Introduction
Plant genetic resources (PGR) are the foundation of our food system as they contain a broad range of genetic diversity potentially useful for plant breeding and crop improvement. To ensure a robust and sustainable food system capable of withstanding climate change, pests, diseases and increasing food demands, the identification, documentation, conservation and utilization of these resources are imperative (Alercia et al. Reference Alercia, Diulgheroff and Mackay2015; FAO 2025). Lack of access to a greater breadth of genetic diversity is now unnecessarily limiting crop improvement (McCouch et al. Reference McCouch, Baute, Bradeen, Bramel, Bretting, Buckler, Burke, Charest, Cloutier, Cole, Dempewolf, Dingkuhn, Feuillet, Gepts, Grattapaglia, Guarino, Jackson, Knapp, Langridge, Lawton-Rauh, Lijua, Lusty, Michael, Myles, Naito, Nelson, Pontarollo, Richards, Rieseberg, Ross-Ibarra, Rounsley, Sackville Hamilton, Schurr, Stein, Tomooka, van der Knaap, van Tassel, Toll, Valls, Varshney, Ward, Wenzl and Zamir2013; IPCC, Reference Field, Barros, Dokken, Mach, Mastrandrea, Bilir, Chatterjee, Ebi, Estrada, Genova, Girma, Kissel, Levy, MacCracken, Mastrandrea and White2014), with the value of PGR dependent upon the availability of relevant descriptive data to aid germplasm selection and its use (Gotor et al. Reference Gotor, Alercia, Rao, Watts and Caraciolo2008).
PGR are conserved using two main strategies and their associated techniques (CBD 1992). Ex situ conservation involves PGR population sampling, transfer to a secure location and potential long-term population maintenance. This is the well-established PGR conservation model whose effectiveness is well recognized and is responsible for over 5.9 million seed accessions in 852 national genebanks across 116 countries, 4 regional genebanks and 13 international genebanks (FAO 2025). In situ conservation is the maintenance of the PGR where they occur, either cultivated on-farm and in-garden or as wild populations in wild and semi-natural habitats (Maxted et al. Reference Maxted, Hunter and Ortiz Rios2020). Under in situ conservation, PGR populations continue to evolve and respond to environmental changes, within their natural habitats or on-farm systems, while retaining their adaptive diversity. The application of in situ conservation techniques focuses on three types of PGR: crop wild relatives (CWR), wild harvested plants (WHP) and landraces (LR; Maxted et al. Reference Maxted, Hunter and Ortiz Rios2020). CWR and WHP conservation takes place within genetic reserves (as defined by Maxted et al. Reference Maxted, Ford-Lloyd, Hawkes, Maxted, Ford-Lloyd and Hawkes1997), these may be within designated protected areas (albeit often protected by passive conservation measures) or other effective area-based conservation measures (OECM), while LR conservation takes place on-farm or within a home garden. These conservation techniques contrast with ex situ conservation, where resources (e.g. CWR, WHP, LR) are collected, moved and stored in another location such as a genebank, field genebank, community seedbank or botanical garden. It is widely accepted that in situ and ex situ implementation actions should be applied in a complementary manner, where one conservation application enhances and backs-up another (FAO 1996). The application of specific in situ PGR conservation techniques has thus far been limited (Maxted et al. Reference Maxted, Amri, Castañeda-Álvarez, Dias, Dulloo, Fielder, Ford-Lloyd, Iriondo, Magos Brehm, Nilsen, Thormann, Vincent, Kell, Maxted, Ehsan Dulloo and Ford-Lloyd2016), but the associated science and its implementation is advancing rapidly, and it is recognized that wider in situ implementation has the potential to at least double the PGR diversity available to breeders and other users (Maxted and Magos Brehm Reference Maxted and Magos Brehm2023). However, a doubling of diversity can only be achieved if methods for conserving PGR in situ are applied in a coherent and effective manner across institutions and across countries, and this is dependent on an effective and appropriate PGR data and informatics foundation.
Lack of access to data and the scarcity or absence of exchange of information as well as decision-support tools, which simplify complex tasks, are key factors affecting the effectiveness of conservation and use (Bioversity International 2007; Maxted et al. Reference Maxted, Hunter and Ortiz Rios2020). Due to the complexity of in situ PGR systems (as described in Maxted et al. Reference Maxted, Adam-Blondon, Aguilar, Barata, Bartha, Bocci, De Paola, Fitzgerald, Fresta, Fusani, Guzzon, Heywood, Holzherr, Holubec, Iriondo Alegria, Labokas, Maggioni, Magos Brehm, Nazri, Palmé, Prohens, Raggi, Rungis, Phillips, Sarikyan, Šuštar Vozlič, Thormann and Zdunic2024), there is a need to progress the development and use of shared data management and analysis tools for in situ resources, particularly in data collection and monitoring. Such data management tools and analysis packages can help significantly in ensuring that different countries operate in comparable manners further facilitating ease of data exchange and integration (Stephenson and Stengel Reference Stephenson and Stengel2020).
One method to help facilitate the collection of appropriate PGR conservation data is through the use of descriptor lists. Descriptor lists are used in data management to provide a standardized approach to documenting data and have been extensively developed by Bioversity International and the FAO since 1976 (Gotor et al. Reference Gotor, Alercia, Rao, Watts and Caraciolo2008) to help describe plant germplasm. PGR descriptor lists were initially developed in the 1970s to describe a minimum set of characteristics for a specific crop and then evolved in the 1990s to include comprehensive lists with a minimum set of highly discriminating descriptors (Gotor et al. Reference Gotor, Alercia, Rao, Watts and Caraciolo2008). Due to long-term implementation of ex situ conservation techniques over the last 100 years, there are well developed systems for data description, management and exchange of passport data with widely accepted standards and descriptors for the minimum data required for ex situ conservation (Gotor et al. Reference Gotor, Alercia, Rao, Watts and Caraciolo2008; Alercia et al. Reference Alercia, Diulgheroff and Mackay2015). Such data infrastructures help to create a rapid, reliable and efficient means for information exchange, storage and retrieval to facilitate the effective conservation and utilization of germplasm (Gotor et al. Reference Gotor, Alercia, Rao, Watts and Caraciolo2008), the latter being a key reason for PGR conservation. The development of the FAO/IPGRI List of Multi-Crop Passport Descriptors (MCPD) in 2001 provided international standards across crops to facilitate germplasm exchange and is now the standard documentation procedure used in ex situ conservation activities (see Alercia et al. Reference Alercia, Diulgheroff and Mackay2015 for the most recent iteration of these descriptors).
Several ex situ PGR descriptor lists already exist for conservation measures and some of these descriptors have potential for dual application for ex situ and in situ conservation. In parallel, specific in situ conservation descriptors have begun to appear. For CWR these include: a Crop Wild Relative Markup Language (CWRML) (Moore et al. Reference Moore, Kell, Iriondo, Ford-Lloyd and Maxted2008); Quality standards for genetic reserve conservation of CWR (Iriondo et al. Reference Iriondo, Maxted, Kell, Ford-Lloyd, Lara-Romero, Labokas, Magos Brehm, Maxted, Dulloo, Ford-Lloyd, Frese, Iriondo and Pinheiro de Carvalho2012); Core CWR descriptors for in situ conservation (Thormann et al. Reference Thormann, Alercia and Dulloo2013); Joining up the dots: a systematic perspective of crop wild relative conservation and use (Maxted et al. Reference Maxted, Amri, Castañeda-Álvarez, Dias, Dulloo, Fielder, Ford-Lloyd, Iriondo, Magos Brehm, Nilsen, Thormann, Vincent, Kell, Maxted, Ehsan Dulloo and Ford-Lloyd2016); Crop wild relative checklist and inventory descriptors v.1 (Bioversity International and University of Birmingham 2017); Occurrence data collation template v.1 (Magos Brehm et al. Reference Magos Brehm, Kell, Thormann, Gaisberger, Dulloo and Maxted2017a); CWR checklist and inventory data template v.1 (Thormann et al. Reference Thormann, Kell, Magos Brehm, Dulloo and Maxted2017); FAO Descriptors for CWR conserved in situ (Alercia et al. Reference Alercia, López, Marsella and Cerutti2022); and Descriptors for uploading in situ CWR passport data to EURISCO (EURISCO 2025). For LR, existing descriptor lists include: Descriptors for in situ LR conservation (Negri et al. Reference Negri, Maxted, Torricelli, Heinonen, Veteläinen and Dias2012); Descriptors for farmers’ knowledge of plants (Bioversity and The Christensen Fund 2009); and in situ LR propagation management and access guidelines (Caproni et al. Reference Caproni, Raggi and Negri2020). Descriptors which are relevant for both CWR and LR are discussed in the “Concept for a possible extension of EURISCO for in situ CWR and on-farm LR data” (Weise et al. Reference Weise, Kreid and Maxted2020). WHP do not currently have any specific existing descriptors for in situ conservation, but it is assumed that these will be equivalent to those used for CWR as they are both wild plant species and in many cases they are the same species, for example, according to Ciancaleoni et al. (Reference Ciancaleoni, Raggi, Barone, Donnini, Gigante, Domina and Negri2021) 94% of Italian WHP species are also CWR when the taxon group approach is applied (Maxted et al. Reference Maxted, Ford-Lloyd, Jury, Kell and Scholten2006). The work on integrating in situ descriptors into EURISCO (van Hintum and Iriondo Reference van Hintum and Iriondo2022; Kotni et al. Reference Kotni, van Hintum, Maggioni, Oppermann and Weise2023; Maxted et al. Reference Maxted, Adam-Blondon, Aguilar, Barata, Bartha, Bocci, De Paola, Fitzgerald, Fresta, Fusani, Guzzon, Heywood, Holzherr, Holubec, Iriondo Alegria, Labokas, Maggioni, Magos Brehm, Nazri, Palmé, Prohens, Raggi, Rungis, Phillips, Sarikyan, Šuštar Vozlič, Thormann and Zdunic2024) is already enabling countries to input and store data on in situ populations of PGR to make these data accessible for users of germplasm. However, although there has already been some work done on the development of in situ descriptors for PGR, there is no single point to access all descriptors and some descriptors, specifically those related to in situ conservation actions, are missing. Such a complete list of descriptors would be applied by those managing in situ conserved PGR, with a subset of these being incorporated in the various platforms (such as EURISCO) that aim at enhancing the use of these resources via sharing the data with potential users.
A further critical characteristic of PGR data and information is that CWR, WHP or LR can be described at two different nested levels: the taxon or LR level, and the population level. The taxonomic or LR data are often linked to the creation of the checklist or inventory and includes information such as: taxon or LR names and authorities, threat status, characterization data and legislative protection. On the other hand, population data is linked to a subset of information, such as population distribution, size, and management as well as observational data and experimental or evaluation data related to the population being monitored (Maxted et al. Reference Maxted, Hunter and Ortiz Rios2020). The taxon and population level distinction are not mutually exclusive, with most PGR data sets containing both taxon and population data.
The aims of this study were (a) to compile a consolidated list of all standards and descriptors related to the documentation and management of in situ conserved taxa and populations of PGR (CWR, WHP and LR); and (b) to review the consolidated list to identify descriptor gaps and, where gaps exist, propose new descriptors.
Methods
Description of the PGR conservation process
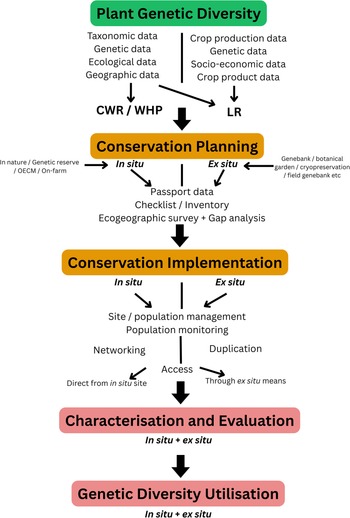
To create a comprehensive descriptor list for PGR conservation, we must first understand how PGR conservation is routinely implemented, describe the process and only then can we locate the data, conduct a gap analysis of current descriptors and fill gaps to generate the consolidated descriptor list. Such a descriptive clarification was recently undertaken by partners within the EU funded PRO GRACE project (https://www.grace-ri.eu/pro-grace), summarized by Maxted et al. (Reference Maxted, Adam-Blondon, Aguilar, Barata, Bartha, Bocci, De Paola, Fitzgerald, Fresta, Fusani, Giuliano, Guzzon, Holzherr, Holubec, Iriondo Alegría, Labokas, Maggioni, Magos Brehm, Palmé, Phillips, Prohens, Raggi, Ralli, Rungis, Sarikyan, Šuštar Vozlič, Thormann and Zdunić2025). They proposed the process is founded on three Principles of PGR Conservation and Use Congruence: (a) long-term, sustainable maintenance of PGR diversity, (b) active conservation and characterization of crop, varietal and related wild taxon diversity using complementary techniques, and (c) conserved resource documentation and availability for utilization within the applicable legislative context. PGR conservation and use is a continuum as reviewed in Maxted et al. (Reference Maxted, Hunter and Ortiz Rios2020, Reference Maxted, Adam-Blondon, Aguilar, Barata, Bartha, Bocci, De Paola, Fitzgerald, Fresta, Fusani, Giuliano, Guzzon, Holzherr, Holubec, Iriondo Alegría, Labokas, Maggioni, Magos Brehm, Palmé, Phillips, Prohens, Raggi, Ralli, Rungis, Sarikyan, Šuštar Vozlič, Thormann and Zdunić2025) and summarized in Figure 1 with three steps:
1. Plant genetic diversity – the base resource;
2. Conservation – the planning and implementation stages;
3. Use – characterization, evaluation and genetic diversity utilization.
Each of these elements can be subdivided and is associated with distinct data types, as listed in Figure 1. Each element will have descriptors associated with it.
Continuum between PGR conservation and use, showing the links between genetic diversity, conservation actions, and utilisation through associated data types and descriptors.

Consolidation of available PGR descriptors in relation to in situ conservation
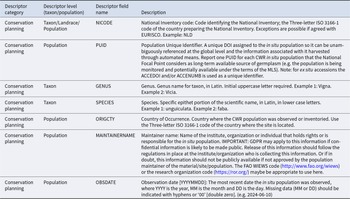
To create a comprehensive descriptor list for the PGR in situ conservation process, all previously developed descriptors for PGR were collated (see Table 1) into one database. These included the Multi-Crop Passport descriptors (Alercia et al. Reference Alercia, Diulgheroff and Mackay2015) which are also equally relevant to in situ conservation activities. Many of the descriptors were duplicated across the references but with slightly different names or descriptions. The duplicates were consolidated by merging descriptors which were similar in terms of their name and/or descriptions, or by keeping the most commonly applied descriptor name or description. All duplicated descriptors were noted so reference back to the original descriptors can be made.
The different categories of descriptors with a list of corresponding references. The references refer to literature where particular descriptors have previously been published and have been included in the consolidated descriptor list. Some descriptors fit into more than one category

The categories of descriptors, illustrated in Figure 1 and Table 1, can be applied at different levels related to the individual taxon, the individual LR or the population as a whole. This differentiation has implications for how data are collected in the field and the use of that data for conservation planning. Below, each descriptor category is explained and the level at which data should be collected is identified.
Conservation planning (taxon, LR and population level):
• Passport data (taxon, LR and population level)
o Data associated with the inventory, population or accession. These help to keep associated data organized and provide a key reference linking datasets to each other. Information includes: national checklist and inventory code; regional inventory code; managing body information (for both in situ and ex situ); population/sample origin; population code, and DOI (Digital Object Identifier) (Alercia et al. Reference Alercia, López, Sackville Hamilton and Marsella2018) information (for both in situ and ex situ).
• Checklist/inventory data (taxon, LR level)
o PGR checklist – Initially a checklist of all relevant PGR within a defined area. A checklist could be produced for a particular PGR (e.g. CWR, WHP or LR), at different levels, such as global, regional, national (the most common), local, or based on a particular area (i.e. a protected area). The checklist contains a list of core taxon information for CWR or WHP, or a list of core LR information, including: taxon/LR ID; taxon name or LR name; family, genus, species, authority of the taxon name; rank, synonyms and a photo or image.
o Annotated checklist – These data are recorded not for individual accessions or populations but are focused on the taxonomic or LR level and usually relate to national level prioritization. The checklist is annotated with information that aids with prioritization, including: related crop information; gene pool or taxon group concept type; related crop/LR use; breeding use, general distribution, socioeconomic data and threat assessment information.
o PGR inventory – The inventory is a subset of the checklist that contains priority taxa/LR identifier and additional data that includes: common name of the taxon or LR; taxon/LR biology details and taxon/LR conservation action information (both in situ and ex situ).
• Ecogeographic survey and gap analysis (taxon, LR and population level)
o Taxonomic, ecological, geographic and genetic information that is gathered as a basis for the ecogeographic survey and the gap analysis. In addition to the data management and passport data descriptors, these data include: detailed location and distribution information of the in situ population; presence within a protected area; ecological and habitat information (such as soil type).
Conservation implementation (population level):
• Site/population management information (population level) – A management plan will be put in place for a site that contains the target CWR, WHP or LR populations. Data generated from this management include: the managing body or maintainer information; management interventions; conservation actions taking place; population sampling and periodic resampling for ex situ backup conservation; characterization and user access.
• Population monitoring (population level) – Once sites or populations are being managed for conservation, then monitoring will need to take place. Time series data generated from this process includes: loss risk of LR; continuity of growing LR; cultivation period; status on-farm; regeneration ex situ; herbivore impacts; vegetation height; negative habitat features; habitat availability; population information (numbers, frequency); threats to population; monitoring of genetic diversity (including genomic and phenotypic data) and habitat conservation activities.
• In situ networking (population level) – In situ conservation networks of sites and populations should be established where possible to ensure the range of diversity of a taxon (as recorded in previous data collection) is efficiently and effectively conserved within a meta-population context. The network can be at a local, national, regional and global scales. Data associated with the creation of a network include: the number of sites in a network; the scale of the network; if samples of the in situ network populations are backed-up and/or conserved ex situ (this information should be harmonized with ex situ information); ex situ backup samples (see below).
• Ex situ conservation and in situ backup (population level) – In situ population collection and safety ex situ duplication are essential for complementary conservation, characterization and to promote utilization. This is distinct from senso stricto ex situ conservation, where a wild or cultivated population is sampled, collected and placed in an ex situ facility, but the original sampled population is not being actively conserved in situ or on-farm. Whether the sampled population is an in situ backup or a normal ex situ sample, existing ex situ descriptors can be used to describe the accession. Ex situ descriptors have been widely developed and utilized (Alercia et al. Reference Alercia, Diulgheroff and Mackay2015) with a recent project producing descriptors that bridge the gap between in situ conservation and ex situ back up (Weise et al. Reference Weise, Kreid and Maxted2020; van Hintum and Iriondo Reference van Hintum and Iriondo2022; EURISCO 2025). Both the in situ and ex situ conserved resource can be backed up in situ. when a sample is taken from an actively conserved in situ population, and ex situ when the sample is duplicated across multiple genebanks. These descriptors incorporate ex situ data which includes information on: ex situ institute; ex situ accession number; and provides information on the in situ population DOI from which the in situ backup sample was collected.
• Access to in situ conserved populations (population level) – Whether the population is conserved in situ or ex situ, it is essential that potential users can access the conserved resource. For in situ resources this could most effectively be provided from the ex situ backup, or less easily from the in situ population. Descriptors for in situ access include the Multilateral System status of the in situ material (Alercia et al. Reference Alercia, López, Marsella and Cerutti2022) and in situ population access. The record needs to clearly show that national or international ABS legislation has been adhered to in relation to the provision of user’s samples from the ex situ backup.
Characterization and evaluation (LR and taxon level):
• Specific characterization and evaluation descriptors have traditionally been developed separately to ex situ conservation descriptors. They have been directly associated with ex situ conservation activities and have been developed for over 115 crops (see the Bioversity Descriptors at https://hdl.handle.net/10568/56589; Alercia Reference Alercia2011). Characterization and evaluation descriptors are crop specific and must be defined for each new crop being utilized (Bioversity International 2007). It is assumed that practical characterization and evaluation is independent of the mode of conservation being used, and descriptors developed in the ex situ context will be equally applicable for in situ conserved CWR, WHP and LR germplasm. Also, practically it seems unlikely that characterization and evaluation will be conducted on in situ populations due the lack of expertise in these assessments of the in situ population maintainers (individual farmers or protected area managers) and therefore it is much more likely that the necessary characterization and evaluation will be carried out on the backup sample of the in situ resource which is held in the genetic resource centre.
Utilization (taxon, LR and population level):
• Sustainable utilization of PGR is a fundamental reason for the conservation of PGR. These resources can be used by plant breeders and farmers (for LR material) and may also have a direct market use as food or medicine. The following data may be collected: any geographical designation or registration (for both LR and WHP); variety registration (for LR); DOI registration; use by breeders; market demands; product use; part of plant used; reasons for growing or harvesting the taxon from the wild and socioeconomic value.
Selection of the descriptors
Using the methodology outlined above, the existing descriptors relevant to in situ conservation (Table 1), were evaluated against the descriptor categories associated with the in situ conservation process (Figure 1). Gaps were identified where there were no existing descriptors covering a particular descriptor category. These new descriptors were generated through a process of consultation and review with experts. New descriptors were given a definition and description to ensure ease of use. Consistent with previous PGR descriptor lists (Alercia et al. Reference Alercia, López, Marsella and Cerutti2022; EURISCO 2025) a ‘Mandatory’ set of descriptors was identified as the minimum amount of data required for the in situ material. Furthermore, a more extensive prioritized subset of descriptors, known as the ‘Core’ descriptor set, was identified to enhance data collection efficiency, as suggested by Thormann et al. (Reference Thormann, Alercia and Dulloo2013), Magos Brehm et al. (Reference Magos Brehm, Kell, Thormann, Maxted and Dulloo2017b), Alercia et al. (Reference Alercia, López, Marsella and Cerutti2022). Finally, a ‘Recommended’ set of descriptors were identified to incorporate the breadth of data that may be possible to collect for the in situ material. This strategy enables recorders to prioritize essential information for each in situ conservation activity, while remaining flexible to include additional relevant data where appropriate. The descriptor lists were then reviewed by a panel of 22 experts from the PRO-GRACE project consortium, and the ECPGR CWR, On-farm Conservation, and Documentation and Information Working Groups representing leading European research and conservation institutions to generate the final recommended set of descriptors.
Results
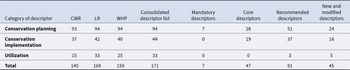
The consolidated descriptor list titled ‘Plant Genetic Resource consolidated in situ descriptors v1’ is available from Harvard Dataverse (Phillips et al. Reference Phillips, Magos Brehm, Adam-Blondon, Avagyan, Clarke, Fresta, Forycka, Gailïte, Gay, Iriondo, Labokas, Palmé, Raggi, Thormann, Weise, Zdunic and Maxted2025). The list comprises 171 descriptors, of which seven are designated as ‘Mandatory descriptors’ (Table 2) and 47 are designated as the ‘Core descriptors’ for in situ conservation of PGR. ‘Mandatory descriptors’ are those which are required to be filled in when collecting and recording PGR. ‘Core descriptors’ are additional highly recommended descriptors which will greatly improve the information available for conservation and use of the PGR. 91 descriptors are designated as ‘Recommended’ to help enable a more thorough assessment of in situ populations. Of the total number of descriptors, 145 relate to CWR, 169 relate to LR and 159 relate to WHP (Table 3). There are 12 descriptors which are unique to LR and one descriptor is unique to WHP. In terms of the different levels of data for each descriptor, 95 descriptors relate to population level data, 82 descriptors relate to taxon level data and 79 relate to LR level data. There are eight descriptors which relate to all three levels of data, primarily descriptors related to conservation planning. The ‘Other’ category includes two options for the user to enter ‘REMARKS’ and ‘COMMENTS’ on other descriptor values. These are excluded from the counts of different descriptors as they are not considered true descriptors because they do not define an attribute, characteristic or measurable trait that is observed.
The seven ‘Mandatory descriptors’ for in situ conservation of PGR

The number of descriptors in each category for: CWR, LR, WHP, core descriptors, recommended descriptors and new and modified descriptors. Counts do not include the ‘Other’ category which allows the user to enter ‘REMARKS’ and ‘COMMENTS’ on other descriptors

The ‘Plant Genetic Resource consolidated in situ descriptors v1.2’ is composed of the following Excel sheets: ‘Explanatory notes’ – a title page displaying information about the descriptor tables and ‘All_descriptors’, showing the full set of 171 descriptors plus information on references related to each descriptor. Table 3 gives an overview of the total number of descriptors identified.
The consolidated list of in situ descriptors (Phillips et al. Reference Phillips, Magos Brehm, Adam-Blondon, Avagyan, Clarke, Fresta, Forycka, Gailïte, Gay, Iriondo, Labokas, Palmé, Raggi, Thormann, Weise, Zdunic and Maxted2025) is extensive and it is not expected that all descriptors will be used by any one PGR institution, project or application. This is why the ‘Mandatory descriptors’ (Table 2) and ‘Core descriptors’ have also been developed (see Table 3 for an overview). It is expected that the user will collect the ‘Mandatory descriptors’ information for either CWR, WHP or LR. The user can select these subsets using the filters provided. The ‘Core descriptors’ provide an opportunity for the user to input more information useful for conservation but are not mandatory (see Phillips et al. Reference Phillips, Magos Brehm, Adam-Blondon, Avagyan, Clarke, Fresta, Forycka, Gailïte, Gay, Iriondo, Labokas, Palmé, Raggi, Thormann, Weise, Zdunic and Maxted2025).
The full descriptor list (Phillips et al. Reference Phillips, Magos Brehm, Adam-Blondon, Avagyan, Clarke, Fresta, Forycka, Gailïte, Gay, Iriondo, Labokas, Palmé, Raggi, Thormann, Weise, Zdunic and Maxted2025) also contains information on which descriptors are useful for the following assessments: Red listing (IUCN Red List Technical Working Group 2018) = 13 descriptors; LR threat assessment (Almeida et al. Reference Almeida, Barata, De Haan, Joshi, Magos Brehm, Yazbek and Maxted2024) = 11 descriptors; inclusion of populations or sites in in situ networks (Maxted et al. Reference Maxted, Amri, Castañeda-Álvarez, Dias, Dulloo, Fielder, Ford-Lloyd, Iriondo, Magos Brehm, Nilsen, Thormann, Vincent, Kell, Maxted, Ehsan Dulloo and Ford-Lloyd2016) = 16 descriptors; and for the establishment of in situ genetic reserves (Iriondo et al. Reference Iriondo, Maxted, Kell, Ford-Lloyd, Lara-Romero, Labokas, Magos Brehm, Maxted, Dulloo, Ford-Lloyd, Frese, Iriondo and Pinheiro de Carvalho2012) = 13 descriptors. The consolidated list of in situ descriptors does not cover the full range of information required for each of these assessment methods.
There were 29 new descriptors generated, with a further 16 descriptors for which either the description or descriptor code was altered from the original to provide more clarity (for a full list of new descriptors see Table S1), therefore resulting in a total of 45 new or modified descriptors (Table 3). Of these new or modified descriptors, one descriptor is a ‘Mandatory descriptor’, eight descriptors are included in the ‘Core descriptor’ list and an additional 26 are included as ‘Recommended descriptors’ (Table S1). Proportionally the number of new descriptors was highest for the conservation planning category, with the utilization category having the lowest number of new descriptors (Table 3). The main type of data required for each descriptor is text, however, some descriptors require numeric data relating to coordinates, a code number (which then relates back to text), or a specific measurement or estimation such as ‘slope angle in degrees’. For each descriptor in the PGR consolidated in situ descriptor list (Phillips et al. Reference Phillips, Magos Brehm, Adam-Blondon, Avagyan, Clarke, Fresta, Forycka, Gailïte, Gay, Iriondo, Labokas, Palmé, Raggi, Thormann, Weise, Zdunic and Maxted2025) there is a clear description for which type of data to input.
Discussion
The PGR consolidated in situ descriptor list presented is built on existing diverse descriptor lists and where gaps were identified further descriptors were added. The number of available descriptor lists in the in situ domain implies that the adoption of these lists has been low as no list has become the standard. We hope that consolidating these lists will create a timely resource to enhance future, practical implementation of PGR in situ conservation at national, regional and global levels, providing a basis for improved information and data management. It is widely accepted that using standardized descriptors avoids unnecessary repetitive data input and input errors, facilitates data exchange, allows multiple uses of the same dataset, reduces any problem of text synonyms, provides greater consistency and automatic checks for data integrity, allows easier comparison of results, and quicker data searching (Gotor et al. Reference Gotor, Alercia, Rao, Watts and Caraciolo2008). Therefore, having a consolidated list of appropriate descriptors will help ensure improved in situ resource conservation and access to PGR materials that are currently not accessible for users.
Of the 171 descriptors, there are seven designated at ‘Mandatory descriptors’ required as the minimum information for in situ conservation of PGR. There are 47 designated as ‘Core descriptors’ which are highly recommended for in situ conservation of PGR, but they are not mandatory. Eight of the ‘Core descriptors’ are newly developed (Phillips et al. Reference Phillips, Magos Brehm, Adam-Blondon, Avagyan, Clarke, Fresta, Forycka, Gailïte, Gay, Iriondo, Labokas, Palmé, Raggi, Thormann, Weise, Zdunic and Maxted2025), with the other descriptors being common across multiple other descriptor lists. Thormann et al. (Reference Thormann, Alercia and Dulloo2013) developed 65 core descriptors for in situ CWR conservation, therefore the number identified here seems realistic. However, Thormann et al. (Reference Thormann, Alercia and Dulloo2013) further reduced the number to 13 for the minimum mandatory descriptors required when exchanging data. In other work on descriptor development for in situ conservation activities of PGR, EURISCO (2025) identified 28 minimum descriptors; Alercia et al. (Reference Alercia, López, Marsella and Cerutti2022) identified 24, including seven mandatory descriptors, and Magos Brehm et al. (Reference Magos Brehm, Kell, Thormann, Gaisberger, Dulloo and Maxted2017a) identified 35. The list of seven ‘Mandatory descriptors’ is aligned to both Alercia et al. (Reference Alercia, López, Marsella and Cerutti2022) and EURISCO (2025), and thus it is expected that this will be the minimum information required for in situ PGR. The likelihood of acquiring the data required for the ‘Mandatory descriptors’ is high as there are only seven descriptors required. If it was not possible to collect this data, the material should still be collected and a clear pathway should be put in place to ensure the ‘Mandatory descriptors’ are completed in a timely manner. Without this information it would not be possible to follow the conservation continuum (Figure 1), thus limiting the effectiveness of in situ conservation and the usefulness of the material for the user. It should therefore be a priority to ensure the information for the ‘Mandatory descriptors’ are collected.
The consolidated descriptor list presented here is expected to be applied across three categories of PGR (CWR, LR and WHP), therefore there will be higher numbers of the ‘Core’ descriptors required than previous descriptor lists which have covered one type of PGR. The descriptor list also includes a ‘Recommended’ descriptor category to enable more thorough data collection to take place. It is expected that this list of descriptors will change over time as they are used in-the-field, so refinement in the number of ‘Mandatory’, ‘Core’ and ‘Recommended’ descriptors may occur at a later point. The mechanism for such a process of descriptor development would need to be identified. At a European level the ECPGR Documentation and Information Working Group, could take on responsibility for coordinating development and use of the descriptors. The list could also be ratified by the Biodiversity Information Standards (TDWG) association, which would also help to ensure the descriptors adhere to the FAIR principles of data sharing (Wilkinson et al. Reference Wilkinson, Dumontier, Aalbersberg, Appleton, Axton, Baak, Blomberg, Boiten, da Silva Santos, Bourne, Bouwman, Brookes, Clark, Crosas, Dillo, Dumon, Edmunds, Evelo, Finkers, Gonzalez-Beltran, Gray, Groth, Goble, Grethe, Heringa, Hoen, Hooft, Kuhn, Kok, Kok, Lusher, Martone, Mons, Packer, Persson, Rocca-Serra, Roos, van Schaik, Sansone, Schultes, Sengstag, Slater, Strawn, Swertz, Thompson, van der Lei, van Mulligen, Velterop, Waagmeester, Wittenburg, Wolstencroft, Zhao and Mons2016, Reference Wilkinson, Sansone, Méndez, David, Dennis, Hecker, Kleemola, Lacagnina, Nikiforova and Castro2022).
The descriptors cover three types of PGR, namely CWR, WHP and LR with many of the descriptors being common across these groups. For example, all the descriptors relating to ‘Conservation Planning’ are applicable to all three components of PGR. This is because ‘Conservation Planning’ data are composed of foundational data around plant breeding systems, taxonomy, common names, etc. and is also why this category has the highest number of descriptors. The 12 descriptors which are unique to LR are composed of a subset of ‘Conservation Implementation’ and ‘Utilization’ descriptors, which were developed by Negri et al. (Reference Negri, Maxted, Torricelli, Heinonen, Veteläinen and Dias2012) and Caproni et al. (Reference Caproni, Raggi and Negri2020). These descriptors represent the unique way in which LRs are maintained by continued farming practices and how they are used not only for their genetic resource potential (like CWR and WHP), but also for their direct market value (also like WHP). The one descriptor which is unique to WHP is the ‘Wild Harvested Plant Continuity’ descriptor which is a new descriptor (Phillips et al. Reference Phillips, Magos Brehm, Adam-Blondon, Avagyan, Clarke, Fresta, Forycka, Gailïte, Gay, Iriondo, Labokas, Palmé, Raggi, Thormann, Weise, Zdunic and Maxted2025). This descriptor was based on a LR descriptor and gathers information on whether the people harvesting the WHP plan to continue doing so. This information will help to determine the conservation needs of the taxa and relative threat, which may change as the wild collection and use of the WHP changes. There are no unique CWR descriptors as they are all shared with WHP, with many of these also shared with LR. This is because both CWR and WHP are components of the natural flora and do not require management by humans. Unlike CWR and WHP, LR populations require management interventions and utilization to continue to exist, this being a key component of their conservation management. This is reflected in the number of ‘Utilization’ descriptors for LR which is twice as many as for CWR. The ‘Utilization’ category has the lowest number of total descriptors which may be because they are generally less developed for in situ conservation, although they are highly relevant for use of PGR. Further development and application of ‘Utilization’ descriptors is required, as data on utilization of resources is not necessarily available when the material is first collected and this study was only focused on the active in situ conservation aspect of PGR.
The descriptors cover data at the levels of the population and taxon or LR. This ensures that the data recorder gathers the appropriate information for the survey they are conducting. In many instances, data at both population and taxon or LR level will be required for a complete assessment of the effective conservation of a particular taxon or LR. Population level data helps to assess the size of a population, and taxon level data helps to determine the bigger picture in terms of conservation actions or its wider distribution (Bevilacqua et al. Reference Bevilacqua, Anderson, Ugland, Somerfield and Terlizzi2021) for that taxon or LR. The descriptors which represent either taxon or LR and population level data, are those related to conservation planning information, such as, the National Inventory Code. This commonality is necessary as either taxon or LR and population level data need to be interlinked and findable across different levels of assessment.
The work in this project was focused on the consolidation and development of conservation descriptors, not on the development of Characterization and Evaluation descriptors. These have traditionally been developed separately to conservation descriptors as they are taxon or variety specific. However, it is worth considering how they might be applied to in situ PGR taxa and LR in the future. In situ characterization of traits of interest for each taxa or LR would help to inform on how the genotype is expressed under the environmental conditions of origin. The benefit of defining taxa specific in situ descriptors is that the connection between conservation and use could be more closely aligned, through the enabling of more targeted in situ conservation efforts of those traits which are relevant for the user community. Clearly, generating such data poses a significant challenge, as the in situ populations must be surveyed precisely when the trait of interest is expressed. In addition, genotype-by-environment interactions complicate comparisons across different populations. Nevertheless, it would be valuable to develop some pilot characterization and evaluation descriptors for the CWR of a few specific crops or for some LR varieties, and to test them in the field.
Currently, the data and information included in EURISCO (Kotni et al. Reference Kotni, van Hintum, Maggioni, Oppermann and Weise2023) which previously related to ex situ accessions only, is being expanded to include the passport data for in situ CWR populations’ data. Should consideration be given to expanding EURISCO beyond its current 28 in situ descriptors proposed by van Hintum and Iriondo (Reference van Hintum and Iriondo2022), to include the additional descriptors generated here? The remit of EURISCO is clearly stated, ‘to provide a European Search Catalogue for Plant Genetic Resources’ (EURISCO 2025), in other words, it is a germplasm search engine to enable potential users to locate and access actively conserved ex situ or in situ PGR resources. It is not a generalized PGR information system and so it is not recommended that the additional descriptors generated here be added to an extended EURISCO. Just as genebank curatorial information is not added to EURISCO, so the equivalent in situ curation (= management and monitoring data) should not be included. However, one of the several distinctions between ex situ and in situ applications is the level of conservation site integration. Genebanks have historically worked well semi-independently, with van Hintum and Wijker (Reference van Hintum and Wijker2024) calling for the wider application of quality management standards in the genebank environment, resulting in the Genebank Managers Network being recently formed with the aim of improving the function and strengthening the cooperation among genebank managers in Europe (ECPGR 2023). The more recent implementation of systematic and formal in situ and on-farm conservation has highlighted the benefit of a more integrated structure that encourages the building of a curatorial (i.e. management related) evidence-based approach; however for such an approach to be implemented effectively, individual in situ/on-farm sites need to be networked, not working in isolation (FAO 2014; Maxted et al. Reference Maxted, Avagyan, Frese, Iriondo, Magos Brehm, Singer and Kell2015, Reference Maxted, Amri, Castañeda-Álvarez, Dias, Dulloo, Fielder, Ford-Lloyd, Iriondo, Magos Brehm, Nilsen, Thormann, Vincent, Kell, Maxted, Ehsan Dulloo and Ford-Lloyd2016, Reference Maxted, Adam-Blondon, Aguilar, Barata, Bartha, Bocci, De Paola, Fitzgerald, Fresta, Fusani, Guzzon, Heywood, Holzherr, Holubec, Iriondo Alegria, Labokas, Maggioni, Magos Brehm, Nazri, Palmé, Prohens, Raggi, Rungis, Phillips, Sarikyan, Šuštar Vozlič, Thormann and Zdunic2024). Elements of such an integrated structure have begun to be established, including the in situ LR best practice evidence-based database (ECPGR 2025a) and the interactive tool for identifying the CWR taxa in European Natura 2000 protected areas (ECPGR 2025b). Such resources are enhancing the value of the ECPGR website and offer a complementary resource to EURISCO that helps maximize conservation efforts of PGR.
All active in situ or on-farm conservation sites will require a site management plan that sets out the prescription for target population management and inevitably this will require and generate significant data as a routine part of population management and monitoring. A question to be addressed is should these data be held entirely within the national PGR network or is there advantage in aggregating these data at the European or global level. There are pros and cons of both approaches, but it is obvious whichever approach is followed, that (a) it would be helpful to debate now the best way forward rather than allow it to evolve ad hoc, and (b) it would be beneficial to have a coherent informatics infrastructure, including tools to aid analysis, at the national, European or global level or all three, to collate population management data over repeated monitoring cycles.
To increase interoperability and ensure that the descriptors contribute to broader biodiversity conservation efforts, alignment with international data standards such as the Biodiversity Information Standards (TDWG) and integration with platforms like GBIF (https://www.gbif.org/) and Genesys PGR (https://www.genesys-pgr.org/) should also be explored. This would enhance visibility, avoid data silos, and facilitate cross-sectoral applications of PGR data. It will still be important to differentiate between purposes of information systems at different levels (e.g. regional, national and conservation network focused) and the different details of information required at each of these levels. These issues, and those relating to which data should be held at what level, require further discussion in the ECPGR Documentation and Information WG, the EURISCO Advisory Committee, and with the CWR and On-farm WGs.
Integration of the consolidated list could be further developed as the basis of a proper ontology, linking terms used in various contexts and allowing automatic analysis of data about in situ PGR conservation activities (Arnaud et al. Reference Arnaud, Laporte, Kim, Aubert, Leonelli, Miro, Cooper, Jaiswal, Kruseman, Shrestha and Buttigieg2020). The basis of an ontology for PGR descriptors has already been developed (Škofič and Dias Reference Škofič and Dias2014) and addition of the PGR consolidated in situ descriptor list would help to expand this ontology. The Crop Ontology (Arnaud et al. Reference Arnaud, Laporte, Kim, Aubert, Leonelli, Miro, Cooper, Jaiswal, Kruseman, Shrestha and Buttigieg2020), which is primarily focused on characterization and evaluation data of crops and has been developed based on the FAIR principles (Wilkinson et al. Reference Wilkinson, Dumontier, Aalbersberg, Appleton, Axton, Baak, Blomberg, Boiten, da Silva Santos, Bourne, Bouwman, Brookes, Clark, Crosas, Dillo, Dumon, Edmunds, Evelo, Finkers, Gonzalez-Beltran, Gray, Groth, Goble, Grethe, Heringa, Hoen, Hooft, Kuhn, Kok, Kok, Lusher, Martone, Mons, Packer, Persson, Rocca-Serra, Roos, van Schaik, Sansone, Schultes, Sengstag, Slater, Strawn, Swertz, Thompson, van der Lei, van Mulligen, Velterop, Waagmeester, Wittenburg, Wolstencroft, Zhao and Mons2016, Reference Wilkinson, Sansone, Méndez, David, Dennis, Hecker, Kleemola, Lacagnina, Nikiforova and Castro2022), should also be used to help refine the PGR consolidated in situ descriptor list and develop it towards a functional ontology. Furthermore, we recommend that the development of a PGR ontology be based on the Open Biological and Biomedical Ontology Foundry (OBO Foundry) principles, to help increase accessibility to conservation communities.
Practical application, specifically of the in situ descriptors (Phillips et al. Reference Phillips, Magos Brehm, Adam-Blondon, Avagyan, Clarke, Fresta, Forycka, Gailïte, Gay, Iriondo, Labokas, Palmé, Raggi, Thormann, Weise, Zdunic and Maxted2025), is also needed to help with the testing, continuous development and updating of the consolidated descriptor list presented here. Although the list is comprehensive, it should be viewed as an initial version to be replaced by improved future iterations. To ensure the widest possible adoption, especially in countries with limited resources or technical infrastructure, a phased or tiered implementation model could be beneficial. Starting with the ‘Mandatory descriptors’ then the ‘Core descriptors’ and allowing for progressive integration of additional data fields, such as the ‘Recommended descriptors’. This would enable flexibility while maintaining core data integrity. With the funding of the ‘Extension of EURISCO for CWR in situ data and preparation of pilot countries’ data sets’ in several countries (Albania, Bulgaria, Cyprus, Czech Republic, Georgia, Germany, Italy, Lithuania, The Netherlands, Poland, Portugal, Romania, Slovenia, Spain and the United Kingdom; https://www.ecpgr.org/working-groups/crop-wild-relatives/cwr-in-eurisco) and the production of the initial iteration of the consolidated descriptor list (Phillips et al. Reference Phillips, Magos Brehm, Adam-Blondon, Avagyan, Clarke, Fresta, Forycka, Gailïte, Gay, Iriondo, Labokas, Palmé, Raggi, Thormann, Weise, Zdunic and Maxted2025), there is a need for further capacity building for users (both in situ and ex situ practitioners) on how to use and apply these descriptors and their value and importance for conservation of PGR. Such training might be offered jointly by the regional ECPGR Documentation and Information, CWR and On-farm WGs and national nodes for PGR conservation.
The descriptor list should continue to be developed in line with the FAIR principles (Wilkinson et al. Reference Wilkinson, Dumontier, Aalbersberg, Appleton, Axton, Baak, Blomberg, Boiten, da Silva Santos, Bourne, Bouwman, Brookes, Clark, Crosas, Dillo, Dumon, Edmunds, Evelo, Finkers, Gonzalez-Beltran, Gray, Groth, Goble, Grethe, Heringa, Hoen, Hooft, Kuhn, Kok, Kok, Lusher, Martone, Mons, Packer, Persson, Rocca-Serra, Roos, van Schaik, Sansone, Schultes, Sengstag, Slater, Strawn, Swertz, Thompson, van der Lei, van Mulligen, Velterop, Waagmeester, Wittenburg, Wolstencroft, Zhao and Mons2016, Reference Wilkinson, Sansone, Méndez, David, Dennis, Hecker, Kleemola, Lacagnina, Nikiforova and Castro2022) and could also be integrated into a network of FAIR germplasm bank databases, in line with the work of Cámara Ballesteros et al. (Reference Cámara Ballesteros, Aguayo Jara, Verykaki, Pastor Del Olmo, Moreno Vázquez, Torres and Wilkinson2024), that would allow greater sharing of data and thus more effective conservation strategies for PGR. Although the Cámara Ballesteros et al. (Reference Cámara Ballesteros, Aguayo Jara, Verykaki, Pastor Del Olmo, Moreno Vázquez, Torres and Wilkinson2024) study focuses on ex situ material, the principles should be explored in their application to in situ conservation methods. Descriptors could also be added into information systems which are being developed for the management of PGR such as GRIN Global Community Edition (GRIN-Global Community Edition | GGCE). In parallel with capacity building, user-friendly digital tools, such as mobile data collection templates, automated validation systems and decision-support dashboards, would help streamline field implementation. These tools are critical to lowering barriers for adoption, ensuring data consistency and reducing the workload for practitioners managing multiple taxa or sites.
Timely resolution is essential, as the practical implementation of in situ CWR population conservation at a broader national and regional level is currently occurring. This urgency is underscored by EC funding for three Horizon Europe research projects: COUSIN – CWR utilization and conservation for sustainable agriculture (https://cousinproject.eu/); PRO-WILD – Protect and promote CWR (https://www.pro-wild.eu); and FRUITDIV – Sustainable agriculture to preserve nature’s treasures (https://fruitdiv.eu). Although each of these three projects focus on a narrow subset of European crops, they all incorporate in situ conservation and recognize the need for accessible CWR information for stakeholders and users. The consolidated in situ descriptor list will help them achieve their aims effectively and efficiently.
Development of the consolidated in situ PGR descriptor list presented here has highlighted the value that the PGR community sees in effective PGR and in situ data management for PGR conservation. The descriptors can be applied at all levels from local and national to regional and global. The process has increased discussion around the appropriate descriptors to use, as well as the process of in situ conservation and how practical it is to conduct this level of data collection. The robustness of the ex situ conservation process and community has helped to enable the development of the in situ descriptors in a pre-emptive rather than reactionary manner. Much work remains to develop robust in situ PGR descriptors. Primarily the descriptors need to be used and adapted by the PGR community to allow further refinement, including the addition of descriptors from other sources. Critically this process of refinement should ensure the descriptors adhere to the ‘FAIR principles’ of Findable, Accessible, Interoperable and Reusable, regarded as the cornerstone of contemporary informatics (Wilkinson et al. Reference Wilkinson, Dumontier, Aalbersberg, Appleton, Axton, Baak, Blomberg, Boiten, da Silva Santos, Bourne, Bouwman, Brookes, Clark, Crosas, Dillo, Dumon, Edmunds, Evelo, Finkers, Gonzalez-Beltran, Gray, Groth, Goble, Grethe, Heringa, Hoen, Hooft, Kuhn, Kok, Kok, Lusher, Martone, Mons, Packer, Persson, Rocca-Serra, Roos, van Schaik, Sansone, Schultes, Sengstag, Slater, Strawn, Swertz, Thompson, van der Lei, van Mulligen, Velterop, Waagmeester, Wittenburg, Wolstencroft, Zhao and Mons2016, Reference Wilkinson, Sansone, Méndez, David, Dennis, Hecker, Kleemola, Lacagnina, Nikiforova and Castro2022) and essential for the effective use of PGR. These processes to improve the consolidated in situ PGR descriptor list will be essential to ensure the continued effective conservation of our most valuable resources.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S1479262126100562.
Acknowledgements
We gratefully acknowledge the support of the Horizon Europe programme, project ‘Promoting a Plant Genetic Resource Community for Europe (PRO-GRACE)’ N. 101094738, in preparation of this manuscript. https://doi.org/10.3030/101094738.
Competing interests
The authors declare none.
OA Funding statement
Open access funding provided by Wageningen University & Research.