Impact Statement
Artificial intelligence (AI) is transforming research across natural and engineering sciences, often in the form of field-specific tools. We argue that by developing AI techniques for supporting the scientific discovery process itself we can take the next step and enable efficient human-AI collaboration for seeking new knowledge. In many cases, the AI methods can be applied across several scientific disciplines, but to encourage researchers to work on general methods we need environments that make it easy and attractive. We introduce the concept of virtual laboratories to facilitate research on domain-agnostic AI assistance for research and development and to make it easy for other scientific disciplines to use these tools.
1. Introduction
Merriam-Webster defines a laboratory as “a place equipped for experimental study in a science or for testing and analysis” or more broadly as “a place providing opportunity for experimentation, observation, or practice in a field of study.” Footnote 1 The definition refers to a physical environment that exists for the purpose of making new discoveries. While laboratory tasks are now frequently carried out on computers, or more and more automated synthesis and measurement devices, the laboratory itself remains surprisingly similar to its 19th century form. In our increasingly digital world, we think it is time for a paradigm shift to virtual laboratories (VLs), following a loose conceptual analogy to virtual machines in computer systems. Continuing with the analogies, a VL is then a digital environment that enables users to carry out research tasks and that encompasses the elements of a classical laboratory as its components.
The starting point for a virtual laboratory is the computational methods and tools that are already an integral part of modern scientific practices. These include computational simulations, digital twins of various instruments, robotic measurement devices, and methods for experimental design, data analysis, and statistical estimation. In most scientific disciplines, physical laboratories already heavily use these computational tools, and research combines computation and real-world experiments. The new digital technologies already provide scale advantages and improve reproducibility and reliability.
In this point of view, we argue, however, that the current tools are not yet sufficient for building virtual laboratories, and two aspects need to be addressed. First, the current toolkit needs to be expanded. The current models and tools are typically field-specific and designed to address specific narrowly defined tasks. What is missing today are tools that would help more efficiently, as well as serve the scientific research process itself, by combining the isolated field-specific models to form a laboratory environment that supports researchers in using the digital tools to address the needs of increasing workflow, tool, and research complexity.
The second step required for reaching the full potential of virtual laboratories is to consider what can be done differently now that the computational backbone exists in different disciplines. Could we develop new types of tools (Figure 1) by thinking across laboratories, and in particular, could we benefit from advances in AI methodology? If the tools were not developed independently in each field but would instead pool the creativity, ingenuity, and resources from a variety of fields, progress would be faster and VLs could become a reality sooner. Such general applicability and acceleration is precisely the promise AI-based tools offer.
AI methods enable generalizing across field-specific virtual laboratories, each using a mixture of field-specific and general methods.
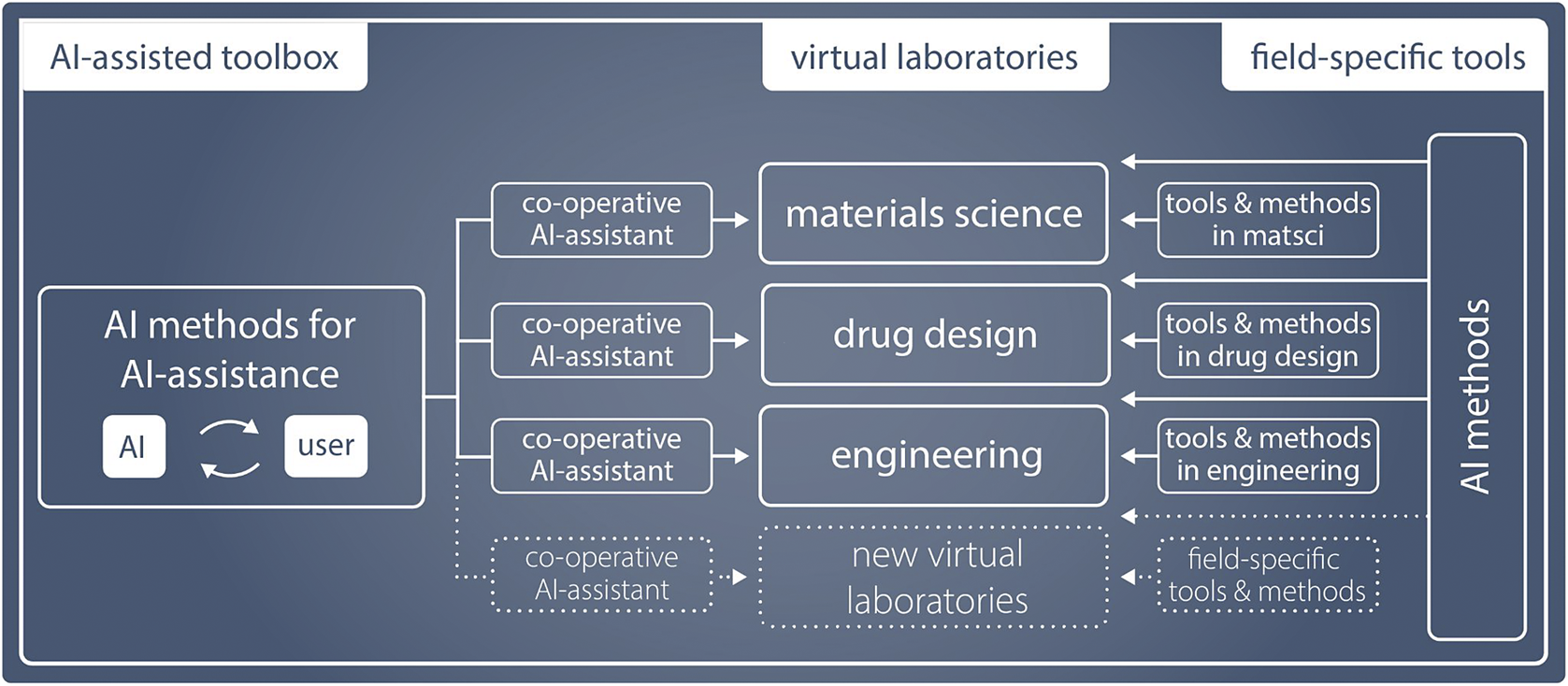
In this paper, we present a vision for AI-assisted virtual laboratories: Digitalization of research and development will move from isolated digital twins to AI-assisted support of the scientific innovation process. In the future, new innovations are made in virtual laboratories, where researchers seamlessly operate with physical and virtual measurements in close collaboration with AI, accelerating the pace and improving the quality of research. The researchers would remain in charge and would continue to develop their research competences, but now assisted by personalized AIs that continuously learn to provide context-dependent support. The virtual laboratories are ideally supported by a common software library that combines field-specific digital twins with domain-agnostic tools for formalizing and assisting the research process.
Virtual laboratories provide a conceptual frame of reference, and in this paper, we outline practical directions for the transition from real to virtual laboratories. This paper is a call for both AI researchers and domain scientists to join forces. Section 2 introduces the main concept of virtual laboratory and outlines the high-level goals and challenges. In Section 3, we present the main actions we think should be taken by different parties. Lastly, Section 4 takes a close look at the current status, by discussing how soon we expect various disciplines to benefit from the efforts and by reviewing the state of emerging VLs in three different fields, outlining for instance how drug design is already largely done in a virtual realm but using field-specific tools.
2. Virtual laboratory concept
2.1. Virtual laboratory
Following our laboratory definition in the introduction, a virtual laboratory (VL) is the in silico equivalent of a physical laboratory. A VL exists primarily in a virtual space, or at least mediates the interaction of stakeholders with the VL remotely through a digital user interface. In practical terms, a VL is a collection of interconnected digital twins and a digital user interface (see Figure 2), operated by a scientist that remains in control but is assisted by AI. In our opinion, AI assistance is a critical VL element, because it facilitates the navigation of the complex VL environment and enhances the research process.
In a virtual laboratory researchers or users perform virtual experiments using digital twins of assets or processes, where the experiments generally refer to any manipulation of the digital twin that may involve also other functions of the rest of the laboratory. They are assisted by AIs, which use digital twins of the researchers (user models) for interactive assistance. The process is grounded in the real world, through use of the physical instruments or assets (called for on-demand by the AI assistants).
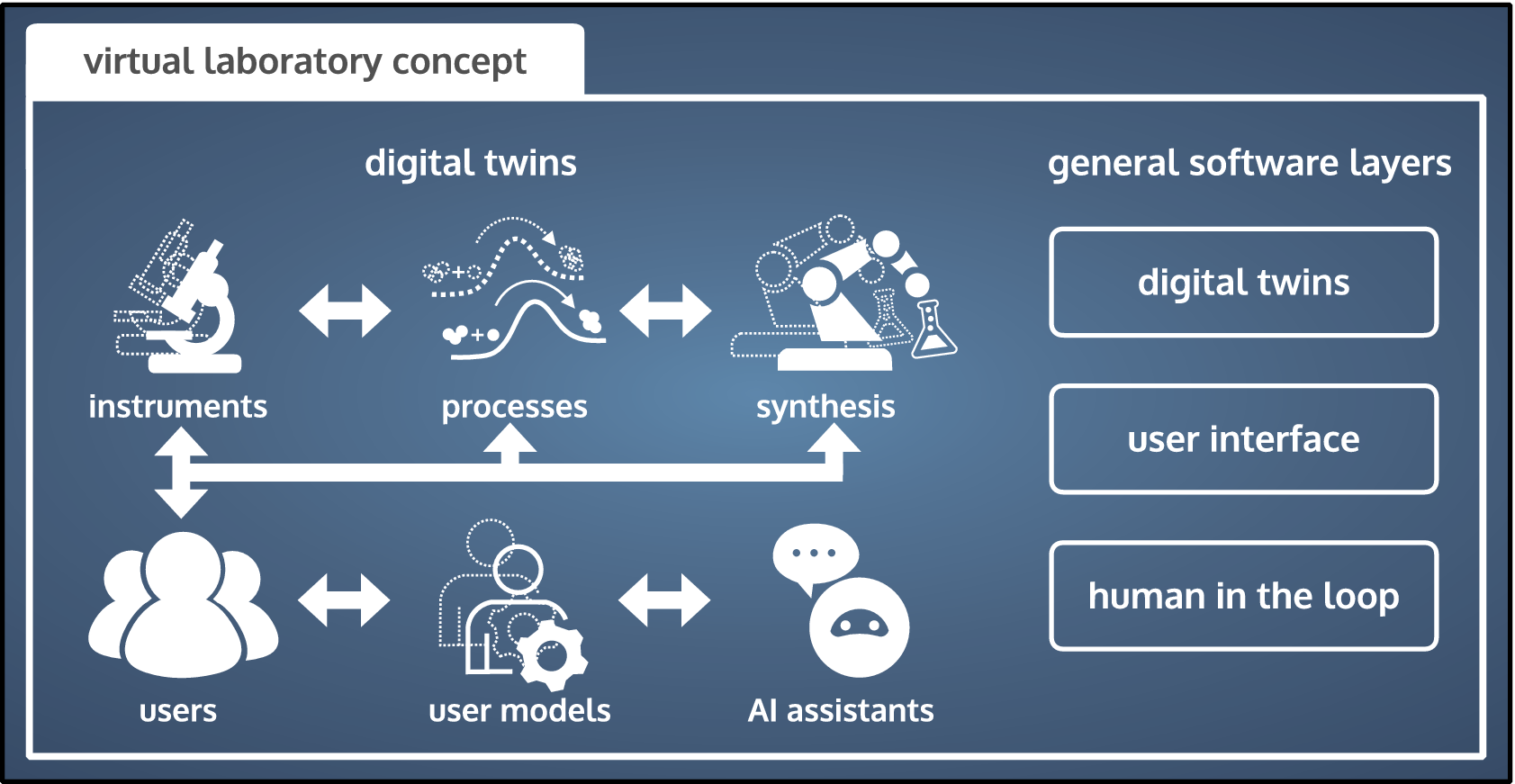
Digital twins are faithful computational representations of real-world entities or processes (Wright and Davidson, Reference Wright and Davidson2020; Blair, Reference Blair2021; Niederer et al., Reference Niederer, Sacks, Girolami and Willcox2021). We here consider a wider definition of digital twins than usual and distinguish between three types: (a) assets, (b) processes, and (c) human interactions. In (a), physical assets is an umbrella term for scientific instruments, measuring devices, and equipment that manufacture goods, fabricate materials, and synthesize substances. In (b), computational models and simulators aim to capture physical or chemical processes. In (c), we refer to user models of human behavior and human-machine interactions. Combined, these three types of digital twins transfer real-world data into the virtual realm, where it is processed by simulators and AI methods.
As implied by the word virtual, one purpose of VLs is to transfer the experimentation and discovery process from the real into the virtual realm. In this virtual mode, users interact with the digital twins instead of their real-world manifestations to derive new knowledge, educate themselves, or receive assistance in complex decision-making. This usually offers significant time and resource savings compared to directly operating in a physical laboratory. The digital twins interact with the real world, when necessary, to stay up-to-date and react to changing conditions. In the real mode, the VL has a direct physical outcome, for example, a material or drug. The VL facilitates, accelerates, or even enables the design and development of the physical outcome.
While digital twins for assets and processes are still largely specific to a field, or even a laboratory, user models and human-machine interactions are already somewhat task-agnostic and more widely deployable. We argue that a field-agnostic virtual laboratory concept should start from these user-related components and develop generalized tools that can be quickly deployed across fields. For example, the chemical models in digital twins for drug design differ from those in construction engineering, but the cognitive models of human sequential decision-making and human biases, employed by collaborative AI assistants, are the same. These models will be useful already in cases where some of the components are not yet digitalized but involve physical measurements or manual work, often resulting in time or cost savings. Similarly, much of the backend machinery (e.g., data passing, digital twin maintenance) is general-purpose. A VL helps in delineating the field-specific components from those that generalize across fields, and thus allows efficient and parallel tool development.
2.2. Elements of digital twins
Although each digital twin serves a specific purpose, current realizations of digital twins share several aspects: (a) live coupling between the physical asset and its digital twin, via multiple streaming data sources originating from live sensing of the physical process, (b) access to additional information about the modeled process, such as geometry, topology, physical laws or 3-D characteristics for physical assets, (c) AI models utilizing the aforementioned data sources and prior knowledge to accurately predict or simulate future states of the physical twin from these, (d) some ability to perform what-if scenarios and counterfactual reasoning over the process, and (e) a decision-making mechanism (typically with a human-in-the-loop) for acting on the asset/process given the model and any what-if reasoning abilities therein.
A digital twin can also be composed of several sub-twins. These sub-twins could be in different physical locations, for example, different real-world laboratories. The VL would integrate (or aggregate) all these sub-twins into one digital twin as illustrated in Figure 3. We are now moving to the realm of decentralized/distributed inference and borrowing statistical and causal strength across experiments. Methodological frameworks such as multioutput/multitask learning (Caruana, Reference Caruana1997), transportability and data fusion (Pearl and Bareinboim, Reference Pearl and Bareinboim2014; Bareinboim and Pearl, Reference Bareinboim and Pearl2016), federated learning (Yang et al., Reference Yang, Liu, Cheng, Kang, Chen and Yu2019), physics-informed ML (Mao et al., Reference Mao, Jagtap and Karniadakis2020), and semi-parametric statistics bridging to traditional numerical methods (Girolami et al., Reference Girolami, Febrianto, Yin and Cirak2021) are central for such Causal DTs and interconnected VLs.
Schematic of digital twins depicting the key information flow and quantities of interest. Several DTs could be aggregated in a VL. Each of the instances could be different instruments combined into one digital twin or different realizations of a device in different labs around the world.
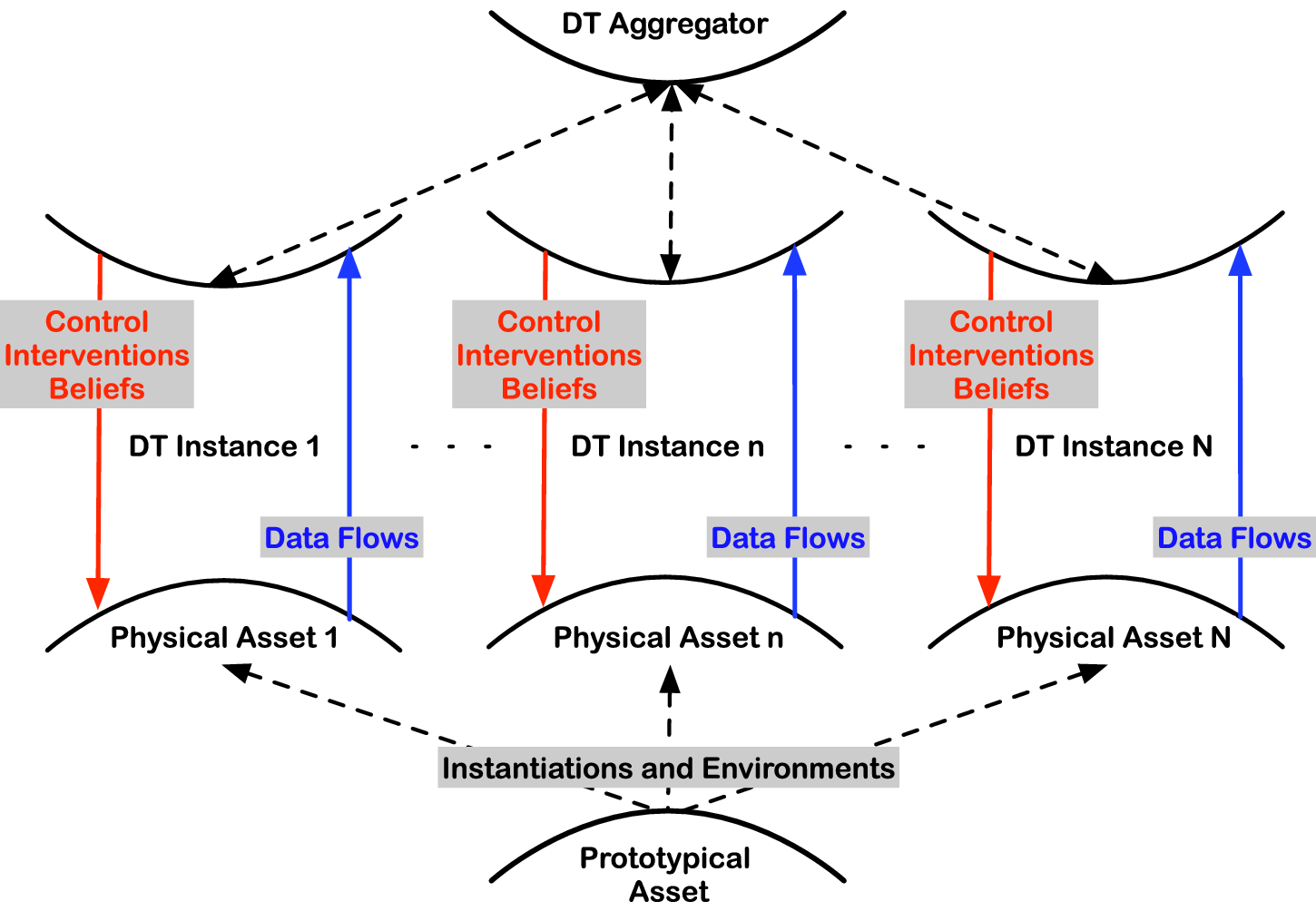
These interconnections are graphically depicted in Figure 3 using the gemini symbol (♊) as a playful abstraction of a digital “twin.” On the bottom hull, the true data-generating process of the real-world component is generating data that is noisily sensed via multiple sensor networks with potentially different characteristics. These are streamed upwards to the digital twin for model-based inference. The model is by construction an approximation of reality, but potential error or uncertainties can be characterized and accounted for, for example, Kleijn and Van der Vaart (Reference Kleijn and Van der Vaart2012) and Dellaporta et al. (Reference Dellaporta, Knoblauch, Damoulas and Briol2022). In turn, the posterior beliefs of the digital twin over the model and/or the model parameters are utilized to compute expectations of functions of interest with respect to these posteriors, and subsequent decision-making or simulation of what-if scenarios is performed leading to actionable interventions back on the physical asset.
2.3. A virtuous cycle of virtual laboratory development
A virtuous cycle is a recurring chain of events with a positive outcome. If we can start such a virtuous cycle in VL development, in which advances in AI and domain-specific knowledge benefit each other, we can increase the pace, quality, and cost-efficiency of scientific research. In this cycle, the VLs are the catalysts for facilitating the interaction and the research environment.
Once kick-started, the virtuous cycle will produce the first success stories. These will trigger an increased interest from both AI researchers and VL users, which, as the cycle progresses, should then result in a self-sustaining community effort.
2.4. Virtual laboratory library: a common software ecosystem
We argue that a key requirement for a successful, virtuous cycle is the increased use of common software components across fields. Such a necessary software ecosystem should be built so that AI advances can be developed independently in a modular fashion and taken immediately into use across many VLs with minimal effort, while making it easy for VLs to include their field-specific digital twins for assets and processes. A collection of modules and tools forms a virtual laboratory library (VLL), and open interfaces allow anyone to either contribute new modules or re-use the existing modules to facilitate concrete VLs in specific fields. The software ecosystem provides the technical realization of the VLs to produce scientific and societal impact. We believe that the development of VLLs is necessary not only for the permeation of VLs across fields, but also for incentivizing VL developers, VL users, and AI researchers to join forces.
3. Toward real virtual laboratories
In this section, we outline the main steps we consider necessary, from a technical perspective, to build VLs and to generalize the concept across disciplines.
3.1. Virtualization
Transferring key components of the scientific method, such as hypothesis generation, experimentation, confirmation, and discovery, from the physical to the virtual setting is the central objective of VLs that enables acceleration, reproducibility, and scalability of research. The primary vehicles for the virtualization of such components are digital twins of assets, processes, and human interactions that are interlinked inside the virtual lab and attempt to address the gap between simulation and real world through their live coupling to the physical twin and emerging frameworks for mispecification, calibration, and interventional consistency, for example, Kennedy and O’Hagan (Reference Kennedy and O’Hagan2001) and Dellaporta et al. (Reference Dellaporta, Knoblauch, Damoulas and Briol2022). Significant resources are already being dedicated to improving the quality and versatility of digital twins as components of the VL and the transition will directly benefit from the results of these efforts, but dedicated research will be needed for virtualization of the research process and the human elements of that.
Many key AI technologies and research areas are necessary for the virtualization process itself: from simulators, emulators, artificial agents, and their data calibration and optimization, to reinforcement learning and robotics for automated measurement devices. Some of these, such as robotics, target the automation of physical measurements while other areas are necessary for exploring and optimizing virtual measurements and for counterfactual reasoning. We note that a large body of AI research including experimental design, Bayesian optimization, reinforcement learning, language models, generative models, synthetic data, causal inference, domain adaptation, transfer learning, probabilistic modeling, probabilistic numerics, uncertainty quantification, and physics-informed ML (
![]() $ \Phi $
-ML) will be central in enabling full virtualization. Any methodological advances will then directly benefit VLs.
$ \Phi $
-ML) will be central in enabling full virtualization. Any methodological advances will then directly benefit VLs.
3.2. Human in the loop
AI tools are predominantly used to automate tasks and supplement or replace human-derived insight with data-driven models. The evolution toward “robot scientists” (Burger et al., Reference Burger, Maffettone, Gusev, Aitchison, Bai, Wang, Li, Alston, Li, Clowes, Rankin, Harris, Sprick and Cooper2020) has been invoked, but in reality, human scientists remain involved, in two ways. They drive the scientific process, by instantiating, designing, and applying AI methods, and they provide knowledge. Through human-in-the-loop machine learning (Monarch, Reference Monarch2021), prior human knowledge could be directly integrated into VLs. Human-in-the-loop methods elicit knowledge from human users to maximally improve AI models with minimal user effort.
Current human-in-the-loop methods are not compatible with the other reason humans are involved—that they drive the research process. The current methods treat humans as passive data sources (e.g., in knowledge elicitation; Mikkola et al. Reference Mikkola, Martin, Chandramouli, Hartmann, Pla, Thomas, Pesonen, Corander, Vehtari, Kaski, Bürkner and Klami2023) instead of active agents. For VLs, we need to develop human-centric AIs and human-AI collaborations (Dafoe et al., Reference Dafoe, Bachrach, Hadfield, Horvitz, Larson and Graepel2021). Multi-agent modeling methods from human-robot interaction (Hadfield-Menell et al., Reference Hadfield-Menell, Russell, Abbeel and Dragan2016) are a start, but work is still needed for formulating assistants which are useful to human scientists while leaving them in full control (Celikok et al., Reference Celikok, Oliehoek and Kaski2022). For this, the assistants will need to infer their user’s intentions, objectives, and goals and then recommend actions in a way they understand—in other words, they would need models of human users to efficiently collaborate with them. In Section 2, we referred to these models as user models or digital twins for human-machine interactions. With user models of scientists, AI assistants will be able to anticipate their actions and aid them in the scientific discovery process.
While dedicated interfaces, for example, for analyzing and interpreting scientific measurements remain the main form of interaction between humans and AIs, recent advances in large language models (LLM) and their dialogue capabilities have made it possible to consider also natural language as the interface. How LLMs can assist with the analysis of scientific literature and scientific writing remain hot topics, but some studies already go further by incorporating LLMs into the scientific discovery process. For instance, discipline-specific LLMs (Luo et al., Reference Luo, Sun, Xia, Qin, Zhang, Poon and Liu2022; Korolev and Protsenko, Reference Korolev and Protsenko2023) can, for example, find relationships between molecules or predict material properties based on written descriptions, blurring the line between a model and communication interface. Research on how to best use these tools for scientific discovery is only starting now (Stokel-Walker and Van Noorden, Reference Stokel-Walker and Van Noorden2023; Van Dis et al., Reference Van Dis, Bollen, Zuidema, van Rooij and Bockting2023), and fundamental questions on their capabilities and limits in supporting complex and long-term tasks remain. VLs are ideal platforms for studying these questions, by providing data on long-term structured interaction between teams of highly skilled human experts and dialogue systems.
In summary, VLs will be a fitting environment for mixed human-AI research teams. Before fully-fledged AI assistants become available, AI tools that give better recommendations would already be beneficial. To reach this point, advances in both AI and human-computer interaction are required.
3.3. Software layer
Building a common software layer for VLs will be critical. We only benefit from up-scaling if multiple VLs can re-use the same underlying software modules, so that AI researchers can easily develop and evaluate their methods for multiple use cases and VL hosts can easily integrate new AI elements.
The VL software layer mediates the scientific process in the virtual realm and provides the link to the physical realm, but should not be specific to any particular laboratory type. The software layer needs to represent digital twins, moderate data flows between digital twins (as well as their physical counterparts), and enable human-AI collaboration. This requires a modular architecture that communicates with domain-specific databases and models, so that all elements of AI assistance and DT operation are provided as independent modules. Common interfaces will be critical, so that both open-source and commercial tools for operating VLs will efficiently re-use the components.
We are not aware of any general VL software development even though many libraries for the individual VL components and the automation of data analysis workflows (Singh, Reference Singh2019; Mölder et al., Reference Mölder, Jablonski, Letcher, Hall, Tomkins-Tinch, Sochat, Forster, Lee, Twardziok, Kanitz, Wilm, Holtgrewe, Rahmann, Nahnsen and Köster2021) already exist (see also Section 4.4). Besides a modular architecture, an emerging software layer should:
-
• allow re-use existing (or future) libraries and help avoid the need to replicate general-purpose algorithms (e.g., machine learning tools or language models).
-
• run on standard cloud architectures and databases.
-
• be free and open-source, but licensed to enable commercial support for broad use in industry and research.
-
• be designed from the start as shared community effort, and eventually establish new standards for information transfer between digital twins and laboratories.
-
• actively support the FAIR (Findable, Accessible, Interoperable, and Reusable) data principles (Wilkinson et al., Reference Wilkinson, Dumontier, Aalbersberg, Appleton, Axton, Baak, Blomberg, Boiten, da Silva Santos, Bourne, Bouwman, Brookes, Clark, Crosas, Dillo, Dumon, Edmunds, Evelo, Finkers, Gonzalez-Beltran, Gray, Groth, Goble, Grethe, Heringa, ’t Hoen, Hooft, Kuhn, Kok, Kok, Lusher, Martone, Mons, Packer, Persson, Rocca-Serra, Roos, van Schaik, Sansone, Schultes, Sengstag, Slater, Strawn, Swertz, Thompson, van der Lei, van Mulligen, Velterop, Waagmeester, Wittenburg, Wolstencroft, Zhao and Mons2016), required for community development and reliable operation.
-
• support compartmentalization of public and private data and models, so that sensitive data and proprietary simulators and digital twins can be excluded from externalized AI development.
-
• make it easy to run virtual instances with no direct connection to the physical laboratory, for external AI research.
3.4. Enabling and encouraging VL research
VLs generate added value from the synergy between AI and research in other domains. To create this synergy, the barrier for contributions from AI researchers and VL domain scientists should be lowered. The common software layer is necessary but not yet sufficient for this. Numerous examples demonstrate a clear benefit from lowering the contribution barrier: ImageNet data (Deng et al., Reference Deng, Dong, Socher, Li, Li and Fei-Fei2009) revolutionized computer vision and MuJoCo (Todorov et al., Reference Todorov, Erez and Tassa2012) and OpenAI gym (Brockman et al., Reference Brockman, Cheung, Pettersson, Schneider, Schulman, Tang and Zaremba2016) reinforcement learning. We need similar success stories for VLs.
A key difference between VLs and the above examples is that VLs are linked also with physical reality and many of the interesting research questions involve humans, as explained in Section 3.2. This introduces additional challenges but we have not identified any immediate show-stoppers that could not be overcome by combining different approaches. Many AI elements can be developed in purely digital laboratories, using simulated human activity if needed. For example, ChemGymRL (ChemGymRL: Chemistry Laboratory Environment)Footnote 2 offers a reinforcement learning environment for a purely virtualized chemical laboratory, Trubucco et al. provide a virtual environment for design problems (Trabucco et al., Reference Trabucco, Kumar, Geng and Levine2021), and some cognitive models of researchers can be trained with non-experts in crowd-sourcing experiments, for instance, models of working memory and decision-making (Brown et al., Reference Brown, Zeidman, Smittenaar, Adams, McNab, Rutledge and Dolan2014).
We believe the most important step toward realizing our vision will be the activation of the research community. Providing computational platforms, theoretical concepts, and individual AI modules is a community effort, both in terms of sufficient resourcing, but also to ensure open standards and broad applicability across different fields. This transformation is best driven by a global open initiative, and we will work toward establishing one. The initiative will bring AI researchers and domain scientists together to design and develop the software platform, to determine incentive structures and funding models for VL hosts, for example, by extending the practices currently in place for data releases, and to work toward key standards. It will also have an important role in increasing awareness of the concept, via a workshop series organized alongside one of the leading AI conferences and challenges designed for steering the efforts of AI researchers, motivated by, for example, the effect the Netflix prize had on recommendation engine research (Bennett et al., Reference Bennett, Lanning and Netflix2007). Besides accelerating the pace of scientific research, we also expect educational synergies, for instance in the form of masters programs centered around the key aspects of VLs, bringing together students working in different disciplines to share knowledge.
4. Current status and future outlook
No fully operational VLs exist today, but significant progress is being made. To make the concept more concrete and to highlight ongoing research and potential outcomes, we first inspect different scientific disciplines from the perspective of how easy the VL concept is to apply in supporting their progress and discuss potential barriers also from non-technical perspectives in Sections 4.1 and 4.2.
Section 4.3 outlines one concrete example of domain-agnostic tools we expect to become available first, and in Section 4.4, we discuss three examples of ongoing research toward VLs in three different scientific disciplines. Each one showcases progress in digital twins of assets and processes, validating that the tools for constructing VLs are starting to be in place, while also revealing that the work toward incorporating AI within the process is happening separately within each field. This is wasteful and limits opportunities for AI research contributions, and highlights how there are clear differences between disciplines in how effectively the digital tools are being integrated into the core discover process. For ease of communication, we use examples from the authors’ research domains, but emphasize that the VL concept is applicable for considerably broader range of fields.
4.1. Which domains will seize the opportunity?
The VL concept aims for maximal generality, but naturally, disciplines will vary in how much they benefit from incorporating external AI development. Table 1 provides a high-level categorization of various domains into four groups in terms of their potential for adopting VLs. The easiest conceptual mapping from current practices to VLs lies within the natural sciences, where empirical or computational observations drive the scientific method. Here we find the early adopters that already use computational simulations and hence will likely drive the development of VLs in the early years (see Section 4.4 for two examples), as well as the stable followers that will be able to re-use many of the same tools, but for whom the transition will take longer due to digitalization being more laborious.
Categorization of VL application domains and their potential
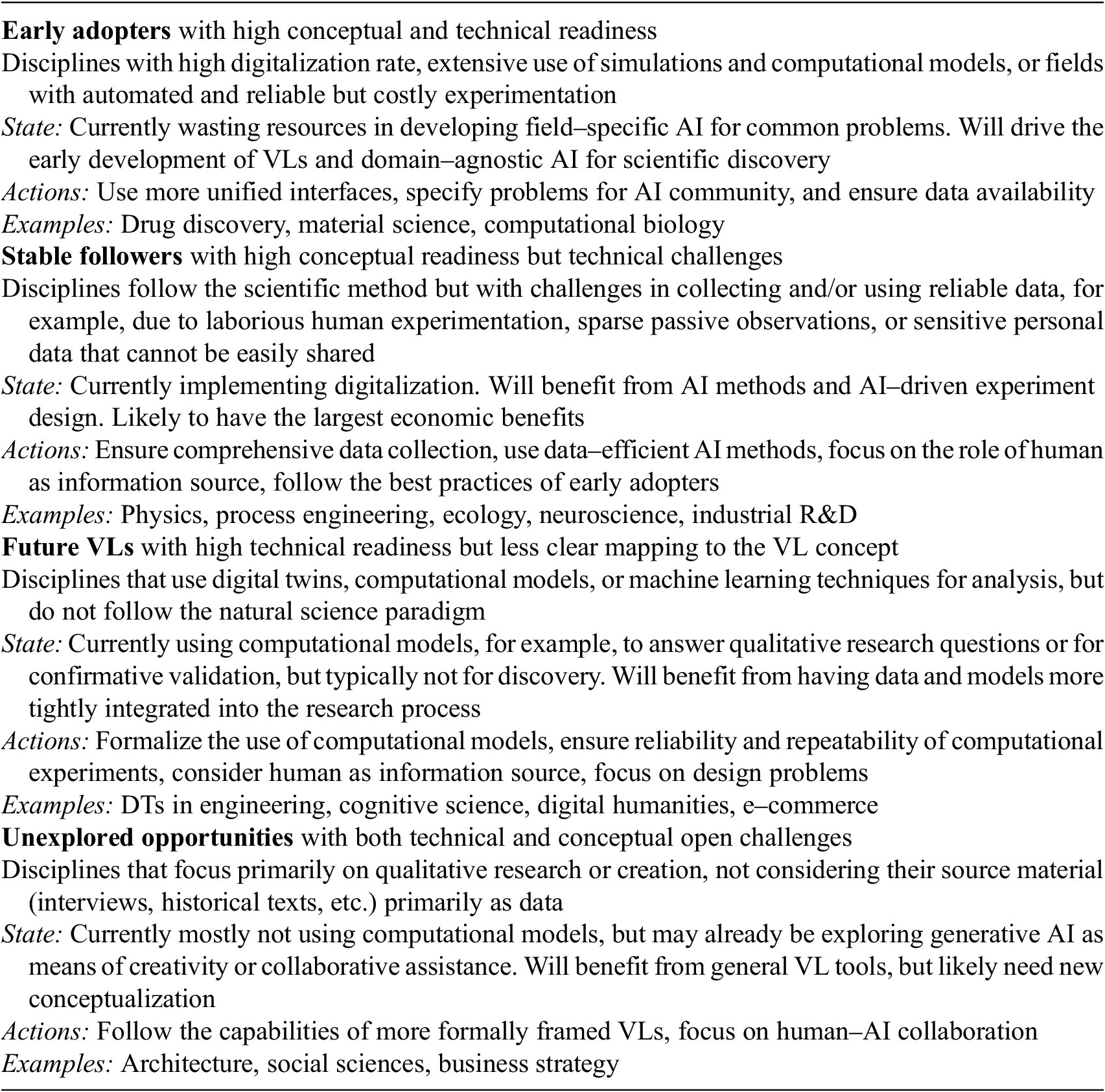
In the other two groups, we anticipate disciplines for which it is less clear how they can best benefit from VLs. These disciplines or deployment areas do not necessarily adhere to the natural science paradigm, but will still benefit from the broad range of domain-agnostic AI tools once the conceptual mapping is clear. We foresee individual examples of future virtual laboratories emerging already in relatively near future; for instance, some formal workflows using computational models for behavioral modeling already exist (Oulasvirta et al., Reference Oulasvirta, Jokinen and Howes2022), and Section 4.4 discusses digital twins in engineering from this perspective. The unexplored opportunities group will likely not benefit from a large proportion of the VL tools, but will in the future be using more AI technologies and is likely to skip the phase of developing field-specific VL tools.
4.2. Challenges
We propose a vision of a feasible transformation and hence focus on the technological enablers and the need for a cultural transformation, and to raise awareness of the opportunities offered by VLs. Naturally, there will be significant challenges as well, spanning across different technical and societal dimensions. Some already identified challenges are briefly discussed below.
4.2.1. Sensitive and personal data
Many VLs need to handle sensitive data, either because of the research subject (e.g., medical research) or because data is collected on the behavior of researchers to improve AI assistance. Running the VL tools in standard secure computing environments is sufficient for ensuring data protection, but explicit effort will be required in ensuring ethical and legal use of the interaction data and the user models; see Sartor et al. (Reference Sartor and Lagioia2020) for a discussion on the European legislation.
4.2.2. Legal aspects, IPR
The complex landscape of AI and intellectual property rights is currently being primarily studied from the perspective of copyright (Samuelson, Reference Samuelson2023). VLs will need to adhere to the established regulation, but additionally, provide opportunities to study, for example, ownership of innovation. By making the roles of different entities (researchers, AI, and data) more systematic and transparent, VLs will play a role in developing responsible innovation practices (Tekic and Füller, Reference Tekic and Füller2023).
4.2.3. Computational and energy cost
Digital twins and computational models can save cost and human time in physical experimentation, but in some cases, the cost of simulations or AI model training is a major factor. However, ML proxies and AI tools for selecting maximally efficient experimental protocols (see Section 4.3) directly address this, and hence VLs are inherently designed to minimize also the energy cost.
4.2.4. Proprietary and legacy components, technical debt
One major technical challenge is support for software and measurement devices with prohibitive licenses. Many important devices, intentionally or unintentionally, only provide a fixed interface and lack support for efficient reading and writing of data. Disciplines, in which the central tools operate in this fashion, may drop down to the stable followers category (Table 1). Any use of AI in these disciplines will require case-specific workarounds or software robotics solutions, resulting in technical debt and high maintenance cost. This problem is particularly apparent for VL integration of such assets. Note, however, that dedicated AI methods, for example, quantifying the uncertainty of black-box models (Sudret et al., Reference Sudret, Marelli and Wiart2017) can help with integrating such legacy components into VLs.
4.2.5. Societal acceptance
Researchers are sometimes conservative and may raise objections on AI involvement, either because of focusing on qualitative conclusions that should be made by humans or because of hesitating on the use of approximations in place of solid evidence. More generally, we often set higher standards for autonomous systems than humans, best exemplified in the case of autonomous cars (Shariff et al., Reference Shariff, Bonnefon and Rahwan2021). These aspects highlight the need for dedicated studies of how the human-AI collaboration is carried out, detailed in Section 3.2. The general public is right now rapidly being exposed to chat-based AI assistance in daily tasks with emerging research on how this transforms the society, and VLs will be an environment to study similar questions in the context of scientific research.
4.3. What generalizes across fields?
The added value of VLs hinges on a sufficiently broad set of components that are usable across fields, that is, can be generalized to new fields. DTs for assets and processes will to a large degree remain domain-specific and we should not expect them all to generalize, but we argue that almost everything else does. First, this holds for the tools needed for creating and maintaining the DTs which are general, as explained in Section 3.1, and it will be important to build on the common advances in these AI technologies. For instance, developing domain-specific solutions for addressing concept drift in measurement data would waste significant effort and the tools would soon get outdated. Second, the DTs for the researcher and the core tools for AI assistance are largely field-independent as explained in Section 3.2, and solutions for helping researchers to interact with the AI assistants would be better developed directly to be generalizable. The VL concept is a necessary enabler for their development and evaluation.
The first demonstrations of added value will come from the early adopters (Table 1) and already active research areas, best exemplified by AI assistance in experimental design that generalizes for all empirical sciences. A significant gap remains between the AI advances and the practical needs of a VL, with no general solutions, for example, for guiding how to best interleave approximate virtual experiments and (often high-cost) empirical experiments while ensuring the user remains in full control, and virtually no research on how to best utilize multiple alternative measurement technologies. Already rudimentary AI assistance for general empirical research setups would be a significant improvement. It would steer the AI development to directions that matter in practice, while opening opportunities for new types of AI, by providing the first systematic and field-agnostic source of data about human interaction in research, kick-starting the virtuous cycle.
4.3.1. Example: AI assistance in optimization
Consider the common task of a researcher iteratively optimizing the parameters of a process to maximize a task-specific quality measure. For instance, a chemist would seek reaction conditions to maximize the yield, or a material scientist would explore the design space to find a material with desired properties. In many cases—especially in the stable follower disciplines—the results of the experiments depend significantly on the competence and practices of the researcher, and irrespective of the field we can develop AI assistance for improving the reliability of experimentation, both on the level of individual measurements and the overall process.
In its simplest form, this task is described in a VL as an iteration that combines a human decision on the next configuration to try with a measurement device or its digital twin to quantify the quality of the configuration, continued until the human decides to terminate the optimization. A natural mode of AI assistance is to supplement the iteration with recommendations for the next configuration (De Peuter and Kaski, Reference De Peuter and Kaski2023), as illustrated also in Figure 4, and typically also for when to terminate. However, formulating the process in a VL enables a much broader form of assistance without notable additional effort from the VL host. For example, the following tasks could all be assisted by stand-alone AI functionality that generalizes for all VLs:
-
• Education and training. AI monitors the choices of the researcher and explains afterward which of the configurations they tried were not ideal, or creates personalized training tasks for a student-centered learning process.
-
• Identification of human biases. Humans search like search algorithms do (Borji and Itti, Reference Borji and Itti2013; Sandholtz et al., Reference Sandholtz, Miyamoto, Bornn and Smith2023). Deviations from algorithmic recommendations can reveal two kinds of human biases: harmful biases preventing efficient work, and useful prior information the researcher uses to solve the problem more efficiently. Both can be used for person-specific assistance.
-
• Evaluation and comparison. Human choices can be analyzed to detect, for example, differences between laboratories or temporal patterns, as a basis for more unified work practices for the field. Data coming from different laboratories can be better combined for meta-analysis or future model training by accounting for the differences.
-
• Team work. AIs can recognize situations in which expertise of a single researcher is not sufficient for making an optimal decision, and recommend, for example, an additional evaluation of the current decision, or assist in combining the expertise of several researchers involved in the process (Song et al., Reference Song, Chen and Yue2019; Mikkola et al., Reference Mikkola, Martinelli, Filstroff and Kaski2023).
-
• From repeated experimentation to DT. Information accumulates throughout the optimization processes and AIs can effectively use this history (Swersky et al., Reference Swersky, Snoek and Adams2013). Accumulated data is needed for building DTs and AI can assist by identifying a minimal set of required additional measurements.
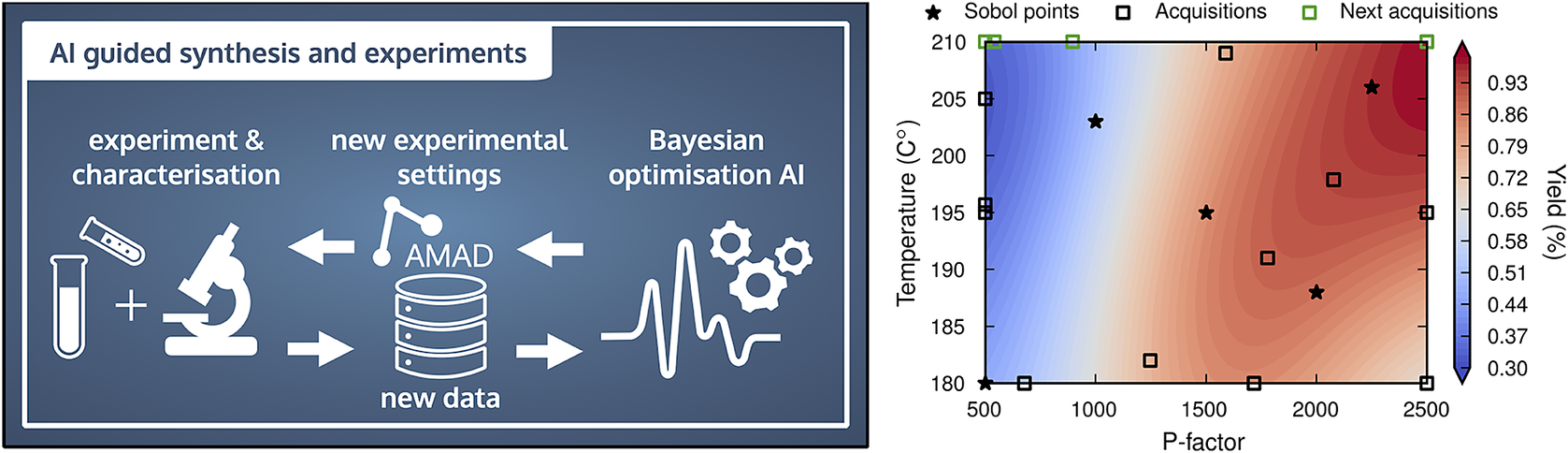
Left: Conceptual illustration of AI-guided materials synthesis and characterization. The Aalto Materials Digitalization (AMAD) Platform (https://www.aalto.fi/en/services/aalto-materials-digitization-platform-amad) facilitates data transfer and collection. Right: Biomaterials example, in which AI guided the extraction and characterization of lignin from birch wood with Bayesian optimization. With very few data points (black and green squares and stars) lignin properties (here the yield) can be correlated to the experimental control variables (here temperature and reactor severity (P-factor).
For many of the tasks in this list, no tools currently exist and research into such tools is only just beginning. VLs would provide a unifying framework to join presently disjoint research and to combine independent data sources for developing methods that generalize across domains.
4.4. Examples of progress toward VLs
4.4.1. Materials science
While no fully-fledged virtual laboratories have emerged in materials science yet, the components are in place and the field is a clear early adopter example. The earliest databases date back to 1965. Their number has risen exponentially since the Materials Genome initiativeFootnote 3 was launched in the United States in 2011 (Himanen et al., Reference Himanen, Geurts, Foster and Rinke2019). Databases evolved via data centers into materials discovery platforms by incorporating data analysis and machine learning tools. The Materials Project (Jain et al., Reference Jain, Ong, Hautier, Chen, Richards, Dacek, Cholia, Gunter, Skinner, Ceder and Persson2013), the Novel Materials Discovery (NOMAD) laboratory (Ghiringhelli et al., Reference Ghiringhelli, Carbogno, Levchenko, Mohamed, Huhs, Lüders, Oliveira and Scheffler2017), and Citrine Informatics (O’Mara et al., Reference O’Mara, Meredig and Michel2016) are prominent examples of such materials discovery platforms and could be viewed as virtual laboratory incubators.
Digital twins are more common in engineering and industry (see Section 4.4.3). They are slowly emerging in materials science, too, with battery development leading the way (Ayerbe et al., Reference Ayerbe, Berecibar, Clark, Franco and Ruhland2021). Ngandjong et al. recently proposed a digital twin of a Li-ion battery manufacturing platform that combines modeling approaches at different scales (Ngandjong et al., Reference Ngandjong, Lombardo, Primo, Chouchane, Shodiev, Arcelus and Franco2021). Thomitzek et al. added a battery cell production digital twin based on digitalization and mechanistic modeling (Thomitzek et al., Reference Thomitzek, Schmidt, Silva, Karaki, Lippke, Krewer, Schröder, Kwade and Herrmann2022). Regarding scientific instruments, Passananti et al. developed a digital twin of a chemical ionization atmospheric pressure interface time-of-flight mass spectrometry (CI-APi-TOF-MS) that facilitates the analysis of molecular cluster formation events in the atmosphere (Passananti et al., Reference Passananti, Zapadinsky, Zanca, Kangasluoma, Myllys, Rissanen, Kurtén, Ehn, Attoui and Vehkamäki2019).
In the Finnish Center for Artificial Intelligence (FCAI), we are developing AI-guided experimentation and synthesis techniques (Jin et al., Reference Jin, Kämäräinen, Rinke, Rojas and Todorović2022; Löfgren et al., Reference Löfgren, Tarasov, Koitto, Rinke, Balakshin and Todorović2022). An example is presented in Figure 4. A Bayesian optimization-based AI requests data from scientists who synthesize and characterize materials. The example shows the extraction of the biopolymer lignin from birch wood and the characterization of the structural properties with 2D nuclear magnetic resonance (NMR) spectroscopy. The data is returned to the AI, which updates its surrogate model of the process and subsequently issues new data requests. Data transfer is mediated by the Aalto Materials Digitalization (AMAD) PlatformFootnote 4 that acts as local data back-end for future virtual laboratories at Aalto University. The lower panel of Figure 4 demonstrates that with relatively few datapoints (i.e., time-consuming synthesis steps), the lignin yield can be maximized. In addition, the surrogate model provides an insightful visualization to the operating scientists of the relation between the extraction (or synthesis) conditions and the lignin (or materials) properties. Such AI-guidance tools are not only the first step toward autonomous experiments or fabrication, and thus the corresponding digital twins, but they also facilitate the collection of data that has traditionally been difficult to digitize due to its acquisition cost (e.g., human, process or computational time and instrument cost).
Bayesian optimization also forms an integral part of self-driving laboratories (SDLs). Such SDLs usually combine robotics, machine learning, and materials chemistry to autonomously synthesize compounds and make materials (Stein and Gregoire, Reference Stein and Gregoire2019; MacLeod et al., Reference MacLeod, Parlane, Morrissey, Häse, Roch, Dettelbach, Moreira, Yunker, Rooney, Deeth, Lai, Ng, Situ, Zhang, Elliott, Haley, Dvorak, Aspuru-Guzik, Hein and Berlinguette2020; Abdel-Latif et al., Reference Abdel-Latif, Epps, Bateni, Han, Reyes and Abolhasani2021; Rooney et al., Reference Rooney, MacLeod, Oldford, Thompson, White, Tungjunyatham, Stankiewicz and Berlinguette2022; Seifrid et al., Reference Seifrid, Pollice, Aguilar-Granda, Chan, Hotta, Ser, Vestfrid, Wu and Aspuru-Guzik2022). SDLs accelerate materials design and exploration and are an important component of future virtual laboratories. SDL concepts and software are already deployed across different materials science domains (Stein and Gregoire, Reference Stein and Gregoire2019; Stein et al., Reference Stein, Sanin, Rahmanian, Zhang, Vogler, Flowers, Fischer, Fuchs, Choudhary and Schroeder2022), medicine, and drug design (Hickman et al., Reference Hickman, Bannigan, Bao, Aspuru-Guzik and Allen2023; Nigam et al., Reference Nigam, Pollice, Tom, Jorner, Thiede, Kundaje and Aspuru-Guzik2022) and thus are an excellent example of the potential for generalization and a common virtual laboratory framework.
Increasingly, software for materials characterization and design is released as open source, such as ChemOS (Roch et al., Reference Roch, Häse, Kreisbeck, Tamayo-Mendoza, Yunker, Hein and Aspuru-Guzik2020), ChemSpectra (Huang et al., Reference Huang, Tremouilhac, Nguyen, Jung and Bräse2021), LiberTem (Clausen et al., Reference Clausen, Weber, Ruzaeva, Migunov, Baburajan, Bahuleyan, Caron, Chandra, Halder, Nord, Müller-Caspary and Dunin-Borkowski2020), elabFTW (A Free and Open Source Electronic Lab Notebook),Footnote 5 FINALES (Vogler et al., Reference Vogler, Busk, Hajiyani, Jørgensen, Safaei, Castelli, Ramirez, Carlsson, Pizzi, Clark, Hanke, Bhowmik and Stein2023), or Chemotion (Electronic Laboratory Notebook & Repository for Research Data)Footnote 6. These components have yet to be cast into a generalized software and ontology framework (Scheffler et al., Reference Scheffler, Aeschlimann, Albrecht, Bereau, Bungartz, Felser, Greiner, Groß, Koch, Kremer, Nagel, Scheidgen, Wöll and Draxl2022) for virtual laboratories as advocated in this article or by Brinson et al. in their recent call for community action on FAIR data (Brinson et al., Reference Brinson, Bartolo, Blaiszik, Elbert, Foster, Strachan and Voorhees2023). In this context, Deagen et al. proposed a materials–information twin tetrahedra (MITT) (Deagen et al., Reference Deagen, Brinson, Vaia and Schadler2022). The term “digital twin” is here used as an analogy between concepts in materials and information science and does not refer to a component of a virtual laboratory. With MITT they advocate a holistic, data-driven approach to materials science, which we believe could be further extended across scientific domains. In a similar vein, Suzuki et al. recently promoted a knowledge transfer from AI applications in pharmaceutical science to materials science through an automated machine learning framework (Suzuki et al., Reference Suzuki, Kurosawa, Marcella, Kanba, Koretaka, Tsuji and Okumura2022).
4.4.2. Drug design
Applying AI to drug design has become very popular in the last 5 years, triggered by the innovations in AI (Chen et al., Reference Chen, Engkvist, Wang, Olivecrona and Blaschke2018). Common application areas are molecular de novo generation, synthetic route predictions, and molecular property predictions. In drug design a starting molecule with typically poor properties is iteratively optimized until a molecule with properties suitable to start clinical trials is identified. The iterative cycle is usually called the Design-Make-Test-Analyze (DMTA) cycle (Figure 5; Plowright et al., Reference Plowright, Johnstone, Kihlberg, Pettersson, Robb and Thompson2012). Due to both clearly defined iterative innovation processes and the growing use of computational experiments drug design is an early adopter.
Drug design is based on iterative cycles of Design-Make-Test-Analyze (DMTA). Within each round, several iterative cycles can be performed in a virtual laboratory (bottom cycle).
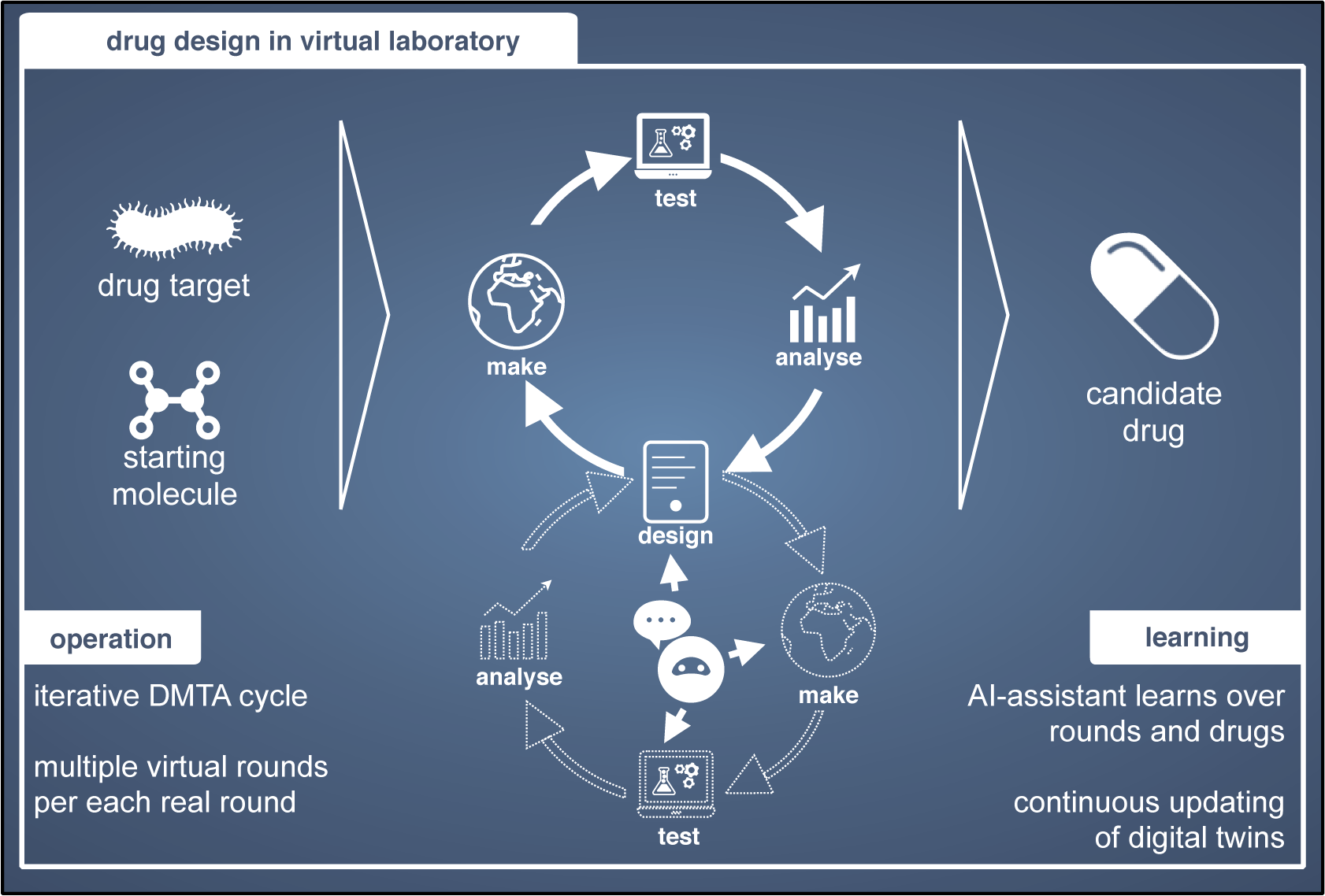
The virtual drug design laboratory will consist of digital twins for the different components in the DMTA cycle. Several of the necessary digital twins are under development. Digital twins are developed for the design part through deep learning-based molecular generation, for the make part through designing synthetic routes by deep learning, and for the test part through developing digital twins for the assays that are used to test the molecules. These aspects are often studied in isolation, and advances in one task sometimes reveal limitations in others. For example, improved generative AI methods for molecule generation (Sadybekov and Katritch, Reference Sadybekov and Katritch2023) call for better screening tools of the resulting, vast candidate sets (Gangwal et al., Reference Gangwal, Ansari, Ahmad, Azad, Kumarasamy, Subramaniyan and Wong2024). An outstanding important research task is to find out how implicit knowledge residing with the scientists can be modeled through human-in-the-loop modeling, so that it can be included in the digital twins of the design and analysis steps. A recent concrete example (Sundin et al., Reference Sundin, Voronov, Xiao, Papadopoulos, Bjerrum, Heinonen, Patronov, Kaski and Engkvist2022) infers implicit user goals from their interaction with the results of a drug synthesis model, providing means for learning the user model agnostic to the specific synthesis model used.
It is important to keep in mind that the virtual laboratory is an approximation of a real drug design laboratory. Virtual molecules are optimized in the virtual laboratory and then actually synthesized and tested in a real laboratory in an iterative manner. An optimal laboratory would combine a virtual laboratory with a fully automated real laboratory. There are several efforts ongoing to create autonomous automation systems for synthesizing and optimizing molecules (Coley et al., Reference Coley, Thomas, Lummiss, Jaworski, Breen, Schultz, Hart, Fishman, Rogers, Gao, Hicklin, Plehiers, Byington, Piotti, Green, Hart, Jamison and Jensen2019) (see also Section 4.4.1). Thus for drug design, virtual and real laboratories need to exist in close collaboration, where as good compounds as possible are proposed by the virtual laboratory, the molecules are then synthesized as efficiently as possible in the laboratory, and the resulting data is fed back to the virtual laboratory. It should be noted that there are components describing the interactions between atoms that would be common for all VLs describing molecules like a material science VL. Examples are molecular dynamics software and atomic force field parameters.
4.4.3. Data-centric engineering
Engineering has recently witnessed a proliferation in data-centric techniques and digital twin development. In many cases constructing the DT is a significant investment and has been a central research focus, whereas the question on how they are best used for scientific or R&D advances remains more open, making the field a future VL example (Table 1). Limited forms of interaction and underuse of AI technologies in DTs have been identified as challenges (Tao et al., Reference Tao, Zhang and Zhang2024), but the necessary components for considering engineering DTs as part of a VL are starting to emerge. For example, recent open software tools for DTs (Tao et al., Reference Tao, Sun, Cheng, Zhu, Liu, Wang, Xu, Hu, Liu, Liu, Sun, Xu, Bao, Xiang and Jin2024) pave way for domain-agnostic DT operations and human-in-the-loop models are already considered in design use cases (Batty, Reference Batty2024).
VLs that enable rapid deployment of DTs will offer tools that generalize across domain-specific instances. The main generalization areas lie at three boundary interfaces: (a) between sensing systems and the DT (e.g., optimal sensor placement and dataset shift tools), (b) between the DT and machine-assisted human decision-making (e.g., AI-assistant interfaces and sequential decision-making tools, and (c) between DT instances (e.g., federated learning and evidence synthesis tools) in a VL that integrates an ecosystem of DTs. Progress in common tools for these aspects will form the basis of VLs that efficiently leverage engineering DTs.
We showcase here two examples of recent engineering DTs developed at the Alan Turing Institute with academic and industrial partners, demonstrating the maturity of the technical enablers and outlining the opportunities a VL would open. The first one is the world’s first 3-D printed steel bridge (MX3D bridge) depicted in Figure 6 (left) and currently situated in Amsterdam, Netherlands. Various sensor networks on the bridge, such as cameras, accelerometers, and load cells, stream live data to its digital twin at the Turing Institute in the UK. The underlying DT model has been developed based on the StatFEM methodology that was recently introduced (Girolami et al., Reference Girolami, Febrianto, Yin and Cirak2021) to formally synthesize observational data and numerical models of its structure, and could be further developed to become a common tool for engineering DTs. Such DTs when embedded in the envisioned VL framework would benefit from evidence integration across physical structures; in this particular case rapidly improving our understanding of 3-D printed steel structures and their stress–strain response under load across environmental and loading conditions.
Left: The 3D-printed steel bridge currently installed in Amsterdam, Netherlands, and its multiple sensing arrays that are streaming live data into the corresponding digital twin in the Turing, UK. Images by Joris Laarman Labs, Thea van den Heuvel, MX3D, and AutoDesk Research. Right: The underground farm in Clapham, London, and its multiple sensing arrays that are streaming live data flows into the corresponding digital twin in The Turing and the University of Cambridge, UK. Images by Rebecca Ward, Flora Roumpani, and Zero Carbon Farms Ltd.

The second example, from the CROP project,Footnote 7 is a digital twin of an underground farm in a tunnel situated in Clapham, London, UK, depicted in Figure 6 (right). This is a hydroponics system with 2 aisles running in parallel in 23 zones and 2 meters long. The underlying DT model utilizes particle filtering for model calibration (Ward et al., Reference Ward, Choudhary, Gregory, Jans-Singh and Girolami2021) and data synthesis, while various environmental measurements and camera footage are live-streamed from sensor networks to monitor crop health, environmental conditions, and yields. The DT already enables the study of a broad range of research questions, from forecasting yields and future conditions to optimizing the placement of crops and the environmental conditions. Within a VL it could be considered more generally as a digital experimentation tool that integrates into the overall research process, likely bringing scientific advantages beyond speed gains such as scenario testing. However, compared to standard measurement devices a DT is considerably more complex in its offering, making the best way of embedding it into a VL an open research question.
5. Conclusion
We introduced the virtual laboratory concept to amalgamate scientific research and R&D in industry with AI technology and AI assistance. We highlighted the benefits of VLs for both research laboratories and AI researchers and outlined key requirements of a common software layer and various research directions to proceed toward VLs. In our opinion, VLs are a community effort and we seek to bring AI researchers and scientists of other domains together to raise awareness for the VL concept and to work together toward realizing VLs.
The goal of transforming research with AI is ambitious and the transformation will not happen fast. The domain scientists are already working toward this direction, as highlighted by the examples in this paper, and hence we conclude our work with words of encouragement for the AI researchers. In short, VLs provide AI researchers means and incentives to contribute to the scientific efforts for solving the grand challenges we are facing. It is hard to think of a subarea of AI that would not be useful for VLs, and hence VLs will provide unique opportunities and cross-fertilization already within AI itself. In many areas, from reinforcement learning to constrained optimization and probabilistic modeling, the current techniques are already clearly sufficient for becoming core elements of VLs. In others, such as causal inference, probabilistic numerics, and domain-agnostic models of researchers, the VLs will provide concrete cases for testing the current solutions and identifying future research directions.
Data availability statement
No specific data was used.
Author contribution
Conceptualization: All authors; Visualization: A.K., T.D., P.R.; Writing—original draft: All authors. All authors approved the final submitted draft.
Funding statement
A.K., P.R., and S.K. were supported by the Research Council of Finland from the Flagship programme: the Finnish Center for Artificial Intelligence FCAI, and from projects 345604 (S.K.), 341732 (S.K.), 341589 (P.R.), 346377 (P.R.), 348180 (P.R.), 352861 (P.R.), and 336019 (A.K.). P.R. acknowledges support from the COST actions CA18234 and CA22154, S.K. from the UKRI Turing AI World-Leading Researcher Fellowship (EP/W002971/1), and T.D. from the UKRI Turing AI Acceleration Fellowship (EP/V02678X/1).
Competing interest
The authors declare no competing interests exist.
Ethical standard
The research meets all ethical guidelines, including adherence to the legal requirements of the study country.










Comments
No Comments have been published for this article.