, and (iii) high stability for dosing and storage (Ali et al., Reference Ali, Lee and Hsu2023). Several bacteriocins have been shown to effectively reduce the abundance of pathogenic intestinal bacteria, for example, the antibacterial peptide secreted by Lactococcus lactis subsp. Lactis strain L2 strongly suppresses certain pathogens, chiefly gram-positive strains, including notably Streptococcus suis (Sanca et al., Reference Sanca, Blanco and Dias2023). Bacteriocin produced by Lactobacillus gasseri LA39 showed promise as a natural substitute for conventional antibiotics in combating diarrhea in mammals through binding to keratin 19 (KRT19) receptors located on intestinal epithelial cells (Hu et al., Reference Hu, Ma and Nie2018). Additionally, supplementation with Microcin J25 (MccJ25), a bacteriocin from fecal Escherichia coli, provided multiple benefits in weaned pigs: boosting growth metrics, reducing diarrheal incidence, mitigating systemic inflammation, strengthening gut barrier integrity, and optimizing microbial populations (Yu et al., Reference Yu, Ding and Li2017).
Clostridium butyricum, a butyrogenic gut commensal, has served as a well-established probiotic for over 50 years (Stoeva et al., Reference Stoeva, Garcia-So and Justice2021). C. butyricum exerts protective effects on intestinal health by restructuring gut microbiota composition (Hagihara et al., Reference Hagihara, Kuroki and Ariyoshi2020), modulating immune function and inflammatory response (Hayashi et al., Reference Hayashi, Sato and Kamada2013), and enhancing intestinal barrier integrity (Long et al., Reference Long, Yang and Li2018). Particularly relevant to animal husbandry, recent studies have demonstrated that C. butyricum can effectively reduce transitional stress in piglets by simultaneously strengthening intestinal immune function and remodeling microbial diversity (Wu et al., Reference Wu, Wang and Lin2023). Beyond these established benefits, C. butyricum produces a diversity of economically valuable metabolites that serve as key mediators of its host benefits (Yang et al., Reference Yang, Shao and Pei2024). C. butyricum and its filtered culture medium, for instance, exhibit therapeutic effects on E. coli-induced endometritis (Mun et al., Reference Mun, Cai and Hu2022), while C. butyricum-derived vesicles (EVs) have shown efficacy in alleviating colitis (Ma et al., Reference Ma, Shen and Lyu2022). However, current research remains limited regarding the effects of antimicrobial peptides/bacteriocins secreted by C. butyricum on weaned piglets.
Microbial perturbation is now recognized as the predominant factor in postweaning diarrhea and concurrent bacteremia in piglets (Gresse et al., Reference Gresse, Chaucheyras-Durand and Fleury2017). Short-chain fatty acids (SCFAs) and bile acids (BAs), key metabolites produced by gut bacteria, are vital for the host’s metabolic processes (Collins et al., Reference Collins, Stine and Bisanz2023; Maslowski et al., Reference Maslowski, Vieira and Ng2009), especially in preserving the mucosal immune system (Sun et al., Reference Sun, Xu and Feng2021). Novel mechanistic studies show SCFAs act through GPR43 receptor engagement, providing a direct pathway from nutritional substrates to microbial-host crosstalk in inflammatory modulation (Macia et al., Reference Macia, Tan and Vieira2015). SCFAs modulate NLRP3 inflammasome activity through GPR43 signaling (Zuo et al., Reference Zuo, Fang and Liu2022). The NLRP3 inflammasome, as the most extensively characterized member of the nucleotide-binding oligomerization domain-like receptor family, serves as a crucial signaling platform that manages the maturation of the pro-inflammatory interleukin (IL)-1 family cytokines, specifically IL-1β and IL-18 (Jo et al., Reference Jo, Kim and Shin2016). Previous evidence showed that BAs suppressed NLRP3 inflammasome activation to ameliorate metabolic disorders (Guo et al., Reference Guo, Xie and Chi2016), and another study also found that Bacillus amyloliquefaciens SC06 could lessen lipopolysaccharide (LPS)-induced liver damage by hindering BA-mediated NLRP3 activation (Wang et al., Reference Wang, Wang and Zhou2024). Building on these findings, we postulate a potential link between the gut microbial metabolites/GPR43-NLRP3 signaling pathway and the integrity of the intestinal mucosal barrier. The role of this pathway in repairing the intestinal mucosal barrier of weaned piglets remains unexplored. Therefore, we propose that C. butyricum and its metabolites mitigate intestinal inflammation through a multi-faceted mechanism. This mechanism integrates the modulation of gut microbiota composition, regulation of microbial metabolites, and coordination of the GPR43-NLRP3 signaling pathway, thereby exerting a comprehensive protective effect on intestinal health.
Thus, this investigation systematically assessed the impact of Clostridium butyricum-derived antimicrobial peptide (CBP) on growth performance and diarrhea incidence in weaned piglets, while simultaneously investigating its modulatory effects on intestinal microbiota and associated metabolites. Utilizing an LPS-induced intestinal injury model, we further investigated the capacity of CBP supplementation to ameliorate gut dysbiosis and mitigate intestinal damage by assessing intestinal morphology, tight junction protein expression, cytokine profiles, microbial composition, and metabolic profiles of SCFAs and BAs. We further aimed to elucidate if CBP’s anti-inflammatory effects against LPS challenge are mediated by gut microbiome-generated metabolites acting through the GPR43-NLRP3 cascade.
Materials and methods
Preparation of antimicrobial peptides produced by C. butyricum
The C. butyricum strain used in this study was an in-house isolate, obtained from the intestinal contents of a healthy weaned piglet. The strain was isolated and maintained under strict anaerobic conditions at 37°C on reinforced clostridial medium (RCM). For identification, genomic DNA was extracted from a pure culture of the isolate. The 16S rRNA gene region was then amplified via polymerase chain reaction (PCR) using the primers 722F (5′-AACACCAGTGGCGAAGGC-3′) and 907R (5′-CCGTCAATTCCTTTRAGTTT-3′). The resulting PCR amplicon was purified and subjected to Sanger sequencing. The strain’s identity was confirmed by conducting a BLAST search of the sequence against the NCBI GenBank database. The isolate was identified as C. butyricum based on >99.9% sequence similarity to reference strains. The 16S rRNA gene sequence of the isolate is available in the Supplementary Information (Supplementary File S1).
Antimicrobial peptides from C. butyricum (CBP) were prepared as follows: bacterial cultures were cultivated in RCM broth at 37°C for a 72 h stretch in an anaerobic environment. To exclude the interference of peptides inherently present in the culture medium, sterile RCM broth (unfermented) was processed as a control under the same conditions (centrifugation, filtration, and ultrafiltration). Following incubation, the cultures were spun down via centrifugation at 7,000 × g for 10 min. Subsequently, the supernatant was passed through 0.22 μm filters and subjected to ultrafiltration through 3 kDa membranes, followed by 0.5 kDa membrane concentration (2450 series; Shanghai Sep-Bio Engineering Co., Ltd, Shanghai, China) to enrich antimicrobial peptides. The concentrated supernatant was then removed, filtered again with a 0.22 μm filter, and concentrated 20-fold via lyophilization (pre-freezing at −40°C, 100 mTorr vacuum pressure, −20°C shelf temperature). To ensure standardized dosing in subsequent functional assays, the total peptide concentration of the final product was determined using a BCA protein assay according to the manufacturer’s instructions, resulting in a stock concentration of 48.53 mg/mL. The final CBP was stored at 4°C.
Peptide identification and characterization by LC-MS/MS
For peptide processing, 100 μL of each sample (including the fermented supernatant and the sterile control) was mixed with 400 μL of methanol and sonicated for 5 min in an ice-water bath to precipitate proteins. The mixture was then centrifuged at 12,000 rpm and 4°C for 10 min, and the resulting supernatant (400 μL) was collected and lyophilized. The dried peptides were reconstituted in 200 μL of Wash Buffer and subsequently desalted using a desalting column, which involved activation, conditioning, equilibration, sample loading, washing, and a two-step elution with 50 μL of Elution Buffer. The combined eluate (100 μL) was dried in a vacuum centrifugal concentrator and reconstituted in Mobile Phase A for analysis. Peptide analysis was performed on a nano-UPLC system (Evosep One) coupled to a timsTOF Pro 2 mass spectrometer (Bruker, Germany) with a nanospray ion source. Approximately 200 ng of total peptide per sample was loaded onto a reversed-phase column (PePSep C18, 1.9 µm, 150 µm × 15 cm) and separated over a 44-min gradient using Mobile Phase A (0.1% formic acid in water) and Mobile Phase B (0.1% formic acid in acetonitrile). The mass spectrometer was operated in DDA PASEF mode with a scan range of 100–1700 m/z, and the collision energy was ramped linearly with ion mobility from 20 eV to 59 eV. The quality and reproducibility of the analysis were ensured by monitoring key metrics, including the stability of retention times, uniformity of peptide elution, high chromatographic resolution, and a stable ion source spray, as evidenced by a smooth total ion chromatogram. To eliminate background interference from the culture medium, the mass spectrometry profiles of the fermented supernatant and the sterile control were compared. Only peptide sequences uniquely identified in the fermented supernatant (intensity > 0) and absent in the sterile control were considered as de novo metabolic products of C. butyricum (Supplementary Table 1 and Supplementary File 6). Subsequently, the identified peptide sequences were subjected to online prediction for further characterization using ProtScale, DeepTMHMM, and AlphaFold3.
Animal experiments
Ethics statement
The animal experiment of this study was approved by the Animal Care and Use Committee guidelines of the Zhejiang Academy of Agricultural Sciences (approval number ZAASLA2025022708).
Animals and experimental design
The experimental design is shown in Figure 1. In the animal experiment, a total of five litters of Jinhua piglets (9.95 ± 2.03 kg body weight) were weaned at 45 days of age. Based on comparable sex, body weight, and litter origin, the piglets (25 male and 25 female) were randomly allocated into two groups with equal gender distribution, each consisting of five replicates with five piglets per replicate: the control group (CON) and the CBP group. The trial lasted for 21 days. Piglets in the CBP group were orally administered 3 mL CBP daily, whereas piglets in the CON group received an equal volume of sterile water. During the experiment, all piglets had free access to water but were fed a restricted diet (restricted feeding can effectively prevent overeating-induced diarrhea in Jinhua pigs). The basal diet, detailed in Table 1, was formulated to meet the nutrient requirements for Jinhua piglets in accordance with the GB/T 2417-2008 standard. The nutrient composition was determined as follows: digestible energy was calculated based on prediction equations from NRC (2012), while all other nutrient levels were determined by chemical analysis. The specific analytical methods used were as follows: crude protein (CP) according to GB/T 6432-2018, crude fiber (CF) according to GB/T 6434-2006, calcium (Ca) according to GB/T 6436-2018, total phosphorus (TP) according to GB/T 6437-2018, and amino acids (Lys, Met + Cys, etc.) according to GB/T 18246-2019.

Figure 1. Experimental design schematic.
Table 1. Ingredients and nutrition levels of the weaned piglet diet

Notes:
1 The piglet concentrated feed H2511 was a custom formulation, composed of the following ingredients (% of the concentrate): soybean meal (35.00), fermented soybean meal (10.00), extruded soybeans (10.00), fish meal (12.00), and plant fat powder (5.00). The formulation was balanced with dicalcium phosphate (6.50), limestone (4.50), sodium chloride (2.50), and a functional additives and amino acid package (9.50) which contained L-Lysine, DL-Methionine, L-Threonine, a premix of essential vitamins and mineral elements, and other functional additives. The guaranteed analysis of the concentrated feed was as follows: crude protein ≥ 32.0%, crude fiber ≤ 4.5%, lysine ≥ 3.8%, calcium 2.8–3.8%, and total phosphorus ≥ 1.5%.
2 All the values in the nutrient composition are measured values, with the exception of digestible energy (DE), which was calculated.
LPS challenge and sample collection
On day 21 of the experiment, a subset of pigs was selected for an acute immune challenge. From each dietary group (CON and CBP), 18 representative pigs were chosen based on a balanced distribution of sex, litter of origin, and body weight (pigs with weights close to their pen average). Subsequently, the 18 selected pigs within the CON group were randomly allocated to receive either an intraperitoneal injection of LPS (100 μg/kg; n = 9) or an equal volume of sterile saline (n = 9). The same random allocation procedure was applied to the 18 selected pigs from the CBP group. For this challenge phase, the individual pig was considered the experimental unit.
Six hours after LPS challenge, piglets were euthanized. Blood was collected from the anterior vena cava into 5 mL vacutainer tubes, then centrifuged at 1,500 × g and 20°C for 15 min to separate serum. The serum was flash-frozen and stored at −80°C. Mid-colon segments (approximately 2 cm long) were dissected and fixed in 4% paraformaldehyde for morphological analysis. Mid-colon digesta were collected and stored at −80°C. Finally, colonic mucosa was scraped into sterile tubes and stored at −80°C. Gall bladders were excised from the carcasses using sterile scalpels, and bile was aspirated with a 20 mL sterile syringe. The collected bile was immediately snap-frozen and stored at −80°C for subsequent targeted metabolomics analysis.
Intestinal morphology
Following a 24-h period of fixation, segments of the colon underwent dehydration, embedding, and sectioning before being stained using hematoxylin and eosin. Crypt depth measurements were then taken according to a prior study (Jin et al., Reference Jin, Wu and Wang2025).
Immunofluorescence analysis
Colonic segments were stained using immunofluorescence for ZO-1, MUC2, Occludin, and Claudin1 protein. Briefly, antibody (ZO-1 21773-1-AP 1:1000, MUC2 GB11344 1:2000, Occludin GB111401 1:2000, Claudin1 GB12032 1:1000) was incubated overnight in a controlled environment of 4℃. Then, the appropriate secondary antibody (TYR-570) was added and allowed to incubate for an hour at room temperature. Following a trio of PBS washes, the slides were treated with DAPI solution for 15 min at room temperature and subsequently stored away from light. Upon completion of immunofluorescent labeling, cellular images were obtained with an inverted Zeiss LSM900 imaging platform.
Colonic mucosal cytokine concentrations
The concentrations of porcine IL-1β, IL-6, IL-10, IL-18, tumor necrosis factor-α (TNF-α), and interferon-γ (IFN-γ) in colonic mucosa samples were quantified using enzyme-linked immunosorbent assay kits from Shanghai Jianglai Biological Technology Co., Ltd (Shanghai, China). The respective catalog numbers were JL21874 (IL-1β), JL21880 (IL-6), JL21886 (IL-10), JL21894 (IL-18), JL21873 (TNF-α), and JL21882 (IFN-γ). For sample preparation, approximately 100 mg of frozen colonic mucosal tissue was homogenized in 900 μL of PBS on ice. The homogenate was then centrifuged at 5,000 × g for 10 min at 4°C, and the resulting supernatant was collected. The total protein concentration of the supernatant was determined using a BCA protein assay kit to normalize the cytokine concentrations. All procedures were performed in strict accordance with the manufacturer’s protocols. The analysis was conducted on six biological replicates per experimental group, and each sample was assayed in technical triplicate to ensure the precision of the results.
SCFA analysis
Sample preparation for SCFA analysis was performed as follows: approximately 0.5 g of each sample was accurately weighed, mixed with 1.5 mL of ultrapure water, and vortexed. The mixture was allowed to stand for 30 min and then centrifuged at 15,000 r/min for 15 min. The supernatant (1 mL) was transferred to a new tube, and 0.2 mL of 25% (m/v) metaphosphoric acid was added. This mixture was incubated at 4°C for 30 min and centrifuged again (15,000 r/min, 15 min). The resulting clear supernatant was filtered through a 0.22 μm membrane into an autosampler vial for GC-MS analysis.
The quantification of SCFAs was conducted using an external standard method. A standard stock solution was prepared using the following standards of chromatographic grade, all purchased from Aladdin (Shanghai, China): acetic acid (Cat# A801294), propionic acid (Cat# P816182), isobutyric acid (Cat# I811668), butyric acid (Cat# B802730), isovaleric acid (Cat# I811830), and valeric acid (Cat# V820439). The stock solution was prepared by dissolving accurately weighed volumes of each standard in ultrapure water and was stored in the dark at 4°C. Working standard solutions were prepared by serially diluting the stock solution with ultrapure water to generate a multi-point calibration curve. For quality control (QC), a blank sample was processed and analyzed with each batch of samples.
The method was validated by determining the limit of detection (LOD) for each analyte. The calculated LODs were as follows: acetic acid (0.18 μg/g), propionic acid (0.73 μg/g), isobutyric acid (1.39 μg/g), butyric acid (0.69 μg/g), isovaleric acid (9.92 μg/g), and valeric acid (3.43 μg/g). The concentrations of SCFAs in the samples were calculated based on the calibration curve, ensuring that they fell within the linear range.
The final concentration of each SCFA in the sample (X) was calculated using the following formula:
where C is the concentration of the SCFA in the test solution (μg/mL) obtained from the calibration curve; V is the initial volume of ultrapure water (1.5 mL); M is the initial mass of the sample (g); and the factor 6/5 corrects for the dilution from adding metaphosphoric acid. The final results were expressed as μg/g of the sample.
Western blot analysis
Total protein was extracted from colon tissues using RIPA buffer (Beyotime, China) supplemented with protease and phosphatase inhibitors. The lysates were centrifuged at 12,000 × g for 10 min at 4°C, and the protein concentration of the supernatant was determined using a BCA protein assay kit (Beyotime, China). For each sample, an equal amount of protein (30 μg) was separated by SDS-PAGE and transferred onto a PVDF membrane (Millipore, USA). The membranes were blocked with 5% skim milk in TBST (Tris-buffered saline with 0.1% Tween-20) for 1 h at room temperature, then incubated overnight at 4°C with the following primary antibodies: anti-NLRP3 (1:1000, Abcam, EPR23094-1), anti-IL-18 (1:1000, Abcam, EPR19956), anti-GPR43 (1:1000, Bioss, bs-23786R), and anti-β-actin (ACTB) (1:1000, Abclonal, AC026). After washing with TBST, the membranes were incubated with an HRP-conjugated Goat anti-Rabbit IgG secondary antibody (1:5000, HUABIO, HA1012) for 1 h. Protein bands were visualized using an Enhanced Chemical Luminescence (ECL) chemiluminescence reagent (Beyotime, China). The band intensities were quantified using ImageJ software (NIH, USA), and the relative expression of target proteins was normalized to β-actin (ACTB). The analysis was performed on three biological replicates per experimental group (n = 3).
16S rRNA gene sequencing
In order to study the bacterial lineup lurking within piglets’ colonic microbiota, we scooped up the contents and pulled out the microbial genomic DNA using the E.Z.N.A.® Stool DNA Kit from Omega Bio-tek. We then cranked up the volume on the 16S rRNA (V4 region, 515F/806R) genes and assembled a sequencing library as per the default primer set and protocol. After that, we cleaned up the sequences with the DADA2 plugin in Qiime2 (version 2020.2), tuning the parameters for pinpoint accuracy. Taxonomic assignment used Qiime2’s Naive Bayes classifier with SILVA v138. And, for the heavy lifting, we hopped over to the Majorbio Cloud Platform’s online service, where bioinformatics analysis is served up for free (find it at https://cloud.majorbio.com/).
BA targeted metabolomics
A targeted metabolomic analysis was performed to quantify 69 BAs, including 37 conjugated and 32 free BAs. A detailed list of the targeted analytes and their corresponding standards is provided in supplementary file 2. For sample preparation, bile samples were first diluted 50-fold. An aliquot of 100 μL of the diluted sample was mixed with 400 μL of an acetonitrile:methanol (1:1, v/v) solution containing a mixture of stable isotope-labeled internal standards (details in supplementary file 3). The mixture was vortexed for 30 s, sonicated for 10 min in an ice-water bath, and then incubated at −40°C for 1 h to precipitate proteins. After centrifugation at 12,500 × g for 15 min at 4°C, the supernatant was collected for analysis. The analysis was conducted using a UHPLC-MS/MS system consisting of a Vanquish UHPLC system coupled to an Orbitrap Exploris 120 mass spectrometer (Thermo Fisher Scientific). Chromatographic separation was achieved on a BEH C18 column (150 × 2.1 mm, 1.7 μm) maintained at 45°C. The mobile phases were (A) 5 mM ammonium acetate in water and (B) acetonitrile. The autosampler was maintained at 4°C, and the injection volume was 1 μL. The mass spectrometer was operated in both positive and negative ion modes with a spray voltage of +3500 V and −3200 V, respectively. The sheath gas and auxiliary gas flow rates were set to 40 and 15 arbitrary units, respectively. The auxiliary gas heater temperature was 350°C, and the capillary temperature was 320°C. QC was ensured by preparing a pooled QC sample from equal aliquots of each sample. The QC sample was injected five times throughout the analytical run to monitor the stability and reproducibility of the system. The average recovery rates for all target compounds were within 81.6–114.6%, and the relative standard deviations were all below 10.5%, as detailed in supplementary file 4. Data processing and analysis followed a rigorous workflow. Prior to sample analysis, the parallel reaction monitoring parameters for each target compound were optimized by infusing individual standard solutions. As most BAs did not yield fragment ions of sufficient intensity, high-resolution parent ions were used for quantification. The specific MS parameters for each analyte are provided in supplementary file 5. Peak identification and integration were performed using the proprietary software BioBud 2.0.3. Absolute quantification was achieved using the internal standard method. Calibration curves were constructed by plotting the peak area ratio of the analyte to the internal standard (y) against the analyte concentration (x, nmol/L). A weighted (1/x) least-squares linear regression was applied. Calibration points with accuracy outside the 80–120% range were excluded. The resulting quantitative data were filtered to retain metabolites with no more than 50% missing values in any single group or across all samples. Remaining missing values were imputed with a random value between 0.1 and 0.5 times the minimum observed value for that metabolite. All subsequent statistical analyses were performed using R software (version 4.3.3).
Statistical analysis
Data were statistically analyzed using SPSS 27.0. All data are presented as mean ± SEM, with statistical significance set at P < 0.05. Fecal scores and diarrhea rate were analyzed using non-parametric methods.
Growth performance and diarrhea assessment
For growth performance and diarrhea assessment, the pen was considered the experimental unit (n = 5). Differences in piglet growth performance metrics (average daily gain [ADG] and feed conversion ratio [F/G]) between the control and CBP groups were assessed using independent samples t-tests. Data normality was confirmed with the Shapiro–Wilk test prior to analysis. The formula for the t-test analysis model is:
 \begin{equation*}t = \frac{{{{\bar X}_1} - {{\bar X}_2}}}{{\sqrt {\frac{{\left( {{n_1} - 1} \right)S_1^2 + \left( {{n_2} - 1} \right)S_2^2}}{{{n_1} + {n_2} - 2}}\left( {\frac{1}{{{n_1}}} + \frac{1}{{{n_2}}}} \right)} }}\end{equation*}
\begin{equation*}t = \frac{{{{\bar X}_1} - {{\bar X}_2}}}{{\sqrt {\frac{{\left( {{n_1} - 1} \right)S_1^2 + \left( {{n_2} - 1} \right)S_2^2}}{{{n_1} + {n_2} - 2}}\left( {\frac{1}{{{n_1}}} + \frac{1}{{{n_2}}}} \right)} }}\end{equation*}where ![]() $\bar X$ is the mean of the samples, S is the variances of the samples, and n is the sample sizes.
$\bar X$ is the mean of the samples, S is the variances of the samples, and n is the sample sizes.
For the fecal score, a non-parametric approach was used. The Mann–Whitney U test was employed to compare the median fecal scores between the two groups. Diarrhea rate was analyzed using the Chi-square test.
Factorial ANOVA for LPS challenge model
For data from the LPS challenge, the individual pig was the experimental unit. The data from the LPS-induced inflammation model, involving the four groups (CON, LPS, CBPLPS, CBP), were analyzed using a 2 × 2 factorial ANOVA to evaluate the main effects of CBP and LPS, as well as their interaction. The statistical model for this analysis is:
 \begin{equation*}y_{ijk} = \mu + {\alpha _i} + {\beta _j} + {\left( {\alpha \beta } \right)_{ij}} + {\epsilon _{ijk}}\end{equation*}
\begin{equation*}y_{ijk} = \mu + {\alpha _i} + {\beta _j} + {\left( {\alpha \beta } \right)_{ij}} + {\epsilon _{ijk}}\end{equation*}where ![]() ${y_{ijk}}$ is the dependent variable (e.g., cytokine levels),
${y_{ijk}}$ is the dependent variable (e.g., cytokine levels), ![]() $\mu $ is the overall mean,
$\mu $ is the overall mean, ![]() ${\alpha _i}$ and
${\alpha _i}$ and ![]() ${\beta _j}$ are the main effects of factor A (CBP treatment) and factor B (LPS challenge), respectively. (αβ)ij represents the interaction effect between CBP and LPS.
${\beta _j}$ are the main effects of factor A (CBP treatment) and factor B (LPS challenge), respectively. (αβ)ij represents the interaction effect between CBP and LPS. ![]() ${\epsilon _{ijk}}$ is the random error term associated with individual animal variation within each treatment group. When a significant interaction effect was detected (P < 0.05), post hoc analysis was performed using Duncan’s multiple range test to determine specific group difference.
${\epsilon _{ijk}}$ is the random error term associated with individual animal variation within each treatment group. When a significant interaction effect was detected (P < 0.05), post hoc analysis was performed using Duncan’s multiple range test to determine specific group difference.
Results
CBP promoted growth performance and reduced diarrhea incidence of piglets
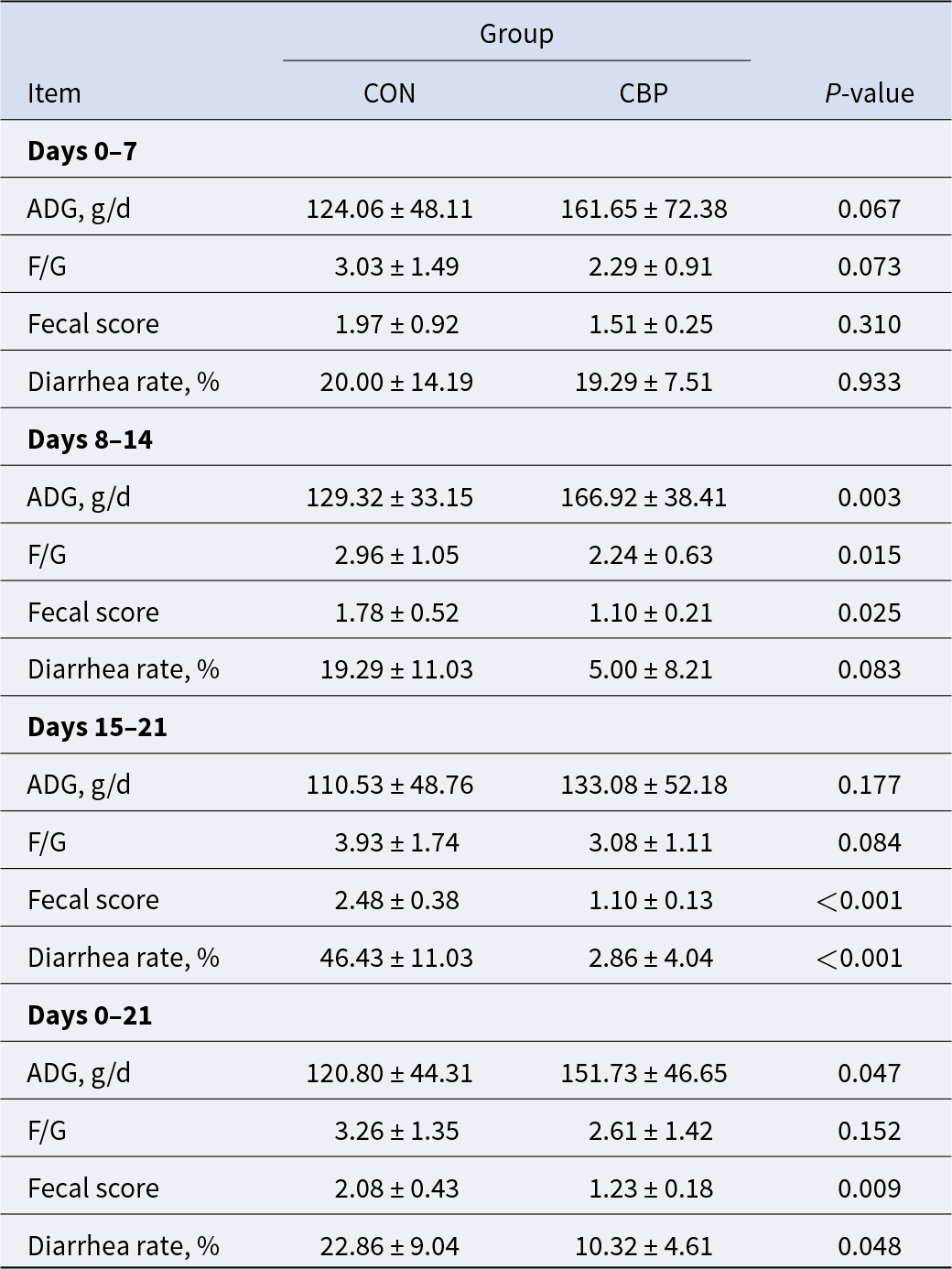
As shown in Table 2, no substantial distinctions in growth performance or diarrhea incidence between the treatment groups over the initial 0–7-day feeding interval (P > 0.05). During the 8–14-day feeding period, piglets received CBP supplementation demonstrated significantly higher ADG and lower F/G compared to the CON group (P < 0.05). CBP administration markedly lowered both diarrhea incidence and fecal scores, with a substantial drop in diarrhea rates observed within the 15–21-day period (P < 0.05) and fecal scores remaining lower throughout the 8–21-day observation period (P < 0.05).
Table 2. Effects of CBP on the growth performance and diarrhea incidence of weaned piglets (n = 25)

CBP ameliorates LPS-challenged intestinal damage in colon of piglets
Histological analysis of colonic tissues demonstrated severe crypt architecture disruption of LPS-treated piglets, confirming successful establishment of colonic injury model (Fig. 2A). Quantitative statistical analysis was performed on the sections. Compared with the LPS group, CBP supplementation significantly attenuated these pathological changes and normalized crypt depth (P < 0.05, Fig. 2B). IF quantification demonstrated significant LPS-induced suppression of intestinal barrier proteins (MUC2, ZO1, OCCLUDIN, CLAUDIN) relative to controls (P < 0.05), which were substantially ameliorated by CBP administration (P < 0.05, Fig. 2C–D).

Figure 2. CBP ameliorates LPS-challenged intestinal damage in colon of piglets. (A) Histopathological analysis of colon tissue using H&E staining (scale bar: 625 μm). (B) Morphometric analysis of colonic crypt depth. (C) IF imaging of MUC2 and tight junction proteins of colon sections (scale bar: 50 μm). (D) Mean fluorescence intensity of MUC2 and tight junction proteins. Data are expressed as mean ± SEM. The data underwent 2 × 2 factorial ANOVA testing, followed by one-way ANOVA and Duncan’s multiple range tests.
CBP attenuated LPS-induced inflammasome activation and pro-inflammatory cytokine levels in the colon
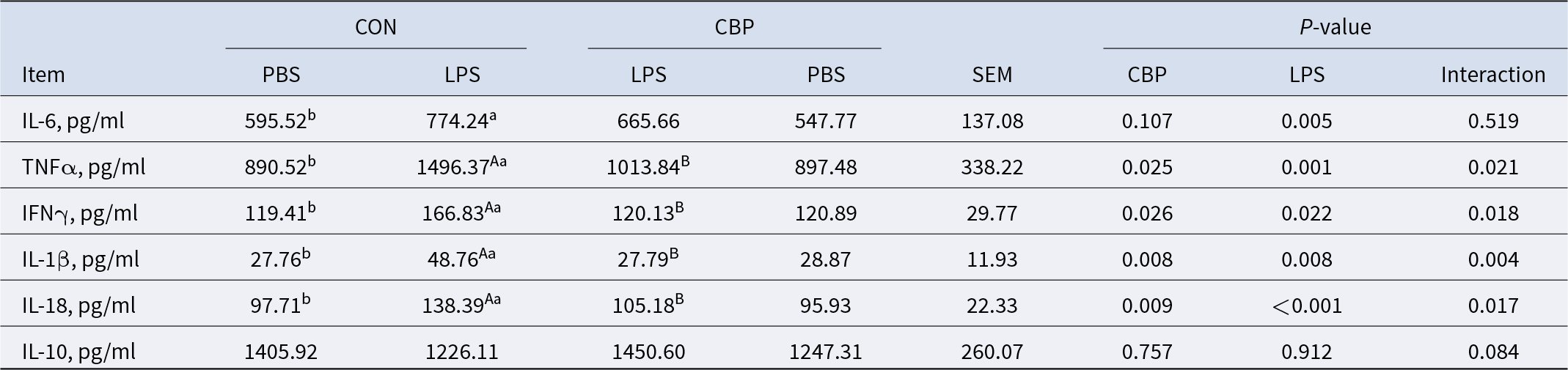
Quantification of inflammatory markers showed significant upregulation of IL-6, IL-1β, IL-18, TNF-α, and IFN-γ in LPS-treated versus CON groups (P < 0.05), while CBP supplementation reversed this pro-inflammatory response, reduced levels of IL-1β, IL-18, TNF-α, and IFN-γ (P < 0.05, Table 3). As the NLRP3 inflammasome plays a pivotal role in inflammation regulation (Zhan et al., Reference Zhan, Li and Xu2022), we assessed NLRP3 pathway protein expression in piglets via western blot analysis. The findings showed that, when compared to the control group, the LPS challenge dramatically increased the expression of NLRP3 and IL-18, as well as both pro- and cleaved IL-1β, while concurrently decreasing the expression of GPR43. Conversely, treatment with CBP effectively attenuated LPS-induced elevations in NLRP3, IL-18, and both precursor and mature forms of IL-1β levels and restored GPR43 expression (P < 0.05, Fig. 3A).

Figure 3. CBP attenuated LPS-induced inflammasome activation in the colon. (A) Western blot analysis of GPR43, NLRP3, IL-18, and IL-1β in colon tissue (n = 3). Data are expressed as mean ± SEM. The data underwent 2 × 2 factorial ANOVA testing, followed by one-way ANOVA and Duncan’s multiple range tests. ACTB (β-actin) was used as the internal loading control.
Table 3. CBP attenuated LPS-induced inflammasome activation and pro-inflammatory cytokine levels in the colon (n = 6)
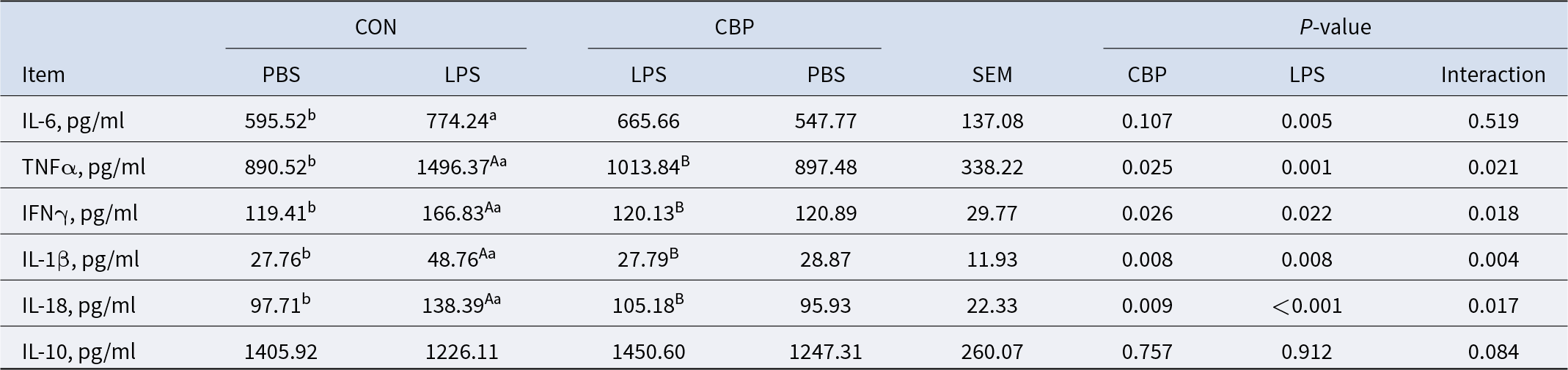
Note: Values with different letters indicate significant differences at P < 0.05. Lowercase letters (a, b) compare PBS vs LPS within the CON group; uppercase letters (A, B) compare CON+LPS vs CBP+LPS groups.
CBP altered the microbial composition of the colon in LPS-challenged piglets
To assess whether microbiota modulation mediates CBP’s anti-inflammatory action in piglet intestines, we performed 16S rRNA sequencing to characterize microbial composition after CBP supplementation and LPS challenge. The CON group contained 1090 unique ASVs, compared to 998 in the LPS group, 931 in the CBP group, and 745 in the CBPLPS group. Notably, the CBP group shared more ASVs with the CON group than that with the LPS group, reflecting structural convergence with CON group microbiota (Fig. 4A). Meanwhile, Principal Coordinate Analysis (PCoA) based on the Bray–Curtis algorithm revealed distinct clustering among groups, indicating that both LPS and CBP induced dynamic changes in the overall structure of intestinal microbiota (P < 0.05, Fig. 4B). Nevertheless, α diversity metrics showed no statistically significant variations among groups (P > 0.05, Fig. 4C). At the phylum level, Bacillota, Bacteroidota, and Spirochaetota were the three dominant phyla (Fig. 4D). At the genus level, Lactobacillus, Coprococcus, Blautia, UCG-008, Faecalibacterium, and Solobacterium were significantly enriched in the CBP group compared with CON group (LEfSe: P < 0.05, LDA > 3.0), moreover, LPS challenge alone significantly enriched 3 bacterial genera, including norank_o_WCHB1-41, p-1088-a5_gut_group and Longibaculum (Fig. 4F). A heatmap of bacterial genera showed clear differences in gut microbiota composition between the different treatments. Among the top 50 genera, CBP supplementation remarkably increased the relative abundances of Lactobacillus, Coprococcus, and Blautia, while decreased the relative abundance of Streptococcus and Escherichia-Shigella. LPS challenge significantly reduced Lachnospira and Escherichia-Shigella abundance but increased norank_o_RF39 and UCG-002 abundance. Additionally, the CBPLPS group showed higher Clostridium, Coprococcus, and Escherichia-Shigella abundance compared to the LPS group (P < 0.05, Fig. 4G–H).

Figure 4. CBP modified the colonic microbial composition in LPS-challenged piglets. (A) Venn diagram of ASV distribution. (B) PCoA of Bray–Curtis distances. (C) α-Diversity indices (Chao1 and Shannon). (D) Phylum-level taxonomic composition. (E) Genus-level taxonomic composition. (F) LDA scores for differentially abundant genera from LEfSe analysis. (G) Heatmap of the 50 most abundant genera. (H) Relative abundance of key differential genera. Values are presented as mean ± SEM (n = 6). P-values for the main effects of CBP and LPS, and their interaction, were determined by 2 × 2 factorial ANOVA. Different letters indicate significant differences among groups (P < 0.05).
CBP regulated the SCFA concentration of the colon in LPS-challenged piglets
SCFAs, key microbial metabolites derived from dietary fiber fermentation, play pivotal roles in immunometabolism regulation. To elucidate their involvement, we quantified colonic SCFA profiles. Results show that the LPS challenge notably lowered acetic, propanoic, butyric, and valeric acid levels in piglets when contrasted with the CON group (P < 0.05), CBP supplementation alleviated the reduction in these SCFAs. Notably, CBP significantly increased butyric acid content (P < 0.05, Table 4).
Table 4. CBP regulated the colonic SCFA levels in LPS-challenged piglets (n = 6)

Note: Values with different letters indicate significant differences at P < 0.05. Lowercase letters (a, b) compare PBS vs LPS within the CON group; uppercase letters (A, B) compare CON+LPS vs CBP+LPS groups.
CBP modulates microbial BA metabolism in LPS-challenged piglets
Given that BAs serve as critical signaling molecules essential for intestinal homeostasis, we conducted a metabolomic analysis of gallbladder BAs applying UPLC-MS to characterize alterations in BA metabolism induced by CBP supplementation and subsequent LPS challenge in piglets. In summary, the primary, secondary, and overall BA counts showed no statistically notable variations across the groups (P > 0.05, Fig. 5A). LPS challenge altered the BA pool composition, which was restored to normal levels by CBP administration (Fig. 5B). Heatmap visualization of average normalized levels for 30 BA compounds (Fig. 5C). Specifically, CBP supplementation markedly elevated the levels of glycodeoxycholic acid, glycohyocholic acid (GHCA), and cholic acid, while decreasing glycohyodeoxycholic acid (GHDCA) levels (P < 0.05). In contrast, LPS challenge dramatically reduced both GHDCA and hyocholic acid (HCA) concentrations, whereas CBP intervention-maintained HCA and taurolithocholic acid (TLCA) at physiological levels (Fig. 5D).

Figure 5. CBP modulates microbial bile acid metabolism in LPS-challenged piglets. (A) The primary, secondary, and total BA levels among groups. (B) The relative proportions of different bile acids within each group. (C) Heatmap displaying standardized bile acid metabolite levels from targeted metabolomic analysis in piglets. (D) Variation in concentrations of significantly altered bile acids between groups (n = 5). Data are expressed as mean ± SEM. The data underwent 2 × 2 factorial ANOVA testing, followed by one-way ANOVA and Duncan’s multiple range tests.
Correlations among differential microbiota, SCFA/BA metabolic profiles, and key metrics of intestinal barrier function
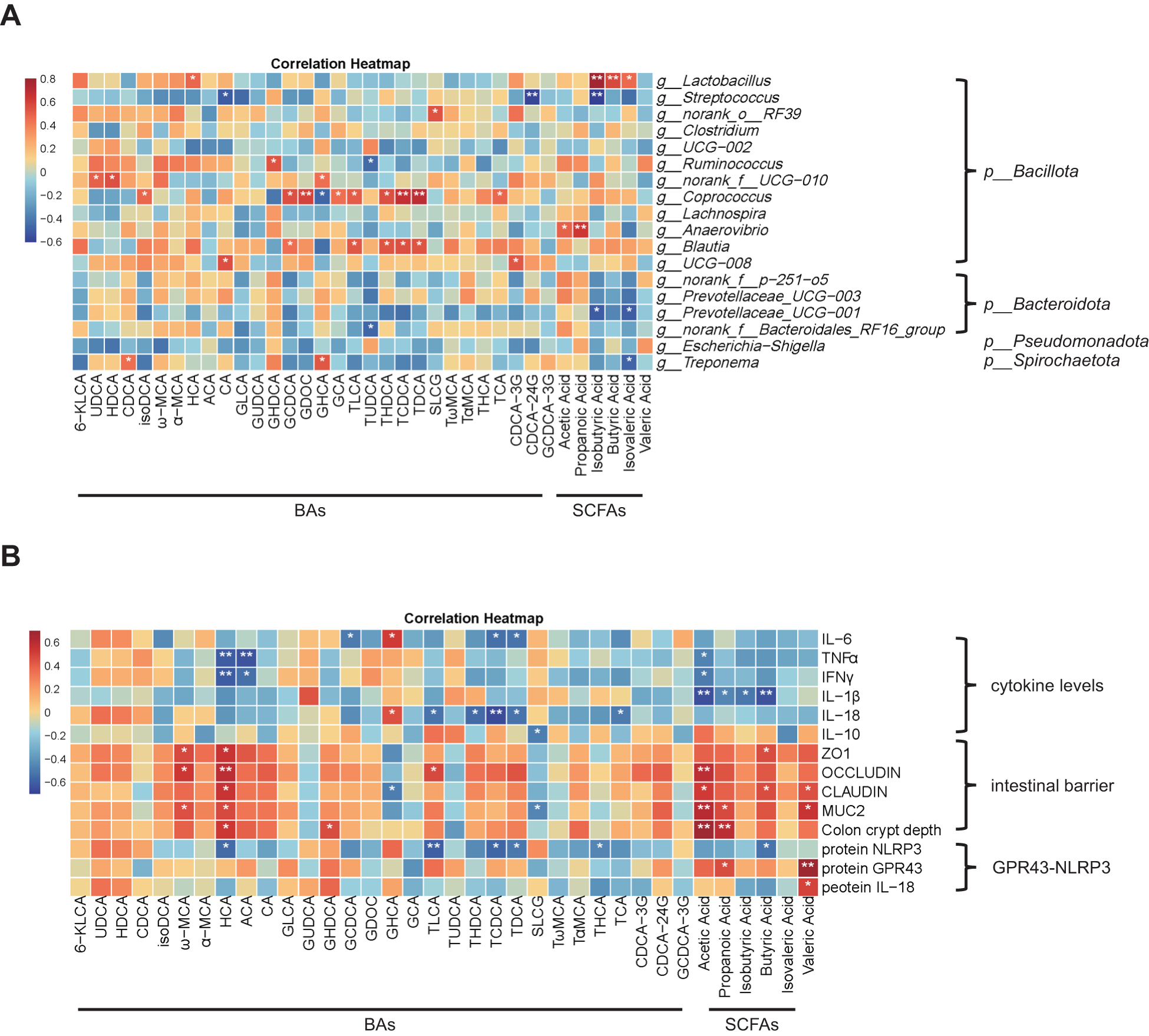
To investigate how gut microbiome signatures and metabolite profiles influence intestinal barrier functionality, we conducted a correlation analysis between colonic microbiota, SCFA/BA metabolic profiles, and key metrics of intestinal barrier function. First, we performed Pearson correlation analysis of microbial-derived SCFA/BA profiling integrated with microbiota characterization to uncover metabolite-mediated regulation networks. The results demonstrated that Lactobacillus abundance was positively correlated with isobutyric, butyric and isovaleric acid, and HCA, while Coprococcus abundance showed positive correlations with Glycodeoxycholic Acid (GDCA) and TLCA but a negative correlation with GHCA (P < 0.05, Fig. 6A). Further analysis revealed significant correlations between SCFA and BA metabolic profiles and key intestinal barrier function markers. Notably, butyric acid showed a negative correlation with IL-1β levels while demonstrating positive correlations with both ZO-1 and claudin-1 expression. Interestingly, HCA was positively associated with intestinal barrier protein expression and crypt depth, yet it also exhibited an inverse relationship with NLRP3 inflammasome activity. Furthermore, both propionic acid and valeric acid were positively correlated with colonic GPR43 expression (P < 0.05, Fig. 6B). Collectively, CBP ameliorates LPS-driven intestinal inflammation by orchestrating microbiota-mediated SCFA/BA metabolic networks, resulting in enhanced mucosal protection and coordinated suppression of inflammatory effectors including the GPR43-NLRP3 pathway.

Figure 6. Correlation among differential microbiota, SCFA/BA metabolic profiles, and key metrics of intestinal barrier function. (A) Correlation between differential microbiota and SCFA/BA metabolic profiles. (B) Correlation between SCFA/BA metabolic profiles and key metrics of intestinal barrier function. Pearson’s correlation analysis was used to assess these associations, with red and blue colors signifying positive and negative correlations, respectively. Asterisks indicate statistical significance (P < 0.05, P < 0.01).
Identification and antimicrobial activity of novel peptides from CBP
The antimicrobial efficacy of the CBP fraction was first validated in vitro using an inhibition zone assay against Escherichia coli K88, Salmonella spp., and Staphylococcus aureus (Figure S1A), which demonstrated a potent inhibitory effect comparable to the ciprofloxacin control (600 μg/mL). To elucidate the molecular basis of this activity, the fraction was subjected to LC-MS/MS analysis. This analysis revealed a profile of several distinct peptide sequences within the active CBP fraction, as detailed in Supplementary Table S1. From these, we pinpointed three as the most prominent candidates based on their signal intensity and sequence characteristics. To provide direct experimental evidence for these three sequences, their secondary mass spectra (MS/MS), confirming the fragmentation patterns used for sequencing, are provided in Supplementary Figure S2A. Our subsequent investigation focused exclusively on these three candidate peptides. First, to explore their potential functions in silico, we computationally analyzed their physicochemical properties, including hydrophilicity, cell-penetrating potential, and predicted tertiary structures (Figure S1B). Second, to provide direct experimental proof of their function, the same three peptides were chemically synthesized to a high purity. A mixture of these synthetic peptides (900 μg/mL) was found to replicate the potent antimicrobial activity of the original natural CBP fraction (Figure S1C). This definitively confirms that the bioactivity is attributable to these specific peptide sequences, rather than to other potential metabolites in the fraction.
Discussion
During weaning, piglets are susceptible to intestinal injury due to immature gut development and abrupt nutritional/environmental changes, leading to growth suppression and revenue losses in porcine farming. The development of antibiotic alternatives to enhance piglet health and prevent intestinal inflammation and diarrhea could therefore provide both health and socioeconomic benefits (Dowarah et al., Reference Dowarah, Verma and Agarwal2017). This study confirms that the CBP supplementation improves piglet growth, reduces diarrhea incidence, and attenuates LPS-mediated intestinal inflammatory responses. Furthermore, our analysis delineated microbiota-BA axis dynamics under CBP treatment and LPS-induced inflammatory conditions, demonstrating that CBP ameliorates microbial dysbiosis and modulates microbiota-derived metabolites. More specifically, CBP supplementation increased the abundance of gut commensal microbes and their metabolites, thereby enhancing barrier function. Collectively, these results demonstrate CBP’s efficacy as an antibiotic-alternative dietary intervention that improves porcine intestinal health of piglet.
The study reveals that supplementing piglets with CBP greatly enhances their growth and gut health before they face the LPS challenge. This is clear from the higher ADG, better feed conversion rates, and lower rates of diarrhea. Past studies have shown that antimicrobial peptides from probiotics can similarly boost the growth of piglets. For example, adding 0, 11, or 16.5 mg/kg of colicin E1 to their diet has lessened the incidence and severity of postweaning diarrhea, improved their growth, and inhibit small intestinal inflammatory gene activation (Cutler et al., Reference Cutler, Lonergan and Cornick2007). This study echoes those findings, showing that CBP can match the growth improvements and diarrhea reductions seen with C. butyricum in piglets (Fu et al., Reference Fu, Wang and Xiao2021; Wu et al., Reference Wu, Wang and Lin2023; Zhang et al., Reference Zhang, Yun and Lai2023). These findings suggest, for the first time, that antimicrobial peptides from C. butyricum have the potential to positively impact the growth and diarrhea rates in weaned pigs.
LPS triggers inflammatory cytokine release, disrupting intestinal architecture, increasing crypt depth, and stimulating intestinal immunity (Yu et al., Reference Yu, Zheng and Guo2024). We provide evidence that CBP administration reverses LPS-triggered gut dysfunction, restoring intestinal integrity and suppressing intestinal inflammation in piglets. The beneficial effects of CBP on maintaining gut barrier health in piglets’ line up with earlier research. Bacteriocins function as antimicrobial signaling peptides, modulating the intricate host–microbe crosstalk. They differentially regulate inflammatory cytokine networks in gut epithelia, functionally tuning immune homeostasis (Teng et al., Reference Teng, Huang and Liu2023). Nisin Z, a lantibiotic churned out by Lactococcus lactis, has been shown to put the brakes on the stimulus-triggered upregulation of ERK1/2 and p38 MAPK signaling pathways. This, in turn, dials down the release of pro-inflammatory cytokines like TNF-α, IL-1β, and IL-6, while simultaneously boosting the release of the anti-inflammatory cytokine IL-10 in LPS-stimulated MCF10A cells (Huang et al., Reference Huang, Teng and Liu2022). Furthermore, Lactobacillus plantarum NCMIB8826, a producer of the class IIb bacteriocin plantaricin EF, enhances tight junction protein expression, thereby strengthening intestinal barrier function. (Heeney et al., Reference Heeney, Zhai and Bendiks2019). It’s also worth noting that some Antimicrobial Peptides (AMPs) can offer protection by tweaking the immune response, which could shed light on the results we’re seeing in this study. These findings demonstrate that CBP can confer protection on the intestinal barrier by immune modulation in weaned pigs.
This research demonstrated that CBP could alleviate intestinal damage in piglets, though the exact mechanism remains unclear. There’s a growing body of evidence suggesting that keeping the gut microbiota in equilibrium is key to intestinal health and preventing diarrhea in piglets (Tang et al., Reference Tang, Xiong and Fang2022). In our study, supplementing with CBP elevated the levels of Lactobacillus and Coprococcus, which helped to correct gut dysbiosis. Furthermore, CBP supplement prevented the LPS-induced reduction in Ruminococcus and Lachnospira abundance, while suppressing the increase in UCG-002. Lactobacillus is crucial for maintaining a balanced gut microbiome and fortifying the immune system in mammals (Tao et al., Reference Tao, Fan and Li2025). A past study showed that incorporating C. butyricum into the diet can boost Lactobacillus populations, thereby stabilizing the cecal microbiome and boosting the growth performance of broiler chickens. Consistent with this, Coprococcus strains exhibiting butyrate synthesis capacity show established connections to gut homeostasis enhancement (Arnold et al., Reference Arnold, Roach and Fabela2021). New research showed that Coprococcus eutactus supplementation in pigs with acute colitis promoted body weight recovery and reduced pro-inflammatory mediators (TNF-α, IL-1β, IL-6) (Yang et al., Reference Yang, Shan and Shi2023). Beyond the enrichment of beneficial genera, we also observed that CBP exerts a targeted inhibitory effect on specific microbial populations. Specifically, CBP supplementation significantly reduced the abundance of genera such as Streptococcus, Bifidobacterium, Prevotellaceae_UCG-001, and Escherichia-Shigella. Furthermore, the CBPLPS group effectively mitigated dysbiosis by suppressing the proliferation of distinct microbes, the abundances of Treponema, norank_f__p-251-o5, Prevotellaceae_UCG-003, and Prevotellaceae_UCG-001 were significantly lower in the CBPLPS group compared to the LPS group. The decrease in Streptococcus abundance likely contributes to the accumulation of HCA and TCLA. Since Streptococcus species are major carriers of bile salt hydrolase (BSH) genes (Song et al., Reference Song, Cai and Lao2019), their inhibition by CBP would naturally reduce intestinal BSH activity. This reduction prevents the rapid deconjugation of TCLA (Sayin et al., Reference Sayin, Wahlstrom and Felin2013) and preserves the primary BA pool, particularly HCA, which is beneficial for metabolic homeostasis (Zheng et al., Reference Zheng, Chen and Jiang2021). In parallel, the suppression of Prevotellaceae_UCG-001 and Treponema reflects an improved gut environment. The reduction of Prevotellaceae_UCG-001, a genus associated with antimicrobial peptide deficiency (Forsyth et al., Reference Forsyth, Shaikh and Engen2024), and Treponema, a stress biomarker in pigs (Nguyen et al., Reference Nguyen, Martinez-Alvaro and Lima2023), suggests that CBP mitigates inflammation and stress-induced dysbiosis. Finally, regarding the increase in butyrate, the inhibition of Escherichia-Shigella plays a key role. By suppressing these facultative anaerobes, CBP alleviates nutrient competition, allowing beneficial butyrate-producing anaerobes to recover and thrive (Baltazar-Diaz et al., Reference Baltazar-Diaz, Gonzalez-Hernandez and Aldana-Ledesma2022). Integrated results demonstrate that CBP administration potentially mitigates enteric inflammation and barrier dysfunction via microbiota rebalancing.
Intestinal microbiota exerts pivotal effects on gut health, primarily through BA/SCFA-mediated metabolic signaling (Wen et al., Reference Wen, Wan and Wu2023). Current studies establish that equilibrated BA dynamics are bidirectionally associated with microbiota composition and activity in farm animals, with direct implications for inflammatory and metabolic dysregulation (Jia and Dong Reference Jia and Dong2024). Moreover, some previous studies have shown that C. butyricum modulates BA composition to exert anti-obesity effects (Liao et al., Reference Liao, Liu and Yao2023). Specifically, the intervention reduces hepatic steatosis in senior layers and corrects metabolic dysregulation while boosting developmental outcomes in growth-restricted suckling pigs (Wang et al., Reference Wang, Wang and Zhang2020; Zhang et al., Reference Zhang, Yun and Lai2023). Our analysis showed that LPS challenge significantly reduced HCA levels, while CBP supplementation effectively maintained them. This is significant, as HCA and its related compounds make up about 76% of the BA pool in pigs, animals known for their remarkable resistance to type 2 diabetes (Zheng et al., Reference Zheng, Chen and Jiang2021). Beyond BAs, SCFAs are also critical anti-inflammatory metabolites (Qin et al., Reference Qin, Yu and Li2023). Consistent with the known role of C. butyricum as a principal producer of butyrate (Stoeva et al., Reference Stoeva, Garcia-So and Justice2021; Wu et al., Reference Wu, Wang and Lin2023), we saw a marked increase in butyrate levels within the colonic after CBP supplementation.
The concept that gut bacterial metabolites can directly regulate the NLRP3 inflammasome is supported by existing literature; for instance, deoxycholic acid has been shown to inhibit NLRP3 activation in other models (Zhao et al., Reference Zhao, Wu and Hao2023a). In line with this, our study identified CBP-mediated NLRP3 inflammasome regulation in the piglet colon, establishing its inhibitory role. Crucially, our correlation analysis provided the direct mechanistic link, butyric acid and HCA were negatively correlated with colonic NLRP3, while propionic acid and valeric acid exhibited a direct positive association with colonic GPR43 expression. These findings strongly suggest that CBP may therapeutically address intestinal inflammation by modulating the microbiome’s metabolic outputs, particularly those targeting the GPR43-NLRP3 axis.
To understand the molecular basis for these correlations, we turn to the established mechanisms linking SCFAs to intestinal immunity. On a mechanistic level, it’s understood that SCFAs largely exert their anti-inflammatory effects through GPR43 (Brooks et al., Reference Brooks, Viardot and Tsakmaki2017). Foundational studies first showed that SCFAs, including butyrate, act as natural ligands for GPR43, a receptor notably expressed in immune cells (Brown et al., Reference Brown, Goldsworthy and Barnes2003). In piglet enteritis, the NLRP3 inflammasome within the intestinal epithelium is over-activated, leading to a surge in pro-inflammatory cytokines like IL-1β and IL-18 (Xia et al., Reference Xia, Zhong and Wu2022). Therefore, targeted NLRP3 regulation emerges as a promising therapeutic avenue for both prophylaxis and treatment of gut inflammation (Coll et al., Reference Coll, Schroder and Pelegrin2022). Indeed, the functional necessity of the GPR43 interaction for mediating NLRP3 inhibition has been confirmed in other models. For instance, a recent study on chronic obstructive pulmonary disease confirmed that the protective action of butyrate against NLRP3 activation is GPR43-dependent, as its effect was abolished when GPR43 was knocked down using siRNA (Zhao et al., Reference Zhao, Tong and Kang2023b). Furthermore, the role of NF-κB as an intermediary is supported by findings that butyrate can block NLRP3-induced pyroptosis through the STING/NF-κB cascade (Tian et al., Reference Tian, Zeng and Tu2023).
We therefore hypothesize that GPR43 activation initiates a signaling cascade that ultimately leads to the inhibition of NF-κB activation. This, in turn, downregulates the transcription of NLRP3, reducing the available pool of NLRP3 protein and thereby suppressing inflammasome assembly and activation. To transition from this hypothesis to a definitive causal mechanism in our model, several future studies are essential. First, in vitro experiments using porcine cell lines with siRNA-mediated knockdown or CRISPR/Cas9-mediated knockout of GPR43 could be performed. Second, the role of NF-κB could be directly assessed by measuring the phosphorylation levels of key pathway components, such as IκBα and p65, following butyrate treatment. Finally, translating these findings in vivo using GPR43 knockout animal models would be crucial to confirm that the protective effects of butyrate are indeed dependent on this receptor. Furthermore, we acknowledge that the present study was designed as a small-scale, mechanism-validation experiment and thus utilized only a single dose of CBP. A critical next step will be to conduct a comprehensive dose–response study to determine the optimal concentration before proceeding to validation in large-scale farming experiments. Together, these future mechanistic and translational investigations will be pivotal in fully elucidating the GPR43-NLRP3 axis and could provide novel therapeutic targets for inflammasome-related diseases.
A primary finding of this work was the identification of several novel peptide sequences within a bioactive CBP fraction (0.5–3 kDa). However, a limitation of the current study is that the observed antimicrobial effects are characteristic of this complex peptide mixture. Consequently, the precise contribution of each individual peptide, or potential synergistic effects between them, remains to be elucidated. The quantification by BCA assay, while useful for standardizing doses of the fraction, reflects the total peptide content and does not allow for a purity assessment or quantification of the specific active molecules. To address this, our future research will prioritize the isolation and characterization of the key active component(s). We plan to utilize methods like reversed-phase HPLC to purify the most abundant or promising peptides identified in our LC-MS/MS analysis. Following purification, the biological activity of each isolated peptide will be individually validated. Furthermore, a more precise quantification method, such as amino acid analysis, will be employed to accurately determine the concentration of the pure, active peptides. This systematic approach will be crucial to definitively link specific sequences to their function, enabling a detailed investigation of their mechanism of action and paving the way for their potential development as therapeutic agents. Furthermore, we acknowledge that a limitation of the present study is the use of a single CBP dose. This study was designed as an initial, mechanism validation experiment to confirm the in vivo efficacy of CBP at a dose previously identified as effective in a preliminary trial. While this approach allowed us to rigorously investigate the underlying physiological mechanisms, it does not establish a full dose–effect relationship. Therefore, a critical next step will be to conduct a comprehensive dose–response study. Such an investigation is essential to determine the optimal concentration that balances maximal efficacy with economic feasibility, which will be crucial before proceeding to validation in large-scale farming experiments and for its potential translational application. Finally, a further limitation of our experimental design is the use of a single time point for post-LPS challenge assessment. This decision was strategic, designed to capture the peak of the acute inflammatory response based on established literature and our preliminary findings, thereby providing a robust “proof-of-concept” for the preventative efficacy of CBP against maximal inflammatory injury. However, we recognize that inflammation is a highly dynamic process. This single-snapshot approach, while effective for its intended purpose, does not allow for the characterization of the full inflammatory cascade, particularly the resolution phase. Therefore, future investigations are being designed to incorporate multiple time points. Such studies will be crucial to elucidate the dynamic effects of CBP, determining whether it only dampens the initial inflammatory spike or also accelerates the return to homeostasis.
Conclusion
This study reveals that CBP significantly enhance growth performance and reduce diarrhea incidence in weaned. The administration of CBP significantly augmented the intestinal barrier function in LPS-induced piglets by optimizing crypt architecture and upregulating the expression of MUC2, as well as tight junction proteins such as ZO1, OCCLUDIN, and CLAUDIN1. Simultaneously, CBP exerted a potent anti-inflammatory effect in the colon, evidenced by the substantial reduction of pro-inflammatory cytokines. This anti-inflammatory action was further manifested by the suppression of NLRP3 inflammasome activation and the restoration of GPR43 expression. Integrated microbiome and metabolomic investigations unveiled that CBP exerted its protective functions via remodeling the gut microbiota, characterized by increased abundance of beneficial microbial taxa, notably Lactobacillus and Coprococcus. Concurrently, the levels of protective metabolites, including HCA and butyrate, were significantly elevated. Correlation analyses confirmed significant associations between these microbiota-derived metabolites and key indicators of intestinal barrier functions. Collectively, these results not only provide strong evidence for the mechanisms of the CBP in enhancing the growth performance and intestinal health of piglets but also illustrate the vast potential of it as an extremely promising alternative to antibiotics.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/anr.2026.10029.
Acknowledgements
This research was financially supported by the National Natural Science Foundation of China (Grant No. 32202682), Zhejiang Provincial Natural Science Foundation of China (Grant No. LQK26C170001), the Key Research and Development Program of Hangzhou (Grant No. 2025SZD1B07) and the “Three Rural Nine Parties” Science and Technology Collaboration Program (Grant No. 2025SNJF041), and the Xianghu Laboratory - Chinese Academy of Agricultural Sciences Special Cooperative Program (Grant No. 2025X1ZN1Z001).
Conflicts of interest
The authors declare that they have no competing interests.