Introduction
The early stages of the plant life cycle, including seed germination and seedling establishment, are key events for seed plant propagation. However, these stages are highly susceptible to environmental conditions, both biotic and abiotic factors. The ability of the seed embryo to restart its metabolic activity in a coordinated and sequential manner is defined as germination vigour (Rajjou et al., Reference Rajjou, Duval, Gallardo, Catusse, Bally, Job and Job2012; Galland et al., Reference Galland, Huguet, Arc, Cueff, Job and Rajjou2014). This is a highly sought trait in agriculture as it is directly correlated to eventual total biomass and yield production. Germination sensu stricto specifically refers to the rehydration of the quiescent and non-dormant dry seed and the subsequent protrusion of the embryonic axis through the surrounding structures, which are driven by localized cell elongation (Bewley and Black Reference Bewley and Black1994; Sliwinska et al., Reference Sliwinska, Bassel and Bewley2009).
Upon imbibition, the seeds release in their surrounding microenvironment, known as the spermosphere, a complex mixture of organic and inorganic molecules (Martins et al., Reference Martins, Medeiros, Lakshmanan and Bais2018; Nelson Reference Nelson2018). The spermosphere is the transient region of microbial and metabolic activities around a germinating seed, which are driven by seed exudate compounds, that exists specifically during the germination sensu stricto phase before transitioning to the rhizosphere. These exudate molecules shape and drive the selection, growth and interactions of microorganisms, which ultimately play an important role in seedling establishment and plant fitness by contributing to the future plant microbiome (Hill et al., Reference Hill, Farrell and Jones2012; Nelson Reference Nelson2018). The composition and functions of plant exudates have been relatively well studied in the phyllosphere (Xu et al., Reference Xu, Zhao, Zhang, Zhang, Wang, Qin, Ke, Qiu, Peijnenburg, Lu and Qian2022) and rhizosphere (Sasse et al., Reference Sasse, Martinoia and Northen2018; Sun et al., Reference Sun, Jiang, Jiang, Wu, Gao and Wang2021). In contrast, the spermosphere is less well described, though this area is advancing and garnering increasing attention, especially with the development of high-throughput techniques (Schiltz et al., Reference Schiltz, Gaillard, Pawlicki-Jullian, Thiombiano, Mesnard and Gontier2015). The seed-derived peptides exuded in the spermosphere are expected to play crucial roles in shaping microbial communities and influencing plant–microbe interactions, as previously described for other plant-derived peptides (Hu et al., Reference Hu, Zhang and Shi2018; Pantigoso et al., Reference Pantigoso, Newberger and Vivanco2022). These peptides can serve as nutrient sources, providing carbon and nitrogen to microorganisms and supporting their growth during seed germination. They may act as signalling molecules, affecting microbial colonization and seed defence. Several studies have demonstrated the release of a diverse range of proteins from seed exudates across various plant species, which are often involved in defence mechanisms (Fuerst et al., Reference Fuerst, Okubara, Anderson and Morris2014; Rocha et al., Reference Rocha, Morais, Oliveira, Oliveira, Sousa, Souza, Moreno, Monteiro-Moreira, de Souza Júnior, de Sá and Vasconcelos2015; Raviv et al., Reference Raviv, Aghajanyan, Granot, Makover, Frenkel, Gutterman and Grafi2017; Hubert et al., Reference Hubert, Marchi, Ly Vu, Tranchant, Tarkowski, Leprince and Buitink2024). Despite the limited focus on the seed peptidome, seeds accumulate significant amounts of storage proteins, serving as reservoirs for potential peptides (Ayala-Niño et al., Reference Ayala-Niño, Rodríguez-Serrano, González-Olivares, Contreras-López, Regal-López and Cepeda-Saez2019). For example, Macadamia integrifolia seeds have been shown to release antimicrobial peptides during germination, illustrating the presence of seed-exuded peptides in the spermosphere (Marcus et al., Reference Marcus, Green, Goulter and Manners1999).
Peptides are polymers of amino acids. Although no definitive molecular size criterion formally distinguishes peptides from proteins, peptides are conventionally defined as chains composed of less than 50 amino acid residues, contrasting with larger protein molecules (Farrokhi et al., Reference Farrokhi, Whitelegge and Brusslan2008; Marmiroli and Maestri, Reference Marmiroli and Maestri2014). Peptide roles in signalling and defence have been documented in the plant (Marmiroli and Maestri, Reference Marmiroli and Maestri2014), but there is literature pertaining to their functions and roles in seeds or their exudation during seed germination. Interestingly, recent advancements in integrated peptidogenomic pipelines have allowed for the large-scale identification of non-conventional peptides (NCPs) in plants, emphasizing their significant roles in essential biological processes (Wang et al., Reference Wang, Tian, Liu, Li, Zhang, Chen, Jia, Zheng, Wu, Chen, Yan and Wu2020).
Our study aims to apply untargeted peptidomics to investigate peptides exuded by germinating seeds in the spermosphere, addressing a significant gap in current research. Seed exudates contain a complex proteinaceous fraction that includes both high- and low-molecular-weight peptides (Scarafoni et al., Reference Scarafoni, Ronchi, Prinsi, Espen, Assante, Venturini and Duranti2013; Rocha et al., Reference Rocha, Morais, Oliveira, Oliveira, Sousa, Souza, Moreno, Monteiro-Moreira, de Souza Júnior, de Sá and Vasconcelos2015). Thus, developing an optimized extraction method is essential for effective peptide isolation. We evaluated various solvents, including a commonly used weak acid, a strong acid, and an acidified organic solvent known for their peptide extraction efficacy (Vitorino et al., Reference Vitorino, Barros, Caseiro, Ferreira and Amado2012; Yu et al., Reference Yu, Canales, Glover, Das, Shi, Liu, Keller, Attie and Li2017; Chen et al., Reference Chen, Hwang, Gleaton, Titus and Hamlin2018; Barashkova and Rogozhin, Reference Barashkova and Rogozhin2020). This assessment aims to identify the most efficient method for capturing a diverse range of peptides from the spermosphere.
To achieve this, we present a novel methodology specifically designed to study exuded peptides during seed germination using common bean seeds. Common bean was chosen as a potential reservoir of bioactive peptides due to their high protein content (Mojica and de Mejía, Reference Mojica and de Mejía2015; Mojica et al., Reference Mojica, Luna-Vital and González de Mejía2017). Our approach involved isolating peptides from the spermosphere using three selected extraction solvents, followed by mass spectrometry for peptide identification. Additionally, we used an online tool to predict various properties of the identified peptides based on their sequences.
Material and methods
Biological material and germination assay
Seeds were harvested in 2020 from the common bean genotype ‘CON’, a French commercial cultivar of Phaseolus vulgaris L. cv. Contender, provided by Vilmorin (https://www.vilmorinmikado.fr/). The seeds were cultivated from the FNAMS field experimental station in Maine-et-Loire (GPS, Global Positioning System coordinates: 47.470532, −0.394526) in the Loire Valley, France. Before testing, the seeds underwent micro-cleaning, and a purity test was performed by GEVES (the French official seed testing organization) in accordance with the International Seed Testing Association (ISTA) recommendations. The thousand seed weight was measured at 286.33 g, and the moisture content was 12.9%. To assess seed vigour, germination tests were conducted at the Institut Jean-Pierre Bourgin (IJPB, INRAE-AgroParisTech) in Versailles, France. Fifty seeds were placed on SARTORIUS Pleated Seed Testing Papers (Grade 20, 119 g/m²) from Fischer Scientific and positioned in polypropylene germination boxes (55 mm × 120 mm × 180 mm, known as GEVES-type boxes, Loire Plastic, France), each containing 60 mL of Milli-Q water. These boxes were incubated in a Phytotron SANYO MLR-359, which was equipped with fluorescent white LED lamps (FL-40SS-W/37) set to maximum brightness (Light Step unit 5) to provide continuous daylight while maintaining a constant temperature of 25°C. Germination was monitored by counting radicle protrusion at 0, 24, 48, 72 and 96 h. The cumulative germination percentage was calculated, and a germination curve was generated using GerminaQuant in R (Lozano-Isla et al., Reference Lozano-Isla, Benites-Alfaro and Pompelli2019). The seed lot exhibited high germination potential, exceeding 99% (Fig. 1A–C).
Workflow of experimental design for spermosphere collection and analysis. (A) Seeds of the genotype ‘CON’ (Phaseolus vulgaris L.), a French commercial cultivar, were harvested in 2020 from the FNAMS field experimental station in Maine-et-Loire, France. (B) Harvested seeds were sent to the National Seed Testing Station (SNES) at GEVES, where they underwent micro-cleaning, purity testing, and sampling according to the ISTA guidelines. (C) Sampled seeds were then sent to the IJPB. Germination tests were conducted in three biological replicates on seed lots of the ‘CON’ genotype harvested in Maine-et-Loire. (D) Spermosphere collection from germinating seeds involved soaking 10 unsterilized seeds from the genotype CON seed lot harvested in Maine-et-Loire. To ensure the reliability of our results, three independent biological repetitions were considered in this study. The total weight of the seeds was measured before placing them in a 50 mL Falcon tube. The seeds were then imbibed with 1.5 times their total weight in sterile Milli-Q water. After verifying that none of the 10 seeds had germinated, the spermosphere was collected 24 h after imbibition. This spermosphere was divided into three Eppendorf tubes (200 μL each) for peptide extraction with different solvents (FA, formic acid: 0.1% or TFA, trifluoroacetic acid: 0.1% or MeOH–H2O–AcOH, methanol:water:acetic acid: 90:9:1 v/v/v). (e) The untargeted peptidomic analyses using mass spectrometry were conducted at the Proteogen platform in Caen (France).

Spermosphere collection
In this study, we assessed the composition of seed-exuded peptides in the spermosphere (Fig. 1D). To ensure the reliability of our results, three independent biological repetitions were considered in this study. To collect the spermosphere, 10 seeds were imbibed for 24 h, meaning at the end of phase II of the germination process, since the first radicle protrusions were observed (Fig. 1C) (Bewley, Reference Bewley1997). For each sample, the collective weight of the 10 seeds was determined. According to previous studies (Mendoza et al., Reference Mendoza, Wiesinger, Lu, Nchimbi-Msolla, Miklas, Kelly and Cichy2018; Vidak et al., Reference Vidak, Lazarević, Javornik, Šatović and Carović-Stanko2022), the absorption capacity of the seeds was measured beforehand and reached a plateau at 110% of their dry mass till radicle protrusion. Therefore, to provide an excess of sterile Milli-Q water (pH 6.5) for sampling the spermosphere without fully submerging the seeds, we determined that using a water mass corresponding to 150% of the dry mass is appropriate for our analysis. Imbibed seeds were shaken at 70 rpm on a platform shaker and kept at 25°C (Novotron INFORS AG CH-4103 Bottmingen) under a light intensity of 1000 lm/m². After checking and finding that none of the 10 seeds had germinated, the water surrounding the seeds, including the spermosphere, was collected after 24 h.
Extraction of peptides from the spermosphere and untargeted LC–MS/MS peptidomic analysis
Peptide extraction
Three biological replicates of the common bean spermosphere were subjected to peptide extraction using three different solvents. In particular, formic acid (FA, 0.1%), trifluoroacetic acid (TFA, 0.1%) and methanol:water:acetic acid (MeOH–H2O–AcOH, 90:9:1 v/v/v) were used separately for peptide solubilization and extraction as previously described (Kline et al., Reference Kline, Pang, Hefta, Opiteck, Kiefer and Scheffler2003; Husson et al., Reference Husson, Clynen, Baggerman, De Loof and Schoofs2005; Hoofnagle et al., Reference Hoofnagle, Whiteaker, Carr, Kuhn, Liu, Massoni, Thomas, Townsend, Zimmerman, Boja, Chen, Crimmins, Davies, Gao, Hiltke, Ketchum, Kinsinger, Mesri, Meyer, Qian, Schoenherr, Scott, Shi, Whiteley, Wrobel, Wu, Ackermann, Aebersold, Barnidge, Bunk, Clarke, Fishman, Grant, Kusebauch, Kushnir, Lowenthal, Moritz, Neubert, Patterson, Rockwood, Rogers, Singh, Van Eyk, Wong, Zhang, Chan, Chen, Ellis, Liebler, Rodland, Rodriguez, Smith, Zhang, Zhang and Paulovich2016). Briefly, 200 μL of spermosphere samples from each biological replicate were distributed into three separate 2 mL Eppendorf tubes for solvent extraction (Fig. 1D). Subsequently, 1.8 mL of each solvent was added separately to the tubes containing the spermosphere samples. The samples with their respective extraction solvents were agitated at 24 rpm on a rotator (Mini LabRoller™ Rotator, LabNet) at 4°C for 30 min. The samples were then centrifuged at 10,000 g to separate the soluble peptides, which were found in the supernatant, from the insoluble components (precipitate). The peptides extracted in the supernatant were subsequently dried using a SpeedVac vacuum concentrator (Thermo Scientific™ SPD 111V).
Nano-LC–MS/MS analysis
The untargeted peptidomic analyses using mass spectrometry were conducted at the Proteogen platform in Caen (France) (Figs. 1E and 2). For nano-LC fragmentation, peptide samples were first desalted and concentrated onto a μC18 Omix (Agilent) before analysis. Dried peptide samples were resuspended in 40 μL of FA (0.1%). The resuspended peptide samples were loaded to a μC18 Omix (Agilent) column for desalting and concentration and subsequently eluted with acetonitrile (80%) containing FA (0.1%). The eluted peptides are dried using a SpeedVac vacuum concentrator. The resulting concentrated peptide samples were then resuspended in 10 μL of 0.1% FA. Finally, 1 μL of the resuspended peptide extract was injected for LC–MS/MS analysis.
Peptidomic analysis workflow. This pipeline illustrates an overview of the steps involved in analysing the peptidome from the spermosphere of germinating common bean seeds.
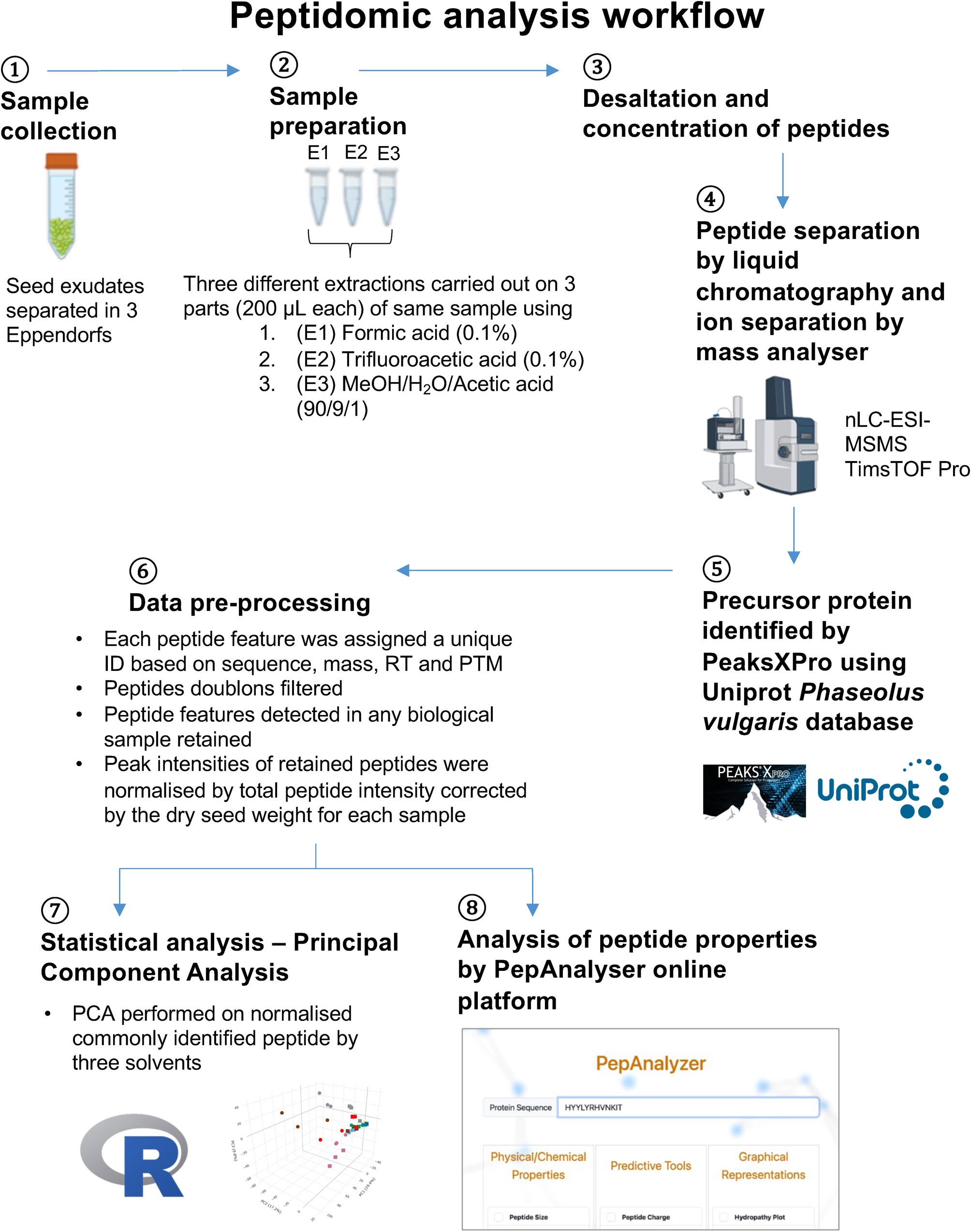
The chromatography step was performed on a NanoElute (Bruker Daltonics) ultra-high-pressure nanoflow chromatography system. Peptide samples were first concentrated onto a C18 PepMap 100 (5 mm × 300 μm i.d.) precolumn (Thermo Scientific) and then separated at 50°C onto a reversed-phase Reprosil column (25 cm × 75 μm i.d.) packed with 1.6 μm C18-coated porous silica beads (IonOpticks). The mobile phases consisted of 0.1% FA, 99.9% water (v/v) (solvent A) and 0.1% FA in 99.9% acetonitrile (v/v) (solvent B). The nanoflow rate was set at 300 nL/min, and the gradient profile was as follows: from 2 to 15% solvent B within 15 min, followed by an increase to 25% solvent B within 10 min and to 37% solvent B within 12 min, followed by a 95% solvent B washing step before reequilibration.
Mass spectrometry experiments were carried out on a Trapped Ion Mobility Spectrometry – Time of Flight Pro mass spectrometer (Bruker Daltonics) with a modified CaptiveSpray Nano Electrospray Ion Source (ESI, Bruker Daltonics). The system was calibrated weekly, and the mass accuracy was better than 1 ppm. A 1600 spray voltage with a capillary temperature of 180°C was typically employed for ionization. MS spectra were acquired in the positive mode in the mass range from 100 to 1700 m/z. In the experiments described here, the mass spectrometer was operated in a parallel accumulation–serial fragmentation (PASEF) mode without the exclusion of single-charged peptides. A number of 10 PASEF MS/MS scans were performed for 1.25 s.
Database searching for peptide identification
Peptide identification was performed using PEAKS XPro software (Bioinformatic Solutions Inc., CA) against an updated version of the UniProt Phaseolus vulgaris database (Proteome ID: UP000000226; The UniProt Consortium, 2022). The variable modifications allowed were as follows: methionine oxidation, acetylation (N-term), deamidation and carbamylation. The parameter ‘no enzyme’ was selected to permit the identification of peptides irrespective of any specific enzyme digestion. The mass accuracy settings were established at 20 ppm and 0.05 Da for MS and MS/MS mode, respectively. These settings were optimized based on the calibration of the mass spectrometer to ensure high-fidelity measurements of peptide masses. Identified peptides were filtered according to a false discovery rate (FDR) of 5%. Additionally, we implemented a redundancy elimination strategy based on proteins being evidenced by the same set or a subset of peptides to ensure the uniqueness of identified proteins.
Data pre-treatment
Unique identifiers were assigned to each detected peptide based on its sequence, mass, retention time (RT) and presence of post-translational modifications (PTMs), namely methionine oxidation, acetylation (N-term), deamidation and carbamylation. Subsequently, redundant peptides were removed.
Statistical analysis
Principal component analysis (PCA) was conducted on the peptidomic dataset using R (v.4.1.1) to assess the relationships among peptides identified across samples. Only commonly identified peptides across all biological repetitions and all solvents (commonly identified peptides across all nine samples) were analysed. Peptide intensities of commonly identified peptides were normalized by the weight of 10 seeds for PCA.
Peptide properties prediction
Following a selection process, the research focused on a specific set of peptides retained for further analysis. To gain insights into their behaviour, the physico-chemical properties of these retained peptides were predicted using PepAnalyser (Rustagi et al., Reference Rustagi, Gupta, Bajaj, Singh and Singh2023). This online tool uses the amino acid sequences of the peptides to predict various characteristics, including molecular weight, isoelectric point (pI), net charge at physiological pH, hydropathicity, half-life in different environments, instability index, aliphatic index and grand average of hydropathicity (GRAVY).
Results
Diversity of distinct peptides in the spermosphere
Using three distinct solvents (FA, TFA, MeOH–H2O–AcOH), a total of 316 unique peptides were identified across three biological replicates of the common bean spermosphere (Supplementary Table S1). Unique codes were assigned to each peptide based on its amino acid sequence, mass, and any PTMs present. The peptide sequences ranged in length from 5 to 29 amino acids, and their masses spanned a range of 546.34 to 2738.31 Da (Fig. 3A, Supplementary Table S1).
Characteristics of peptides identified in the spermosphere of common bean of genotype CON. (A) A total of 316 distinct peptides, with lengths ranging from 5 to 29 amino acids, were identified across the three biological repetitions using three solvents: formic acid (FA) at 0.1%, trifluoroacetic acid (TFA) at 0.1%, and a methanol–water–acetic acid mixture (90:9:1 v/v/v). (B) The distribution of identified peptides with PTMs, including methionine oxidation, acetylation (N-term), deamidation, and carbamylation, among the total of 316 identified peptides indicated that 31 peptides exhibited at least one PTM. (C) Percentage distribution of the various types of PTMs among the 31 identified peptides.
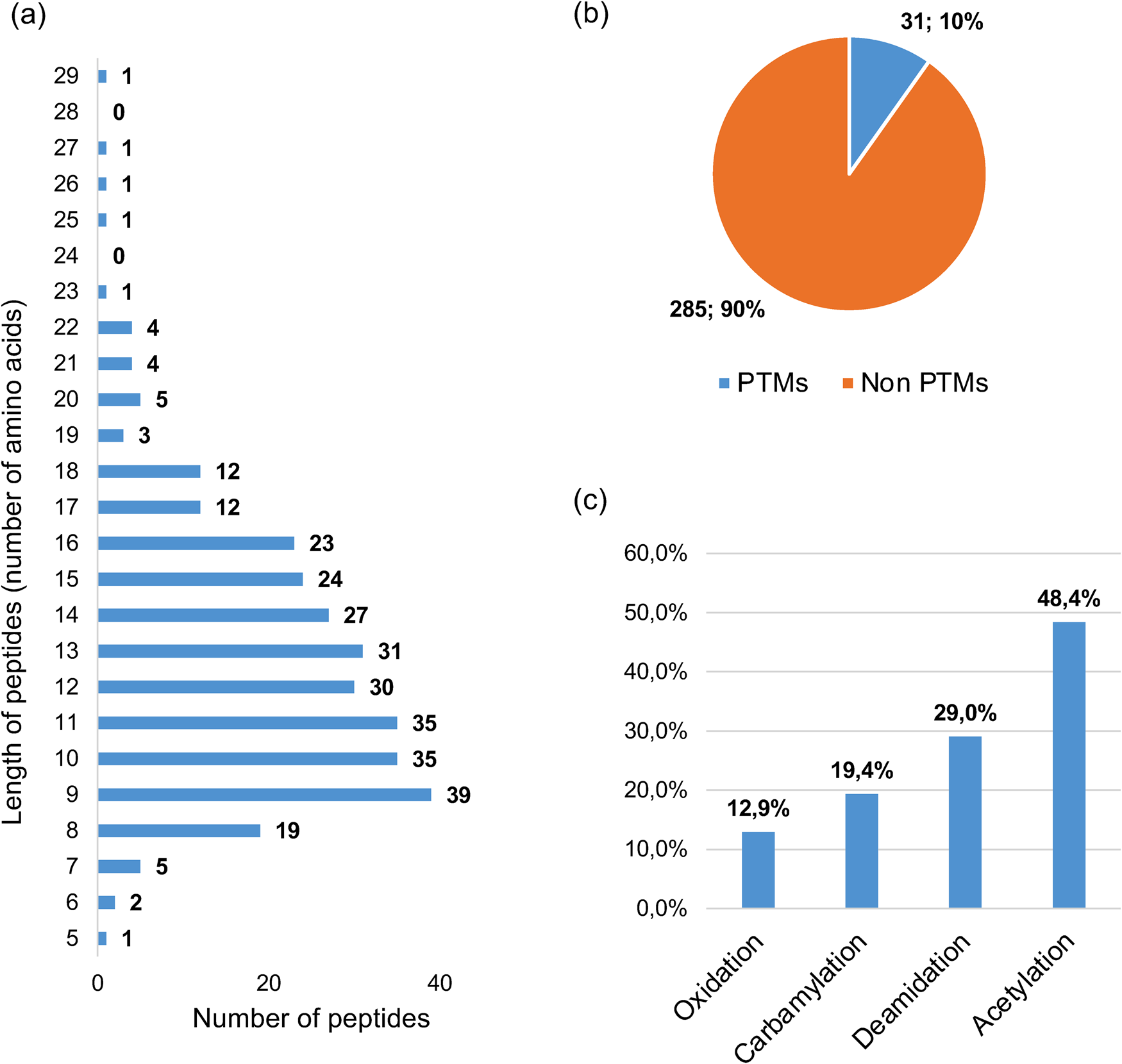
The analysis of identified peptides revealed a distinct distribution in their lengths with peptides composed of 9, 10 or 11 amino acids being the most represented. The vast majority (98%) of all identified peptides fell within a broader range of 6–22 amino acids (Fig. 3A).
As illustrated in Fig. 3B, 31 out of the 316 identified peptides in the germinating common bean spermosphere (approximately 10%) exhibited PTMs. Among the four PTMs investigated on the identified peptides, acetylation (48.4%) was the most abundant modification, followed by deamidation (29.0%), carbamylation (19.4%) and methionine oxidation (12.9%) (Fig. 3C).
Solvent choice impacts peptide identified
The choice of solvent significantly impacted peptide yield during extraction from the spermosphere. Among the three solvents tested (FA, TFA and MeOH–H2O–AcOH), FA emerged as the most efficient solvent, accounting for 85% of the identified peptides. MeOH–H2O–AcOH and TFA exhibited moderate efficiency, extracting 64 and 54% of the peptides, respectively (Fig. 4A).
Comparison of extraction solvents for peptide extraction. (A) Proportion of the total peptides identified by using the different extraction solvents (FA, formic acid: 0.1% or TFA, trifluoroacetic acid: 0.1% or MeOH–H2O–AcOH, methanol:water:acetic acid: 90:9:1 v/v/v) represented as percentages. (B) The repeatability of each solvent is represented as the percentage of commonly identified peptides across the three biological repetitions for each solvent, respectively. The commonly identified peptides across the three biological repetitions for each solvent were represented as a proportion of the total identified peptides from the solvents, respectively. (C) Comparison of the extraction solvents for the identification of peptides. The commonly identified peptides across the three biological repetitions for each solvent, respectively, were used to compute the Venn diagram. (D) PCA of commonly identified peptides across any three biological repetitions. PCA was applied to the normalized peptide intensity data. The figure illustrates the distribution of samples in the reduced-dimensional space defined by the first two principal components (PC1 and PC2). Each point represents an individual sample, and colours indicate the different solvents.
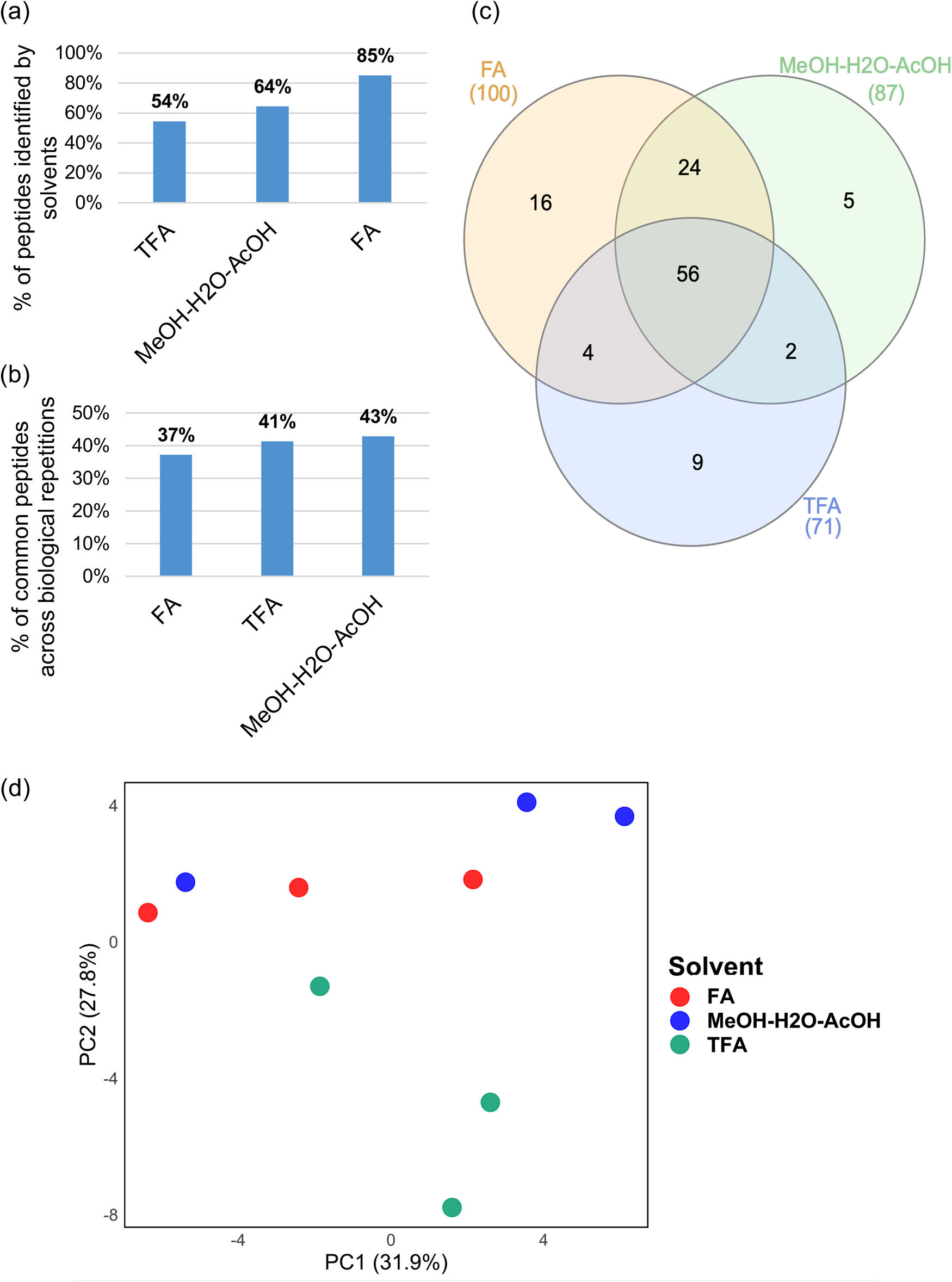
To account for biological variation, our experimental design incorporated three biological replicates for each solvent used in peptide extraction. The repeatability of peptide identification across these replicates was evaluated by analysing commonly identified peptides. Notably, using FA, 269 different peptides were identified, with 100 peptides consistently detected in all three biological replicates. Using MeOH–H2O–AcOH, we identified 203 peptides in the samples and 87 of them were consistently identified across the three biological repetitions. Lastly, 172 peptides were identified with TFA in the samples, with 71 peptides being consistently identified across the three biological repetitions. For each solvent, we determined the ratio of consistently identified peptides across the three biological repetitions to the total number of peptides identified by the solvent expressed in percentage to evaluate the repeatability in peptide extraction and identification (Fig. 4B).
We compared the consistently identified peptides across the three biological repetitions (116 peptides) for each solvent by means of a Venn diagram (Heberle et al., Reference Heberle, Meirelles, da Silva, Telles and Minghim2015). We revealed 56 commonly identified peptides by the three different solvents (Fig. 4C). Unique peptides were also identified according to each solvent. FA, MeOH–H2O–AcOH and TFA were responsible for the identification of 16, 5 and 9 unique peptides, respectively.
PCA based on peptide intensity data of the 56 commonly identified peptides grouped samples according to extraction solvents (Fig. 4D). Analysis of variance (ANOVA) of peptide intensities of the 56 common peptides across the three distinct solvents revealed only four differential peptides (FDR, P-value 0.05) (Supplementary Table S2).
Characteristics of identified peptides by the three different solvents
Using PepAnalyzer (http://www.iksmbrlabdu.in/), an online user-friendly tool that predicts several properties of a known peptide sequence (Rustagi et al., Reference Rustagi, Gupta, Bajaj, Singh and Singh2023), we identified several properties pertaining to the 316 identified peptides (Supplementary Table S1, Fig. 5).
Predicted properties of 316 identified peptides in the spermosphere. The physico-chemical properties of peptides identified in the spermosphere were predicted based on their amino acid sequence using PepAnalyser (Rustagi et al., Reference Rustagi, Gupta, Bajaj, Singh and Singh2023). Each property is shown as a percentage of the total dataset, giving insights into the relative prevalence of these properties among the peptides identified in the spermosphere. (A) Distribution of peptide stability. (B) Distribution of peptide polarity. (C) Distribution of peptide-binding potential. (D) Distribution of peptide aromaticity. (E) Comparison of extraction solvents (FA, formic acid: 0.1% or TFA, trifluoroacetic acid: 0.1% or MeOH–H2O–AcOH, methanol:water:acetic acid: 90:9:1 v/v/v) on predicted peptide properties.

The stability of the peptides was evaluated using an algorithm developed by Guruprasad et al. (Reference Guruprasad, Reddy and Pandit1990) to calculate an instability index (Supplementary Table S1). According to this algorithm, a score below 40 suggests a stable peptide, while values above 40 indicate potential instability. In accordance with this criterion, our results (Fig. 5A) indicate that 75% of the peptides were predicted as stable.
Kyte and Doolittle's (Reference Kyte and Doolittle1982) method was used to assess peptide polarity through the grand average of hydropathy (GRAVY) index (Supplementary Table S1). This method assigns a hydropathy value to each amino acid in the sequence. To assess local preferences, it calculates the average hydropathy over a window of a predefined size that slides along the sequence from the N-terminus to the C-terminus. Peptides with a positive GRAVY index are predicted to be more hydrophilic (polar), while negative values indicate a more hydrophobic character. Our analysis revealed that a majority (70%) of the identified peptides exhibited a preference for hydrophilic interactions (Fig. 5B).
The propensity of the peptides to interact with proteins or membranes was evaluated using the Boman index (Supplementary Table S1). This index, also known as the potential protein interaction index, was developed by Boman (Reference Boman2003) to estimate binding potential. It is an estimate of the potential of a peptide to bind to membranes or other proteins as receptors (Boman et al. Reference Boman, Wade, Boman, Wåhlin and Merrifield1989; Boman Reference Boman2003). The Boman index is determined by summing the solubility values of all residues in a sequence and then normalizing this sum by dividing it by the number of residues. A binding potential of greater than 2.48 indicates that a peptide has a strong binding potential (Boman Reference Boman2003; Karnati et al., Reference Karnati, Gonuguntala, Barbadikar, Mishra, Jha, Prakasham, Chilumula, Shaik, Pesari, Sundaram and Chinnaswami2022). Only a small proportion of 316 identified peptides (18%) were defined as having a high binding potential (Fig. 5C).
Aromacity describes whether a peptide sequence contains at least one of the aromatic amino acids (phenylalanine, tryptophan or tyrosine). The 316 peptides identified by the three distinct solvents (Supplementary Table S1) had a higher proportion of aromatic peptides (62%) (Fig. 5D).
We compared the properties of peptides extracted by different solvents by focusing on the peptides commonly identified in all three biological replicates for each solvent (FA–100, MeOH–H2O–AcOH–87 and TFA–71). We expressed the number of peptides with specific properties (aromaticity, binding potential, polarity and stability) as a proportion of the total commonly identified peptides for each solvent (Fig. 5E).
Discussion
Peptides are increasingly recognized for their diverse bioactivities, playing crucial roles in plant–microbe interactions, signalling, defence mechanisms and nutrition (Hill et al., Reference Hill, Farrell and Jones2012; Hu et al., Reference Hu, Zhang and Shi2018; Li et al., Reference Li, Hu, Jian, Xie and Yang2021; Pantigoso et al., Reference Pantigoso, Newberger and Vivanco2022; Furumizu and Shinohara, Reference Furumizu and Shinohara2024). Despite this growing interest, research on peptides exuded by germinating seeds, particularly within the spermosphere, remains limited. The spermosphere represents a highly dynamic microenvironment where plant–microbe interactions can significantly influence seed vigour and protection. The need for further functional studies on seed-derived peptides in this context is crucial, given their potential to improve seed resilience in the face of varying biotic and abiotic stresses (Schiltz et al., Reference Schiltz, Gaillard, Pawlicki-Jullian, Thiombiano, Mesnard and Gontier2015; Feng et al., Reference Feng, Zhu, Xue, Chen and Yu2023). Given the complex composition of seed exudates and the diverse physico-chemical characteristics of peptides, peptide isolation poses significant challenges, necessitating optimized extraction methods (Farrokhi et al., Reference Farrokhi, Whitelegge and Brusslan2008; Finoulst et al., Reference Finoulst, Pinkse, Van Dongen and Verhaert2011; Pinkse et al., Reference Pinkse, Evaristo, Pieterse, Yu and Verhaert2014; Schiltz et al., Reference Schiltz, Gaillard, Pawlicki-Jullian, Thiombiano, Mesnard and Gontier2015). This study represents a novel attempt to address these challenges by extracting and analysing seed-derived peptides from the spermosphere in an untargeted manner.
In the broader context of plant peptide extraction, solvents are generally classified into two categories: water-based solutions, such as acids and buffers, and organic-based solutions, like ethanol–water mixtures (Barashkova and Rogozhin, Reference Barashkova and Rogozhin2020). In this study, we employed two commonly used water-based acidic solvents, TFA and FA, along with an acidified organic solvent, MeOH–H₂O–AcOH. These solvents are widely used for their capacity to selectively precipitate and extract peptides in a single step (Vitorino et al., Reference Vitorino, Barros, Caseiro, Ferreira and Amado2012; Yu et al., Reference Yu, Canales, Glover, Das, Shi, Liu, Keller, Attie and Li2017; Chen et al., Reference Chen, Hwang, Gleaton, Titus and Hamlin2018). Given their differing properties, we hypothesized that each solvent would influence the nature of the extracted peptides. Among the 316 peptides identified, FA demonstrated the highest extraction efficiency, yet each solvent uniquely identified specific peptides. While the solvents efficiently identified a shared set of 56 peptides, no consistent physico-chemical characteristics were favoured by any particular solvent (Fig. 5E). A PCA of peptide peak intensities further revealed differences in the extraction efficiency of the common peptides, underscoring the significant influence of solvent selection (Fig. 4D).
A notable limitation of our experimental design was the use of a small seed sample size (10 seeds per biological replicate), which likely limited our ability to effectively minimize biological variability (Barashkova and Rogozhin, Reference Barashkova and Rogozhin2020). This, combined with technical variability associated with solvent extraction, may have influenced the reproducibility of the results. To better account for biological variability, future studies should increase the number of seeds per sample and examine multiple genotypes to capture a broader spectrum of peptide diversity (Vitorino et al., Reference Vitorino, Barros, Caseiro, Ferreira and Amado2012). Moreover, given the dynamic nature of the spermosphere, focusing on a single time point (24 h after imbibition) may have limited our ability to capture the full temporal dynamics of peptide release. It is also important to note that factors such as the seed's water content prior to imbibition, as well as the temperature during imbibition, significantly influence both the quantity and composition of seed exudates, particularly in larger seeds like beans (Schroth et al., Reference Schroth, Weinhold and Hayman1966; Toledo et al., Reference Toledo, Cavariani, França-Neto and Nakagawa2010). Studying the peptidome at multiple time points and diverse environments would provide a more comprehensive understanding of the changes in peptide composition throughout seed germination (Nelson, Reference Nelson2004; Schiltz et al., Reference Schiltz, Gaillard, Pawlicki-Jullian, Thiombiano, Mesnard and Gontier2015).
Our untargeted peptidome analysis successfully identified peptides ranging from 5 to 29 amino acids in length. However, a well-recognized challenge in mass spectrometry-based peptidomics is the identification of very short peptides (<5 amino acids) (Nongonierma and FitzGerald, Reference Nongonierma and FitzGerald2017). These peptides are difficult to detect due to poor fragmentation patterns and the limited structural diversity that complicates their assignment (Fricker, Reference Fricker2015; Hollebrands et al., Reference Hollebrands, Hageman, van de Sande, Albada and Janssen2023). Moreover, existing protein databases often exclude short peptides, increasing the likelihood of false discoveries (Miller and Smith, Reference Miller and Smith2023). To overcome these challenges, future research should consider incorporating De Novo sequencing, specialized RT prediction models or alternative workflows better suited for short peptide detection and identification (Piovesana et al., Reference Piovesana, Capriotti, Cavaliere, La Barbera, Montone, Zenezini Chiozzi and Laganà2018; Hollebrands et al., Reference Hollebrands, Hageman, van de Sande, Albada and Janssen2023).
Our methodology not only allowed for the identification of peptide sequences but also provided insights into their predicted properties, such as stability, polarity and aromaticity (Rustagi et al., Reference Rustagi, Gupta, Bajaj, Singh and Singh2023). These characteristics suggest significant potential for biological activity, aligning with recent research into the functional roles of peptides. The findings of Wang et al. (Reference Wang, Tian, Liu, Li, Zhang, Chen, Jia, Zheng, Wu, Chen, Yan and Wu2020) further underscore the potential functional diversity of peptides in plants, indicating that many NCPs may play significant roles in key biological processes. This includes the proposed functions of seed-derived peptides in influencing and shaping microbial communities within the spermosphere.
Conclusion
Our study represents a pioneering effort to develop a methodology for analysing peptide diversity in the seed spermosphere, focusing on establishing an untargeted approach to detect the presence of peptides and evaluate the impact of different solvents on extraction efficiency. This work lays the groundwork for future investigations by demonstrating the importance of solvent choice in peptide recovery and characterization. For more comprehensive studies, future research should consider pooling peptides extracted with various solvents and including a broader range of genotypes to capture a wider diversity of peptides. Experimental protocols and peptidomic analysis pipelines should be tailored to address specific biological questions, such as the roles of these peptides in plant–microbe interactions and their potential applications in seed technology. Furthermore, although factors such as sample size, temporal dynamics, genetic diversity and environmental seed conditions were not evaluated in this study, they are crucial considerations for optimizing peptide analysis outcomes and should be informed by the specific biological context of future investigations.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0960258525000030.
Data availability statement
The data supporting the findings of this study are openly available in online repositories. Peptidomics data are available on iProX at https://www.iprox.cn/ and are referenced under the Project ID: IPX0008926001.
Author contributions
C.S., B.B., C.B., and B.C. conducted experiments. C.S., B.B., S.P., and L.R. analysed the results. C.S. and L.R. wrote the original draft preparation. C.S., B.B., M.C., and L.R. wrote, reviewed and edited the manuscript. All authors have read and agreed to the published version of the manuscript.
Funding statement
This work was part of the 3rd Programme for Future Investments (France2030). It is operated by the SUCSEED project (ANR-20-PCPA-0009) and funded by the ‘Growing and Protecting Crops Differently’ French Priority Research Program (PPR-CPA), part of the national investment plan operated by the French National Research Agency (ANR). The IJPB benefits from the support of Saclay Plant Sciences – SPS (ANR-17-EUR-0007).
Competing interests
The authors declare no conflicts of interest.







