INTRODUCTION
The first infections due to penicillin-intermediate isolates appeared at The Children's Hospital of Alabama in 1981, with serotypes/groups 6, 14, 19, and 23 predominating [Reference Willett, Dillon and Gray1]. The frequency and level of resistance was relatively low until the late 1980s and early 1990s, when more fully penicillin-resistant and multi-resistant clones emerged and spread in many parts of the world [Reference Doern2]. By the time the 7-valent pneumococcal conjugate vaccine (PCV7) was introduced, eight mainly international clones accounted for most resistant isolates from the USA [Reference Gherardi3]. Looking in more detail at serogroup 6, two international clones (Spain6B-2 and South Africa6B-8) were common in the USA at the time of PCV7 introduction, but many other clones were known to be present at least back to 1997 [Reference Gertz4]. In the current work we focus on serogroup 6 isolates from our collections of older isolates, with the goal of looking for ancestors of the clones that came into play around the time of the introduction of the PCV7 vaccine.
Prior to the early 2000s, the major evolutionary stress on this human pathogen was the widespread use of penicillin, beginning in the late 1940s. ‘Epidemic’ serotypes – 1, 3, 5, 7 – became less frequent, probably because early treatment broke the cycle of spread from sick individuals to susceptible healthy contacts [Reference Feikin and Klugman5]. These types were gradually replaced with previously well-known ‘childhood’ serotypes/groups – 6, 14, 19, 23 – that were less virulent but more successful colonizers and more often associated with infections in children [Reference Feikin and Klugman5, Reference Nemir, Andrews and Vinograd6]. The types selected for the PCV7 vaccine were the most common and those most often associated with penicillin and multiple resistance, including serotypes 4, 6B, 9 V, 14, 18C, 19F, and 23F. Within a few years of implementing the vaccine in 2000 in the USA, rates of penicillin resistance and of invasive disease, as well as otitis media due to vaccine types, were in decline and have remained low. In Massachusetts the replacement of vaccine serotypes with non-vaccine serotypes in colonization and infection had reached equilibrium by about 2007 [Reference Hanage7, Reference Wroe8]. Although it may not be possible to determine exactly when and how such changes occurred, we presume there were disturbances to the pneumococcal ecological niche due to vaccine-induced immunological changes within the population. Some of the replacement clones probably emerged by expansion from among less frequent non-vaccine serotypes already present. In a few instances vaccine serotype clones may have become ‘escape variants’ by switching their capsule genes to produce non-vaccine capsule types [Reference Hanage9].
Although PCV7 reduced carriage and infection of these isolates, resistance was widespread among many serotypes not in the vaccine. For example, a multi-resistant serotype 6A clone was already present and expanding in Israel before the use of PCV7, and it became a major replacement strain after the vaccine was implemented in 2006 [Reference Porat10]. Penicillin-sensitive isolates were not driven out by prevalent resistant clones: they co-existed and remained a reservoir of potential replacement isolates [Reference Wroe8, Reference Colijn11]. Serotype 6C, which was not described until 2007 [Reference Park12], was present in some areas, although unrecognized. In the PCV7 decade, serotype 6C has increased in prevalence throughout the USA [Reference Green13] and has essentially replaced other serogroup 6 isolates in invasive disease in Native Americans [Reference Millar14] and in the general population [Reference Carvalho15]. Included among the isolates from our older collections was the first serotype 6C strain to be identified by Park and colleagues in studies defining this new serotype [Reference Park12].
Little is known about the length of time individual clones persist in the human environment or about the selective factors that are likely to contribute to modifications of individual clones and their long-term stability. We examined our older collections of serogroup 6 isolates from the 1970s, at the beginning of the worldwide spread of penicillin resistance and over a decade before PCV7 vaccine. We used pulsed-field gel electrophoresis (PFGE) to look for genetic relationships within clonal groups. Multi-locus sequence typing (MLST) was used to further define these groups and trace genetic lineages based on ‘housekeeping’ genes that remain quite stable over time [Reference Feil16]. We compared the 1970s isolates with a collection from around the world in the decade of the 1990s and with a collection from the Centers for Disease Control and Prevention (CDC) covering the same period in the USA. Although our 1970s isolates were from a small geographical area, Birmingham, Alabama, we found many isolates that were related to those of importance in later decades.
METHODS
Bacterial isolates
We examined a total of 188 S. pneumoniae, all serogroup 6, representing isolates isolated in Alabama in the 1970s and from US and worldwide collections in the 1990s. Sixty-six were from collections in the Departments of Pediatrics and Microbiology at the University of Alabama at Birmingham, isolated between 1975 and 1980. These were all paediatric isolates, from a prospective epidemiological study of pneumococcal colonization and infection in children followed prospectively from birth [Reference Gray and Dillon17] and from clinical cultures at The Children's Hospital of Alabama [Reference Gray and Dillon18, Reference Gray and Dillon19]. The 1990s isolates were from two sources: 37 were from the Active Bacterial Core (ABC) surveillance programme of the CDC [Reference Gertz4, Reference Beall20] and 85 were from 33 different countries isolated between 1989 and 1998 selected to represent the seven major serogroup 6 clonal complexes (CCs) in Robinson et al. 2002 [Reference Mavroidi21, Reference Robinson22].
Lyophilized cells were rehydrated with 200 μl Todd–Hewitt broth (Difco, USA) with 0·5% yeast extract and isolated on blood agar. Single colony subcultures were screened for susceptibility to penicillin using 1 μg oxacillin disks (susceptible defined as zone diameter ⩾20 mm) [Reference Jette and Sinave23]. Penicillin sensitivity was further determined by the agar gradient method (E test, AB Biodisk, USA) with intermediate isolates defined by a minimal inhibitory concentration (MIC) between 0·06 and 2·0 μg/ml, according to the 2008 revised breakpoints for non-meningeal isolates [Reference Weinstein, Klugman and Jones24].
Serotyping
Capsular serotyping was performed by a latex bead-based flow cytometric immunoassay [Reference Park, Briles and Nahm25]. Cross-checking for serotypes 6A and 6B was done by inhibition enzyme-linked immunosorbent assay (ELISA), as described previously [Reference Sun, Hwang and Nahm26]. This test used two monoclonal antibodies: Hyp6BM8 recognizes both serotypes 6B and 6A, and Hyp6BM1 is specific for 6B alone. Pneumococcal lysates were prepared for inhibition assays, as described previously [Reference Hollingshead27]. The 50% inhibition at 405 nm wavelength optical density was calculated using DeltaSoft v. 2.22F microplate analysis software (BioMetallics, USA). Lysates capable of inhibiting over 50% of the reactivity with Hyp6BM1 were considered to be serotype 6B. Lysates showing no inhibition of Hyp6BM1 were further tested with Hyp6BM8, and those showing 50% inhibition of Hyp6BM8 were considered to be serotype 6A. One isolate, 2197, originally identified as 6A, was later found to be 6C. For selected isolates, the capsular serotype determined by inhibition ELISA was confirmed by sequencing of the wciP alleles, as described previously [Reference Mavroidi21].
Genotyping
DNA extraction [Reference Payne28], PFGE [Reference McEllistrem29], and MLST [Reference Gertz4, Reference Enright and Spratt30] were performed as described previously. Included in PFGE was a reference set of serogroup 6 isolates from previous studies [Reference Gertz4, Reference Robinson22] and penicillin-resistant isolates from the Pneumococcal Molecular Epidemiology Network (PMEN) [Reference McGee31]. All of the isolates were examined by PFGE. MLST was used to confirm clonal relationships of 23 isolates selected from the various PFGE patterns. The seven ‘housekeeping’ genes sequenced for MLST were those described in Enright & Spratt [Reference Enright and Spratt30] and used for the S. pneumoniae database at the website http://www.mlst.net. MLST sequence types (STs) were assigned after verification by the database curator.
Data analysis
Cluster analysis of PFGE was performed using BioNumerics (Applied Maths, USA). The BioNumerics software was used to make UPGMA (unweighted pair-group method with arithmetic means) dendrograms of fragment patterns with the Dice coefficient [Reference Gertz4]. For MLST, the allele number at each locus was assigned at the MLST website (http://www.mlst.net). eBURST analysis was used to assess genetic lineages and to predict putative ancestral STs (http://eburst.mlst.net) [Reference Feil16]. The term CC is used here to denote an eBURST group in which all isolates share five or more alleles with another isolate in the cluster. The putative ancestor or ‘founder’ of each CC is the strain with the greatest number of related isolates with STs differing at a single allele, termed a single locus variant [Reference Feil32]. The eBURST algorithm also combines CCs into larger ‘groups’ of more broadly related isolates. Two large groups are illustrated in Figure 2, and all groups are listed in Table 1. Persistent CCs are defined as isolates from different decades having the same ST and/or PFGE pattern and falling into the same eBURST grouping.
Table 1. Description of pneumococcal isolates
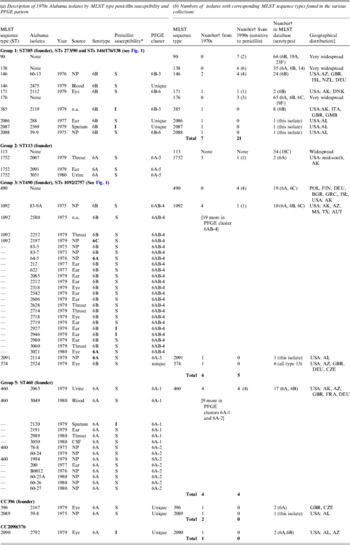
CSF, Cerebrospinal fluid; n.a., not available; NP, nasopharyngeal.
* S, sensitive; I, intermediate.
† Number of strains in the given category.
‡ Country abbreviations: AUT. Australia; BGR, Bulgaria; CZE, Czech Republic; DEU, Germany; DNK, Denmark; FIN, Finland; FRA, France; GBR, Great Brittan; GMB, The Gambia; GRC, Greece; ISL, Israel; POL, Poland. USA, states: AL, Alabama; AZ, Arizona; MS, Mississippi; TX, Texas.
RESULTS
PFGE and MLST
The 66 1970s isolates from Alabama fell into five PFGE clusters and 15 unique patterns, as shown in the dendrogram in Figure 1. Selected isolates of each PFGE cluster or pattern were found to represent 14 STs by MLST, including six new STs. For practical purposes, isolates of the same serotype and PFGE pattern were presumed to have the same ST, as listed in Table 1 a. As previously validated [Reference Gertz4], the agreement between serotype/PFGE and MLST was excellent.
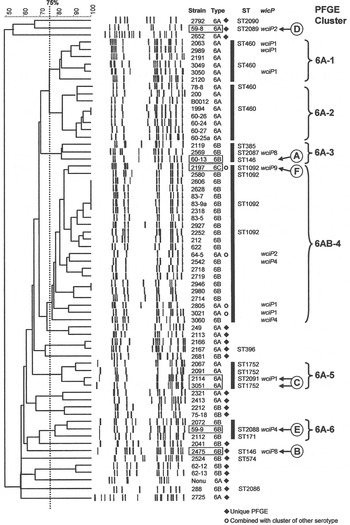
Fig. 1. Pulsed-field gel electrophoresis (PFGE) dendrogram of 1970s isolates listed by name and serotype. Clusters share a Dice coefficient >75% (dotted line). Sequence types (STs) and wciP allele number results are given.
All of the 1970s isolates are listed in the Table 1 a, according to their relationship to the larger eBURST groups described below. Table 1 b shows the number of isolates by ST in the collections from the 1970s and 1990s and in the entire MLST database, as of 2012. The seven groups and CCs included in the table are among the 10 most common eBURST groups that are comprised primarily of serogroup 6 isolates in the MLST database. Other groups are seldom seen in the USA. Supplementary Table S1 lists all 109 isolates typed by MLST, including 23 of the 1970s Alabama isolates and 86 of the 1990s isolates from the ABC surveillance and ‘worldwide’ collections, which were MLST-typed in previous studies [Reference Gertz4, Reference Mavroidi21].
eBURST analysis
An eBURST analysis [Reference Feil33, Reference Spratt, Hanage, Brueggemann, Tuomanen, Mitchell, Morrison and Spratt34] was performed using the 109 isolates typed by MLST run as a set and also together with all serogroup 6 STs in the S. pneumoniae MLST database (www.spneumoniae.mlst.net), as of February 2012. There were 32 eBURST CCs, 12 of which had four or more STs and 20 of which included only two or three STs. There were also 76 singletons, which shared fewer than five alleles with any other isolate. This confirms that a broad diversity of genetic backgrounds was represented among the serogroup 6 isolates. The six largest eBURST groups had the same the CCs as were derived by a different MLST scheme in an earlier study [Reference Robinson22].
Group 1 is the largest of the major eBURST groups that contained 1970s isolates (Fig. 2 a). This group included many other serotypes, as illustrated in the shaded areas of the inset. Serogroup 6 isolates connect at CC385 and radiate in two directions. Our group 1 isolates from the 1970s, shown with STs enclosed in rectangular boxes in Figure 2 a, were seen in CC385 and ranged almost through to CC138. None radiated in the other direction towards CC90. Only two of five 1970s STs in group 1 were still present in the 1990s (Table 1 b). It would be surprising to find all of the STs present in both time periods due to the rate of evolution. Moreover, the number of isolates included here is relatively small and might not be sufficiently sensitive to detect the sum of pneumococcal diversity. These CCs, along with CC176, are the largest groups worldwide, and in the USA they continued to account for 6–11% of the invasive type 6B isolates collected from children aged <5 years during 1999 and 2001 [Reference Gertz4, Reference Beall20].
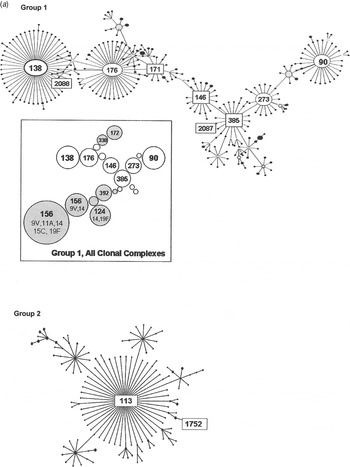
Fig. 2a. Serogroup 6 clonal complexes (CCs) in groups 1 and 2. eBURST analysis of serogroup 6 strains. The set included 23 isolates from the 1970s, 44 from the global collection, 42 ‘control’ strains from the USA, and all serogroup 6 isolates in the MLST database. Numbers in rectangular boxes denote CCs represented in our 1970s collection; other CCs are enclosed by ellipses. The inset shows all CCs in group 1, with CCs dominated by other serotypes in shaded ovals.
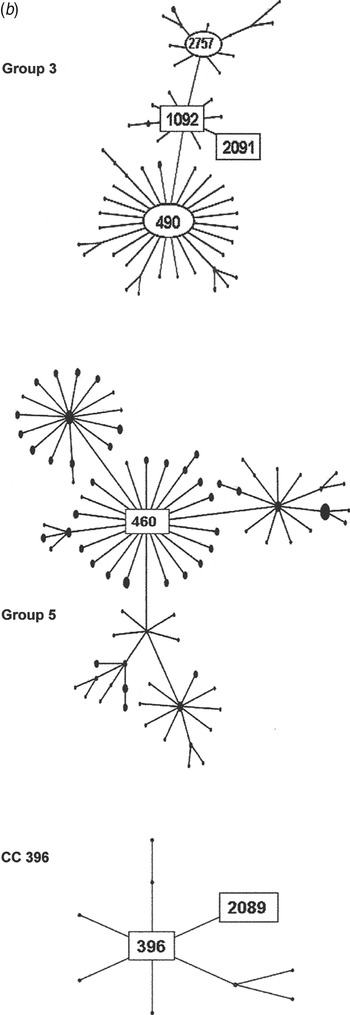
Fig. 2b. Serogroup 6 clonal complexes (CCs) in groups 3, 5, and CC396. eBURST analysis of serogroup 6 strains. The set included 23 isolates from the 1970s, 44 from the global collection, 42 ‘control’ strains from the USA, and all serogroup 6 isolates in the MLST database. Numbers in rectangular boxes denote CCs represented in our 1970s collection; other CCs are enclosed by ellipses.
Group 2 is a large group centred on founder ST113, which includes 34 mostly serotype 18C isolates. Our ST1752 isolates were located on the periphery of this group, sandwiched in between serotype 19F (ST3890) and another serotype 6A (ST2786) as their nearest neighbours. ST1752 has been found only in the US mid-south and Alaska.
The group 3 isolates from the 1970s were mostly ST1092 and comprised our largest PFGE cluster (6AB-4). ST1092 formed a major node related to the large CC490, which contains a mix of serotypes 6A, 6B and 6C. Of particular interest was our strain 2197, which was isolated in 1979. This strain prompted the search for a new serotype and was the first serotype 6C to be identified [Reference Park35]. The ST2091 a single locus variant of 1092 was also isolated in 1979 as was serotype 6A, whereas most ST1092 were serotype 6B (Fig. 2b ).
Group 5 was founded by ST460, the centre of a large CC consisting mainly of serotype 6A surrounded by CCs of serotypes 10A, 23F, and 35F. All of our 1970 isolates were serotype 6A, ST460, and were represented by two PFGE clusters (6A-1 and 6A2). ST460 isolates have persisted over the decades. This and related STs accounted for 22–29% of invasive type 6A isolates collected from children aged <5 years in the USA during 1999–2002, although the number of type 6A isolates from this population showed a marked decline during 2001 and 2002 [Reference Gertz4, Reference Beall20].
Serotype changes within CCs
The capsular serotype was confirmed by sequencing of the wciP alleles for group 14 isolates, which included serotypes 6A, 6B, and 6C (mainly ST1092 in PFGE cluster 6AB-4). When serotypes change within a CC, it may be due to recombinational replacements of wciP or a number of capsule genes or to the large capsule region, without affecting the ST. All WciP proteins in our serotype 6A isolates and in the single serotype 6C had an alanine at amino acid 135, whereas those in serotype 6B had a serine at this position, as we have previously observed [Reference Mavroidi21].
Penicillin susceptibility
The 1970s collection was dominated by penicillin-susceptible isolates. Penicillin-intermediate isolates did not emerge among serious infections in Alabama until 1981 [Reference Willett, Dillon and Gray1]. Eight of the 1970s isolates (from middle ear, eye, and asymptomatic colonization) harboured intermediate penicillin resistance. Penicillin non-susceptibility was probably already widespread by the late 1970s, since penicillin-intermediate isolates were found in four STs and distributed in three of the most prevalent groups and CCs (ST385, ST460, ST1092 ST2090).
DISCUSSION
Our collections of serotype 6 pneumococci provided us with a way of going back in time prior to PCV7 introduction and before widespread penicillin resistance to look for lineages that were common in later surveys. It is an obvious limitation that we did not have a later set of isolates from our own geographical area for comparison. Nevertheless, we observed that nearly half of the STs found in the 1970s continued to colonize and cause infection over the ensuing decades. Given that our isolates were from a relatively small population, it is remarkable that we were able to identify five of the most abundant clonal lineages seen in later years in the USA and 40 other countries.
Our largest PFGE cluster fell into group 2, consisting mainly of ST1092 but not the founder, ST490, which was more prominent in Europe. The first serotype 6C to be identified was among the ST1092 isolates, along with two of our earliest penicillin-intermediate isolates. ST1092 does not appear to have expanded significantly, although it has shown up in the 10 isolates in the MLST database from elsewhere in the USA, in Native American populations, and Australia. ST1092 includes examples of penicillin-sensitive and -resistant serotype 6A, 6B, and 6C isolates, suggesting that it provides a durable genetic background for strain survival and diversity [Reference Jacobs36–Reference Zhuo38].
Our next largest group was made up of ST460 isolates in two PFGE clusters, 6A-1 and 6A-2. ST460 is the founder of group 5, which has persisted through the 1990s [Reference Beall20] and at least to 2007 (see MLST database at http://spneumoniae.mlst.net/). It is found widely in the USA and Europe, but as with our 1970s isolates, there are only a few examples of penicillin resistance in later isolates in the MLST database. Group 1 isolates were more diverse, with six STs mainly related to the founder, ST385. Few isolates were seen at the outer reaches of the eBURST map towards CC138 or CC90, which became more prominent in the 1990s, especially after the influx of international clones such as Spain6B-2 (ST90).
By focusing on serogroup 6 we were able to observe the persistence of some pneumococcal clones over time, despite the net-like evolution that is predicted due to the high rates of horizontal gene transfer, serotype switching, and acquisition of antibiotic resistance [Reference Feil33, Reference Spratt, Hanage, Brueggemann, Tuomanen, Mitchell, Morrison and Spratt34, Reference Spratt and Maiden39]. MLST was especially helpful in assessing the genotypic relationship of isolates not epidemiologically linked because of separation in time and geography. While it is quite possible that serogroup 6 clones and CCs are not typical of all S. pneumoniae, limiting the study to the one serogroup kept the number of lineages small enough to allow detection of related clones or CCs. Some of the implications with regard to diversity and persistence may extend to other pneumococcal serogroups, such as 14, 19, and 23, but perhaps not to serotypes/groups, such as 1, 3, 4, and 7, which are represented by very few STs. The ‘childhood’ types are very commonly carried and therefore exposed to more and varied evolutionary stresses, especially antibiotics and vaccines. Paediatric serotypes types have increased in prevalence in older adults for various reasons including the selective pressure of antibiotics [Reference Feikin40].
All but six of the 1970s isolates were susceptible to penicillin. There was no clear case for penicillin resistance contributing to the persistence of any ST in these isolates. We did not see any ST376 isolates in the 1970s, but one strain was a single locus variant (ST2090) that was penicillin intermediate. In the 1990s collections we found six ST376, all of which were penicillin intermediate or penicillin resistant (Table 1 b). It can be speculated that ST376 and its relatives might have some predisposition to genetic exchange or acquisition of resistance factors in general.
eBURST groups that are widespread enough to be detected on different continents are also found to contain isolates of either serotypes 6A or 6B, indicating either horizontal gene exchange or mutations in the capsule locus [Reference Gertz4]. In our 1970s and 1990s collections we found that such instances were best explained by the horizontal spread of a portion of the cps locus, as previously suggested [Reference Mavroidi21]. It appears that conversion between serotypes 6A or 6C and 6B can occur freely in nature, and serotype 6C isolates are represented by diverse backgrounds as judged by their many different STs [Reference Green13, Reference Jacobs36, Reference Zhuo38]. The PCV7 vaccine contains type 6B but confers some cross-protection against 6A, which is not in PCV7. The prevalence of 6C was probably low prior to the vaccines, as suggested by our finding of only one isolate in our 1970s collection. Serotype 6C increased after PCV7 came into use [Reference Nahm41] and has gone on to replace 6A and 6B in some populations [Reference Millar14, Reference Carvalho15]. The inclusion of both serotypes 6A and 6B in PCV13 vaccine may further this effect. However, PCV13 vaccine appears to confer cross-functional antibody against serotype 6C [Reference Cooper42] and to protect against serotype 6C colonization [Reference Cohen43]. It remains to be seen if the prevalence of 6C declines as PCV13 comes into more widespread use.
Pneumococci evolve relatively rapidly in response to environmental changes, as has been noted, responding to antibiotic pressures and to new vaccines with such strategies as acquisition of antibiotic resistance and serotype switching [Reference Hanage9, Reference Croucher44]. Exactly how forces promote the selection and expansion remains the subject of investigation and debate. We have also seen the emergence of replacement serotypes, such as 6C and 19A. In some instances this might represent the expansion of lineages already in the community found at relatively low frequencies in asymptomatic carriage and even less frequently in disease. Other instances suggest the direct importation of a ‘foreign’ strain, such as the Spain6B-2 clone, which was imported to Iceland in 1989. This international clone became dominant in Iceland for several years, but over time it lost its resistance or died out in areas where the use of antibiotics was reduced [Reference Vilhelmsson, Tomasz and Kristinsson45].
In conclusion, we found ancestors of some of the clones that came into play around the time of the introduction of the PCV7 vaccine, showing that CCs present in the 1970s were detected globally in the 1990s and beyond. This finding suggests that successful lineages are capable of both persistence and transmissibility, resulting in their widespread distribution. At the time this study was conducted there were few isolates from the 1970s and too few to discern important evolutionary relationships. When Danish investigators re-examined both recent and archived 6A isolates from Denmark, they found three of eight older isolates dating from 1962, 1968, and 1987 that proved to be 6C with various unrelated STs [Reference Lambertsen and Kerrn46]. As more isolates from earlier periods are added to the MLST database, we expect to uncover additional insights into the evolution of the resourceful pneumococcus.
SUPPLEMENTARY MATERIAL
For supplementary material accompanying this paper visit http://dx.doi.org/10.1017/S0950268814000508.
ACKNOWLEDGEMENTS
This work was supported by NIH/NIAID grant R01 AI053749 (Hollingshead) and a subcontract from the Alliance for the Prudent Use of Antibiotics (APUA) under NIH/NIAID grant 5 U24 AI050139-03 (Levy/Feldgarden). The UAB Genomics Core Facility was funded in part by a grant from the Health Services Foundation of the University of Alabama at Birmingham. We acknowledge the use of the pneumococcal MLST database which is located at Imperial College London and is funded by the Wellcome Trust.
We thank Susan Hollingshead for her mentorship in developing the study plan, and we are grateful to D. Angeliki Mavroidi, whose prior work had suggested the possibility for this study. Pat Coan provided assistance in PCR, ELISA, and susceptibility testing; and Moon Nahm provided antibodies used in serotyping. We also thank Ken Waites and Lynn Duffy for advice and skilled help with resistance typing of the isolates. We thank Bernard Beall and Bob Gertz for providing the CDC isolates and for assistance with PFGE.
DECLARATION OF INTEREST
None.






